Bioinformatic Analysis from a Descriptive Profile of miRNAs in Chronic Migraine
Abstract
1. Introduction
2. Results
2.1. miRs Profile Expression in Patients with Chronic Migraine
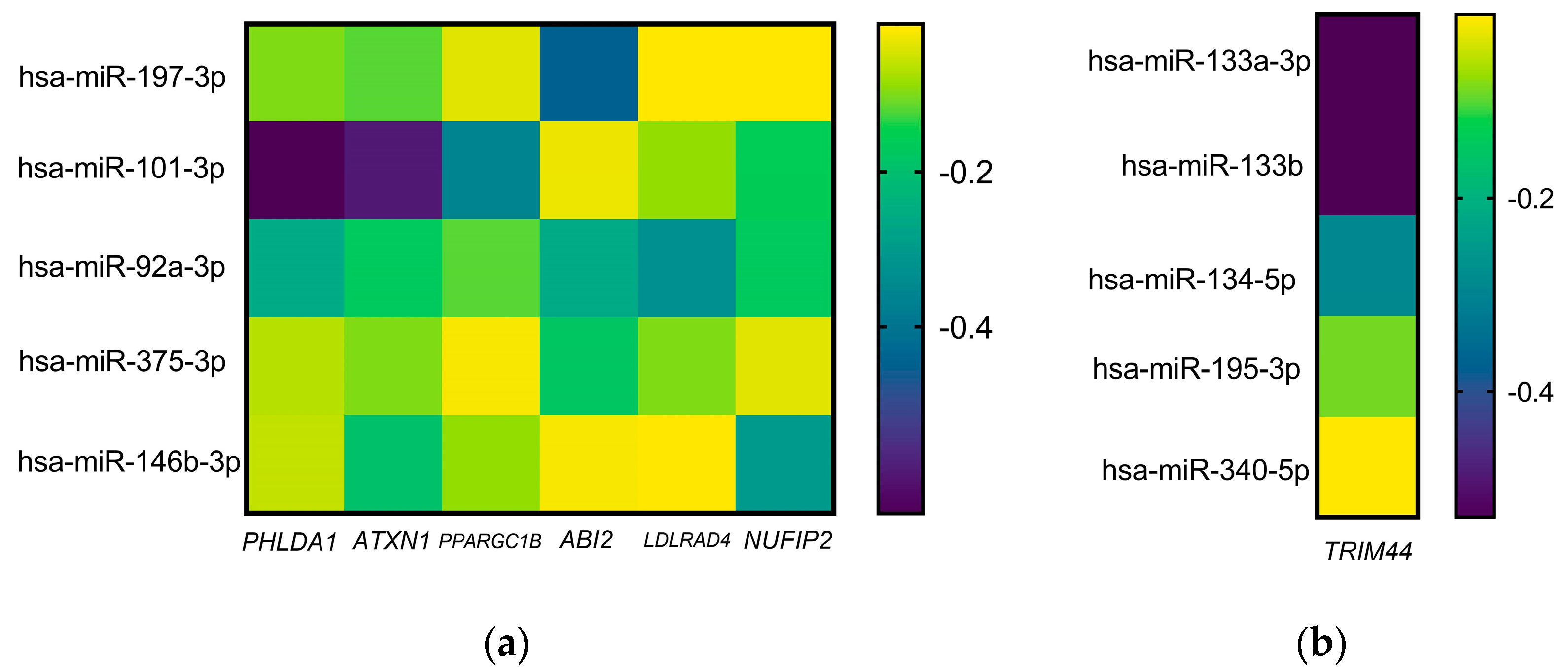
2.2. miR-Targets Interaction in Patients with Chronic Migraine
2.3. Signalling Pathways in miR-Targets Profile in Chronic Migraine
2.4. Relationship among Chronic Migraine miRs and Other Pathologies
3. Discussion
4. Materials and Methods
4.1. Patient Selection and Blood Sample Collection
4.2. Extraction and Reverse Transcription of MicroRNAs from Blood Plasma
4.3. MicroRNA Expression
4.4. Bioinformatic Analysis of the Results
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Stovner, L.J.; Hagen, K.; Linde, M.; Steiner, T.J. The global prevalence of headache: An update, with analysis of the influences of methodological factors on prevalence estimates. J. Headache Pain 2022, 23, 34. [Google Scholar] [CrossRef] [PubMed]
- Aczel, T.; Benczik, B.; Agg, B.; Kortesi, T.; Urban, P.; Bauer, W.; Gyenesei, A.; Tuka, B.; Tajti, J.; Ferdinandy, P.; et al. Disease- and headache-specific microRNA signatures and their predicted mRNA targets in peripheral blood mononuclear cells in migraineurs: Role of inflammatory signalling and oxidative stress. J. Headache Pain 2022, 23, 113. [Google Scholar] [CrossRef] [PubMed]
- Ran, Y.; Yin, Z.; Lian, Y.; Xu, Y.; Li, Y.; Liu, J.; Gu, Q.; Yan, F.; Ge, Z.; Lian, Y.; et al. Gradually shifting clinical phenomics in migraine spectrum: A cross-sectional, multicenter study of 5438 patients. J. Headache Pain 2022, 23, 89. [Google Scholar] [CrossRef] [PubMed]
- Camara, M.S.; Bujanda, M.M.; Iriarte, M.M. Epigenetic changes in headache. Neurologia (Engl. Ed.) 2021, 36, 369–376. [Google Scholar]
- Treiber, T.; Treiber, N.; Meister, G. Regulation of microRNA biogenesis and its crosstalk with other cellular pathways. Nat. Rev. Mol. Cell Biol. 2019, 20, 5–20. [Google Scholar] [CrossRef]
- Gallelli, L.; Cione, E.; Caroleo, M.C.; Carotenuto, M.; Lagana, P.; Siniscalchi, A.; Guidetti, V. microRNAs to Monitor Pain-migraine and Drug Treatment. Microrna 2017, 6, 152–156. [Google Scholar] [CrossRef]
- O‘Brien, J.; Hayder, H.; Zayed, Y.; Peng, C. Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Front. Endocrinol. 2018, 9, 402. [Google Scholar] [CrossRef]
- Xiang, W.; Jiang, L.; Zhou, Y.; Li, Z.; Zhao, Q.; Wu, T.; Cao, Y.; Zhou, J. The lncRNA Ftx/miR-382-5p/Nrg1 axis improves the inflammation response of microglia and spinal cord injury repair. Neurochem. Int. 2021, 143, 104929. [Google Scholar] [CrossRef]
- Greco, R.; De Icco, R.; Demartini, C.; Zanaboni, A.M.; Tumelero, E.; Sances, G.; Allena, M.; Tassorelli, C. Plasma levels of CGRP and expression of specific microRNAs in blood cells of episodic and chronic migraine subjects: Towards the identification of a panel of peripheral biomarkers of migraine? J. Headache Pain 2020, 21, 122. [Google Scholar] [CrossRef]
- Ornello, R.; Zelli, V.; Compagnoni, C.; Caponnetto, V.; De Matteis, E.; Tiseo, C.; Tessitore, A.; Sacco, S. MicroRNA profiling in women with migraine: Effects of CGRP-targeting treatment. J. Headache Pain 2024, 25, 80. [Google Scholar] [CrossRef]
- Han, C.; Yan, P.; He, T.; Cheng, J.; Zheng, W.; Zheng, L.T.; Zhen, X. PHLDA1 promotes microglia-mediated neuroinflammation via regulating K63-linked ubiquitination of TRAF6. Brain Behav. Immun. 2020, 88, 640–653. [Google Scholar] [CrossRef] [PubMed]
- Fearon, A.E.; Carter, E.P.; Clayton, N.S.; Wilkes, E.H.; Baker, A.M.; Kapitonova, E.; Bakhouche, B.A.; Tanner, Y.; Wang, J.; Gadaleta, E.; et al. PHLDA1 Mediates Drug Resistance in Receptor Tyrosine Kinase-Driven Cancer. Cell Rep. 2018, 22, 2469–2481. [Google Scholar] [CrossRef] [PubMed]
- Handler, H.P.; Duvick, L.; Mitchell, J.S.; Cvetanovic, M.; Reighard, M.; Soles, A.; Mather, K.B.; Rainwater, O.; Serres, S.; Nichols-Meade, T.; et al. Decreasing mutant ATXN1 nuclear localization improves a spectrum of SCA1-like phenotypes and brain region transcriptomic profiles. Neuron 2023, 111, 493–507.e6. [Google Scholar] [CrossRef]
- Lou, L.Q.; Zhou, W.Q.; Song, X.; Chen, Z. Elevation of hsa-miR-7-5p level mediated by CtBP1-p300-AP1 complex targets ATXN1 to trigger NF-kappaB-dependent inflammation response. J. Mol. Med. 2023, 101, 223–235. [Google Scholar] [CrossRef] [PubMed]
- Merriman, T.; Terkeltaub, R. PPARGC1B: Insight into the expression of the gouty inflammation phenotype: PPARGC1B and gouty inflammation. Rheumatology 2017, 56, 323–325. [Google Scholar] [CrossRef][Green Version]
- Jensen, C.C.; Clements, A.N.; Liou, H.; Ball, L.E.; Bethard, J.R.; Langlais, P.R.; Toth, R.K.; Chauhan, S.S.; Casillas, A.L.; Daulat, S.R.; et al. PIM1 phosphorylates ABI2 to enhance actin dynamics and promote tumor invasion. J. Cell Biol. 2023, 222, e202208136. [Google Scholar] [CrossRef]
- Ichigotani, Y.; Fujii, K.; Hamaguchi, M.; Matsuda, S. In search of a function for the E3B1/Abi2/Argbp1/NESH family (Review). Int. J. Mol. Med. 2002, 9, 591–595. [Google Scholar] [CrossRef]
- Liu, Z.; Huo, X.; Zhao, S.; Yang, J.; Shi, W.; Jing, L.; Li, W.; Li, Y.; Ma, L.; Gao, Y.; et al. Low density lipoprotein receptor class A domain containing 4 (LDLRAD4) promotes tumorigenesis of hepatic cancer cells. Exp. Cell Res. 2017, 360, 189–198. [Google Scholar] [CrossRef] [PubMed]
- Jin, F.; Li, L.; Hao, Y.; Tang, L.; Wang, Y.; He, Z. Identification of Candidate Blood mRNA Biomarkers in Intracerebral Hemorrhage Using Integrated Microarray and Weighted Gene Co-expression Network Analysis. Front. Genet. 2021, 12, 707713. [Google Scholar] [CrossRef]
- Jia, J.; Wang, F.; Bhujabal, Z.; Peters, R.; Mudd, M.; Duque, T.; Allers, L.; Javed, R.; Salemi, M.; Behrends, C.; et al. Membrane Atg8ylation, stress granule formation, and MTOR regulation during lysosomal damage. Autophagy 2023, 19, 1893–1895. [Google Scholar] [CrossRef]
- Liu, L.; Liu, M.; Zhang, D.; Song, Z.; Zhang, H. DHT inhibits REDOX damage and neuroinflammation to reduce PND occurrence in aged mice via mmu_circ_0001442/miR-125a-3p/NUFIP2 axis. Brain Behav. 2023, 13, e3180. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Dong, C.; Yue, X.; Ge, P.; Zheng, G.; Ye, Z.; Pan, B. Silencing of TRIM44 Inhibits Inflammation and Alleviates Traumatic Brain Injury in Rats by Downregulating TLR4-NF-kappaB Signaling. Neuroimmunomodulation 2022, 29, 439–449. [Google Scholar] [CrossRef] [PubMed]
- Gazerani, P. Current Evidence on Potential Uses of MicroRNA Biomarkers for Migraine: From Diagnosis to Treatment. Mol. Diagn. Ther. 2019, 23, 681–694. [Google Scholar] [CrossRef]
- Olesen, J. Headache Classification Committee of the International Headache Society (IHS) The International Classification of Headache Disorders, 3rd edition. Cephalalgia 2018, 38, 1–211. [Google Scholar]
- Liu, L.; Qi, W.; Wang, Y.; Ni, X.; Gao, S.; Zhou, Z.; Chen, D.; He, Z.; Sun, M.; Wang, Z.; et al. Circulating exosomal microRNA profiles in migraine patients receiving acupuncture treatment: A placebo-controlled clinical trial. Front. Mol. Neurosci. 2022, 15, 1098766. [Google Scholar]
- Gallelli, L.; Cione, E.; Peltrone, F.; Siviglia, S.; Verano, A.; Chirchiglia, D.; Zampogna, S.; Guidetti, V.; Sammartino, L.; Montana, A.; et al. Hsa-miR-34a-5p and hsa-miR-375 as Biomarkers for Monitoring the Effects of Drug Treatment for Migraine Pain in Children and Adolescents: A Pilot Study. J. Clin. Med. 2019, 8, 928. [Google Scholar] [CrossRef]
- Tafuri, E.; Santovito, D.; de Nardis, V.; Marcantonio, P.; Paganelli, C.; Affaitati, G.; Bucci, M.; Mezzetti, A.; Giamberardino, M.A.; Cipollone, F. MicroRNA profiling in migraine without aura: Pilot study. Ann. Med. 2015, 47, 468–473. [Google Scholar] [CrossRef]
- Cheng, C.Y.; Chen, S.P.; Liao, Y.C.; Fuh, J.L.; Wang, Y.F.; Wang, S.J. Elevated circulating endothelial-specific microRNAs in migraine patients: A pilot study. Cephalalgia 2018, 38, 1585–1591. [Google Scholar] [CrossRef]
- Zobdeh, F.; Eremenko, I.I.; Akan, M.A.; Tarasov, V.V.; Chubarev, V.N.; Schioth, H.B.; Mwinyi, J. The Epigenetics of Migraine. Int. J. Mol. Sci. 2023, 24, 9127. [Google Scholar] [CrossRef]
- Gallardo, V.J.; Gomez-Galvan, J.B.; Asskour, L.; Torres-Ferrus, M.; Alpuente, A.; Caronna, E.; Pozo-Rosich, P. A study of differential microRNA expression profile in migraine: The microMIG exploratory study. J. Headache Pain 2023, 24, 11. [Google Scholar] [CrossRef]
- Li, M.P.; Hu, Y.D.; Hu, X.L.; Zhang, Y.J.; Yang, Y.L.; Jiang, C.; Tang, J.; Chen, X.P. MiRNAs and miRNA Polymorphisms Modify Drug Response. Int. J. Environ. Res. Public. Health 2016, 13, 1096. [Google Scholar] [CrossRef]
- Andersen, H.H.; Duroux, M.; Gazerani, P. Serum MicroRNA Signatures in Migraineurs During Attacks and in Pain-Free Periods. Mol. Neurobiol. 2016, 53, 1494–1500. [Google Scholar] [CrossRef]
- Gao, F.; Chen, X.; Xu, B.; Luo, Z.; Liang, Y.; Fang, S.; Li, M.; Wang, X.; Lin, X. Inhibition of MicroRNA-92 alleviates atherogenesis by regulation of macrophage polarization through targeting KLF4. J. Cardiol. 2022, 79, 432–438. [Google Scholar] [CrossRef] [PubMed]
- Tian, X.; Liu, J.; Jia, W.; Jiang, P.; Cheng, Z.; Zhang, Y.; Li, J.; Liu, X.; Tian, C. MiR-197-3p affects angiogenesis and inflammation of endothelial cells by targeting CXCR2/COX2 axis. Am. J. Transl. Res. 2022, 14, 4666–4677. [Google Scholar] [PubMed]
- Zhao, X.; Li, S.; Wang, Z.; Bai, N.; Feng, Y. miR-101-3p negatively regulates inflammation in systemic lupus erythematosus via MAPK1 targeting and inhibition of the NF-kappaB pathway. Mol. Med. Rep. 2021, 23, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Guo, F.; Xu, L. Downregulation of microRNA-101-3p participates in systemic lupus erythematosus progression via negatively regulating HDAC9. J. Cell. Biochem. 2020, 121, 4310–4320. [Google Scholar] [CrossRef]
- Artimovic, P.; Spakova, I.; Macejkova, E.; Pribulova, T.; Rabajdova, M.; Marekova, M.; Zavacka, M. The ability of microRNAs to regulate the immune response in ischemia/reperfusion inflammatory pathways. Genes. Immun. 2024, 25, 277–296. [Google Scholar] [CrossRef]
- Xu, P.; Xin, J.; Song, L.; Chen, Y.; Ma, J.; Liu, L.; Qi, Z.; Pan, X.; Zhou, S. Serum miR-133 as a Potential Biomarker in Acute Cerebral Infarction Patients. Clin. Lab. 2020, 66. [Google Scholar] [CrossRef]
- Dai, Z.; Chu, H.; Ma, J.; Yan, Y.; Zhang, X.; Liang, Y. The Regulatory Mechanisms and Therapeutic Potential of MicroRNAs: From Chronic Pain to Morphine Tolerance. Front. Mol. Neurosci. 2018, 11, 80. [Google Scholar] [CrossRef]
- Ni, J.; Gao, Y.; Gong, S.; Guo, S.; Hisamitsu, T.; Jiang, X. Regulation of mu-opioid type 1 receptors by microRNA134 in dorsal root ganglion neurons following peripheral inflammation. Eur. J. Pain. 2013, 17, 313–323. [Google Scholar] [CrossRef]
- Yunta, M.; Nieto-Diaz, M.; Esteban, F.J.; Caballero-Lopez, M.; Navarro-Ruiz, R.; Reigada, D.; Pita-Thomas, D.W.; del Aguila, A.; Munoz-Galdeano, T.; Maza, R.M. MicroRNA dysregulation in the spinal cord following traumatic injury. PLoS ONE 2012, 7, e34534. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Tao, R.; Wang, J.; Xia, L. Upregulation of miR-375 level ameliorates morphine analgesic tolerance in mouse dorsal root ganglia by inhibiting the JAK2/STAT3 pathway. J. Pain Res. 2017, 10, 1279–1287. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Li, P.; Ni, Y.; Zhao, J.; Liu, Z. Decreased microRNA-125a-3p contributes to upregulation of p38 MAPK in rat trigeminal ganglions with orofacial inflammatory pain. PLoS ONE 2014, 9, e111594. [Google Scholar] [CrossRef]

| Gene | Associated Conditions | Regulator miRs |
|---|---|---|
| PHLDA1 | Neuroinflammation [11], Drug resistance [12] | 197-3p, 101-3p, 92-3p, 375, 146b-3p |
| ATXN1 | Spinocerebellar ataxia type 1 [13] Inflammation via NF-κB regulation [14] | 197-3p, 101-3p, 92-3p, 375, 146b-3p |
| PPARGC1B | Gouty inflammation by NLRP3-inflammasome [15] | 197-3p, 101-3p, 92-3p, 375, 146b-3p |
| ABI2 | Tumor promotion and invasion [16,17] | 197-3p, 101-3p, 92-3p, 375, 146b-3p |
| LDLRAD4 | Metastasis [18], Intracerebral hemorrhage [19] | 197-3p, 101-3p, 92-3p, 375, 146b-3p |
| NUFIP2 | Lysosomal damage [20], Neuroinflammation [21] | 197-3p, 101-3p, 92-3p, 375, 146b-3p |
| TRIM44 | Inflammation [22] | 133a, 133b, 134, 195-3p, 340 |
| miRs | Pathway | Hits (P) |
|---|---|---|
| hsa-miR-197-3p | Nervous system development | 970 (6.45 × 10−6) |
| Neuronal differentiation | 552 (0.001) | |
| Neuronal projection | 477 (0.001) | |
| Neuronal system | 191 (0.002) | |
| Neurogenesis | 645 (0.002) | |
| hsa-miR-101-3p | Nervous system development | 950 (9.43 × 10−7) |
| Neuronal differentiation | 441 (1.22 × 10−4) | |
| Neuronal projection | 208 (9.80 × 10−6) | |
| Neurogenesis | 645 (1.67 × 10−5) | |
| Immune system processes | 100 (1.12 × 10−5) | |
| Central nervous system vasculogenesis | 2 (0.023) | |
| hsa-miR-92a-3p | Nervous system development | 883 (4.12 × 10−5) |
| Neuronal differentiation | 509 (2.87 × 10−4) | |
| Neuronal projection | 430 (0.010) | |
| Neurogenesis | 609 (4.23 × 10−5) | |
| Immune system processes | 258 (0.023) | |
| Blood vessel development | 277 (0.009) | |
| hsa-miR-375-3p | Nervous system development | 792 (3.23 × 10−7) |
| Neuronal differentiation | 441 (9.59 × 10−4) | |
| Neurogenesis | 523 (5.37 × 10−4) | |
| Blood vessel development | 239 (0.025) | |
| hsa-miR-146b-3p | Nervous system development | 886 (1.36 × 10−4) |
| Neuronal differentiation | 513 (4.34 × 10−4) | |
| Neuronal projection | 456 (7.68 × 10−5) | |
| Neurogenesis | 612 (1.36 × 10−4) | |
| Blood vessel morphogenesis | 242 (0.028) | |
| hsa-miR-133a-3p | Nervous system development | 745 (1.01 × 10−7) |
| Neuronal differentiation | 424 (6.55 × 10−5) | |
| Neuronal projection | 360 (9.13 × 10−4) | |
| Neurogenesis | 502 (1.89 × 10−5) | |
| Immune system development | 19 (0.003) | |
| Immune system processes | 34 (0.009) | |
| Vascular development | 12 (0.031) | |
| Blood vessel morphogenesis | 12 (0.025) | |
| hsa-miR-133b | Nervous system development | 745 (1.01 × 10−7) |
| Neuronal differentiation | 424 (6.55 × 10−5) | |
| Neuronal projection | 360 (9.13 × 10−4) | |
| Neurogenesis | 14 (0.026) | |
| Immune system development | 12 (0.007) | |
| Immune system processes | 22 (0.021) | |
| Vascular development | 12 (9.29 × 10−4) | |
| Blood vessel development | 12 (7.13 × 10−4) | |
| hsa-miR-134-5p | Nervous system development | 773 (0.002) |
| Neuronal projection | 402 (1.77 × 10−4) | |
| Neurogenesis | 512 (0.036) | |
| Vascular development | 261 (0.007) | |
| Blood vessel development | 250 (0.011) | |
| Blood vessel morphogenesis | 223 (0.009) | |
| hsa-miR-195-3p | Nervous system development | 959 (2.50 × 10−11) |
| Neuronal differentiation | 544 (3.35 × 10−7) | |
| Neuronal projection | 445 (0.010) | |
| Neurogenesis | 637 (5.54 × 10−7) | |
| Immune system development | 380 (0.009) | |
| Blood vessel development | 304 (7.85 × 10−6) | |
| Blood vessel morphogenesis | 267 (3.10 × 10−5) | |
| hsa-miR-340-5p | Nervous system development | 701 (1.02 × 10−4) |
| Neuronal differentiation | 695 (6.98 × 10−7) | |
| Neurogenesis | 826 (1.10 × 10−7) | |
| Immune system development | 505 (5.11 × 10−4) | |
| Cytokine signaling | 8 (0.009) | |
| Vascular development | 226 (0.027) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tovar-Cuevas, A.J.; Rosales Gómez, R.C.; Martín-Márquez, B.T.; Peña Dueñas, N.A.; Sandoval-García, F.; Guzmán Ornelas, M.O.; Chávez Tostado, M.; Hernández Corona, D.M.; Corona Meraz, F.-I. Bioinformatic Analysis from a Descriptive Profile of miRNAs in Chronic Migraine. Int. J. Mol. Sci. 2024, 25, 10491. https://doi.org/10.3390/ijms251910491
Tovar-Cuevas AJ, Rosales Gómez RC, Martín-Márquez BT, Peña Dueñas NA, Sandoval-García F, Guzmán Ornelas MO, Chávez Tostado M, Hernández Corona DM, Corona Meraz F-I. Bioinformatic Analysis from a Descriptive Profile of miRNAs in Chronic Migraine. International Journal of Molecular Sciences. 2024; 25(19):10491. https://doi.org/10.3390/ijms251910491
Chicago/Turabian StyleTovar-Cuevas, Alvaro Jovanny, Roberto Carlos Rosales Gómez, Beatriz Teresita Martín-Márquez, Nathan Alejandro Peña Dueñas, Flavio Sandoval-García, Milton Omar Guzmán Ornelas, Mariana Chávez Tostado, Diana Mercedes Hernández Corona, and Fernanda-Isadora Corona Meraz. 2024. "Bioinformatic Analysis from a Descriptive Profile of miRNAs in Chronic Migraine" International Journal of Molecular Sciences 25, no. 19: 10491. https://doi.org/10.3390/ijms251910491
APA StyleTovar-Cuevas, A. J., Rosales Gómez, R. C., Martín-Márquez, B. T., Peña Dueñas, N. A., Sandoval-García, F., Guzmán Ornelas, M. O., Chávez Tostado, M., Hernández Corona, D. M., & Corona Meraz, F.-I. (2024). Bioinformatic Analysis from a Descriptive Profile of miRNAs in Chronic Migraine. International Journal of Molecular Sciences, 25(19), 10491. https://doi.org/10.3390/ijms251910491








