Baicalin Ameliorates Depression-like Behaviors via Inhibiting Neuroinflammation and Apoptosis in Mice
Abstract
:1. Introduction
2. Results
2.1. Structural Information and Druggability of Baicalin
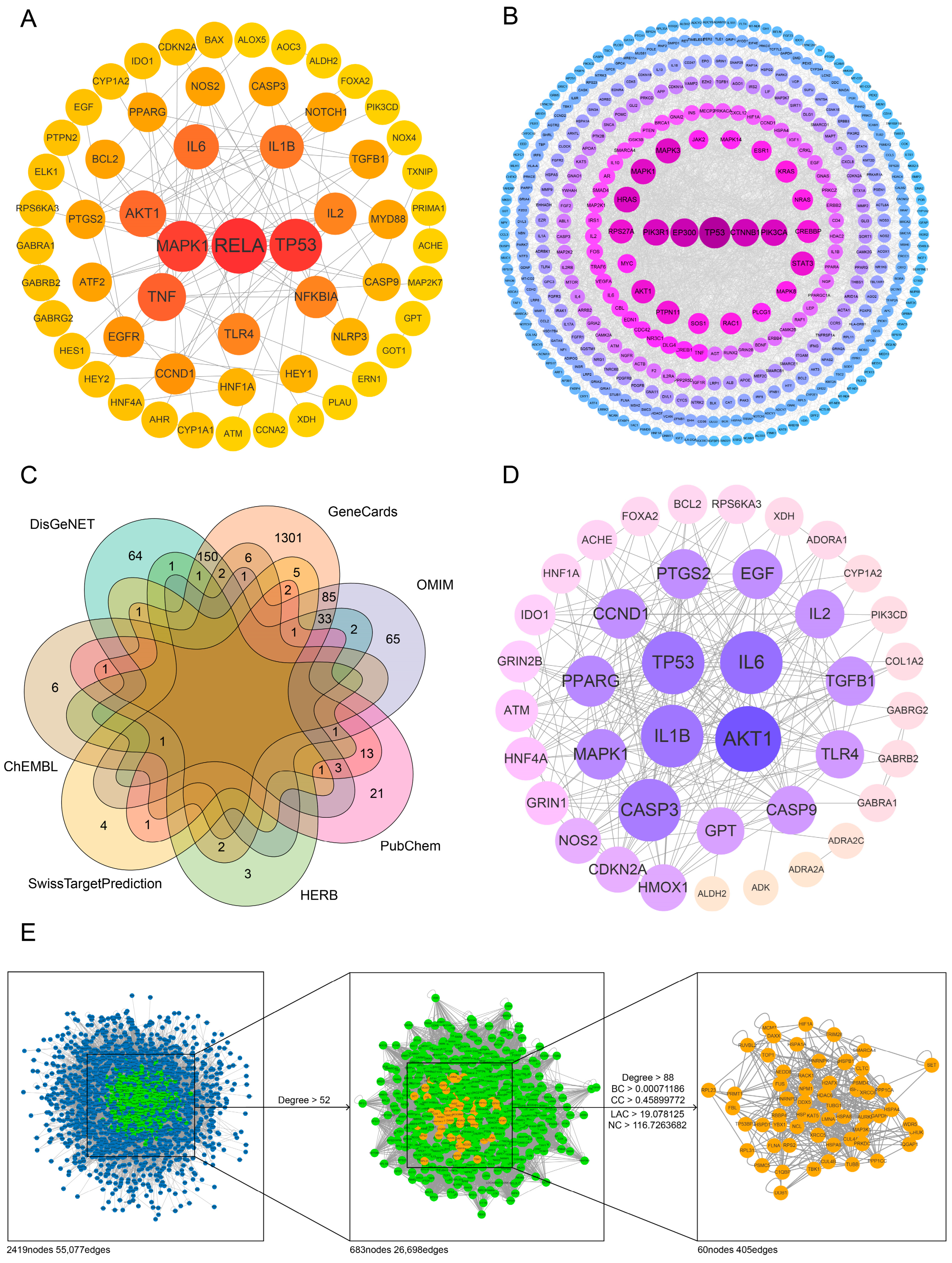
2.2. Network Construction Revealed the Potential Targets of the Antidepressant Effect of Baicalin and Their Interactions
2.2.1. Target Network for Baicalin
2.2.2. Target Network for Depression
2.2.3. Target Network for Antidepressant Effect of Baicalin
2.3. The Peripheral Genes and Secondary Mechanisms of Baicalin’s Antidepressant Effect Were Explored by Topology Analysis of Network Structure
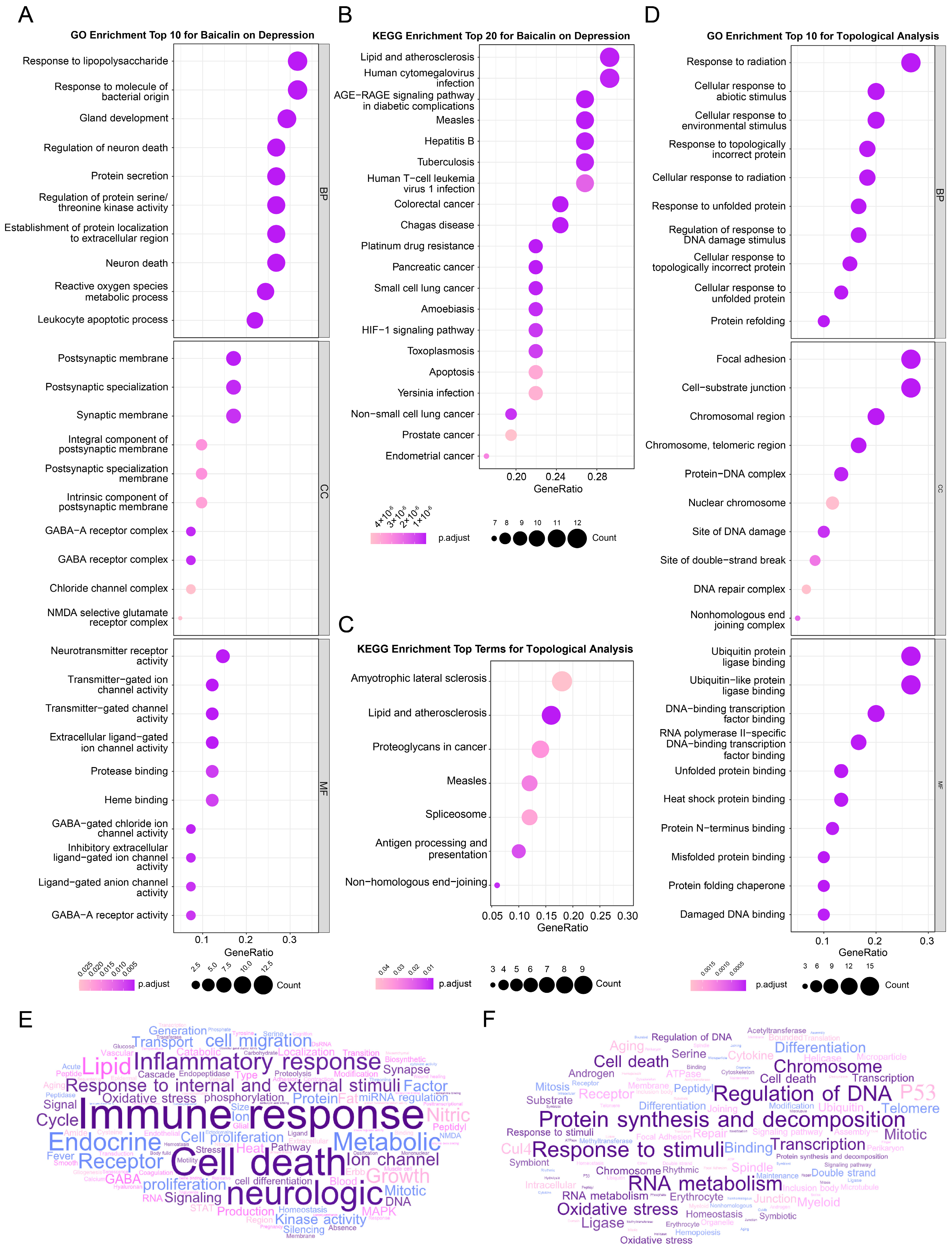
2.4. Enrichment Analysis Clarified the Physiological Mechanism of Baicalin’s Antidepressant Effect
2.4.1. Enrichment and Analysis of Baicalin’s Antidepressant Targets
2.4.2. Enrichment Analysis of Peripheral Genes
2.5. The Results of Molecular Docking Suggest the Possibility of the Interaction between Baicalin and the Target Proteins
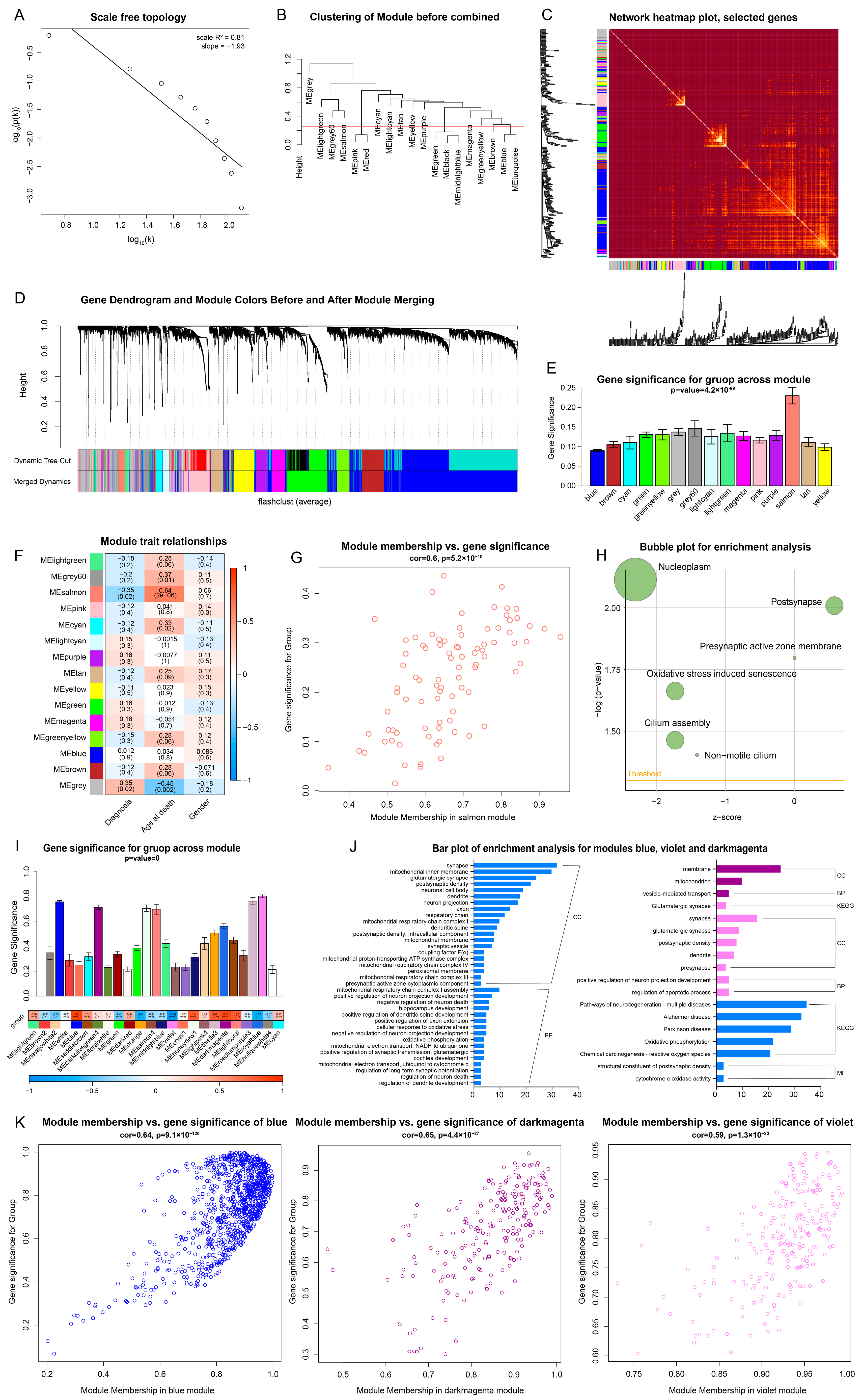
2.6. WGCNA Analysis and Expression Difference Analysis Verified the Effect of Baicalin on Depression
2.6.1. Multiple Pathways in the Hippocampus of Patients with Depression Have Changed
2.6.2. Multiple Pathways in the Hippocampus of Mice with the Treatment of Baicalin Have Changed
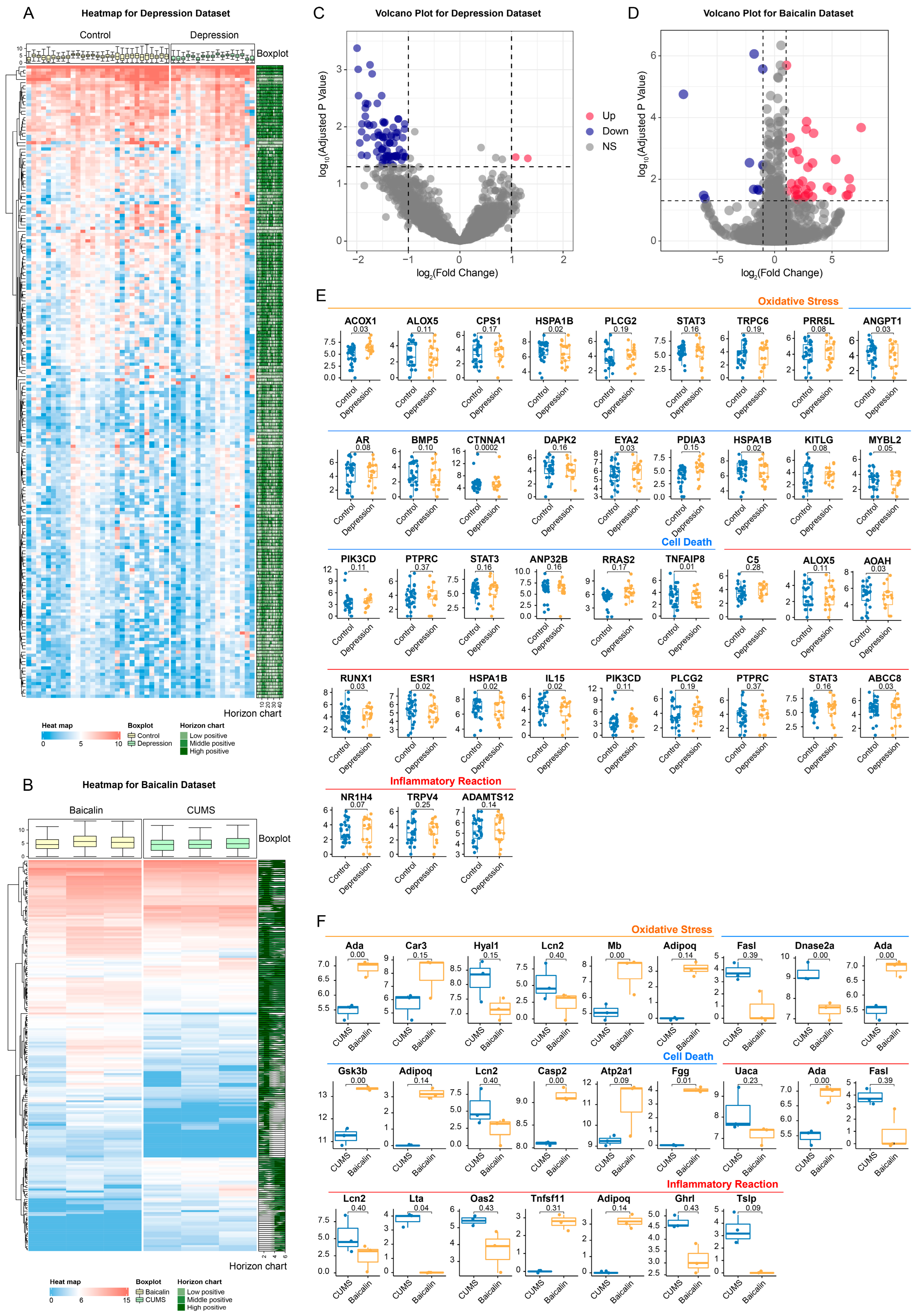
2.6.3. Changes in Gene Expression Level Occur in the Hippocampus of Patients with Depression
2.6.4. Baicalin Can Play an Antidepressant Role Possibly by Regulating the Gene Expression Level
2.7. Animal Experiments Confirmed the Antidepressant Effect of Baicalin
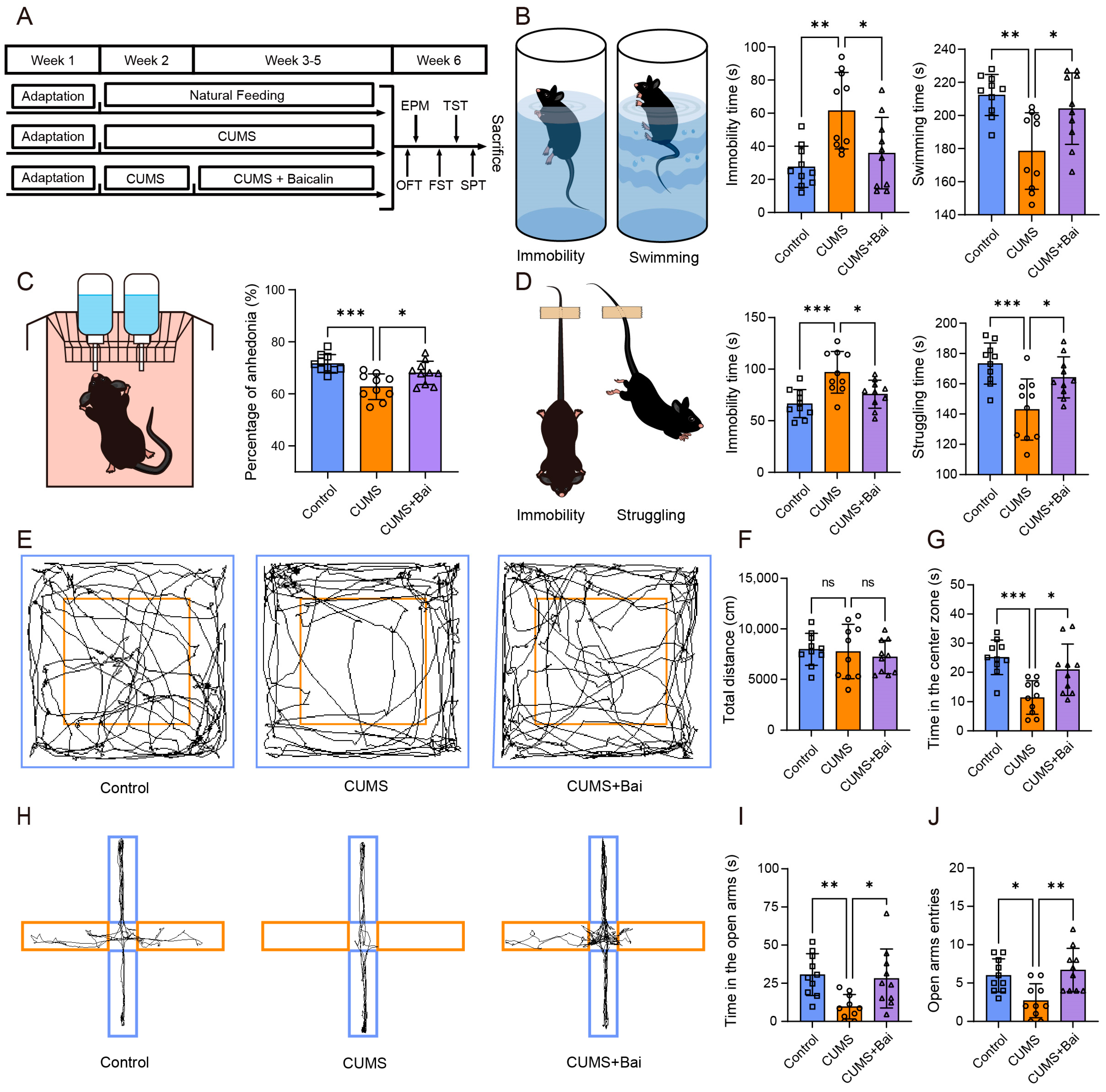
2.7.1. Baicalin Improved the Behavioral Disorder of Depressed Mice Induced by CUMS
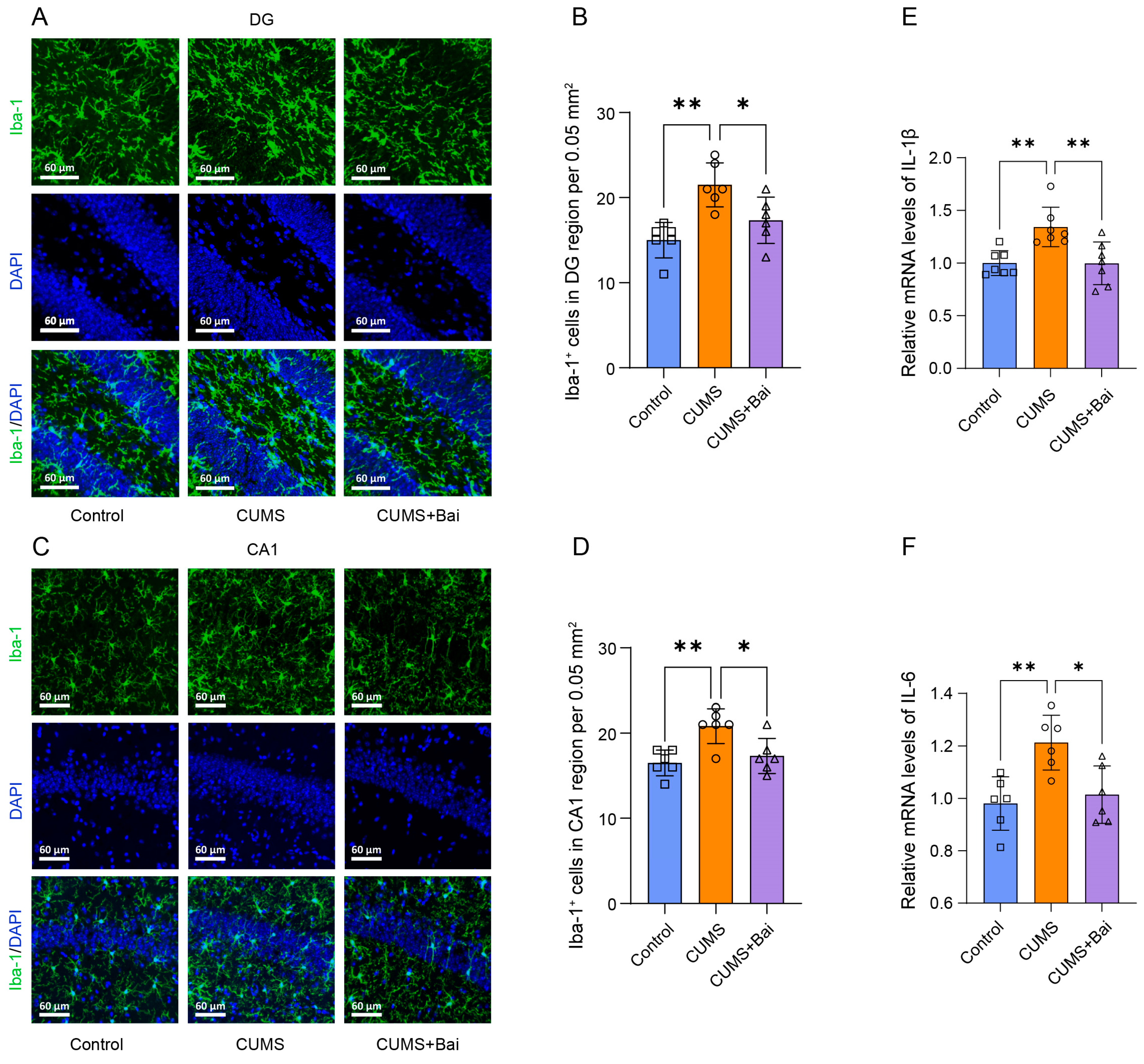
2.7.2. Baicalin Exerted Antidepressant Effects by Inhibiting Neuroinflammation
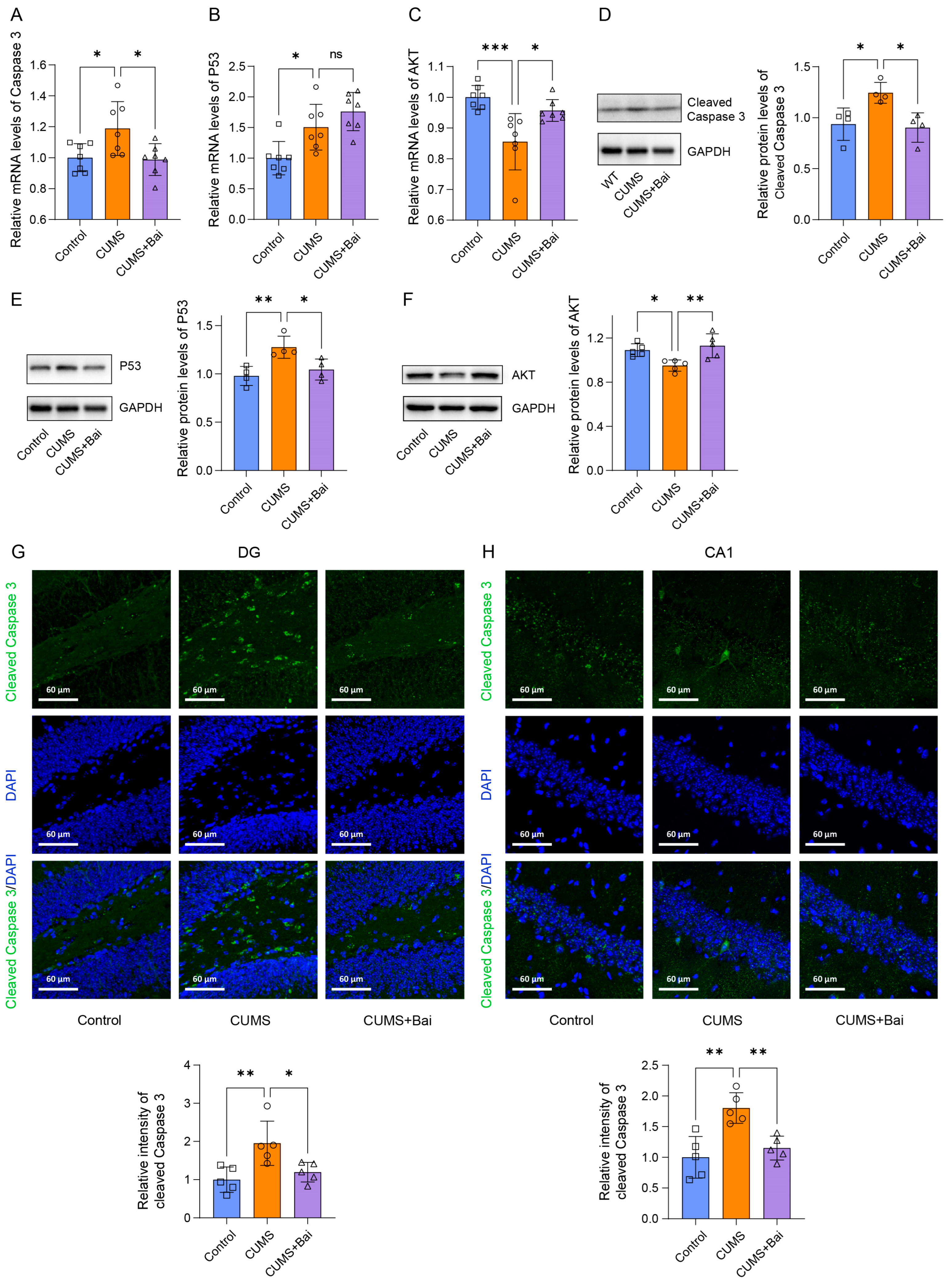
2.7.3. Baicalin Exerted Antidepressant Effects by Inhibiting Apoptosis
3. Discussion
4. Materials and Methods
4.1. Data Preparation
4.1.1. Structural Information and Druggability Evaluation of Baicalin
4.1.2. Targets Prediction for Baicalin
4.1.3. Identification of Targets Associated with Depression
4.1.4. Acquirement of Potential Targets for Baicalin’s Antidepressant Effect
4.1.5. Selection and Preprocessing of High-Throughput Datasets
4.2. Network Construction and Topological Analysis
4.2.1. Target Networks for Baicalin and Depression
4.2.2. Construction of Network for Target Genes of Baicalin on Depression
4.2.3. Topology Analysis
4.3. Enrichment Analysis
4.4. Molecular Docking
4.5. Differential Gene Expression Analysis
4.6. Weighted Gene Co-Expression Network Analysis
4.7. Animals
4.8. Drugs
4.9. Chronic Unpredicted Mild Stress (CUMS) Model and Intragastrical Administration
4.10. Behavioral Tests
4.10.1. Sucrose Preference Test
4.10.2. Forced Swimming Test
4.10.3. Tail Suspension Test
4.10.4. Open Field Test
4.10.5. Elevated Plus Maze Test
4.11. Reverse Transcription PCR (RT-PCR) and Real-Time Quantitative PCR
4.12. Western Blot Analysis
4.13. Immunofluorescence Staining
4.14. Statistical Analysis
4.15. Ethics Approval and Informed Consent
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Smith, K. Mental health: A world of depression. Nature 2014, 515, 181. [Google Scholar] [CrossRef]
- Dattani, S.; Ritchie, H.; Roser, M. Mental Health. Available online: https://ourworldindata.org/mental-health (accessed on 10 May 2022).
- Bueno-Notivol, J.; Gracia-Garcia, P.; Olaya, B.; Lasheras, I.; Lopez-Anton, R.; Santabarbara, J. Prevalence of depression during the COVID-19 outbreak: A meta-analysis of community-based studies. Int. J. Clin. Health Psychol. 2021, 21, 100196. [Google Scholar] [CrossRef]
- World Health Organization. Depression and Other Common Mental Disorders: Global Health Estimates; WHO: Geneva, Switzerland, 2017. [Google Scholar]
- Lively, S.; Schlichter, L.C. Microglia Responses to Pro-inflammatory Stimuli (LPS, IFNgamma+TNFalpha) and Reprogramming by Resolving Cytokines (IL-4, IL-10). Front. Cell Neurosci. 2018, 12, 215. [Google Scholar] [CrossRef]
- Raison, C.L.; Dantzer, R.; Kelley, K.W.; Lawson, M.A.; Woolwine, B.J.; Vogt, G.; Spivey, J.R.; Saito, K.; Miller, A.H. CSF concentrations of brain tryptophan and kynurenines during immune stimulation with IFN-alpha: Relationship to CNS immune responses and depression. Mol. Psychiatry 2010, 15, 393–403. [Google Scholar] [CrossRef]
- Schwarcz, R.; Stone, T.W. The kynurenine pathway and the brain: Challenges, controversies and promises. Neuropharmacology 2017, 112, 237–247. [Google Scholar] [CrossRef]
- Pariante, C.M. Why are depressed patients inflamed? A reflection on 20 years of research on depression, glucocorticoid resistance and inflammation. Eur. Neuropsychopharmacol. 2017, 27, 554–559. [Google Scholar] [CrossRef]
- Dantzer, R. Neuroimmune Interactions: From the Brain to the Immune System and Vice Versa. Physiol. Rev. 2018, 98, 477–504. [Google Scholar] [CrossRef]
- Pandya, C.D.; Howell, K.R.; Pillai, A. Antioxidants as potential therapeutics for neuropsychiatric disorders. Prog. Neuropsychopharmacol. Biol. Psychiatry 2013, 46, 214–223. [Google Scholar] [CrossRef]
- Black, C.N.; Bot, M.; Scheffer, P.G.; Cuijpers, P.; Penninx, B.W. Is depression associated with increased oxidative stress? A systematic review and meta-analysis. Psychoneuroendocrinology 2015, 51, 164–175. [Google Scholar] [CrossRef]
- Ekdahl, C.T.; Claasen, J.H.; Bonde, S.; Kokaia, Z.; Lindvall, O. Inflammation is detrimental for neurogenesis in adult brain. Proc. Natl. Acad. Sci. USA 2003, 100, 13632–13637. [Google Scholar] [CrossRef]
- Kubera, M.; Obuchowicz, E.; Goehler, L.; Brzeszcz, J.; Maes, M. In animal models, psychosocial stress-induced (neuro)inflammation, apoptosis and reduced neurogenesis are associated to the onset of depression. Prog. Neuropsychopharmacol. Biol. Psychiatry 2011, 35, 744–759. [Google Scholar] [CrossRef]
- Wooff, Y.; Man, S.M.; Aggio-Bruce, R.; Natoli, R.; Fernando, N. IL-1 Family Members Mediate Cell Death, Inflammation and Angiogenesis in Retinal Degenerative Diseases. Front. Immunol. 2019, 10, 1618. [Google Scholar] [CrossRef]
- Bhatt, S.; Nagappa, A.N.; Patil, C.R. Role of oxidative stress in depression. Drug Discov. Today 2020, 25, 1270–1276. [Google Scholar] [CrossRef]
- Sabella, D. Antidepressant Medications. Am. J. Nurs. 2018, 118, 52–59. [Google Scholar] [CrossRef]
- Li, C.; Huang, J.; Cheng, Y.C.; Zhang, Y.W. Traditional Chinese Medicine in Depression Treatment: From Molecules to Systems. Front. Pharmacol. 2020, 11, 586. [Google Scholar] [CrossRef]
- Zhang, Y.; Long, Y.; Yu, S.; Li, D.; Yang, M.; Guan, Y.; Zhang, D.; Wan, J.; Liu, S.; Shi, A.; et al. Natural volatile oils derived from herbal medicines: A promising therapy way for treating depressive disorder. Pharmacol. Res. 2021, 164, 105376. [Google Scholar] [CrossRef]
- Sowndhararajan, K.; Deepa, P.; Kim, M.; Park, S.J.; Kim, S. Neuroprotective and Cognitive Enhancement Potentials of Baicalin: A Review. Brain Sci. 2018, 8, 104. [Google Scholar] [CrossRef]
- Huang, T.; Liu, Y.; Zhang, C. Pharmacokinetics and Bioavailability Enhancement of Baicalin: A Review. Eur. J. Drug Metab. Pharmacokinet. 2019, 44, 159–168. [Google Scholar] [CrossRef]
- Li, Y.; Song, K.; Zhang, H.; Yuan, M.; An, N.; Wei, Y.; Wang, L.; Sun, Y.; Xing, Y.; Gao, Y. Anti-inflammatory and immunomodulatory effects of baicalin in cerebrovascular and neurological disorders. Brain Res. Bull. 2020, 164, 314–324. [Google Scholar] [CrossRef]
- Guo, L.T.; Wang, S.Q.; Su, J.; Xu, L.X.; Ji, Z.Y.; Zhang, R.Y.; Zhao, Q.W.; Ma, Z.Q.; Deng, X.Y.; Ma, S.P. Baicalin ameliorates neuroinflammation-induced depressive-like behavior through inhibition of toll-like receptor 4 expression via the PI3K/AKT/FoxO1 pathway. J. Neuroinflamm. 2019, 16, 95. [Google Scholar] [CrossRef]
- Lu, Y.; Sun, G.; Yang, F.; Guan, Z.; Zhang, Z.; Zhao, J.; Liu, Y.; Chu, L.; Pei, L. Baicalin regulates depression behavior in mice exposed to chronic mild stress via the Rac/LIMK/cofilin pathway. Biomed. Pharmacother. 2019, 116, 109054. [Google Scholar] [CrossRef]
- Jia, Z.; Yang, J.; Cao, Z.; Zhao, J.; Zhang, J.; Lu, Y.; Chu, L.; Zhang, S.; Chen, Y.; Pei, L. Baicalin ameliorates chronic unpredictable mild stress-induced depression through the BDNF/ERK/CREB signaling pathway. Behav. Brain Res. 2021, 414, 113463. [Google Scholar] [CrossRef]
- Li, S.; Zhang, B. Traditional Chinese medicine network pharmacology: Theory, methodology and application. Chin. J. Nat. Med. 2013, 11, 110–120. [Google Scholar] [CrossRef]
- Casas, A.I.; Hassan, A.A.; Larsen, S.J.; Gomez-Rangel, V.; Elbatreek, M.; Kleikers, P.W.M.; Guney, E.; Egea, J.; Lopez, M.G.; Baumbach, J.; et al. From single drug targets to synergistic network pharmacology in ischemic stroke. Proc. Natl. Acad. Sci. USA 2019, 116, 7129–7136. [Google Scholar] [CrossRef]
- Li, J.; Zhao, P.; Tian, Y.; Li, K.; Zhang, L.; Guan, Q.; Mei, X.; Qin, Y. The Anti-Inflammatory Effect of a Combination of Five Compounds from Five Chinese Herbal Medicines Used in the Treatment of COPD. Front. Pharmacol. 2021, 12, 709702. [Google Scholar] [CrossRef]
- Berk, M.; Williams, L.J.; Jacka, F.N.; O’Neil, A.; Pasco, J.A.; Moylan, S.; Allen, N.B.; Stuart, A.L.; Hayley, A.C.; Byrne, M.L.; et al. So depression is an inflammatory disease, but where does the inflammation come from? BMC Med. 2013, 11, 200. [Google Scholar] [CrossRef]
- Cummings, C.M.; Caporino, N.E.; Kendall, P.C. Comorbidity of anxiety and depression in children and adolescents: 20 years after. Psychol. Bull. 2014, 140, 816–845. [Google Scholar] [CrossRef]
- Lenze, E.J. Comorbidity of depression and anxiety in the elderly. Curr. Psychiatry Rep. 2003, 5, 62–67. [Google Scholar] [CrossRef]
- Brunet, A.; Bonni, A.; Zigmond, M.J.; Lin, M.Z.; Juo, P.; Hu, L.S.; Anderson, M.J.; Arden, K.C.; Blenis, J.; Greenberg, M.E. Akt promotes cell survival by phosphorylating and inhibiting a Forkhead transcription factor. Cell 1999, 96, 857–868. [Google Scholar] [CrossRef]
- Mihara, M.; Hashizume, M.; Yoshida, H.; Suzuki, M.; Shiina, M. IL-6/IL-6 receptor system and its role in physiological and pathological conditions. Clin. Sci. 2012, 122, 143–159. [Google Scholar] [CrossRef]
- Rose-John, S. IL-6 trans-signaling via the soluble IL-6 receptor: Importance for the pro-inflammatory activities of IL-6. Int. J. Biol. Sci. 2012, 8, 1237–1247. [Google Scholar] [CrossRef]
- Campbell, I.L.; Abraham, C.R.; Masliah, E.; Kemper, P.; Inglis, J.D.; Oldstone, M.B.; Mucke, L. Neurologic disease induced in transgenic mice by cerebral overexpression of interleukin 6. Proc. Natl. Acad. Sci. USA 1993, 90, 10061–10065. [Google Scholar] [CrossRef]
- Roohi, E.; Jaafari, N.; Hashemian, F. On inflammatory hypothesis of depression: What is the role of IL-6 in the middle of the chaos? J. Neuroinflamm. 2021, 18, 45. [Google Scholar] [CrossRef]
- Levine, J.; Barak, Y.; Chengappa, K.N.; Rapoport, A.; Rebey, M.; Barak, V. Cerebrospinal cytokine levels in patients with acute depression. Neuropsychobiology 1999, 40, 171–176. [Google Scholar] [CrossRef]
- Tsai, S.J. Effects of interleukin-1beta polymorphisms on brain function and behavior in healthy and psychiatric disease conditions. Cytokine Growth Factor Rev. 2017, 37, 89–97. [Google Scholar] [CrossRef]
- Owen, B.M.; Eccleston, D.; Ferrier, I.N.; Young, A.H. Raised levels of plasma interleukin-1beta in major and postviral depression. Acta Psychiatr. Scand. 2001, 103, 226–228. [Google Scholar] [CrossRef]
- Mahmood, S.; Evinova, A.; Skerenova, M.; Ondrejka, I.; Lehotsky, J. Association of EGF, IGFBP-3 and TP53 Gene Polymorphisms with Major Depressive Disorder in Slovak Population. Cent. Eur. J. Public Health 2016, 24, 223–230. [Google Scholar] [CrossRef]
- Ranjan, A.; Iwakuma, T. Non-Canonical Cell Death Induced by p53. Int. J. Mol. Sci. 2016, 17, 2068. [Google Scholar] [CrossRef]
- Bakunina, N.; Pariante, C.M.; Zunszain, P.A. Immune mechanisms linked to depression via oxidative stress and neuroprogression. Immunology 2015, 144, 365–373. [Google Scholar] [CrossRef]
- Arantes-Goncalves, F.; Coelho, R. Depression and treatment. Apoptosis, neuroplasticity and antidepressants. Acta Med. Port. 2006, 19, 9–20. [Google Scholar]
- Spencer, S.L.; Sorger, P.K. Measuring and modeling apoptosis in single cells. Cell 2011, 144, 926–939. [Google Scholar] [CrossRef]
- Saveanu, R.V.; Nemeroff, C.B. Etiology of depression: Genetic and environmental factors. Psychiatr. Clin. N. Am. 2012, 35, 51–71. [Google Scholar] [CrossRef]
- Morris, G.M.; Lim-Wilby, M. Molecular docking. Methods Mol. Biol. 2008, 443, 365–382. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Langfelder, P.; Horvath, S. WGCNA: An R package for weighted correlation network analysis. BMC Bioinform. 2008, 9, 559. [Google Scholar] [CrossRef]
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 2001, 46, 3–26. [Google Scholar] [CrossRef]
- Veber, D.F.; Johnson, S.R.; Cheng, H.Y.; Smith, B.R.; Ward, K.W.; Kopple, K.D. Molecular properties that influence the oral bioavailability of drug candidates. J. Med. Chem. 2002, 45, 2615–2623. [Google Scholar] [CrossRef]
- Shen, M.; Tian, S.; Li, Y.; Li, Q.; Xu, X.; Wang, J.; Hou, T. Drug-likeness analysis of traditional Chinese medicines: 1. property distributions of drug-like compounds, non-drug-like compounds and natural compounds from traditional Chinese medicines. J. Cheminform. 2012, 4, 31. [Google Scholar] [CrossRef]
- Tao, W.; Xu, X.; Wang, X.; Li, B.; Wang, Y.; Li, Y.; Yang, L. Network pharmacology-based prediction of the active ingredients and potential targets of Chinese herbal Radix Curcumae formula for application to cardiovascular disease. J. Ethnopharmacol. 2013, 145, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Fang, S.; Dong, L.; Liu, L.; Guo, J.; Zhao, L.; Zhang, J.; Bu, D.; Liu, X.; Huo, P.; Cao, W.; et al. HERB: A high-throughput experiment- and reference-guided database of traditional Chinese medicine. Nucleic Acids Res. 2021, 49, D1197–D1206. [Google Scholar] [CrossRef] [PubMed]
- Cao, X.; Zao, X.; Xue, B.; Chen, H.; Zhang, J.; Li, S.; Li, X.; Zhu, S.; Guo, R.; Li, X.; et al. The mechanism of TiaoGanYiPi formula for treating chronic hepatitis B by network pharmacology and molecular docking verification. Sci. Rep. 2021, 11, 8402. [Google Scholar] [CrossRef]
- Dusa, A. venn: Draw Venn Diagrams, R Package Version 1.10. 2021. Available online: https://github.com/dusadrian/venn (accessed on 22 September 2024).
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Martin, A.; Ochagavia, M.E.; Rabasa, L.C.; Miranda, J.; Fernandez-de-Cossio, J.; Bringas, R. BisoGenet: A new tool for gene network building, visualization and analysis. BMC Bioinform. 2010, 11, 91. [Google Scholar] [CrossRef]
- Liu, X.; Hong, Z.; Liu, J.; Lin, Y.; Rodriguez-Paton, A.; Zou, Q.; Zeng, X. Computational methods for identifying the critical nodes in biological networks. Brief Bioinform. 2020, 21, 486–497. [Google Scholar] [CrossRef]
- Wu, T.; Hu, E.; Xu, S.; Chen, M.; Guo, P.; Dai, Z.; Feng, T.; Zhou, L.; Tang, W.; Zhan, L.; et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation 2021, 2, 100141. [Google Scholar] [CrossRef]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene ontology: Tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef] [PubMed]
- Gene Ontology Consortium. The Gene Ontology project in 2008. Nucleic Acids Res. 2008, 36, D440–D444. [Google Scholar] [CrossRef]
- Forli, S.; Huey, R.; Pique, M.E.; Sanner, M.F.; Goodsell, D.S.; Olson, A.J. Computational protein-ligand docking and virtual drug screening with the AutoDock suite. Nat. Protoc. 2016, 11, 905–919. [Google Scholar] [CrossRef]
- Huo, M.; Ma, L.; Liu, G. Exploring the mechanism of Yixinyin for myocardial infarction by weighted co-expression network and molecular docking. Sci. Rep. 2021, 11, 22567. [Google Scholar] [CrossRef]
- Laskowski, R.A.; Swindells, M.B. LigPlot+: Multiple ligand-protein interaction diagrams for drug discovery. J. Chem. Inf. Model. 2011, 51, 2778–2786. [Google Scholar] [CrossRef] [PubMed]
- Schrodinger, LLC. The AxPyMOL Molecular Graphics Plugin for Microsoft PowerPoint, Version 1.8; Schrodinger, LLC: New York, NY, USA, 2015. [Google Scholar]
- Schrodinger, LLC. The JyMOL Molecular Graphics Development Component, Version 1.8; Schrodinger, LLC: New York, NY, USA, 2015. [Google Scholar]
- Schrodinger, LLC. The PyMOL Molecular Graphics System, Version 1.8; Schrodinger, LLC: New York, NY, USA, 2015. [Google Scholar]
- Zhou, W.; Wu, J.; Zhang, J.; Liu, X.; Guo, S.; Jia, S.; Zhang, X.; Zhu, Y.; Wang, M. Integrated bioinformatics analysis to decipher molecular mechanism of compound Kushen injection for esophageal cancer by combining WGCNA with network pharmacology. Sci. Rep. 2020, 10, 12745. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Luo, J.; Liu, Z.; Liu, X.; Ma, Y.; Zhang, B.; Chen, Y.; Li, X.; Feng, Z.; Yang, N.; et al. Identification of hub genes in colorectal cancer based on weighted gene co-expression network analysis and clinical data from The Cancer Genome Atlas. Biosci. Rep. 2021, 41, BSR20211280. [Google Scholar] [CrossRef]
- Fu, Y.J.; Xu, B.; Huang, S.W.; Luo, X.; Deng, X.L.; Luo, S.; Liu, C.; Wang, Q.; Chen, J.Y.; Zhou, L. Baicalin prevents LPS-induced activation of TLR4/NF-κB p65 pathway and inflammation in mice via inhibiting the expression of CD14. Acta Pharmacol. Sin. 2021, 42, 88–96. [Google Scholar] [CrossRef] [PubMed]
- Ma, H.; Wang, W.; Xu, S.; Wang, L.; Wang, X. Potassium 2-(1-hydroxypentyl)-benzoate improves depressive-like behaviors in rat model. Acta Pharm. Sin. B 2018, 8, 881–888. [Google Scholar] [CrossRef]
- Li, Y.; Fan, C.; Wang, C.; Wang, L.; Yi, Y.; Mao, X.; Chen, X.; Lan, T.; Wang, W.; Yu, S.Y. Stress-induced reduction of Na(+)/H(+) exchanger isoform 1 promotes maladaptation of neuroplasticity and exacerbates depressive behaviors. Sci. Adv. 2022, 8, eadd7063. [Google Scholar] [CrossRef]
- Secoli, S.R.; Teixeira, N.A. Chronic prenatal stress affects development and behavioral depression in rats. Stress 1998, 2, 273–280. [Google Scholar] [CrossRef]
- Porsolt, R.D.; Le Pichon, M.; Jalfre, M. Depression: A new animal model sensitive to antidepressant treatments. Nature 1977, 266, 730–732. [Google Scholar] [CrossRef]
- Cryan, J.F.; Mombereau, C.; Vassout, A. The tail suspension test as a model for assessing antidepressant activity: Review of pharmacological and genetic studies in mice. Neurosci. Biobehav. Rev. 2005, 29, 571–625. [Google Scholar] [CrossRef]
- Zhang, M.; Liu, Y.; Zhao, M.; Tang, W.; Wang, X.; Dong, Z.; Yu, S. Depression and anxiety behaviour in a rat model of chronic migraine. J. Headache Pain 2017, 18, 27. [Google Scholar] [CrossRef]


| Parameter | Value | Reference |
|---|---|---|
| Hydrogen bond donor count | 6 | ≤5 |
| Hydrogen bond acceptor count | 11 | ≤10 |
| Molecular weight | 446.4 | <500 |
| Octanol/water partition coefficient | 1.1 | ≤5 |
| Rotatable bond count | 4 | ≤10 |
| Polar surface area (Å2) | 183 | ≤140 |
| Oral bioavailability (%) | 29.53 | ≥20 |
| Drug-likeness | 0.77 | ≥0.1 |
| Gene Symbol | ASPL | Betweenness Centrality | Closeness Centrality | Clustering Coefficient | Degree |
|---|---|---|---|---|---|
| RELA | 2.0625 | 0.36635005 | 0.48484848 | 0.24166667 | 16 |
| TP53 | 2.04166667 | 0.34916413 | 0.48979592 | 0.11428571 | 15 |
| MAPK1 | 2.20833333 | 0.21595323 | 0.45283019 | 0.16483516 | 14 |
| TNF | 2.60416667 | 0.09610351 | 0.384 | 0.26666667 | 10 |
| AKT1 | 2.5 | 0.24545128 | 0.4 | 0.08333333 | 9 |
| IL6 | 2.41666667 | 0.03947357 | 0.4137931 | 0.46428571 | 8 |
| IL1B | 2.77083333 | 0.02122383 | 0.36090226 | 0.47619048 | 7 |
| IL2 | 2.375 | 0.06283983 | 0.42105263 | 0.46666667 | 6 |
| TLR4 | 2.83333333 | 0.04262707 | 0.35294118 | 0.53333333 | 6 |
| NFKBIA | 2.39583333 | 0.02804162 | 0.4173913 | 0.46666667 | 6 |
| CCND1 | 2.52083333 | 0.01892942 | 0.39669421 | 0.3 | 5 |
| EGFR | 2.70833333 | 0.1214539 | 0.36923077 | 0.2 | 5 |
| NOTCH1 | 3.35416667 | 0.12012411 | 0.29813665 | 0.33333333 | 4 |
| CASP3 | 2.64583333 | 0.00757979 | 0.37795276 | 0.66666667 | 4 |
| NOS2 | 2.60416667 | 0.03900709 | 0.384 | 0.5 | 4 |
| BCL2 | 2.64583333 | 0.0114805 | 0.37795276 | 0.5 | 4 |
| PTGS2 | 2.6875 | 0.04794622 | 0.37209302 | 0.16666667 | 4 |
| ATF2 | 2.60416667 | 0.00143322 | 0.384 | 0.66666667 | 4 |
| MYD88 | 2.91666667 | 0.00088652 | 0.34285714 | 0.66666667 | 4 |
| TGFB1 | 2.72916667 | 0.00150709 | 0.36641221 | 0.66666667 | 4 |
| Gene Symbol | ASPL | Betweenness Centrality | Closeness Centrality | Clustering Coefficient | Degree |
|---|---|---|---|---|---|
| TP53 | 2.64510914 | 0.09478941 | 0.37805623 | 0.06731985 | 113 |
| EP300 | 2.6523848 | 0.05607056 | 0.3770192 | 0.10060606 | 100 |
| CTNNB1 | 2.64349232 | 0.06465836 | 0.37828746 | 0.08157895 | 96 |
| PIK3R1 | 2.74777688 | 0.01763209 | 0.36393057 | 0.16117216 | 91 |
| PIK3CA | 2.72190784 | 0.02173176 | 0.36738937 | 0.16434676 | 91 |
| HRAS | 2.74454325 | 0.02687518 | 0.36435935 | 0.16317671 | 89 |
| MAPK1 | 2.75101051 | 0.01976236 | 0.36350279 | 0.14883721 | 86 |
| MAPK3 | 2.75505255 | 0.01753599 | 0.36296948 | 0.15376197 | 86 |
| STAT3 | 2.77041229 | 0.02476242 | 0.36095711 | 0.15253165 | 80 |
| RPS27A | 2.84236055 | 0.06752715 | 0.35182025 | 0.0691334 | 79 |
| AKT1 | 2.66370251 | 0.04034818 | 0.3754173 | 0.12720613 | 78 |
| PTPN11 | 2.87146322 | 0.01752892 | 0.3482545 | 0.19824561 | 76 |
| CREBBP | 2.71544058 | 0.03602726 | 0.36826436 | 0.12699 | 74 |
| KRAS | 2.80679062 | 0.01429623 | 0.3562788 | 0.20372671 | 70 |
| RAC1 | 2.89814066 | 0.02085928 | 0.34504881 | 0.15364355 | 68 |
| SOS1 | 2.91835085 | 0.00537091 | 0.34265928 | 0.26686508 | 64 |
| NRAS | 2.94017785 | 0.00885878 | 0.34011548 | 0.2202381 | 64 |
| MAPK8 | 2.82538399 | 0.02066826 | 0.35393419 | 0.12544803 | 63 |
| ESR1 | 2.74454325 | 0.03216523 | 0.36435935 | 0.15901639 | 61 |
| JAK2 | 2.95149555 | 0.01048452 | 0.33881128 | 0.1954023 | 58 |
| Gene Symbol | ASPL | Betweenness Centrality | Closeness Centrality | Clustering Coefficient | Degree |
|---|---|---|---|---|---|
| AKT1 | 1.30769231 | 0.18065866 | 0.76470588 | 0.45238095 | 28 |
| IL6 | 1.53846154 | 0.05851599 | 0.65 | 0.55797101 | 24 |
| IL1B | 1.48717949 | 0.07172935 | 0.67241379 | 0.56126482 | 23 |
| TP53 | 1.58974359 | 0.03567395 | 0.62903226 | 0.60474308 | 23 |
| CASP3 | 1.53846154 | 0.03535266 | 0.65 | 0.64069264 | 22 |
| MAPK1 | 1.56410256 | 0.06271286 | 0.63934426 | 0.58421053 | 20 |
| PPARG | 1.66666667 | 0.02191758 | 0.6 | 0.68947368 | 20 |
| PTGS2 | 1.69230769 | 0.00928826 | 0.59090909 | 0.76608187 | 19 |
| CCND1 | 1.71794872 | 0.01594669 | 0.58208955 | 0.67251462 | 19 |
| EGF | 1.71794872 | 0.01524371 | 0.58208955 | 0.70175439 | 19 |
| TGFB1 | 1.71794872 | 0.01121916 | 0.58208955 | 0.75163399 | 18 |
| IL2 | 1.71794872 | 0.01244353 | 0.58208955 | 0.76470588 | 18 |
| CASP9 | 1.74358974 | 0.00703711 | 0.57352941 | 0.79411765 | 17 |
| TLR4 | 1.74358974 | 0.01616254 | 0.57352941 | 0.72058824 | 17 |
| GPT | 1.76923077 | 0.0714995 | 0.56521739 | 0.675 | 16 |
| HMOX1 | 1.79487179 | 0.00105304 | 0.55714286 | 0.93333333 | 15 |
| NOS2 | 1.69230769 | 0.0320215 | 0.59090909 | 0.8021978 | 14 |
| CDKN2A | 1.84615385 | 0.00177688 | 0.54166667 | 0.87912088 | 14 |
| GRIN1 | 1.79487179 | 0.17178252 | 0.55714286 | 0.25454545 | 11 |
| HNF4A | 1.92307692 | 0.01964939 | 0.52 | 0.65454545 | 11 |
| Target Gene | Specific Primer Sequences |
|---|---|
| IL-1β | forward primer: 5′-AAGATGAAGGGCTGCTTCCAAACC-3′ reverse primer: 5′-ATACTGCCTGCCTGAAGCTCTTGT-3′ |
| IL-6 | forward primer: 5′-TGATCGGTCCCAACAAGGA-3′ reverse primer: 5′-CCACTTGTTGGCTTATGTT-3′ |
| Caspase 3 | forward primer: 5′-GGAGCTTGGAACGCGAAGAA-3′ reverse primer: 5′-ACACAAGCCCATTTCAGGGT-3′ |
| AKT | forward primer: 5′-GTTTTGTTTCTCGGATGCGCT-3′ reverse primer: 5′-CATGGTCGCGTCAGTCCTTA-3′ |
| P53 | forward primer: 5′-GGCGTAAACGCTTCGAGATG-3′ reverse primer: 5′-CTTCAGGTAGCTGGAGTGAGC-3′ |
| GAPDH | forward primer: 5′-TCTCTGCTCCTCCCTGTTC-3′ reverse primer: 5′-ACACCGACCTTCACCATCT-3′ |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yi, Y.; Liu, G.; Li, Y.; Wang, C.; Zhang, B.; Lou, H.; Yu, S. Baicalin Ameliorates Depression-like Behaviors via Inhibiting Neuroinflammation and Apoptosis in Mice. Int. J. Mol. Sci. 2024, 25, 10259. https://doi.org/10.3390/ijms251910259
Yi Y, Liu G, Li Y, Wang C, Zhang B, Lou H, Yu S. Baicalin Ameliorates Depression-like Behaviors via Inhibiting Neuroinflammation and Apoptosis in Mice. International Journal of Molecular Sciences. 2024; 25(19):10259. https://doi.org/10.3390/ijms251910259
Chicago/Turabian StyleYi, Yuhang, Guiyu Liu, Ye Li, Changmin Wang, Bin Zhang, Haiyan Lou, and Shuyan Yu. 2024. "Baicalin Ameliorates Depression-like Behaviors via Inhibiting Neuroinflammation and Apoptosis in Mice" International Journal of Molecular Sciences 25, no. 19: 10259. https://doi.org/10.3390/ijms251910259






