Profiling Reduced Expression of Contractile and Mitochondrial mRNAs in the Human Sinoatrial Node vs. Right Atrium and Predicting Their Suppressed Expression by Transcription Factors and/or microRNAs
Abstract
1. Introduction
2. Results
2.1. Differential mRNA Expression in the Human Sinus Node and Atrial Muscle
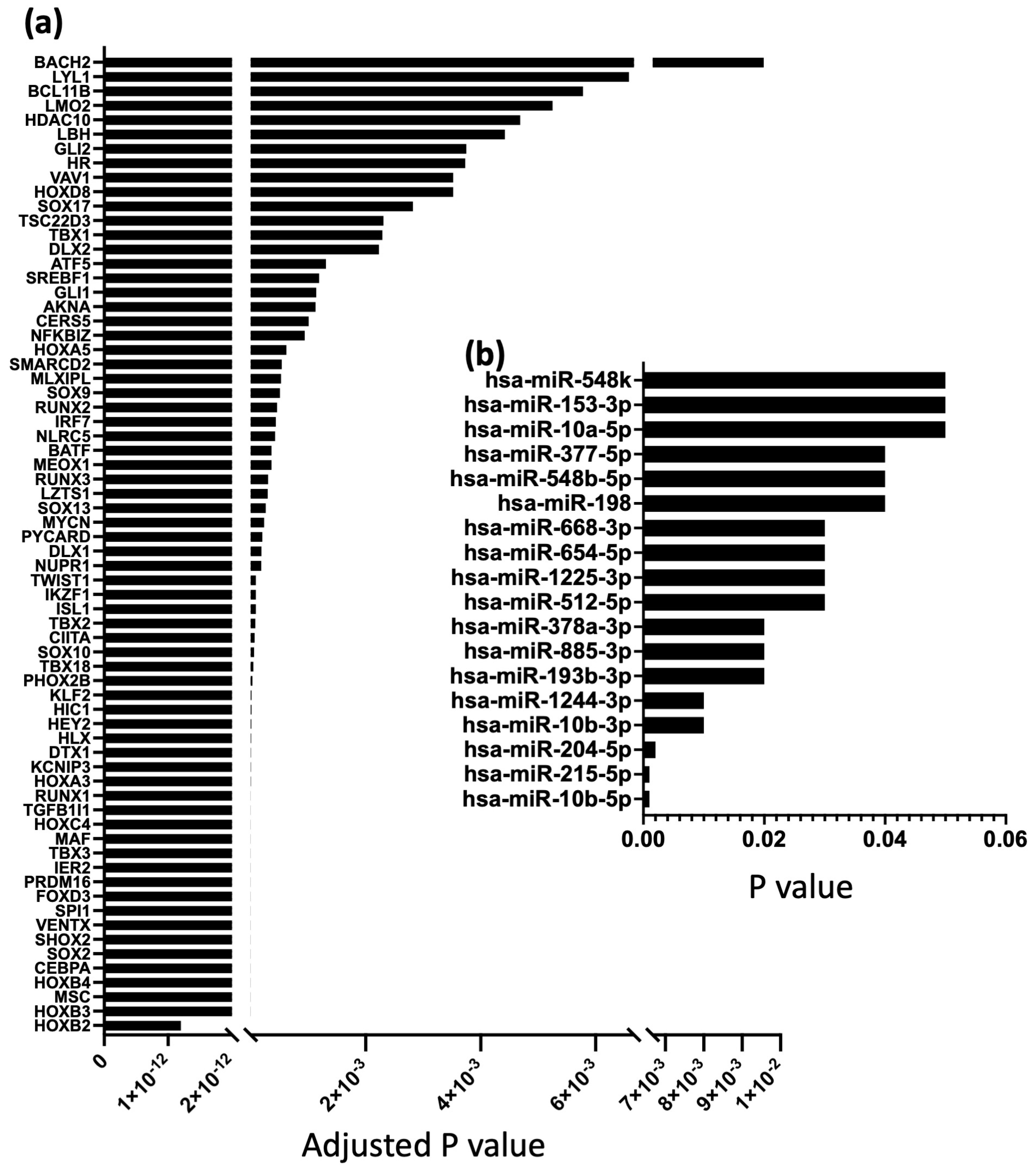
2.2. Differential TFs and miRs Expression in the Human SN and Atrial Muscle
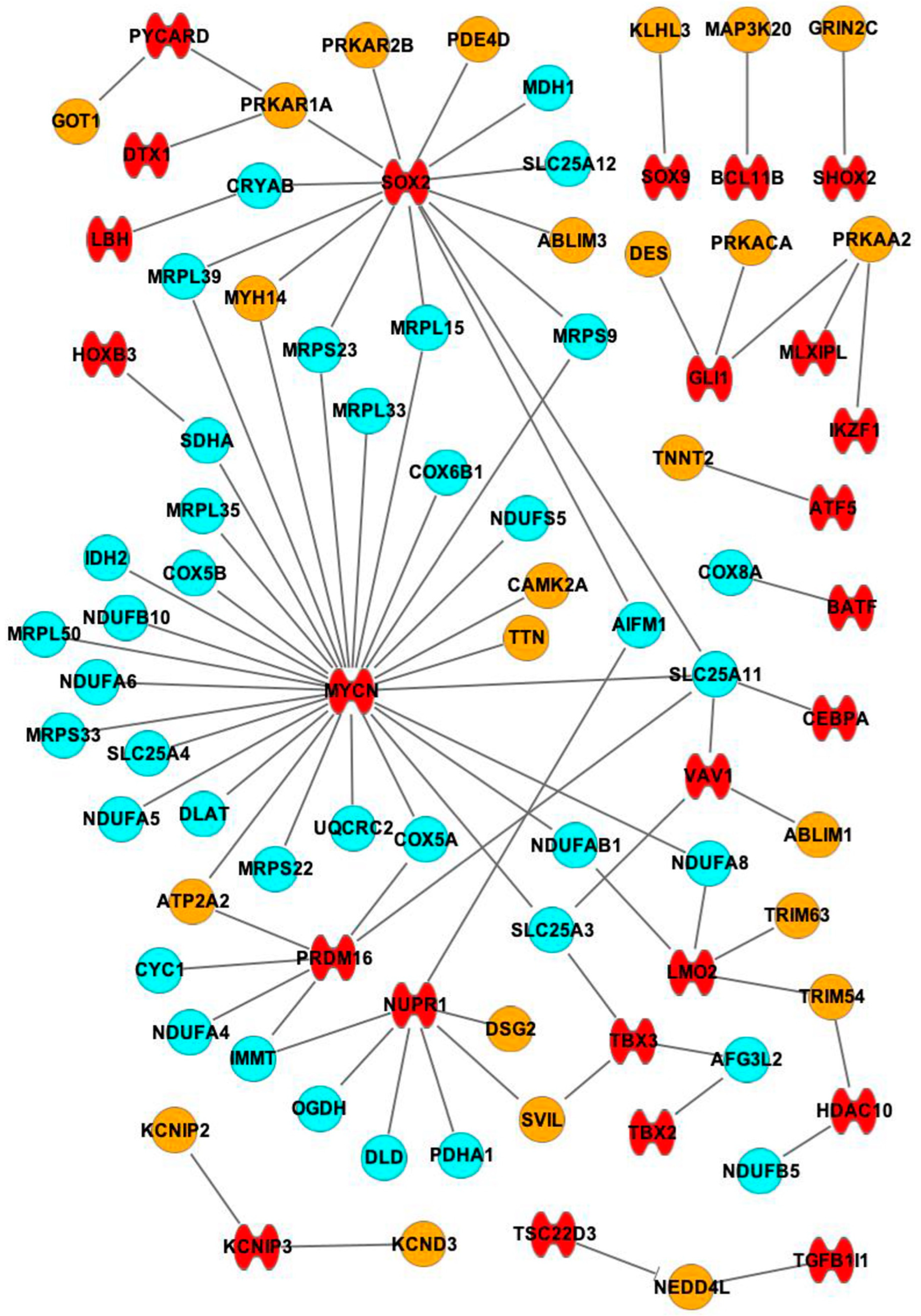
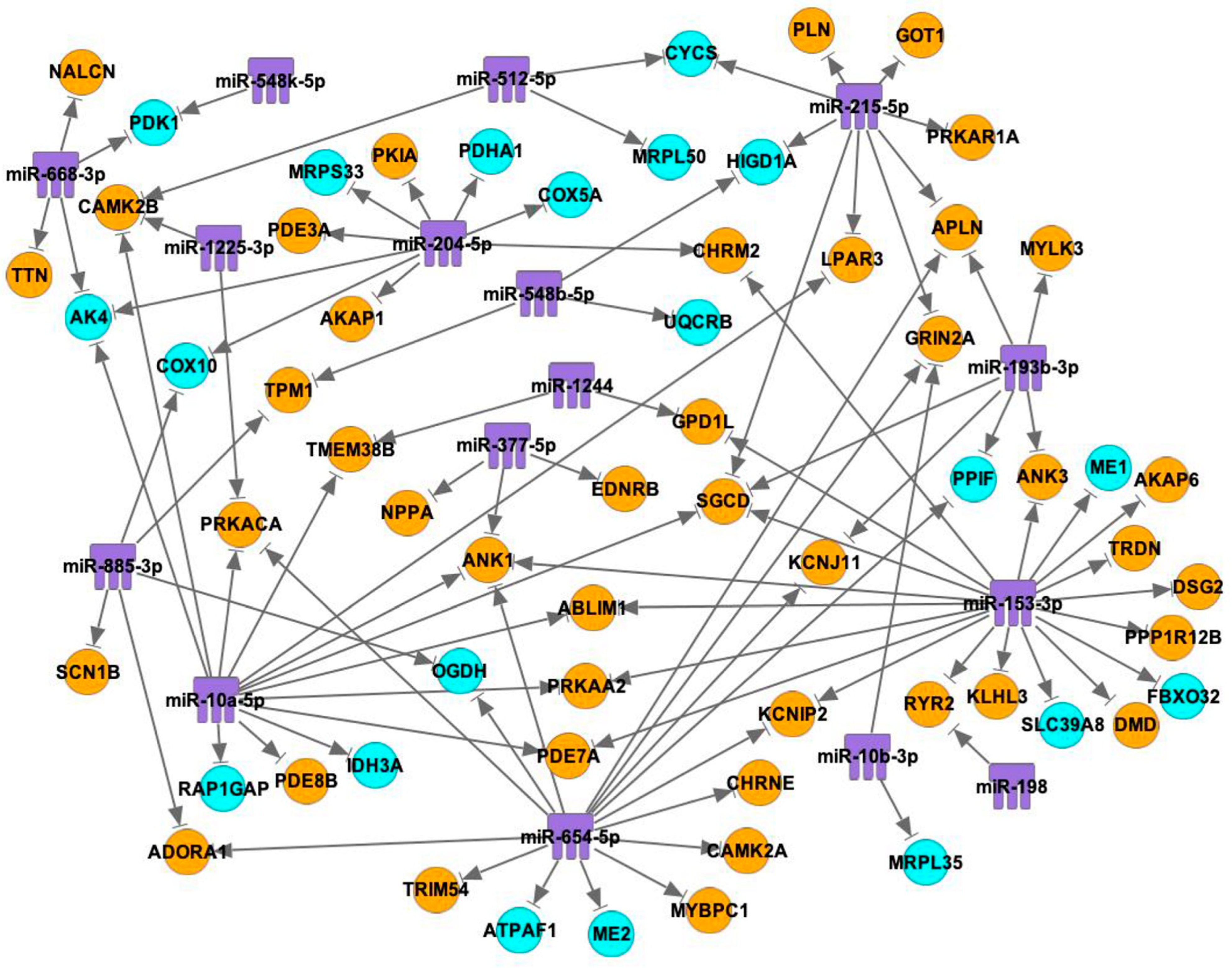
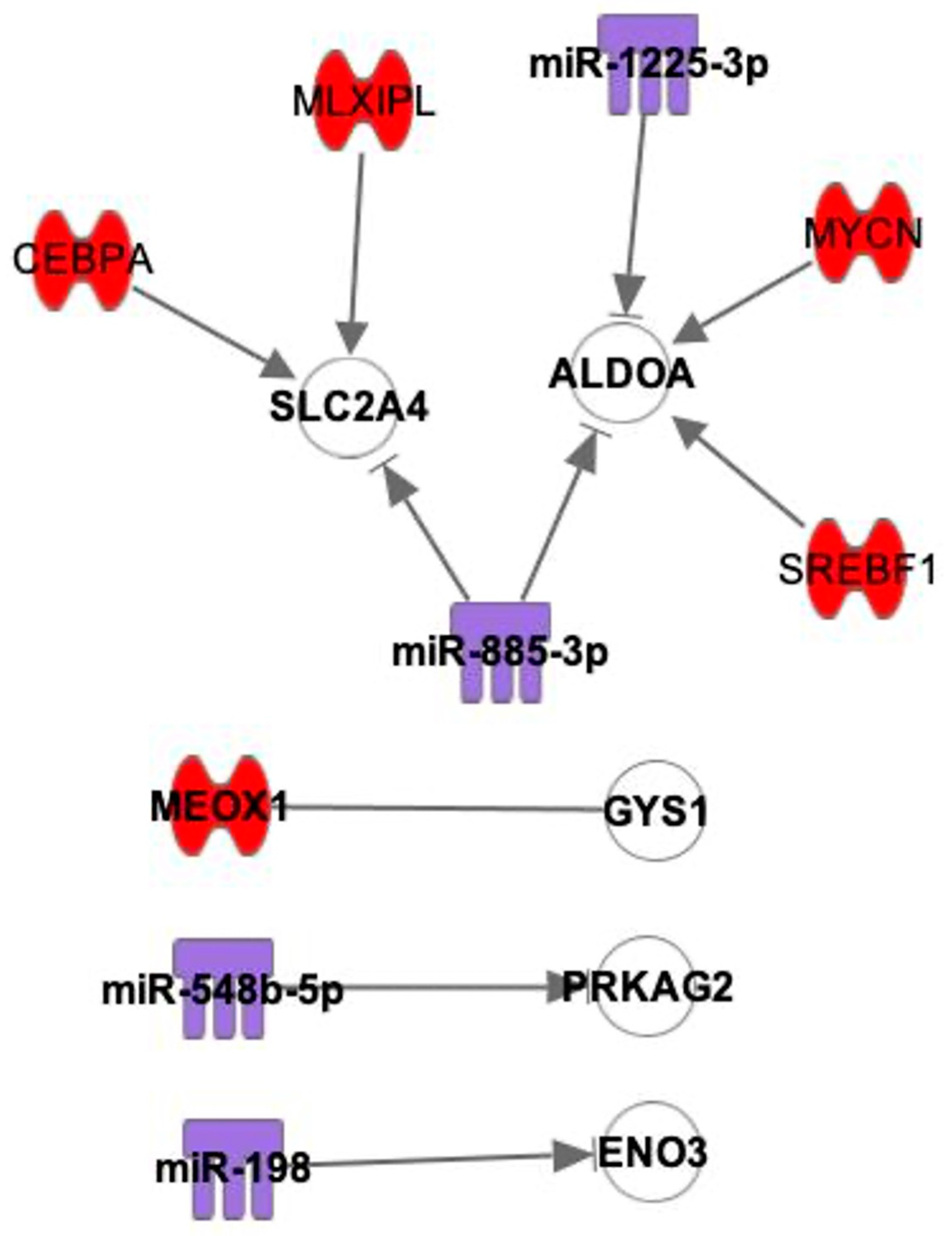
2.3. Interactions between mRNAs and miRNAs, TFs
3. Discussion
4. Materials and Methods
4.1. Next-Generation Sequencing for mRNA, Bioinformatic Analysis for TFs and qPCR for miRs
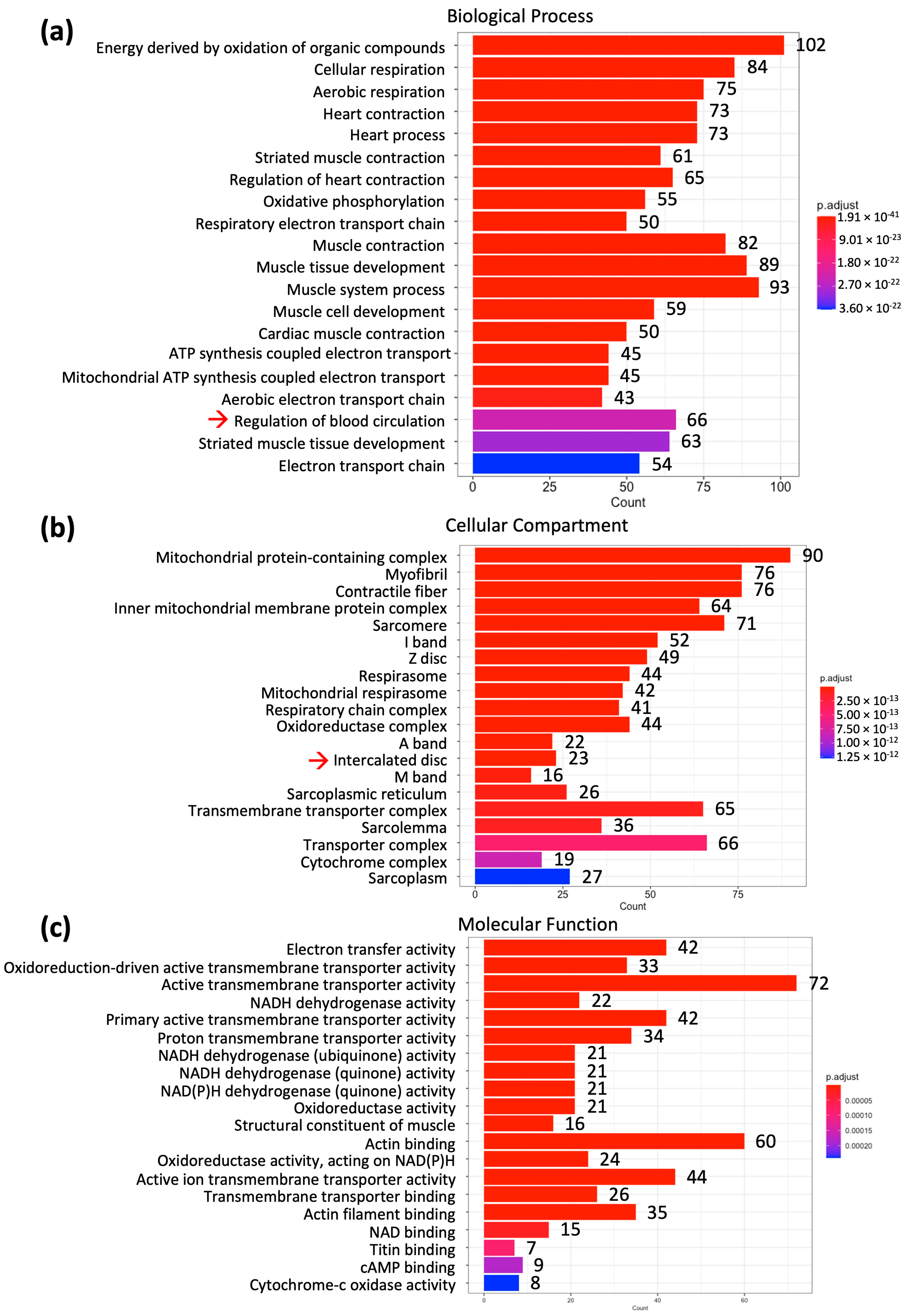
4.2. Bioinformatic Analysis to Identify the mRNAs That Relate to Contractile Function, Mitochondrial Function and Glycogen Metabolism
4.2.1. RStudio Analysis
4.2.2. Canonical Pathway Analysis in IPA
4.2.3. Contractile Function, Mitochondrial Function, and Glycogen Metabolism Identification
4.3. Creating and Analysing mRNA, TF and miR Interactions/Networks in IPA
4.4. Binding Site Prediction between mRNAs and miRs
4.5. Heat Maps, Graphs, and Statistical Analysis
5. Limitations
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Stephenson, R.S.; Atkinson, A.; Kottas, P.; Perde, F.; Jafarzadeh, F.; Bateman, M.; Iaizzo, P.A.; Zhao, J.; Zhang, H.; Anderson, R.H.; et al. High resolution 3-Dimensional imaging of the human cardiac conduction system from microanatomy to mathematical modeling. Sci. Rep. 2017, 7, 7188. [Google Scholar] [CrossRef] [PubMed]
- Dobrzynski, H.; Anderson, R.H.; Atkinson, A.; Borbas, Z.; D’Souza, A.; Fraser, J.F.; Inada, S.; Logantha, S.J.; Monfredi, O.; Morris, G.M.; et al. Structure, function and clinical relevance of the cardiac conduction system, including the atrioventricular ring and outflow tract tissues. Pharmacol. Ther. 2013, 139, 260–288. [Google Scholar] [CrossRef]
- Chandler, N.J.; Greener, I.D.; Tellez, J.O.; Inada, S.; Musa, H.; Molenaar, P.; Difrancesco, D.; Baruscotti, M.; Longhi, R.; Anderson, R.H.; et al. Molecular architecture of the human sinus node: Insights into the function of the cardiac pacemaker. Circulation 2009, 119, 1562–1575. [Google Scholar] [CrossRef]
- Boyett, M.R.; Honjo, H.; Kodama, I. The sinoatrial node, a heterogeneous pacemaker structure. Cardiovasc. Res. 2000, 47, 658–687. [Google Scholar] [CrossRef] [PubMed]
- James, T.N.; Sherf, L.; Fine, G.; Morales, A.R. Comparative ultrastructure of the sinus node in man and dog. Circulation 1966, 34, 139–163. [Google Scholar] [CrossRef]
- Calvet-Marquez, S.; Domenech-Mateu, J.M. Presence or absence of glycogen-beta particles in sinoatrial node cells. Acta Anat. 1994, 150, 267–273. [Google Scholar] [CrossRef] [PubMed]
- Gu, J.M.; Grijalva, S.I.; Fernandez, N.; Kim, E.; Foster, D.B.; Cho, H.C. Induced cardiac pacemaker cells survive metabolic stress owing to their low metabolic demand. Exp. Mol. Med. 2019, 51, 1–12. [Google Scholar] [CrossRef]
- Akazawa, H.; Komuro, I. Roles of cardiac transcription factors in cardiac hypertrophy. Circ. Res. 2003, 92, 1079–1088. [Google Scholar] [CrossRef]
- Mommersteeg, M.T.; Hoogaars, W.M.; Prall, O.W.; de Gier-de Vries, C.; Wiese, C.; Clout, D.E.; Papaioannou, V.E.; Brown, N.A.; Harvey, R.P.; Moorman, A.F.; et al. Molecular pathway for the localized formation of the sinoatrial node. Circ. Res. 2007, 100, 354–362. [Google Scholar] [CrossRef]
- Christoffels, V.M.; Smits, G.J.; Kispert, A.; Moorman, A.F. Development of the pacemaker tissues of the heart. Circ. Res. 2010, 106, 240–254. [Google Scholar] [CrossRef]
- Blaschke, R.J.; Hahurij, N.D.; Kuijper, S.; Just, S.; Wisse, L.J.; Deissler, K.; Maxelon, T.; Anastassiadis, K.; Spitzer, J.; Hardt, S.E.; et al. Targeted mutation reveals essential functions of the homeodomain transcription factor Shox2 in sinoatrial and pacemaking development. Circulation 2007, 115, 1830–1838. [Google Scholar] [CrossRef] [PubMed]
- Aminu, A.J.; Petkova, M.; Atkinson, A.J.; Yanni, J.; Morris, A.D.; Simms, R.T.; Chen, W.; Yin, Z.; Kuniewicz, M.; Holda, M.K.; et al. Further insights into the molecular complexity of the human sinus node—The role of ‘novel’ transcription factors and microRNAs. Prog. Biophys. Mol. Biol. 2021, 166, 86–104. [Google Scholar] [CrossRef] [PubMed]
- Ambros, V. The functions of animal microRNAs. Nature 2004, 431, 350–355. [Google Scholar] [CrossRef]
- Lewis, B.P.; Shih, I.H.; Jones-Rhoades, M.W.; Bartel, D.P.; Burge, C.B. Prediction of mammalian microRNA targets. Cell 2003, 115, 787–798. [Google Scholar] [CrossRef] [PubMed]
- Grimson, A.; Farh, K.K.; Johnston, W.K.; Garrett-Engele, P.; Lim, L.P.; Bartel, D.P. MicroRNA targeting specificity in mammals: Determinants beyond seed pairing. Mol. Cell 2007, 27, 91–105. [Google Scholar] [CrossRef] [PubMed]
- Asjad, E.; Dobrzynski, H. MicroRNAs: Midfielders of Cardiac Health, Disease and Treatment. Int. J. Mol. Sci. 2023, 24, 16207. [Google Scholar] [CrossRef]
- Petkova, M.; Atkinson, A.J.; Yanni, J.; Stuart, L.; Aminu, A.J.; Ivanova, A.D.; Pustovit, K.B.; Geragthy, C.; Feather, A.; Li, N.; et al. Identification of Key Small Non-Coding MicroRNAs Controlling Pacemaker Mechanisms in the Human Sinus Node. J. Am. Heart Assoc. 2020, 9, e016590. [Google Scholar] [CrossRef]
- Bers, D.M. Cardiac excitation-contraction coupling. Nature 2002, 415, 198–205. [Google Scholar] [CrossRef]
- Lei, Z.; Wahlquist, C.; El Azzouzi, H.; Deddens, J.C.; Kuster, D.; van Mil, A.; Rojas-Munoz, A.; Huibers, M.M.; Mercola, M.; de Weger, R.; et al. miR-132/212 Impairs Cardiomyocytes Contractility in the Failing Heart by Suppressing SERCA2a. Front. Cardiovasc. Med. 2021, 8, 592362. [Google Scholar] [CrossRef]
- van Rooij, E.; Sutherland, L.B.; Qi, X.; Richardson, J.A.; Hill, J.; Olson, E.N. Control of stress-dependent cardiac growth and gene expression by a microRNA. Science 2007, 316, 575–579. [Google Scholar] [CrossRef] [PubMed]
- Wahlquist, C.; Jeong, D.; Rojas-Munoz, A.; Kho, C.; Lee, A.; Mitsuyama, S.; van Mil, A.; Park, W.J.; Sluijter, J.P.; Doevendans, P.A.; et al. Inhibition of miR-25 improves cardiac contractility in the failing heart. Nature 2014, 508, 531–535. [Google Scholar] [CrossRef]
- Paavola, J.; Alakoski, T.; Ulvila, J.; Kilpio, T.; Siren, J.; Perttunen, S.; Narumanchi, S.; Wang, H.; Lin, R.; Porvari, K.; et al. Vezf1 regulates cardiac structure and contractile function. EBioMedicine 2020, 51, 102608. [Google Scholar] [CrossRef] [PubMed]
- Dasgupta, N.; Peng, Y.; Tan, Z.; Ciraolo, G.; Wang, D.; Li, R. miRNAs in mtDNA-less cell mitochondria. Cell Death Discov. 2015, 1, 15004. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.Q.; Wang, S.Q.; Chen, Y.; Fu, L.Y.; Xu, Y.N.; Li, L.; Tao, L.; Shen, X.C. MicroRNAs Regulating Mitochondrial Function in Cardiac Diseases. Front. Pharmacol. 2021, 12, 663322. [Google Scholar] [CrossRef]
- Kumar, A.A.; Kelly, D.P.; Chirinos, J.A. Mitochondrial Dysfunction in Heart Failure With Preserved Ejection Fraction. Circulation 2019, 139, 1435–1450. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.; Hu, J.; Zhang, X.; Li, C.; Zuo, Y.; Xie, S.; Zhang, Z.; Zhu, S. Neuropeptide Y Induces Cardiomyocyte Hypertrophy via Attenuating miR-29a-3p in Neonatal Rat Cardiomyocytes. Protein Pept. Lett. 2020, 27, 878–887. [Google Scholar] [CrossRef] [PubMed]
- Hathaway, Q.A.; Pinti, M.V.; Durr, A.J.; Waris, S.; Shepherd, D.L.; Hollander, J.M. Regulating microRNA expression: At the heart of diabetes mellitus and the mitochondrion. Am. J. Physiol. Heart Circ. Physiol. 2018, 314, H293–H310. [Google Scholar] [CrossRef] [PubMed]
- Bandiera, S.; Mategot, R.; Girard, M.; Demongeot, J.; Henrion-Caude, A. MitomiRs delineating the intracellular localization of microRNAs at mitochondria. Free Radic. Biol. Med. 2013, 64, 12–19. [Google Scholar] [CrossRef]
- Pinti, M.V.; Hathaway, Q.A.; Hollander, J.M. Role of microRNA in metabolic shift during heart failure. Am. J. Physiol. Heart Circ. Physiol. 2017, 312, H33–H45. [Google Scholar] [CrossRef]
- Shepherd, D.L.; Hathaway, Q.A.; Pinti, M.V.; Nichols, C.E.; Durr, A.J.; Sreekumar, S.; Hughes, K.M.; Stine, S.M.; Martinez, I.; Hollander, J.M. Exploring the mitochondrial microRNA import pathway through Polynucleotide Phosphorylase (PNPase). J. Mol. Cell. Cardiol. 2017, 110, 15–25. [Google Scholar] [CrossRef]
- Das, S.; Bedja, D.; Campbell, N.; Dunkerly, B.; Chenna, V.; Maitra, A.; Steenbergen, C. miR-181c regulates the mitochondrial genome, bioenergetics, and propensity for heart failure in vivo. PLoS ONE 2014, 9, e96820. [Google Scholar] [CrossRef] [PubMed]
- Kang, D.; Kim, S.H.; Hamasaki, N. Mitochondrial transcription factor A (TFAM): Roles in maintenance of mtDNA and cellular functions. Mitochondrion 2007, 7, 39–44. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.; Aryal, S.; Geng, X.; Zhou, X.; Fast, V.G.; Zhang, J.; Lu, R.; Zhou, Y. TBX20 Improves Contractility and Mitochondrial Function During Direct Human Cardiac Reprogramming. Circulation 2022, 146, 1518–1536. [Google Scholar] [CrossRef]
- Lin, Y.K.; Cheng, C.C.; Tsai, M.C.; Wu, P.Y.; Chen, Y.A.; Chen, Y.C.; Chen, S.A.; Chen, Y.J. Mitochondrial dysfunction on sinoatrial node and pulmonary vein electrophysiological activities. Exp. Ther. Med. 2017, 13, 2486–2492. [Google Scholar] [CrossRef]
- Schotten, U.; Ausma, J.; Stellbrink, C.; Sabatschus, I.; Vogel, M.; Frechen, D.; Schoendube, F.; Hanrath, P.; Allessie, M.A. Cellular mechanisms of depressed atrial contractility in patients with chronic atrial fibrillation. Circulation 2001, 103, 691–698. [Google Scholar] [CrossRef]
- Ren, L.; Gopireddy, R.R.; Perkins, G.; Zhang, H.; Timofeyev, V.; Lyu, Y.; Diloretto, D.A.; Trinh, P.; Sirish, P.; Overton, J.L.; et al. Disruption of mitochondria-sarcoplasmic reticulum microdomain connectomics contributes to sinus node dysfunction in heart failure. Proc. Natl. Acad. Sci. USA 2022, 119, e2206708119. [Google Scholar] [CrossRef] [PubMed]
- Steinberg, C.; Roston, T.M.; van der Werf, C.; Sanatani, S.; Chen, S.R.W.; Wilde, A.A.M.; Krahn, A.D. RYR2-ryanodinopathies: From calcium overload to calcium deficiency. Europace 2023, 25, euad156. [Google Scholar] [CrossRef]
- Faggioni, M.; van der Werf, C.; Knollmann, B.C. Sinus node dysfunction in catecholaminergic polymorphic ventricular tachycardia: Risk factor and potential therapeutic target? Trends Cardiovasc. Med. 2014, 24, 273–278. [Google Scholar] [CrossRef]
- Chandler, N.; Aslanidi, O.; Buckley, D.; Inada, S.; Birchall, S.; Atkinson, A.; Kirk, D.; Monfredi, O.; Molenaar, P.; Anderson, R.; et al. Computer three-dimensional anatomical reconstruction of the human sinus node and a novel paranodal area. Anat. Rec. 2011, 294, 970–979. [Google Scholar] [CrossRef]
- Shen, Y.; Chen, M.; Gu, W.; Wan, J.; Cheng, Z.; Shen, K.; Zhang, W.; He, J.; Wang, Y.; Deng, X. The molecular mechanism of cardiac injury in SARS-CoV-2 infection: Focus on mitochondrial dysfunction. J. Infect. Public Health 2023, 16, 746–753. [Google Scholar] [CrossRef]
- Harmelink, C.; Peng, Y.; DeBenedittis, P.; Chen, H.; Shou, W.; Jiao, K. Myocardial Mycn is essential for mouse ventricular wall morphogenesis. Dev. Biol. 2013, 373, 53–63. [Google Scholar] [CrossRef] [PubMed]
- Rao, J.; Pfeiffer, M.J.; Frank, S.; Adachi, K.; Piccini, I.; Quaranta, R.; Arauzo-Bravo, M.; Schwarz, J.; Schade, D.; Leidel, S.; et al. Stepwise Clearance of Repressive Roadblocks Drives Cardiac Induction in Human ESCs. Cell Stem Cell 2016, 18, 341–353. [Google Scholar] [CrossRef] [PubMed]
- Gironella, M.; Malicet, C.; Cano, C.; Sandi, M.J.; Hamidi, T.; Tauil, R.M.; Baston, M.; Valaco, P.; Moreno, S.; Lopez, F.; et al. p8/nupr1 regulates DNA-repair activity after double-strand gamma irradiation-induced DNA damage. J. Cell. Physiol. 2009, 221, 594–602. [Google Scholar] [CrossRef] [PubMed]
- Martin, T.A.; Li, A.X.; Sanders, A.J.; Ye, L.; Frewer, K.; Hargest, R.; Jiang, W.G. NUPR1 and its potential role in cancer and pathological conditions (Review). Int. J. Oncol. 2021, 58, 21. [Google Scholar] [CrossRef]
- Wu, T.; Liang, Z.; Zhang, Z.; Liu, C.; Zhang, L.; Gu, Y.; Peterson, K.L.; Evans, S.M.; Fu, X.D.; Chen, J. PRDM16 Is a Compact Myocardium-Enriched Transcription Factor Required to Maintain Compact Myocardial Cardiomyocyte Identity in Left Ventricle. Circulation 2022, 145, 586–602. [Google Scholar] [CrossRef]
- Arndt, A.K.; Schafer, S.; Drenckhahn, J.D.; Sabeh, M.K.; Plovie, E.R.; Caliebe, A.; Klopocki, E.; Musso, G.; Werdich, A.A.; Kalwa, H.; et al. Fine mapping of the 1p36 deletion syndrome identifies mutation of PRDM16 as a cause of cardiomyopathy. Am. J. Hum. Genet. 2013, 93, 67–77. [Google Scholar] [CrossRef]
- Long, P.A.; Evans, J.M.; Olson, T.M. Diagnostic Yield of Whole Exome Sequencing in Pediatric Dilated Cardiomyopathy. J. Cardiovasc. Dev. Dis. 2017, 4, 11. [Google Scholar] [CrossRef]
- Wang, T.; Zhai, M.; Xu, S.; Ponnusamy, M.; Huang, Y.; Liu, C.Y.; Wang, M.; Shan, C.; Shan, P.P.; Gao, X.Q.; et al. NFATc3-dependent expression of miR-153-3p promotes mitochondrial fragmentation in cardiac hypertrophy by impairing mitofusin-1 expression. Theranostics 2020, 10, 553–566. [Google Scholar] [CrossRef]
- Fang, Y.; Shi, C.; Manduchi, E.; Civelek, M.; Davies, P.F. MicroRNA-10a regulation of proinflammatory phenotype in athero-susceptible endothelium in vivo and in vitro. Proc. Natl. Acad. Sci. USA 2010, 107, 13450–13455. [Google Scholar] [CrossRef] [PubMed]
- Wei, Y.; Corbalan-Campos, J.; Gurung, R.; Natarelli, L.; Zhu, M.; Exner, N.; Erhard, F.; Greulich, F.; Geissler, C.; Uhlenhaut, N.H.; et al. Dicer in Macrophages Prevents Atherosclerosis by Promoting Mitochondrial Oxidative Metabolism. Circulation 2018, 138, 2007–2020. [Google Scholar] [CrossRef]
- Latimer, M.N.; Williams, L.J.; Shanmugan, G.; Carpenter, B.J.; Lazar, M.A.; Dierickx, P.; Young, M.E. Cardiomyocyte-specific disruption of the circadian BMAL1-REV-ERBalpha/beta regulatory network impacts distinct miRNA species in the murine heart. Commun. Biol. 2023, 6, 1149. [Google Scholar] [CrossRef] [PubMed]
- Singh, D.D.; Kim, Y.; Choi, S.A.; Han, I.; Yadav, D.K. Clinical Significance of MicroRNAs, Long Non-Coding RNAs, and CircRNAs in Cardiovascular Diseases. Cells 2023, 12, 1629. [Google Scholar] [CrossRef] [PubMed]
- Anderson, C.M.; Hu, J.; Thomas, R.; Gainous, T.B.; Celona, B.; Sinha, T.; Dickel, D.E.; Heidt, A.B.; Xu, S.M.; Bruneau, B.G.; et al. Cooperative activation of cardiac transcription through myocardin bridging of paired MEF2 sites. Development 2017, 144, 1235–1241. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Wang, W.E.; Zhang, X.; Li, Y.; Chen, B.; Liu, C.; Ai, X.; Zhang, X.; Tian, Y.; Zhang, C.; et al. Cardiomyocyte PKA Ablation Enhances Basal Contractility While Eliminates Cardiac beta-Adrenergic Response Without Adverse Effects on the Heart. Circ. Res. 2019, 124, 1760–1777. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Xia, P.; Chen, J.; Bandettini, W.P.; Kirschner, L.S.; Stratakis, C.A.; Cheng, Z. PRKAR1A deficiency impedes hypertrophy and reduces heart size. Physiol. Rep. 2020, 8, e14405. [Google Scholar] [CrossRef]
- Zhang, J.; Wang, Y.; Liu, X.; Dagda, R.K.; Zhang, Y. How AMPK and PKA Interplay to Regulate Mitochondrial Function and Survival in Models of Ischemia and Diabetes. Oxid. Med. Cell. Longev. 2017, 2017, 4353510. [Google Scholar] [CrossRef]
- Gade, P.; Kalvakolanu, D.V. Chromatin immunoprecipitation assay as a tool for analyzing transcription factor activity. Methods Mol. Biol. 2012, 809, 85–104. [Google Scholar]
- Aminu, A.; Petkova, M.; Chen, W.; Yin, Z.; Kuzmin, V.; Atkinson, A.; Dobrzynski, H. MiR-486-3p and MiR-938—Important Inhibitors of Pacemaking Ion Channels and/or Markers of Immune Cells. Appl. Sci. 2021, 11, 11366. [Google Scholar] [CrossRef]

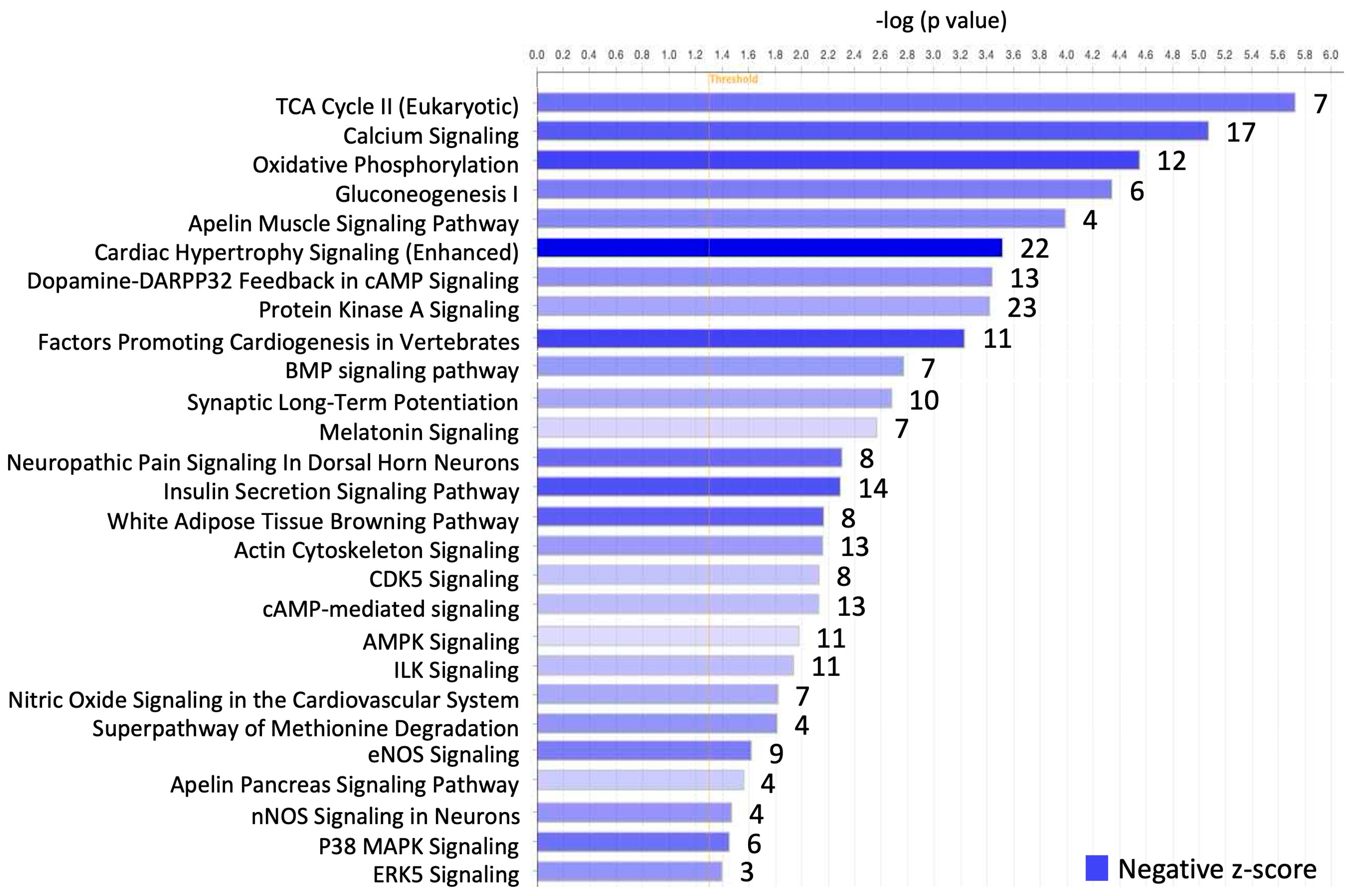

| Pathway (Number of Genes Involved) | Gene |
|---|---|
| TCA Cycle II (Eukaryotic) (7) | ACO2, CS, FH, IDH3A, MDH1, OGDH, SDHA |
| Calcium Signaling (17) | ATP2A2, CAMK2A, CAMK2B, CASQ2, CHRNE, GRIN2A, GRIN2C, MYH6, MYH14, MYL4, PNCK, PRKAG2, PRKAR2B, RCAN2, RYR2, TNNT2, TRDN |
| Oxidative Phosphorylation (12) | ATPAF1, COX10, COX5A, COX6A2, CYCS, NDUFA4, NDUFA5, NDUFAB1, NDUFS1, SDHA, UQCRB, UQCRC2 |
| Gluconeogenesis I (6) | ALDOA, ENO3, MDH1, ME1, ME2, PGAM2 |
| Apelin Muscle Signaling Pathway (4) | APLN, PRKAA2, PRKAG2, SLC2A4 |
| Cardiac Hypertrophy Signaling (Enhanced) (22) | ATP2A2, CAMK2A, CAMK2B, HSPB3, HSPB7, IFNLR1, IL5RA, MAP3K9, MAP3K20, MAPKAPK3, NPPA, PDE3A, PDE7A, PDE8B, PDK1, PLCL1, PLN, PRKAG2, PRKAR2B, RASD1, RPS6KA5, RYR2 |
| Dopamine-DARPP32 Feedback in cAMP Signaling (13) | ATP2A2, GRIN2A, GRIN2C, KCNJ11, PLCL1, PPM1L, PPP1R3A, PPP1R3C, PPP2R2C, PPP2R3A, PRKAG2, PRKAR2B, PRKG2 |
| Protein Kinase A Signaling (23) | AKAP1, AKAP3, AKAP6, CAMK2A, CAMK2B, EYA1, MYL4, MYLK3, NTN1, PDE3A, PDE7A, PDE8B, PHKG1, PLCL1, PLN, PPP1R3A, PPP1R3C, PRKAG2, PRKAR2B, PTPRQ, PYGM, RYR2, TTN |
| Factors Promoting Cardiogenesis in Vertebrates (11) | BMP7, BMP10, BMP8A, CAMK2A, CAMK2B, GJA5, MYH6, NPPA, PLCL1, SCN5A, TBX5 |
| BMP signaling pathway (7) | BMP7, BMP8A, BMP10, PRKAG2, PRKAR2B, RASD1, SOSTDC1 |
| Synaptic Long Term Potentiation (10) | CAMK2A, CAMK2B, GRIN2A, GRIN2C, PLCL1, PPP1R3A, PPP1R3C, PRKAG2, PRKAR2B, RASD1 |
| Melatonin Signaling (7) | CAMK2A, CAMK2B, PLCL1, PRKAG2, PRKAR2B, RORC, SLC2A4 |
| Neuropathic Pain Signaling In Dorsal Horn Neurons (8) | CAMK2A, CAMK2B, GRIN2A, GRIN2C, KCNH2, PLCL1, PRKAG2, PRKAR2B |
| Insulin Secretion Signaling Pathway (14) | CAMK2A, CAMK2B, DLAT, GIPR, KCNJ11, NALCN, PDHA1, PLCL1, PRKAG2, PRKAR2B, RPS6KA5, RYR2, SLC2A4, STAT4 |
| White Adipose Tissue Browning Pathway (8) | BMP7, FNDC5, NPPA, PRKAA2, PRKAG2, PRKAR2B, PRKG2, VEGFA |
| Actin Cytoskeleton Signaling (13) | ACTN2, EGF, FGF11, FGF12, MYH6, MYH14, MYL4, MYLK3, PAK6, PIP5K1B, PPP1R12B, RASD1, TTN |
| CDK5 Signaling (8) | PPM1L, PPP1R3A, PPP1R3C, PPP2R2C, PPP2R3A, PRKAG2, PRKAR2B, RASD1 |
| cAMP-mediated signaling (13) | ADORA1, AKAP1, AKAP3, AKAP6, CAMK2A, CAMK2B, CHRM2, PDE3A, PDE7A, PDE8B, PKIA, PRKAR2B, RAP1GAP |
| AMPK Signaling (11) | AK4, CHRM2, CHRNE, CKM, PPM1L, PPP2R2C, PPP2R3A, PRKAA2, PRKAG2, PRKAR2B, SLC2A4 |
| ILK Signaling (11) | ACTN2, DSP, ITGB6, MYH6, MYH14, MYL4, PPM1L, PPP2R2C, PPP2R3A, RPS6KA5, VEGFA |
| Nitric Oxide Signaling in the Cardiovascular System (7) | ATP2A2, PLN, PRKAG2, PRKAR2B, PRKG2, RYR2, VEGFA |
| Superpathway of Methionine Degradation (4) | GOT1, GOT2, PCCB, PRMT8 |
| eNOS Signaling (9) | AQP4, AQP7, CHRM2, CHRNE, LPAR3, PRKAA2, PRKAG2, PRKAR2B, VEGFA |
| Apelin Pancreas Signaling Pathway (4) | APLN, PRKAA2, PRKAG2, PRKAR2B |
| nNOS Signaling in Neurons (4) | CAMK2A, GRIN2A, GRIN2C, RASD1 |
| P38 MAPK Signaling (6) | HSPB3, HSPB7, MAPKAPK3, PLA2G12B, RPS6KA5, TIFA |
| ERK5 Signaling (3) | RASD1, RPS6KA5, WNK1 |
| mRNA | SN Mean ± SEM | RA Mean ± SEM | Log2fold Change | Adjusted p Value |
|---|---|---|---|---|
| NPPA | 40,366.13 ± 11,840.28 | 587,003.64 ± 233,459.53 | −3.80 | 3.17 × 10−47 |
| GRIN2C | 150.49 ± 41.54 | 1505.33 ± 206.64 | −3.40 | 1.32 × 10−29 |
| GRIN2A | 34.41 ± 9.15 | 395.65 ± 63.35 | −3.61 | 1.20 × 10−21 |
| LPAR3 | 239.30 ± 78.64 | 1563.00 ± 250.50 | −2.87 | 2.03 × 10−19 |
| ENO3 | 947.30 ± 433.60 | 5786.00 ± 672.30 | −2.89 | 9.77 × 10−18 |
| TMEM38B | 435.93 ± 66.81 | 2180.97 ± 137.96 | −2.37 | 5.96 × 10−17 |
| FBXO32 | 1823.69 ± 622.47 | 8064.94 ± 2141.13 | −2.22 | 3.57 × 10−16 |
| RYR2 | 6201.72 ± 2293.58 | 43,858.17 ± 21,127.23 | −2.77 | 1.82 × 10−15 |
| CHRNE | 532.36 ± 146.12 | 2493.47 ± 302.70 | −2.33 | 2.01 × 10−14 |
| MYLK3 | 2917.00 ± 966.30 | 16,247.00 ± 1172.00 | −2.65 | 5.87 × 10−14 |
| PDE3A | 1669.00 ± 133.00 | 6510.00 ± 506.40 | −1.98 | 9.75 × 10−14 |
| ME2 | 1040.19 ± 75.53 | 4288.91 ± 471.40 | −2.05 | 1.07 × 10−13 |
| PPP1R12B | 8186.14 ± 2305.97 | 37,247.20 ± 6630.09 | −2.25 | 6.54 × 10−13 |
| RAP1GAP | 107.50 ± 12.52 | 544.70 ± 21.50 | −2.37 | 7.62 × 10−13 |
| KCNIP2 | 1871.96 ± 431.95 | 9718.33 ± 735.63 | −2.47 | 1.86 × 10−12 |
| SCN1B | 458.62 ± 60.01 | 1785.74 ± 293.19 | −1.96 | 4.18 × 10−12 |
| PDE8B | 506.40 ± 113.30 | 1798.00 ± 188.20 | −1.91 | 2.64 × 10−10 |
| ATP2A2 | 32,967.84 ± 8525.27 | 140,042.55 ± 11,888.16 | −2.20 | 3.15 × 10−10 |
| SGCD | 1118.50 ± 130.93 | 4005.43 ± 752.98 | −1.83 | 4.79 × 10−10 |
| ANK1 | 326.65 ± 72.56 | 1302.83 ± 51.25 | −2.09 | 5.94 × 10−10 |
| PDK1 | 435.31 ± 85.02 | 1691.11 ± 137.84 | −2.02 | 1.09 × 10−9 |
| NEDD4L | 548.57 ± 131.29 | 2313.96 ± 493.04 | −2.13 | 1.11 × 10−9 |
| UQCRB | 8496.29 ± 1355.26 | 31,581.55 ± 7034.10 | −1.85 | 1.68 × 10−9 |
| PKIA | 957.30 ± 178.40 | 3342.00 ± 294.50 | −1.85 | 2.51 × 10−9 |
| MYBPC1 | 80.81 ± 14.79 | 424.38 ± 67.97 | −2.41 | 4.76 × 10−9 |
| TRIM63 | 1263.61 ± 398.03 | 5687.17 ± 1969.60 | −2.19 | 5.19 × 10−9 |
| PRKAA2 | 952.70 ± 259.80 | 4127.00 ± 1218.00 | −2.16 | 5.44 × 10−9 |
| ME1 | 645.39 ± 142.07 | 2306.13 ± 46.86 | −1.92 | 6.80 × 10−9 |
| SLC39A8 | 726.23 ± 149.47 | 2476.79 ± 221.81 | −1.83 | 1.51 × 10−8 |
| PRKAG2 | 3090.00 ± 572.40 | 106,57.00 ± 858.00 | −1.84 | 1.52 × 10−8 |
| MYH14 | 1172.51 ± 230.82 | 3694.07 ± 614.29 | −1.69 | 1.72 × 10−8 |
| DSG2 | 1180.31 ± 381.92 | 4273.57 ± 852.77 | −2.03 | 2.55 × 10−8 |
| DMD | 2411.16 ± 277.11 | 8679.83 ± 1901.66 | −1.81 | 4.42 × 10−8 |
| PRKAR2B | 791.19 ± 108.73 | 2418.93 ± 79.89 | −1.65 | 4.55 × 10−8 |
| NDUFA5 | 2768.78 ± 345.70 | 8071.65 ± 861.85 | −1.56 | 5.13 × 10−8 |
| DLAT | 1198.44 ± 194.96 | 3902.53 ± 187.84 | −1.75 | 5.18 × 10−8 |
| TTN | 27,820.20 ± 8702.96 | 154,512.67 ± 72,399.52 | −2.38 | 5.39 × 10−8 |
| CHRM2 | 1265.95 ± 525.92 | 3877.67 ± 1095.15 | −1.83 | 6.68 × 10−8 |
| GOT1 | 4504.00 ± 1167.00 | 15,507.00 ± 642.50 | −1.90 | 7.20 × 10−8 |
| MAP3K20 | 5446.00 ± 1082.00 | 16,464.00 ± 3505.00 | −1.60 | 9.15 × 10−8 |
| AKAP1 | 1824.00 ± 512.30 | 5030.00 ± 656.60 | −1.56 | 1.23 × 10−7 |
| TRDN | 2415.45 ± 701.59 | 10,011.69 ± 2499.98 | −2.15 | 1.65 × 10−7 |
| ABLIM3 | 2310.55 ± 372.12 | 6165.94 ± 1169.24 | −1.42 | 1.95 × 10−7 |
| KCNJ11 | 90.50 ± 26.67 | 370.59 ± 97.45 | −2.04 | 2.16 × 10−7 |
| CYCS | 4494.10 ± 1257.61 | 14,511.32 ± 1868.14 | −1.78 | 3.26 × 10−7 |
| IDH3A | 2512.14 ± 569.16 | 7355.81 ± 423.98 | −1.63 | 5.48 × 10−7 |
| SDHA | 4783.13 ± 765.73 | 14,099.54 ± 1017.44 | −1.60 | 5.76 × 10−7 |
| CAMK2B | 886.45 ± 242.81 | 2844.18 ± 389.22 | −1.81 | 6.27 × 10−7 |
| ALDOA | 21,888.38 ± 4209.53 | 69,812.69 ± 10,672.80 | −1.71 | 6.44 × 10−7 |
| COX10 | 370.51 ± 82.92 | 1177.80 ± 41.61 | −1.76 | 9.40 × 10−7 |
| COX5A | 3850.64 ± 932.14 | 12,170.01 ± 2045.61 | −1.70 | 2.30 × 10−6 |
| OGDH | 4144.09 ± 905.90 | 11,254.98 ± 1322.87 | −1.51 | 2.43 × 10−6 |
| ATPAF1 | 2336.00 ± 496.90 | 6454.00 ± 15.82 | −1.55 | 2.71 × 10−6 |
| NALCN | 200.50 ± 33.82 | 580.40 ± 38.73 | −1.58 | 2.88 × 10−6 |
| PDE7A | 454.50 ± 93.88 | 1775.00 ± 711.30 | −1.80 | 2.91 × 10−6 |
| ABLIM1 | 8707.20 ± 1976.33 | 23,835.54 ± 8276.75 | −1.39 | 3.61 × 10−6 |
| PDHA1 | 4045.38 ± 917.17 | 11,743.03 ± 669.07 | −1.63 | 4.49 × 10−6 |
| NDUFA4 | 11,401.81 ± 2311.16 | 34,441.74 ± 5244.19 | −1.63 | 5.67 × 10−6 |
| PLN | 13,987.76 ± 3665.84 | 42,146.81 ± 4579.06 | −1.70 | 5.89 × 10−6 |
| SLC25A4 | 14,584.36 ± 4389.42 | 46,121.40 ± 4482.91 | −1.80 | 5.98 × 10−6 |
| UQCRC2 | 5516.37 ± 839.18 | 15,418.20 ± 1675.86 | −1.51 | 8.95 × 10−6 |
| MDH1 | 9211.00 ± 2270.00 | 28,563.00 ± 4373.00 | −1.70 | 1.04 × 10−5 |
| APLN | 53.12 ± 20.70 | 193.86 ± 52.80 | −2.08 | 1.16 × 10−5 |
| AK4 | 954.20 ± 245.80 | 2598.00 ± 608.00 | −1.51 | 1.52 × 10−5 |
| TRIM54 | 1293.40 ± 455.02 | 3632.34 ± 413.85 | −1.75 | 1.71 × 10−5 |
| AKAP6 | 1628.06 ± 423.62 | 4406.32 ± 259.99 | −1.57 | 1.72 × 10−5 |
| ANK3 | 2195.19 ± 135.79 | 5698.77 ± 854.85 | −1.36 | 2.24 × 10−5 |
| IDH2 | 2813.86 ± 680.68 | 7679.94 ± 545.31 | −1.54 | 2.29 × 10−5 |
| SLC2A4 | 1083.62 ± 376.68 | 3169.26 ± 913.33 | −1.63 | 2.40 × 10−5 |
| ADORA1 | 202.32 ± 75.96 | 563.38 ± 53.83 | −1.72 | 3.06 × 10−5 |
| NDUFAB1 | 2853.73 ± 666.19 | 8491.52 ± 1971.77 | −1.56 | 3.21 × 10−5 |
| IMMT | 2548.38 ± 457.59 | 6293.75 ± 181.62 | −1.37 | 3.55 × 10−5 |
| PRKACA | 3651.82 ± 592.69 | 8568.10 ± 302.59 | −1.28 | 3.68 × 10−5 |
| AIFM1 | 1250.79 ± 272.32 | 3128.84 ± 75.60 | −1.41 | 6.67 × 10−5 |
| TNNT2 | 52,996.68 ± 15,892.94 | 144,599.19 ± 21,446.60 | −1.56 | 1.10 × 10−4 |
| SLC25A12 | 1658.49 ± 208.14 | 3801.41 ± 191.81 | −1.23 | 1.12 × 10−4 |
| KLHL3 | 583.44 ± 155.63 | 1617.75 ± 772.65 | −1.28 | 1.14 × 10−4 |
| TPM1 | 59,923.00 ± 20,212.99 | 153,480.87 ± 31,542.18 | −1.45 | 1.26 × 10−4 |
| MRPL35 | 1054.26 ± 163.38 | 2374.22 ± 25.60 | −1.22 | 1.46 × 10−4 |
| KCND3 | 409.54 ± 78.90 | 1311.74 ± 614.24 | −1.42 | 1.97 × 10−4 |
| GPD1L | 4931.18 ± 1450.70 | 10,273.33 ± 1593.37 | −1.19 | 2.06 × 10−4 |
| NDUFA8 | 1784.11 ± 388.01 | 4248.67 ± 601.70 | −1.30 | 2.44 × 10−4 |
| NDUFB5 | 2470.69 ± 347.08 | 5857.23 ± 754.88 | −1.26 | 2.71 × 10−4 |
| PDE4D | 934.41 ± 155.71 | 2037.22 ± 476.12 | −1.09 | 2.87 × 10−4 |
| PRKAR1A | 7776.36 ± 550.02 | 15,506.23 ± 1274.24 | −1.01 | 3.71 × 10−4 |
| CYC1 | 3470.36 ± 625.10 | 8433.44 ± 1428.91 | −1.29 | 4.35 × 10−4 |
| EDNRB | 1215.00 ± 114.40 | 2583.00 ± 176.70 | −1.11 | 4.69 × 10−4 |
| GYS1 | 1783.00 ± 630.40 | 3749.00 ± 805.40 | −1.21 | 5.40 × 10−4 |
| MRPS23 | 674.46 ± 103.97 | 1396.07 ± 128.12 | −1.08 | 5.40 × 10−4 |
| HIGD1A | 1351.36 ± 282.95 | 2904.10 ± 537.75 | −1.12 | 5.62 × 10−4 |
| MRPL33 | 1310.67 ± 180.02 | 2891.31 ± 306.91 | −1.16 | 5.85 × 10−4 |
| SLC25A11 | 1854.97 ± 336.97 | 4194.34 ± 509.14 | −1.21 | 9.34 × 10−4 |
| MRPL39 | 449.21 ± 36.86 | 985.84 ± 149.48 | −1.12 | 9.68 × 10−4 |
| NDUFA6 | 2675.74 ± 698.20 | 6039.42 ± 1259.69 | −1.20 | 1.04 × 10−3 |
| COX5B | 6125.33 ± 1447.09 | 14,888.12 ± 3592.31 | −1.26 | 1.10 × 10−3 |
| DLD | 2161.55 ± 418.65 | 4964.17 ± 623.10 | −1.25 | 1.20 × 10−3 |
| MRPL50 | 540.20 ± 47.38 | 1071.58 ± 32.52 | −1.01 | 1.23 × 10−3 |
| COX8A | 1709.46 ± 377.55 | 4097.35 ± 964.90 | −1.23 | 1.29 × 10−3 |
| AFG3L2 | 1471.45 ± 191.94 | 2892.52 ± 40.73 | −1.01 | 1.30 × 10−3 |
| CAMK2A | 226.40 ± 89.79 | 539.50 ± 31.36 | −1.54 | 1.43 × 10−3 |
| COX6B1 | 4041.13 ± 1030.85 | 9143.47 ± 2151.20 | −1.18 | 1.55 × 10−3 |
| MRPS22 | 683.43 ± 37.07 | 1432.12 ± 201.58 | −1.05 | 1.57 × 10−3 |
| NDUFB10 | 3759.25 ± 798.24 | 8448.57 ± 1735.75 | −1.16 | 1.73 × 10−3 |
| SLC25A3 | 10,661.69 ± 1643.67 | 22,876.10 ± 3179.36 | −1.12 | 2.08 × 10−3 |
| NDUFS5 | 4811.13 ± 932.38 | 9909.52 ± 2177.33 | −1.02 | 2.92 × 10−3 |
| PPIF | 1692.49 ± 512.50 | 3511.49 ± 61.05 | −1.24 | 3.34 × 10−3 |
| MRPS33 | 721.80 ± 49.51 | 1461.57 ± 210.66 | −1.01 | 3.43 × 10−3 |
| CRYAB | 42,557.08 ± 15,964.14 | 84,832.79 ± 15,603.49 | −1.13 | 3.44 × 10−3 |
| SVIL | 6089.43 ± 1387.26 | 12,895.94 ± 3926.73 | −1.04 | 3.67 × 10−3 |
| MRPS9 | 865.43 ± 123.52 | 1804.79 ± 260.93 | −1.07 | 4.47 × 10−3 |
| DES | 112,042.00 ± 40,572.00 | 202,808.00 ± 17,832.00 | −1.14 | 7.18 × 10−3 |
| MRPL15 | 1129.54 ± 234.79 | 2254.08 ± 289.96 | −1.04 | 7.61 × 10−3 |
| TF (Number of Interactions) | mRNA Target |
|---|---|
| MYCN (29) | MRPL15, MRPL33, MRPS23, MRPS9, MRPL39, SDHA, MRPL35, NDUFS5, COX5B, COX5A, IDH2, NDUFB10, MRPL50, NDUFA6, MRPS33, SLC25A4, NDUFA5, DLAT, SLC25A11, MRPS22, UQCRC2, COX6B1, SLC25A3, NDUFAB1, NDUFA8, TTN, ATP2A2, CAMK2A, MYH14 |
| SOX2 (14) | SLC25A12, MDH1, CRYAB, SLC25A11, MRPL39, MRPS9, MRPS23, MRPL15, AIFM1, ABLIM3, MYH14, PRKAR2B, PRKAR1A, PDE4D |
| NUPR1 (7) | IMMT, DLD, PDHA1, OGDH, AIFM1, DSG2, SVIL |
| PRDM16 (5) | CYC1, IMMT, NDUFA4, SLC25A11, ATP2A2 |
| LMO2 (4) | NDUFA8, NDUFAB1, TRIM54, TRIM63 |
| GLI1 (3) | DES, PRKAA2, PRKACA |
| TBX3 (3) | SLC25A3, AFG3L2, SVIL |
| VAV1 (3) | SLC25A11, SLC25A3, ABLIM1 |
| HDAC10 (2) | NDUFB5, TRIM54 |
| KCNIP3 (2) | KCNIP2, KCND3 |
| PYCARD (2) | GOT1, PRKAR1A |
| CEBPA (1) | SLC25A11 |
| IKZF1 (1) | PRKAA2 |
| TGFB1I1 (1) | NEDD4L |
| ATF5 (1) | TNNT2 |
| BCL11B (1) | MAP3K20 |
| BATF (1) | COX8A |
| DTX1 (1) | PRKAR1A |
| HOXB3 (1) | SDHA |
| LBH (1) | CRYAB |
| MLXIPL (1) | PRKAA2 |
| SHOX2 (1) | GRIN2C |
| SOX9 (1) | KLHL3 |
| TBX2 (1) | AFG3L2 |
| TSC22D3 (1) | NEDD4L |
| miRNA (Number of Interactions) | mRNA of Predicted Target | Number of Binding Sites for Each miRNA on Predicted Target (RNA22) | Number of Binding Sites for Each miRNA on Predicted Target (TargetScanHuman) |
|---|---|---|---|
| hsa-miR-153-3p (19) | ANK3 | 0 | 2 |
| TRDN | 0 | 1 | |
| DSG2 | 1 | 1 | |
| PPP1R12B | 0 | 1 | |
| DMD | 0 | 1 | |
| KLHL3 | 1 | 3 | |
| RYR2 | 0 | 2 | |
| KCNIP2 | 0 | 1 | |
| PDE7A | 0 | 2 | |
| PRKAA2 | 1 | 1 | |
| SGCD | 1 | 1 | |
| AKAP6 | 0 | 1 | |
| ABLIM1 | 1 | 1 | |
| ANK1 | 0 | 1 | |
| CHRM2 | 0 | 1 | |
| GPD1L | 0 | 1 | |
| FBXO32 | 1 | 1 | |
| SLC39A8 | 0 | 1 | |
| ME1 | 1 | 1 | |
| hsa-miR-654-5p (15) | TRIM54 | 1 | 0 |
| MYBPC1 | 1 | 0 | |
| CAMK2A | 3 | 0 | |
| CHRNE | 1 | 0 | |
| KCNIP2 | 1 | 0 | |
| KCNJ11 | 2 | 0 | |
| APLN | 5 | 0 | |
| GRIN2A | 8 | 0 | |
| ADORA1 | 1 | 0 | |
| ANK1 | 2 | 0 | |
| PRKACA | 1 | 0 | |
| OGDH | 4 | 0 | |
| ATPAF1 | 4 | 0 | |
| ME2 | 4 | 0 | |
| PPIF | 4 | 0 | |
| hsa-miR-10a-5p (13) | PRKAA2 | 0 | 2 |
| PDE7A | 0 | 2 | |
| PDE8B | 0 | 3 | |
| CAMK2B | 0 | 2 | |
| TMEM38B | 0 | 4 | |
| PRKACA | 0 | 1 | |
| LPAR3 | 0 | 2 | |
| ANK1 | 0 | 2 | |
| ABLIM1 | 0 | 1 | |
| SGCD | 0 | 1 | |
| IDH3A | 0 | 1 | |
| RAP1GAP | 4 | 1 | |
| AK4 | 0 | 7 | |
| hsa-miR-204-5p (9) | AKAP1 | 1 | 2 |
| CHRM2 | 0 | 7 | |
| PDE3A | 1 | 2 | |
| PKIA | 0 | 2 | |
| MRPS33 | 2 | 0 | |
| AK4 | 2 | 4 | |
| COX10 | 0 | 1 | |
| PDHA1 | 0 | 1 | |
| COX5A | 0 | 2 | |
| hsa-miR-215-5p (9) | PLN | 0 | 2 |
| LPAR3 | 0 | 2 | |
| GRIN2A | 2 | 2 | |
| SGCD | 0 | 2 | |
| PRKAR1A | 0 | 2 | |
| APLN | 0 | 1 | |
| GOT1 | 0 | 2 | |
| HIGD1A | 0 | 2 | |
| CYCS | 1 | 3 | |
| hsa-miR-193b-3p (6) | MYLK3 | 0 | 3 |
| ANK3 | 0 | 2 | |
| APLN | 0 | 1 | |
| KCNJ11 | 0 | 1 | |
| SGCD | 0 | 2 | |
| PPIF | 0 | 2 | |
| hsa-miR-885-3p (5) | SCN1B | 3 | 0 |
| TPM1 | 2 | 0 | |
| ADORA1 | 5 | 0 | |
| COX10 | 3 | 0 | |
| OGDH | 3 | 0 | |
| hsa-miR-668-3p (4) | TTN | 1 | 1 |
| NALCN | 2 | 0 | |
| PDK1 | 4 | 0 | |
| AK4 | 4 | 1 | |
| hsa-miR-548b-5p (3) | TPM1 | 1 | 0 |
| HIGD1A | 2 | 0 | |
| UQCRB | 10 | 0 | |
| hsa-miR-512-5p (3) | CAMK2B | 2 | 0 |
| CYCS | 2 | 0 | |
| MRPL50 | 1 | 0 | |
| hsa-miR-377-5p (3) | EDNRB | 1 | 0 |
| ANK1 | 3 | 0 | |
| NPPA | 1 | 0 | |
| hsa-miR-1225-3p (2) | CAMK2B | 2 | 0 |
| PRKACA | 2 | 0 | |
| hsa-miR-1244 (2) | GPD1L | 1 | 0 |
| TMEM38B | 1 | 0 | |
| hsa-miR-10b-3P (2) | GRIN2A | 2 | 0 |
| MRPL35 | 1 | 0 | |
| hsa-miR-198 (1) | RYR2 | 1 | 0 |
| hsa-miR-548k (1) | PDK1 | 2 | 0 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, W.; Aminu, A.J.; Yin, Z.; Karaesmen, I.; Atkinson, A.J.; Kuniewicz, M.; Holda, M.; Walocha, J.; Perde, F.; Molenaar, P.; et al. Profiling Reduced Expression of Contractile and Mitochondrial mRNAs in the Human Sinoatrial Node vs. Right Atrium and Predicting Their Suppressed Expression by Transcription Factors and/or microRNAs. Int. J. Mol. Sci. 2024, 25, 10402. https://doi.org/10.3390/ijms251910402
Chen W, Aminu AJ, Yin Z, Karaesmen I, Atkinson AJ, Kuniewicz M, Holda M, Walocha J, Perde F, Molenaar P, et al. Profiling Reduced Expression of Contractile and Mitochondrial mRNAs in the Human Sinoatrial Node vs. Right Atrium and Predicting Their Suppressed Expression by Transcription Factors and/or microRNAs. International Journal of Molecular Sciences. 2024; 25(19):10402. https://doi.org/10.3390/ijms251910402
Chicago/Turabian StyleChen, Weixuan, Abimbola J. Aminu, Zeyuan Yin, Irem Karaesmen, Andrew J. Atkinson, Marcin Kuniewicz, Mateusz Holda, Jerzy Walocha, Filip Perde, Peter Molenaar, and et al. 2024. "Profiling Reduced Expression of Contractile and Mitochondrial mRNAs in the Human Sinoatrial Node vs. Right Atrium and Predicting Their Suppressed Expression by Transcription Factors and/or microRNAs" International Journal of Molecular Sciences 25, no. 19: 10402. https://doi.org/10.3390/ijms251910402
APA StyleChen, W., Aminu, A. J., Yin, Z., Karaesmen, I., Atkinson, A. J., Kuniewicz, M., Holda, M., Walocha, J., Perde, F., Molenaar, P., & Dobrzynski, H. (2024). Profiling Reduced Expression of Contractile and Mitochondrial mRNAs in the Human Sinoatrial Node vs. Right Atrium and Predicting Their Suppressed Expression by Transcription Factors and/or microRNAs. International Journal of Molecular Sciences, 25(19), 10402. https://doi.org/10.3390/ijms251910402






