Influence of the Chemical Properties of Cereal Grains on the Structure and Metabolism of the Bacteriome of Rhyzopertha dominica (F.) and Its Development: A Cause–Effect Analysis
Abstract
:1. Introduction
2. Results
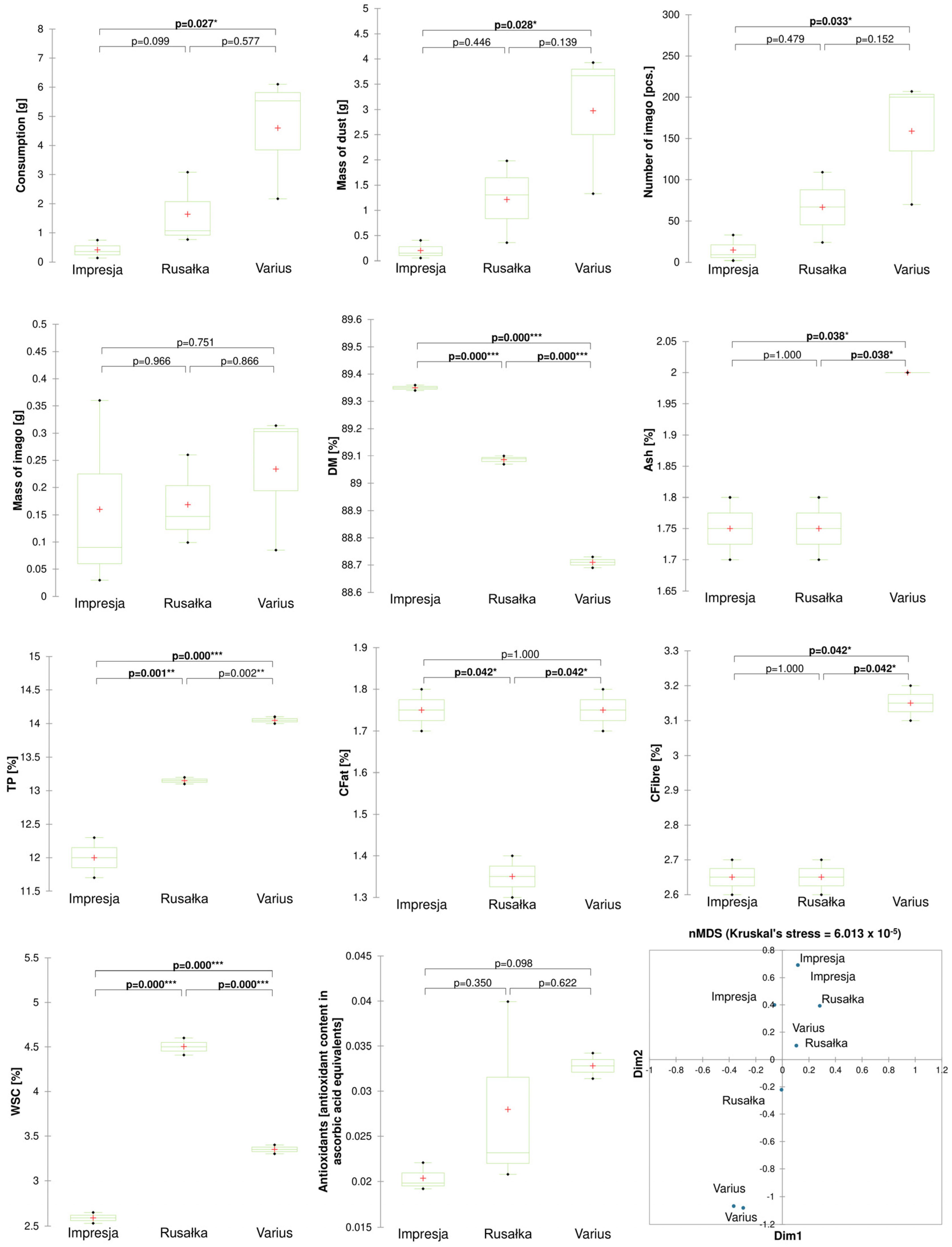
2.1. Relationship between the Development of R. dominica and Chemical Parameters of Cereal Grains
2.2. Analysis of the Bacteriobiome of R. dominica
2.3. Metabolism of the R. dominica Bacteriobiome
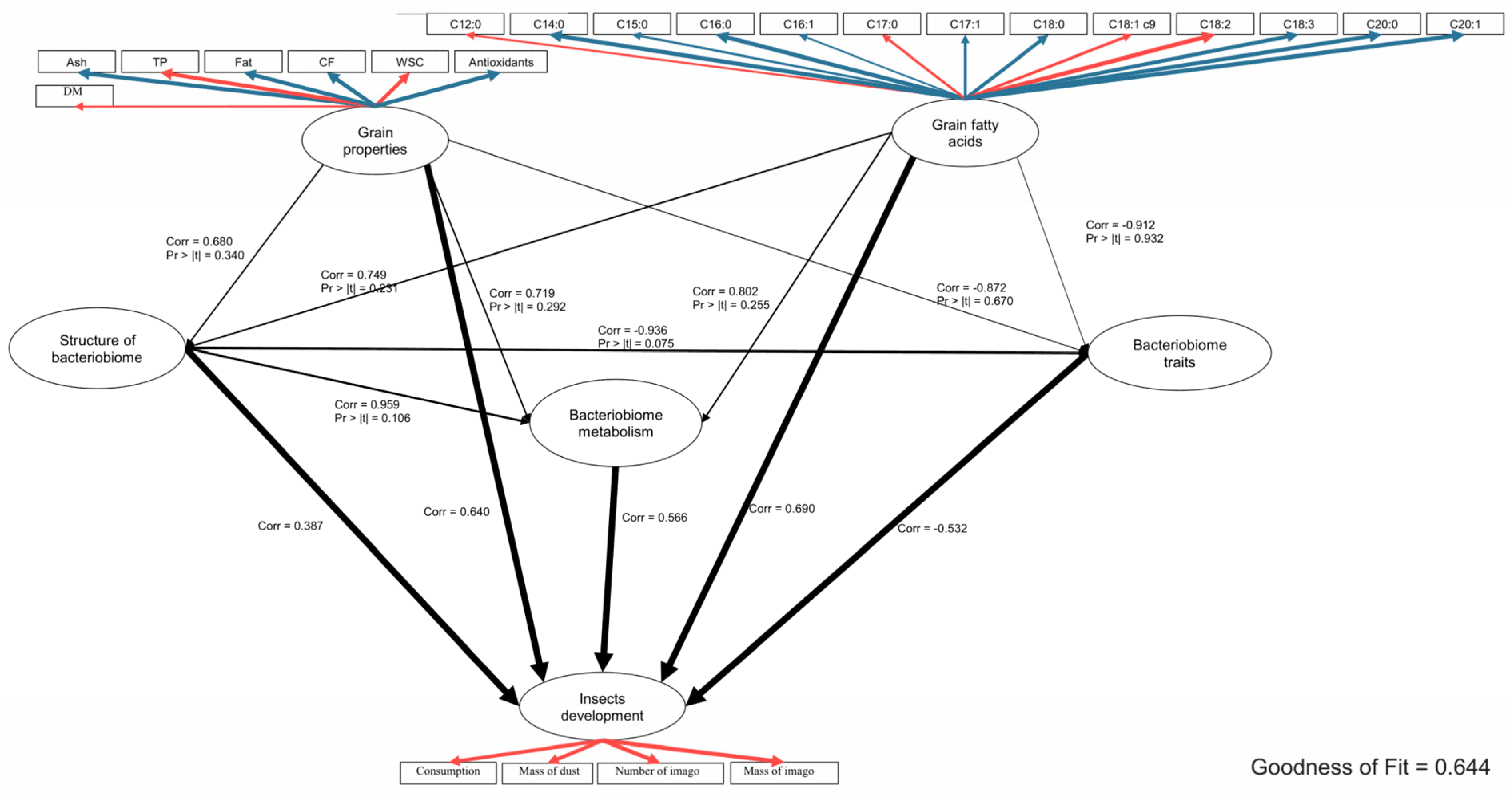
2.4. Global Analysis of the Relationships between the Tested Parameters
3. Discussion
4. Materials and Methods
4.1. Experimental Design
4.2. Experiment with Different Feed Variants
4.3. Chemical Analyses of Grains
4.4. Biochemical Properties of Insects
4.5. Statistical Calculations
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Oppert, B.; Muszewska, A.; Steczkiewicz, K.; Šatović-Vukšić, E.; Plohl, M.; Fabrick, J.A.; Vinokurov, K.S.; Koloniuk, I.; Johnston, J.S.; Smith, T.P.L.; et al. The Genome of Rhyzopertha dominica (Fab.) (Coleoptera: Bostrichidae): Adaptation for Success. Genes 2022, 13, 446. [Google Scholar] [CrossRef] [PubMed]
- Ren, Y.; Wang, T.; Wang, C.; D’Isita, I.; Hu, Q.; Germinara, G.S.; Cao, Y. Population Development of Rhyzopertha dominica (F.) (Coleoptera: Bostrichidae) on Different Stored Products. Entomol. Res. 2023, 53, 359–366. [Google Scholar] [CrossRef]
- Phillips, T.W.; Throne, J.E. Biorational Approaches to Managing Stored-Product Insects. Annu. Rev. Entomol. 2010, 55, 375–397. [Google Scholar] [CrossRef] [PubMed]
- Desta, A.G.; Yeshitila, B.H. The Distribution of Insect Pests and the Associated Loss of Stored Sorghum in the Kena District of Konso Zone, South-Western Ethiopia. PLoS ONE 2024, 19, e0295833. [Google Scholar] [CrossRef] [PubMed]
- Mesterházy, Á.; Oláh, J.; Popp, J. Losses in the Grain Supply Chain: Causes and Solutions. Sustainability 2020, 12, 2342. [Google Scholar] [CrossRef]
- Daglish, G.J.; Nayak, M.K. Prevalence of Resistance to Deltamethrin in Rhyzopertha dominica (F.) in Eastern Australia. J. Stored Prod. Res. 2018, 78, 45–49. [Google Scholar] [CrossRef]
- Kosewska, O.; Przemieniecki, S.W.; Nietupski, M. The Effect of Antibiotics on Bacteriome of Sitophilus Oryzae and Rhyzopertha Dominica as a Factor Determining the Success of Foraging: A Chance for Antibiotic Therapy in Grain Stores. Appl. Sci. 2023, 13, 1576. [Google Scholar] [CrossRef]
- Klejdysz, T.; Nawrot, J. First Record of Outdoor Occurrence of Stored-Product Coleopterans in Arable Landscape in Poland. J. Plant Prot. Res. 2010, 50, 551–553. [Google Scholar] [CrossRef]
- Ebadollahi, A.; Borzoui, E. Growth Performance and Digestive Enzymes Activity of Rhyzopertha dominica (F.) (Coleoptera: Bostrichidae) Feeding on Six Rice Cultivars. J. Stored Prod. Res. 2019, 82, 48–53. [Google Scholar] [CrossRef]
- Wilkes Walburn, J.; Wemheuer, B.; Thomas, T.; Copeland, E.; O’Connor, W.; Booth, M.; Fielder, S.; Egan, S. Diet and Diet-associated Bacteria Shape Early Microbiome Development in Yellowtail Kingfish (Seriola lalandi). Microb. Biotechnol. 2019, 12, 275–288. [Google Scholar] [CrossRef]
- Dillon, R.J.; Dillon, V.M. The Gut Bacteria of Insects: Nonpathogenic Interactions. Annu. Rev. Entomol. 2004, 49, 71–92. [Google Scholar] [CrossRef] [PubMed]
- Engel, P.; Moran, N.A. The Gut Microbiota of Insects—Diversity in Structure and Function. FEMS Microbiol. Rev. 2013, 37, 699–735. [Google Scholar] [CrossRef] [PubMed]
- Shamjana, U.; Vasu, D.A.; Hembrom, P.S.; Nayak, K.; Grace, T. The Role of Insect Gut Microbiota in Host Fitness, Detoxification and Nutrient Supplementation. Antonie Van Leeuwenhoek 2024, 117, 71. [Google Scholar] [CrossRef] [PubMed]
- Martignoni, M.M.; Kolodny, O. Microbiome Transfer from Native to Invasive Species May Increase Invasion Risk and Shorten Invasion Lag Statements and Declarations. bioRxiv 2023. [Google Scholar] [CrossRef]
- Kolodny, O.; Callahan, B.J.; Douglas, A.E. The Role of the Microbiome in Host Evolution. Philos. Trans. R. Soc. B Biol. Sci. 2020, 375, 20190588. [Google Scholar] [CrossRef]
- Wielkopolan, B.; Krawczyk, K.; Szabelska-Beręsewicz, A.; Obrępalska-Stęplowska, A. The Structure of the Cereal Leaf Beetle (Oulema melanopus) Microbiome Depends on the Insect’s Developmental Stage, Host Plant, and Origin. Sci. Rep. 2021, 11, 20496. [Google Scholar] [CrossRef]
- Fontaine, S.S.; Mineo, P.M.; Kohl, K.D. Experimental Manipulation of Microbiota Reduces Host Thermal Tolerance and Fitness under Heat Stress in a Vertebrate Ectotherm. Nat. Ecol. Evol. 2022, 6, 405–417. [Google Scholar] [CrossRef]
- Kikuchi, Y.; Hayatsu, M.; Hosokawa, T.; Nagayama, A.; Tago, K.; Fukatsu, T. Symbiont-Mediated Insecticide Resistance. Proc. Natl. Acad. Sci. USA 2012, 109, 8618–8622. [Google Scholar] [CrossRef]
- Muhammad, Z.M.; Tahir, M.; Mudassar, J.; Fatima, S.; Muhammad, A.R. Muhammad A Biology and Management of Stored Products’ Insect Pest Rhyzopertha dominica (Fab.) (Coleoptera: Bostrichidae). Int. J. Biosci. (IJB) 2015, 7, 78–93. [Google Scholar] [CrossRef]
- Montagna, M.; Chouaia, B.; Mazza, G.; Prosdocimi, E.M.; Crotti, E.; Mereghetti, V.; Vacchini, V.; Giorgi, A.; De Biase, A.; Longo, S.; et al. Effects of the Diet on the Microbiota of the Red Palm Weevil (Coleoptera: Dryophthoridae). PLoS ONE 2015, 10, e0117439. [Google Scholar] [CrossRef]
- Xue, D.; Chen, T.; Li, Q.; Yang, Y.; Wu, Y. Microbiota Composition of Allopatric Laboratory and Wild Populations of Rhyzopertha dominica. J. Stored Prod. Res. 2023, 104, 102202. [Google Scholar]
- Kešnerová, L.; Mars, R.A.T.; Ellegaard, K.M.; Troilo, M.; Sauer, U.; Engel, P. Disentangling Metabolic Functions of Bacteria in the Honey Bee Gut. PLoS Biol. 2017, 15, e2003467. [Google Scholar]
- Nietupski, M.; Kwiatkowski, J.; Kosewska, A. Physicochemical Properties of Achenes of Buckwheat Cultivars Affecting the Development of Grain Weevil (Sitophilus granarius L.) and Lesser Grain Borer (Rhyzopertha dominica F.). Zemdirb.-Agric. 2017, 104, 311–320. [Google Scholar]
- Perisic, V.; Perisic, V.; Vukajlović, F.N.; Pesic, S.B. Feeding Preferences and Progeny Production of Rhyzopertha dominica (Fabricius 1792) (Coleoptera: Bostrichidae) in Small Grains. Biol. Nyssana 2018, 9, 55–61. [Google Scholar]
- Kosewska, O.; Przemieniecki, S.W.; Koronkiewicz, S.; Nietupski, M. Effect of Different Chemical Properties of Cereal Grains on the Foraging and Microbiome of the Rice Weevil (Sitophilus oryzae L.). Int. Agrophys. 2024, 38, 165–176. [Google Scholar]
- Lorini, I.; Galley, D.J. Deltamethrin Resistance in Rhyzopertha dominica (F.) (Coleoptera: Bostrichidae), a Pest of Stored Grain in Brazil. J. Stored Prod. Res. 1999, 35, 37–45. [Google Scholar]
- Edde, P.A. A Review of the Biology and Control of Rhyzopertha dominica (F.) the Lesser Grain Borer. J. Stored Prod. Res. 2012, 48, 1–18. [Google Scholar]
- Mansour, K. On the Intracellular Microorganisms of Some Bostrychid Beetles. J. Cell Sci. 1934, s2-77, 243–253. [Google Scholar] [CrossRef]
- Okude, G.; Koga, R.; Hayashi, T.; Nishide, Y.; Meng, X.-Y.; Nikoh, N.; Miyanoshita, A.; Fukatsu, T. Novel Bacteriocyte-Associated Pleomorphic Symbiont of the Grain Pest Beetle Rhyzopertha dominica (Coleoptera: Bostrichidae). Zool. Lett. 2017, 3, 13. [Google Scholar]
- Kordan, B.; Nietupski, M.; Ludwiczak, E.; Gabryś, B.; Cabaj, R. Selected Cultivar-Specific Parameters of Wheat Grain as Factors Influencing Intensity of Development of Grain Weevil Sitophilus granarius (L.). Agriculture 2023, 13, 1492. [Google Scholar] [CrossRef]
- Cinco-Moroyoqui, F.J.; Rosas-Burgos, E.C.; Borboa-Flores, J.; Cortez-Rocha, M.O. α-Amylase Activity of Rhyzopertha dominica (Coleoptera: Bostrichidae) Reared on Several Wheat Varieties and Its Inhibition with Kernel Extracts. J. Econ. Entomol. 2006, 99, 2146–2150. [Google Scholar] [CrossRef] [PubMed]
- Mariey, S.A.; Mohamed, E.N.M.; Nasr, G.M.; Ahmed, K.R.; Elsamahy, B.E. Comparative Efficacy of Phenol Concentration against Rhyzopertha dominica (F.) in Some Hulled Barley Cultivars Productivity and Grain Quality. J. Glob. Ecol. Environ. 2023, 18, 1–13. [Google Scholar] [CrossRef]
- Nietupski, M.; Ludwiczak, E.; Cabaj, R.; Purwin, C.; Kordan, B. Fatty Acids Present in Wheat Kernels Influence the Development of the Grain Weevil (Sitophilus granarius L.). Insects 2021, 12, 806. [Google Scholar] [CrossRef]
- Kordan, B.; Skrajda-Brdak, M.; Tańska, M.; Konopka, I.; Cabaj, R.; Załuski, D. Phenolic and Lipophilic Compounds of Wheat Grain as Factors Affecting Susceptibility to Infestation by Granary Weevil (Sitophilus granarius L.). J. Appl. Bot. Food Qual. 2019, 92, 64–72. [Google Scholar]
- Kosewska, O.; Nietupski, M.; Koronkiewicz, S.; Przemieniecki, S.W. The Chemical Grain Composition of Wheat and Barley Affects the Development of the Lesser Grain Borer (Rhyzopertha dominica F.) and the Rice Weevil (Sitophilus oryzae L.). J. Plant Prot. Res. 2024. [Google Scholar] [CrossRef]
- Grundmann, C.O.; Guzman, J.; Vilcinskas, A.; Pupo, M.T. The Insect Microbiome Is a Vast Source of Bioactive Small Molecules. Nat. Prod. Rep. 2024, 41, 935–967. [Google Scholar] [CrossRef]
- Gołębiowski, M.; Maliński, E.; Boguś, M.I.; Kumirska, J.; Stepnowski, P. The Cuticular Fatty Acids of Calliphora Vicina, Dendrolimus Pini and Galleria Mellonella Larvae and Their Role in Resistance to Fungal Infection. Insect Biochem. Mol. Biol. 2008, 38, 619–627. [Google Scholar] [CrossRef]
- Walterson, A.M.; Stavrinides, J. Pantoea: Insights into a Highly Versatile and Diverse Genus within the Enterobacteriaceae. FEMS Microbiol. Rev. 2015, 39, 968–984. [Google Scholar] [CrossRef]
- Elston, K.M.; Leonard, S.P.; Geng, P.; Bialik, S.B.; Robinson, E.; Barrick, J.E. Engineering Insects from the Endosymbiont Out. Trends Microbiol. 2022, 30, 79–96. [Google Scholar] [CrossRef]
- Mondal, S.; Somani, J.; Roy, S.; Babu, A.; Pandey, A.K. Insect Microbial Symbionts: Ecology, Interactions, and Biological Significance. Microorganisms 2023, 11, 2665. [Google Scholar] [CrossRef]
- Brunetti, M.; Magoga, G.; Gionechetti, F.; De Biase, A.; Montagna, M. Does Diet Breadth Affect the Complexity of the Phytophagous Insect Microbiota? The Case Study of Chrysomelidae. Environ. Microbiol. 2022, 24, 3565–3579. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Cobas, A.E.; Maiques, E.; Angelova, A.; Carrasco, P.; Moya, A.; Latorre, A. Diet Shapes the Gut Microbiota of the Omnivorous Cockroach Blattella germanica. FEMS Microbiol. Ecol. 2015, 91, fiv022. [Google Scholar] [CrossRef] [PubMed]
- Chouaia, B.; Goda, N.; Mazza, G.; Alali, S.; Florian, F.; Gionechetti, F.; Callegari, M.; Gonella, E.; Magoga, G.; Fusi, M.; et al. Developmental Stages and Gut Microenvironments Influence Gut Microbiota Dynamics in the Invasive Beetle Popillia japonica Newman (Coleoptera: Scarabaeidae). Environ. Microbiol. 2019, 21, 4343–4359. [Google Scholar] [CrossRef] [PubMed]
- Muturi, E.J.; Dunlap, C.; Ramirez, J.L.; Rooney, A.P.; Kim, C.-H. Host Blood Meal Source Has a Strong Impact on Gut Microbiota of Aedes Aegypti. FEMS Microbiol. Ecol. 2018, 95, fiy213. [Google Scholar] [CrossRef] [PubMed]
- Kiefer, J.S.T.; Bauer, E.; Okude, G.; Fukatsu, T.; Kaltenpoth, M.; Engl, T. Cuticle Supplementation and Nitrogen Recycling by a Dual Bacterial Symbiosis in a Family of Xylophagous Beetles. ISME J. 2023, 17, 1029–1039. [Google Scholar] [CrossRef] [PubMed]
- Le Boulch, M.; Déhais, P.; Combes, S.; Pascal, G. The MACADAM Database: A MetAboliC PAthways DAtabase for Microbial Groups for Mining Potential Metabolic Capacities of Archaeal and Bacterial Taxonomic Groups. Available online: https://macadam.toulouse.inra.fr/ (accessed on 13 July 2024).
- Metacyc. Available online: https://metacyc.org/pathway?orgid=META&id=PWY-5703 (accessed on 13 July 2024).
- Metacyc. Available online: https://metacyc.org/pathway?orgid=META&id=PWY-5704 (accessed on 13 July 2024).
- Jing, T.-Z.; Qi, F.-H.; Wang, Z.-Y. Most Dominant Roles of Insect Gut Bacteria: Digestion, Detoxification, or Essential Nutrient Provision? Microbiome 2020, 8, 38. [Google Scholar] [CrossRef] [PubMed]
- Rupawate, P.S.; Roylawar, P.; Khandagale, K.; Gawande, S.; Ade, A.B.; Jaiswal, D.K.; Borgave, S. Role of Gut Symbionts of Insect Pests: A Novel Target for Insect-Pest Control. Front. Microbiol. 2023, 14, 1146390. [Google Scholar] [CrossRef]
- McCutcheon, J.P.; Moran, N.A. Functional Convergence in Reduced Genomes of Bacterial Symbionts Spanning 200 My of Evolution. Genome Biol. Evol. 2010, 2, 708–718. [Google Scholar] [CrossRef]
- Coolen, S.; Rogowska-van der Molen, M.; Welte, C.U. The Secret Life of Insect-Associated Microbes and How They Shape Insect–Plant Interactions. FEMS Microbiol. Ecol. 2022, 98, fiac083. [Google Scholar] [CrossRef]
- Nagel, R.; Turrini, P.C.G.; Nett, R.S.; Leach, J.E.; Verdier, V.; Van Sluys, M.; Peters, R.J. An operon for production of bioactive gibberellin A4 phytohormone with wide distribution in the bacterial rice leaf streak pathogen Xanthomonas oryzae pv. oryzicola. New Phytol. 2017, 214, 1260–1266. [Google Scholar] [CrossRef]
- COBORU Research Centre for Cultivar Testing. Cultivars Data. Available online: https://www.coboru.gov.pl/en/search (accessed on 13 July 2024).
- Golebiowska, Z. The Feeding and Fecundity of Sitophilus granarius (L.), Sitophilus orvzae (L.) and Rhyzopertha dominica (F.) in Wheat Grain. J. Stored Prod. Res. 1969, 5, 143–155. [Google Scholar] [CrossRef]
- Arthur Thomas, T. An Automated Procedure for the Determination of Soluble Carbohydrates in Herbage. J. Sci. Food Agric. 1977, 28, 639–642. [Google Scholar] [CrossRef]
- Benzie, I.F.F.; Strain, J.J. The Ferric Reducing Ability of Plasma (FRAP) as a Measure of “Antioxidant Power”: The FRAP Assay. Anal. Biochem. 1996, 239, 70–76. [Google Scholar] [CrossRef] [PubMed]
- Przemieniecki, S.W.; Kosewska, A.; Kosewska, O.; Purwin, C.; Lipiński, K.; Ciesielski, S. Polyethylene, Polystyrene and Lignocellulose Wastes as Mealworm (Tenebrio molitor L.) Diets and Their Impact on the Breeding Condition, Biometric Parameters, Metabolism, and Digestive Microbiome. Sci. Total Environ. 2022, 832, 154758. [Google Scholar] [CrossRef] [PubMed]
- Hammer, D.A.T.; Ryan, P.D.; Hammer, Ø.; Harper, D.A.T. Past: Paleontological Statistics Software Package for Education and Data Analysis. Palaeontol. Electron. 2001, 4. Available online: http://palaeo-electronica.org/2001_1/past/issue1_01.htm (accessed on 31 July 2024).
- Przemieniecki, S.W.; Damszel, M.; Ciesielski, S.; Kubiak, K.; Mastalerz, J.; Sierota, Z.; Gorczyca, A. Bacterial Microbiome in Armillaria Ostoyae Rhizomorphs Inhabiting the Root Zone during Progressively Dying Scots Pine. Appl. Soil Ecol. 2021, 164, 103929. [Google Scholar] [CrossRef]
- Louca, S.; Parfrey, L.W.; Doebeli, M. Decoupling Function and Taxonomy in the Global Ocean Microbiome. Science (1979) 2016, 353, 1272–1277. [Google Scholar] [CrossRef]
- Lumivero XLSTAT Basic Solutions. Available online: https://www.xlstat.com/en/solutions/basic (accessed on 13 July 2024).
- Jacomy, M.; Venturini, T.; Heymann, S.; Bastian, M. ForceAtlas2, a Continuous Graph Layout Algorithm for Handy Network Visualization Designed for the Gephi Software. PLoS ONE 2014, 9, e98679. [Google Scholar] [CrossRef]
- Bastian, M.; Heymann, S.; Jacomy, M. Gephi: An Open Source Software for Exploring and Manipulating Networks. Proc. Int. AAAI Conf. Web Soc. Media 2009, 3, 361–362. [Google Scholar] [CrossRef]





| Generation | Generation I | Generation II | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Species of the | Background without Feed | Wheat | Barley | Wheat | Barley | ||||||||
| Cultivar | Impresja | Rusałka | Varius | Trofeum | Radek | Ismena | Impresja | Rusałka | Varius | Trofeum | Radek | Ismena | |
| g__Pantoea | 23.3 | 18.9 | 18.6 | 24.2 | 29.2 | 12.9 | 12.3 | 19.7 | 35.3 | 36.0 | 28.0 | 10.8 | 10.1 |
| g__Ralstonia | 21.6 | 14.3 | 11.4 | 27.7 | 8.9 | 6.9 | 22.8 | 19.9 | 3.3 | 11.0 | 19.2 | 5.1 | 15.6 |
| g__Staphylococcus | 6.0 | 20.3 | 28.7 | 0.0 | 14.5 | 54.9 | 0.7 | 0.0 | 0.0 | 0.0 | 0.0 | 7.8 | 0.4 |
| g__Pseudomonas | 9.3 | 17.1 | 9.3 | 11.0 | 10.1 | 6.4 | 11.2 | 6.3 | 5.4 | 8.8 | 10.8 | 5.5 | 22.6 |
| g__Stenotrophomonas | 10.8 | 6.4 | 7.9 | 11.4 | 0.9 | 5.7 | 25.3 | 22.3 | 4.7 | 6.1 | 1.2 | 1.8 | 5.4 |
| g__Massilia | 7.6 | 5.3 | 4.8 | 10.3 | 8.5 | 2.2 | 7.4 | 9.5 | 5.2 | 12.4 | 9.9 | 1.6 | 8.1 |
| g__Acinetobacter | 0.0 | 0.3 | 0.5 | 0.6 | 12.8 | 0.8 | 1.4 | 1.4 | 21.1 | 7.4 | 7.1 | 4.0 | 13.8 |
| g__Serratia | 0.0 | 0.4 | 0.7 | 1.7 | 6.4 | 0.6 | 0.0 | 10.3 | 17.7 | 9.1 | 14.1 | 0.7 | 11.1 |
| g__Xanthomonas | 0.0 | 3.2 | 8.8 | 6.5 | 0.7 | 4.9 | 7.2 | 2.3 | 0.0 | 2.3 | 1.8 | 14.3 | 0.0 |
| g__Sphingomonas | 6.6 | 4.1 | 2.5 | 3.0 | 1.2 | 1.1 | 2.2 | 3.2 | 2.2 | 2.0 | 3.1 | 1.2 | 1.2 |
| g__Candidatus_Sulcia | 0.0 | 0.0 | 0.0 | 0.0 | 0.5 | 0.0 | 0.6 | 0.0 | 0.0 | 0.0 | 0.0 | 19.6 | 6.4 |
| g__Sphingobium | 4.4 | 2.5 | 0.6 | 0.6 | 0.4 | 1.3 | 1.0 | 1.4 | 1.1 | 1.2 | 0.7 | 0.4 | 0.9 |
| o__Enterobacterales | 7.9 | 2.2 | 4.5 | 0.3 | 0.0 | 0.6 | 3.6 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| g__Caulobacter | 0.0 | 1.8 | 0.3 | 1.4 | 0.5 | 0.7 | 0.7 | 0.5 | 0.7 | 1.3 | 0.9 | 0.0 | 0.4 |
| g__Paenibacillus | 0.0 | 0.5 | 0.3 | 0.8 | 0.7 | 0.0 | 1.1 | 0.8 | 1.0 | 1.0 | 1.1 | 0.0 | 0.7 |
| g__Brevibacterium | 0.0 | 0.0 | 0.0 | 0.0 | 3.1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| g__Bosea | 0.0 | 0.7 | 0.0 | 0.0 | 0.2 | 0.0 | 0.2 | 0.3 | 0.3 | 0.4 | 0.3 | 0.4 | 0.6 |
| g__Rhizobium | 0.0 | 0.3 | 0.0 | 0.0 | 0.0 | 0.7 | 0.6 | 0.0 | 0.5 | 0.3 | 0.5 | 0.0 | 0.3 |
| g__Variovorax | 0.0 | 0.0 | 0.0 | 0.0 | 0.2 | 0.0 | 0.0 | 0.0 | 0.7 | 0.2 | 0.8 | 0.0 | 0.9 |
| g__Burkholderia | 2.7 | 0.0 | 0.0 | 0.0 | 0.3 | 0.0 | 0.0 | 1.0 | 0.2 | 0.5 | 0.0 | 0.0 | 0.4 |
| g__Bacillus | 0.0 | 0.8 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 4.2 | 0.9 |
| g__Cupriavidus | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.4 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| g__Methylobacterium | 0.0 | 0.4 | 0.0 | 0.3 | 0.2 | 0.0 | 0.0 | 1.2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| g__Afipia | 0.0 | 0.6 | 0.2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| g__Escherichia-Shigella | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 2.4 | 0.0 |
| g__Corynebacterium | 0.0 | 0.0 | 0.4 | 0.2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.2 |
| g__Brevundimonas | 0.0 | 0.0 | 0.4 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.5 | 0.0 | 0.0 |
| g__Alcaligenes | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 5.1 | 0.0 |
| g__Sporosarcina | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.7 | 0.0 |
| f__Comamonadaceae | 0.0 | 0.0 | 0.0 | 0.0 | 0.2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.3 | 0.0 | 0.0 | 0.0 |
| g__Conchiformibius | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.4 | 0.0 |
| g__Ancylobacter | 0.0 | 0.0 | 0.0 | 0.0 | 0.4 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| g__Aureimonas | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.3 | 0.0 |
| g__Lysobacter | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.4 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| f__Xanthobacteraceae | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.5 | 0.0 | 0.0 | 0.0 | 0.0 |
| g__Microvirga | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 4.7 | 0.0 |
| g__Pseudarthrobacter | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 4.7 | 0.0 |
| g__Candidatus_Berkiella | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.8 | 0.0 |
| g__Moraxella | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.7 | 0.0 |
| g__Kocuria | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| Eudominant | Dominant | Subdominant | Occasional | Casual | |||||||||
| >10.01% | 5.1–10.0% | 2.1–5.0% | 1.1–2.0% | <1.0% | |||||||||
| Dominance_D | 0.1832 ab | 0.2185 ab | 0.2104 ab | 0.2398 a | 0.6804 a | 0.2033 ab | 0.1595 ab | 0.2579 a | 0.1986 ab | 0.2256 ab | 0.2341 ab | 0.1322 b | |
| Shannon_H | 2.0590 ab | 1.7480 b | 1.8110 b | 1.8460 b | 0.7871 b | 1.8510 b | 2.0770 ab | 1.7390 b | 2.0090 ab | 1.7700 ab | 1.9490 ab | 2.2490 a | |
| Evenness_e^H/S | 0.5848 ab | 0.5223 ab | 0.5982 ab | 0.5278 b | 0.2632 b | 0.5292 ab | 0.6445 a | 0.4831 b | 0.4973 ab | 0.5703 ab | 0.5866 ab | 0.4989 ab | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kosewska, O.; Przemieniecki, S.W.; Nietupski, M. Influence of the Chemical Properties of Cereal Grains on the Structure and Metabolism of the Bacteriome of Rhyzopertha dominica (F.) and Its Development: A Cause–Effect Analysis. Int. J. Mol. Sci. 2024, 25, 10130. https://doi.org/10.3390/ijms251810130
Kosewska O, Przemieniecki SW, Nietupski M. Influence of the Chemical Properties of Cereal Grains on the Structure and Metabolism of the Bacteriome of Rhyzopertha dominica (F.) and Its Development: A Cause–Effect Analysis. International Journal of Molecular Sciences. 2024; 25(18):10130. https://doi.org/10.3390/ijms251810130
Chicago/Turabian StyleKosewska, Olga, Sebastian Wojciech Przemieniecki, and Mariusz Nietupski. 2024. "Influence of the Chemical Properties of Cereal Grains on the Structure and Metabolism of the Bacteriome of Rhyzopertha dominica (F.) and Its Development: A Cause–Effect Analysis" International Journal of Molecular Sciences 25, no. 18: 10130. https://doi.org/10.3390/ijms251810130







