Identification and Functional Analysis of Ras-Related Associated with Diabetes Gene (rrad) in Edwardsiella piscicida-Resistant Individuals of Japanese Flounder (Paralichthys olivaceus)
Abstract
1. Introduction
2. Results
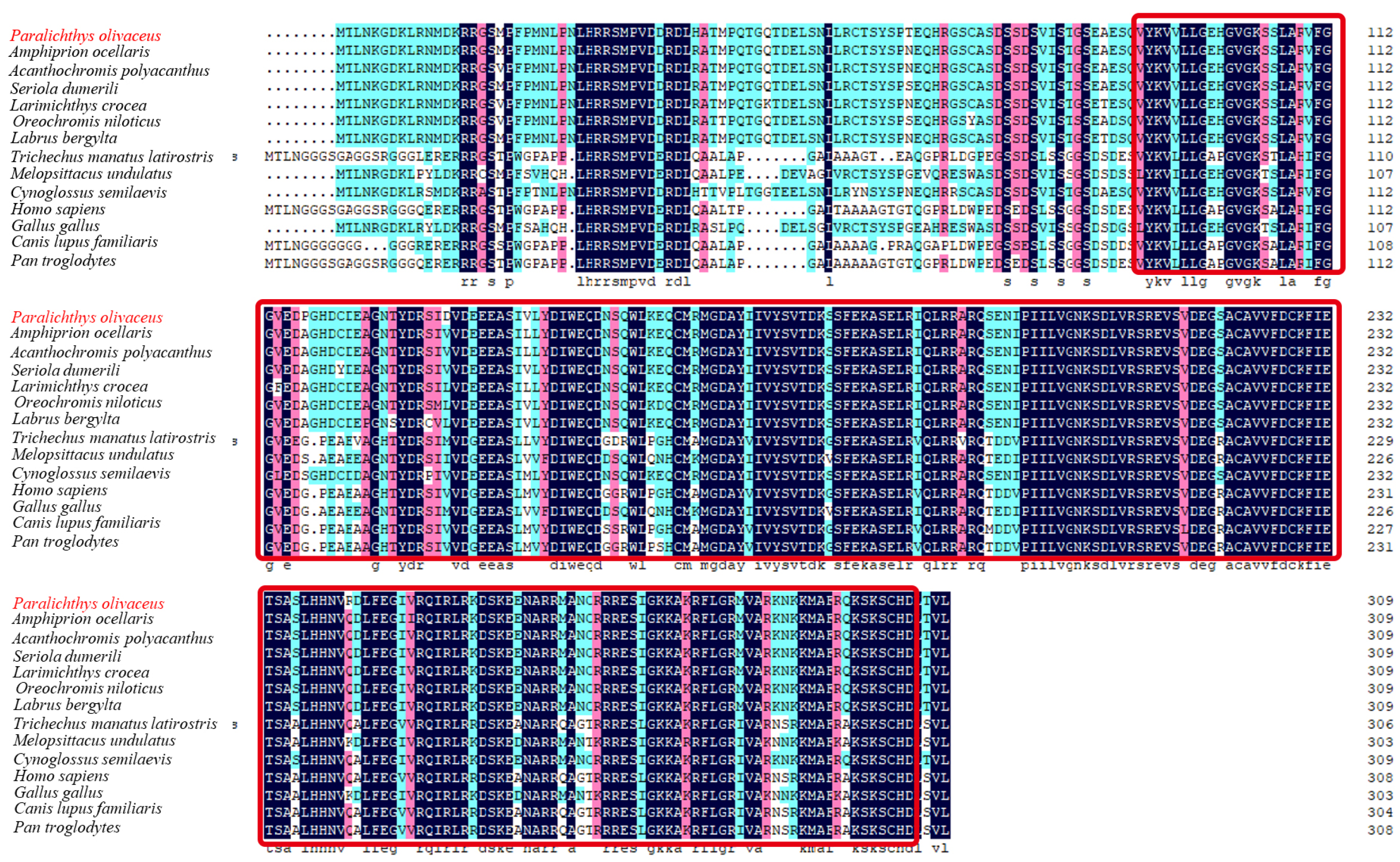
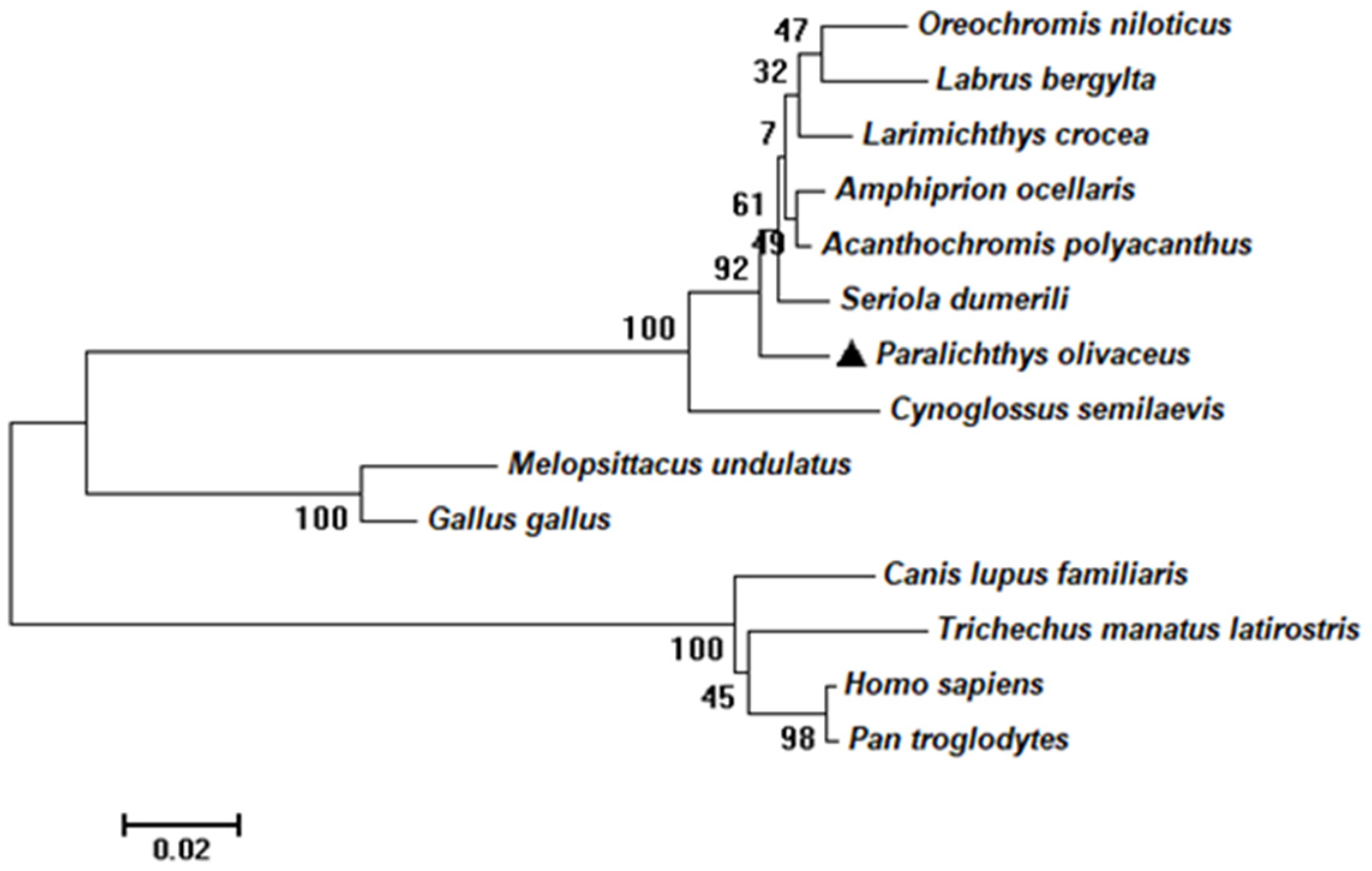
2.1. Sequence and Characteristics of rrad in P. olivaceus
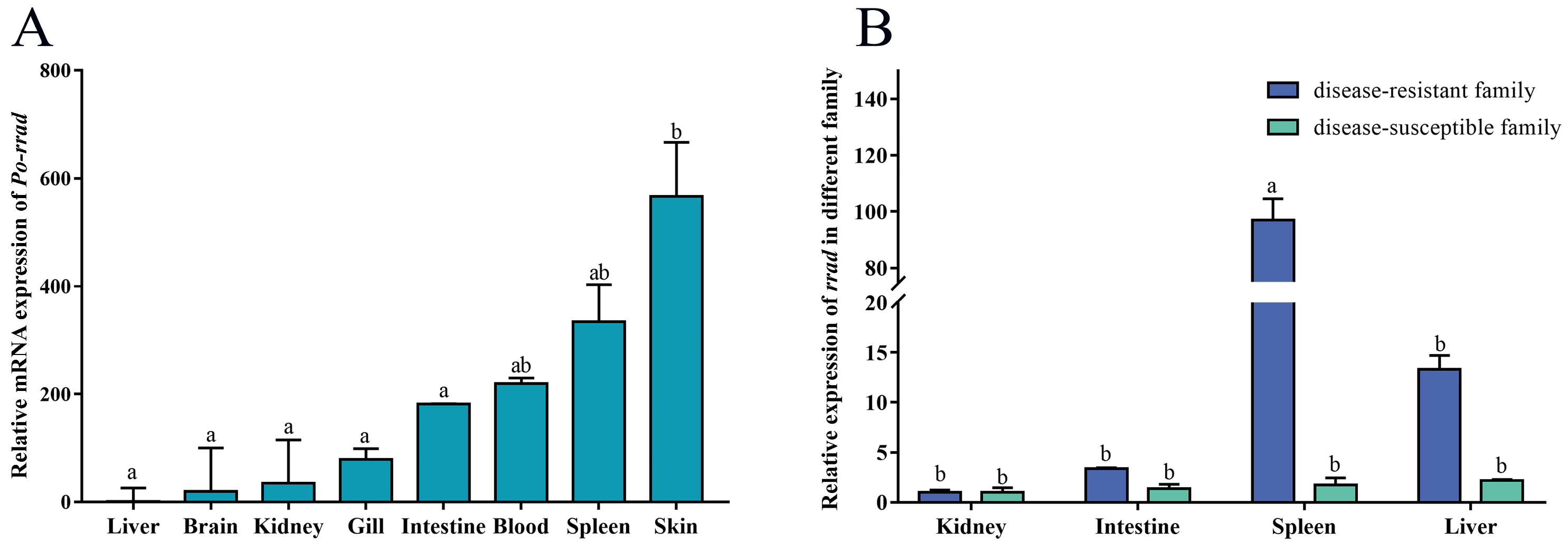
2.2. Tissue Expression Distribution of rrad
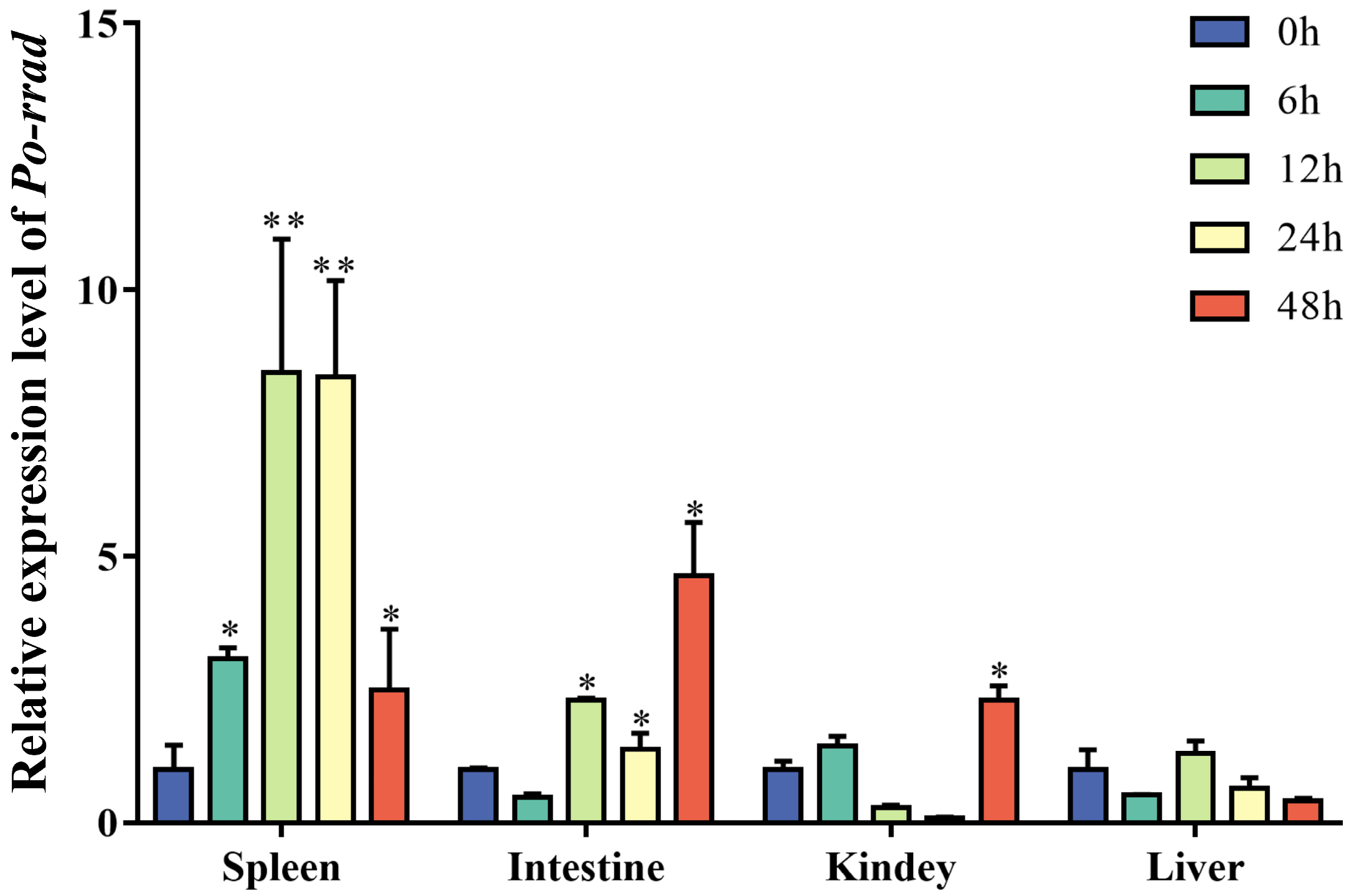
2.3. Expression of rrad in Immune-Related Tissue after E. piscicida Infection
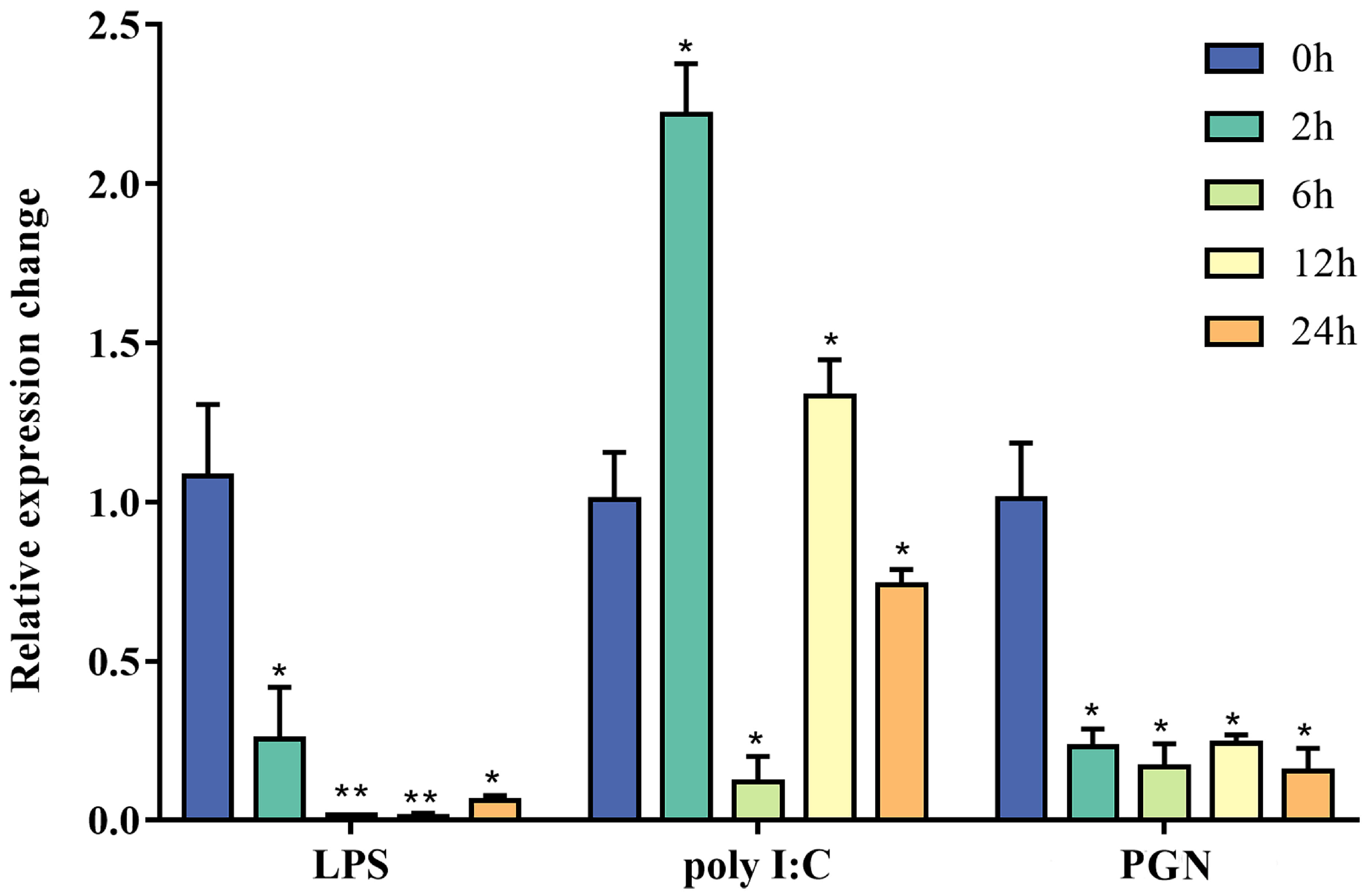
2.4. In Vitro Stimulation with PAMPs
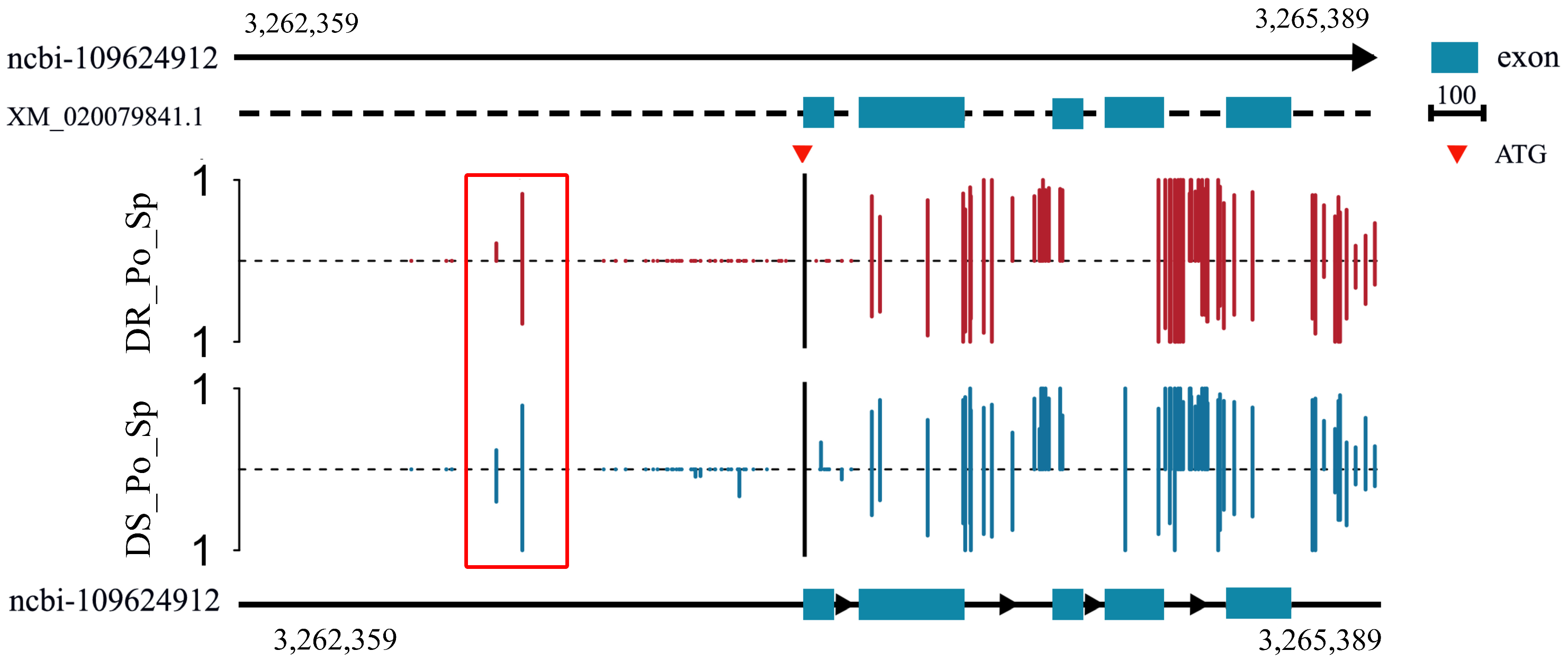
2.5. Methylation Analysis of rrad in the Spleen
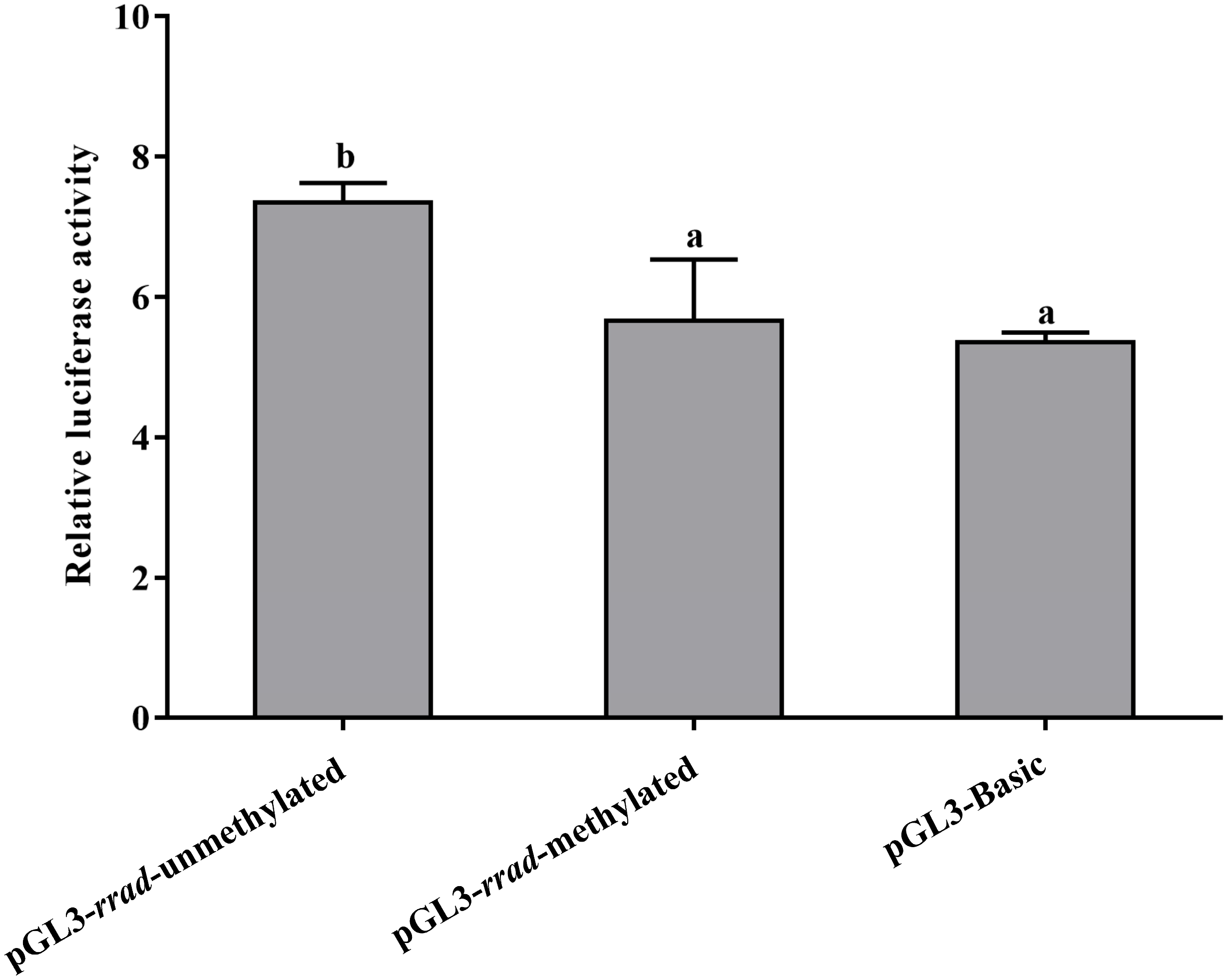
2.6. Functional Characterization of rrad Promoter
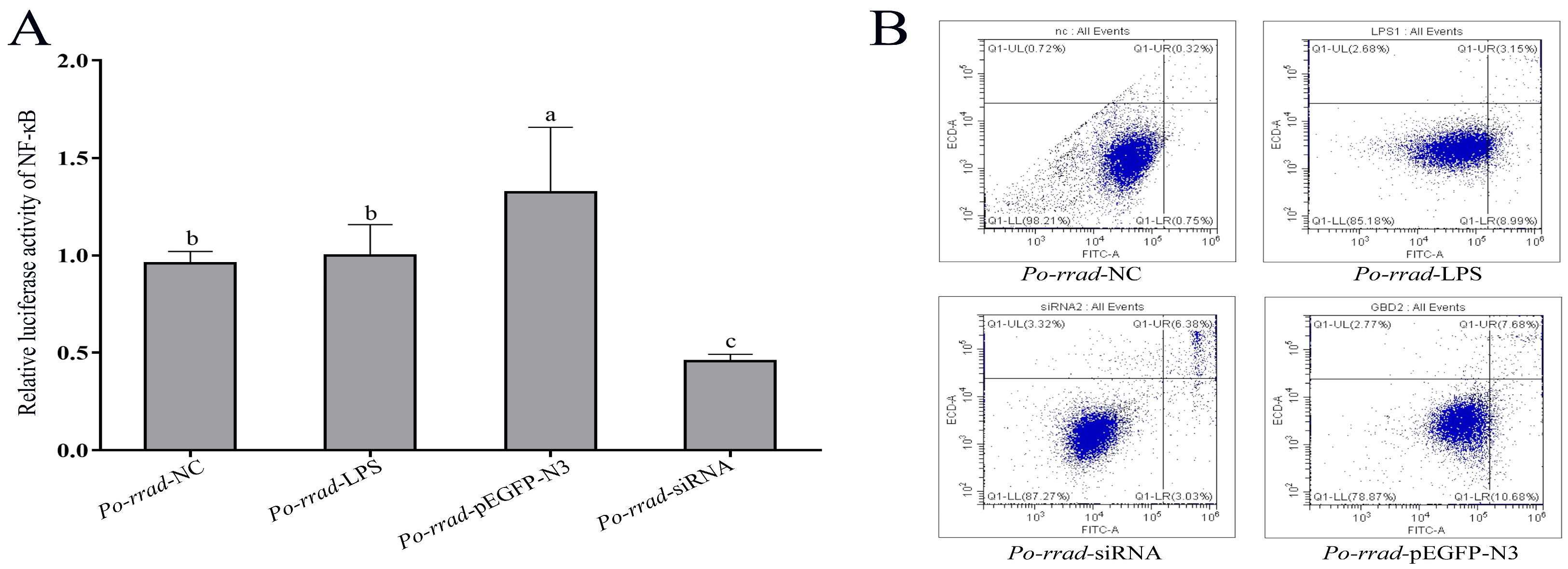
2.7. Luciferase Activity and Flow Cytometry Analyses
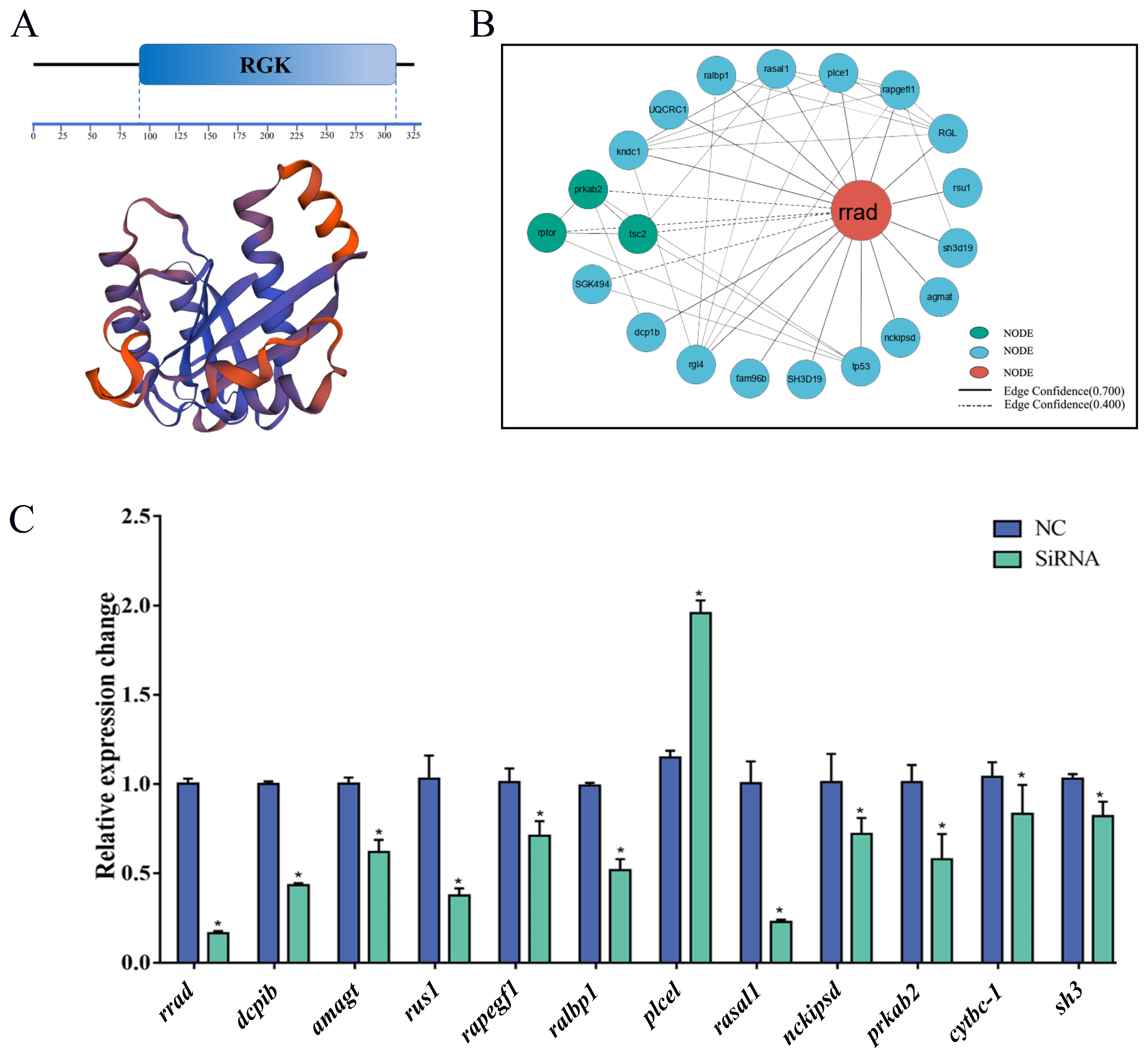
2.8. Protein–Protein Interaction (PPI) Network of rrad
2.9. Effect of the Knockdown of rrad and Other Related Genes via siRNA Transfection
3. Discussion
4. Materials and Methods
4.1. Fish Sample Preparation
4.2. Gene Cloning and cDNA Synthesis
4.3. Bioinformatic Analysis
4.4. Quantitative Real-Time PCR (qPCR) Analysis
4.5. Effect of rrad on the Survival Integrity of Target Bacteria
4.6. DNA Methylation Status of rrad in Different Individuals
4.7. rrad Promoter Activity Analysis
4.8. SiRNA Design and rrad Knockdown in Gill Cell Line of P. olivaceus
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Reynet, C.; Kahn, C.R. Rad: A member of the Ras family overexpressed in muscle of type II diabetic humans. Science 1993, 262, 1441–1444. [Google Scholar] [CrossRef] [PubMed]
- Downward, J. Regulatory mechanisms for ras proteins. Bioessays News Rev. Mol. Cell. Dev. Biol. 1992, 14, 177. [Google Scholar] [CrossRef] [PubMed]
- Tisdale, E.J.; Bourne, J.R.; Khosravifar, R.; Der, C.J.; Balch, W.E. GTP-binding mutants of rab1 and rab2 are potent inhibitors of vesicular transport from the endoplasmic reticulum to the Golgi complex. J. Cell Biol. 1992, 119, 749–761. [Google Scholar] [CrossRef] [PubMed]
- Ridley, A.J.; Hall, A. The small GTP-binding protein rho regulates the assembly of focal adhesions and actin stress fibers in response to growth factors. Cell 1992, 70, 389–399. [Google Scholar] [CrossRef]
- Bos, J.L. The ras gene family and human carcinogenesis. Mutat. Res./Rev. Genet. Toxicol. 1988, 195, 255–271. [Google Scholar] [CrossRef]
- Knobbe, C.B.; Reifenberger, J.; Reifenberger, G. Mutation analysis of the Ras pathway genes NRAS, HRAS, KRAS and BRAF in glioblastomas. Acta Neuropathol. 2004, 108, 467. [Google Scholar] [CrossRef]
- Béguin, P.; Mahalakshmi, R.N.; Nagashima, K.; Cher, D.H.; Takahashi, A.; Yamada, Y.; Seino, Y.; Hunziker, W. 14-3-3 and calmodulin control subcellular distribution of Kir/Gem and its regulation of cell shape and calcium channel activity. J. Cell Sci. 2005, 118, 1923–1934. [Google Scholar] [CrossRef][Green Version]
- Moyers, J.S.; Bilan, P.J.; Zhu, J.; Kahn, C.R. Rad and Rad-related GTPases interact with calmodulin and calmodulin-dependent protein kinase II. J. Biol. Chem. 1997, 272, 11832–11839. [Google Scholar] [CrossRef]
- Zhu, J.; Bilan, P.J.; Moyers, J.S.; Antonetti, D.A.; Kahn, C.R. Rad, a novel Ras-related GTPase, interacts with skeletal muscle β-tropomyosin. J. Biol. Chem. 1996, 271, 768–773. [Google Scholar] [CrossRef]
- Karin, M. Nuclear factor-κB in cancer development and progression. Nature 2006, 441, 431–436. [Google Scholar] [CrossRef]
- Sarkar, F.H.; Li, Y.; Wang, Z.; Kong, D. NF-kappaB signaling pathway and its therapeutic implications in human diseases. Int. Rev. Immunol. 2008, 27, 293–319. [Google Scholar] [CrossRef] [PubMed]
- Yeom, S.-Y.; Nam, D.-H.; Park, C. RRAD promotes EGFR-mediated STAT3 activation and induces temozolomide resistance of malignant glioblastoma. Mol. Cancer Ther. 2014, 13, 3049–3061. [Google Scholar] [CrossRef] [PubMed]
- Fajardo, C.A.; Guedan, S.; Rojas, L.A.; Moreno, R.; Arias-Badia, M.; de Sostoa, J.; June, C.H.; Alemany, R. Oncolytic adenoviral delivery of an EGFR-targeting T-cell engager improves antitumor efficacy. Cancer Res. 2017, 77, 2052–2063. [Google Scholar] [CrossRef] [PubMed]
- Hardbower, D.M.; A Coburn, L.; Asim, M.; Singh, K.; Sierra, J.C.; Barry, D.P.; Gobert, A.P.; Piazuelo, M.B.; Washington, M.K.; Wilson, K.T. EGFR-mediated macrophage activation promotes colitis-associated tumorigenesis. Oncogene 2017, 36, 3807–3819. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Zhang, Y.; Liu, S.; Li, C.; Sun, L.; Bao, L.; Feng, J.; Liu, Z. Analysis of 52 Rab GTPases from channel catfish and their involvement in immune responses after bacterial infections. Dev. Comp. Immunol. 2014, 45, 21–34. [Google Scholar] [CrossRef]
- Zhao, Z.-X.; Cao, D.-C.; Xu, J.; Xu, R.; Li, J.-T.; Zhang, Y.; Xu, P.; Sun, X.-W. Diversification of the duplicated Rab1a genes in a hypoxia-tolerant fish, common carp (Cyprinus carpio). Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 2015, 188, 54–62. [Google Scholar] [CrossRef]
- Han, F.; Wang, X.; Yang, Q.; Cai, M.; Wang, Z.Y. Characterization of a RacGTPase up-regulated in the large yellow croaker Pseudosciaena crocea immunity. Fish Shellfish Immunol. 2011, 30, 501–508. [Google Scholar] [CrossRef]
- Bai, H.; Zhou, T.; Zhao, J.; Chen, B.; Pu, F.; Bai, Y.; Wu, Y.; Chen, L.; Shi, Y.; Ke, Q. Transcriptome analysis reveals the temporal gene expression patterns in skin of large yellow croaker (Larimichthys crocea) in response to Cryptocaryon irritans infection. Fish Shellfish Immunol. 2020, 99, 462–472. [Google Scholar] [CrossRef]
- Hsiao, B.-Y.; Chang, T.K.; Wu, I.T.; Chen, M.Y. Rad GTPase inhibits the NFκB pathway through interacting with RelA/p65 to impede its DNA binding and target gene transactivation. Cell. Signal. 2014, 26, 1437–1444. [Google Scholar] [CrossRef]
- Yamamoto, M.; Takeda, K. Role of nuclear IκB proteins in the regulation of host immune responses. J. Infect. Chemother. 2008, 14, 265–269. [Google Scholar] [CrossRef]
- Zhang, M.; Xiao, Z.Z.; Sun, L. Overexpression of NF-κB inhibitor alpha in Cynoglossus semilaevis impairs pathogen-induced immune response. Dev. Comp. Immunol. 2012, 36, 253–257. [Google Scholar] [CrossRef] [PubMed]
- Bourne, H.R.; Sanders, D.A.; McCormick, F. The GTPase superfamily: Conserved structure and molecular mechanism. Nature 1991, 349, 117–127. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, G.; Mao, F.; Li, X.; Liu, Q.; Chen, L.; Lv, L.; Wang, X.; Wu, J.; Dai, W.; et al. Ras-induced Epigenetic Inactivation of the RRAD (Ras-related Associated with Diabetes) Gene Promotes Glucose Uptake in a Human Ovarian Cancer Model. J. Biol. Chem. 2014, 289, 14225–14238. [Google Scholar] [CrossRef] [PubMed]
- Mo, Y.; Midorikawa, K.; Zhang, Z.; Zhou, X.; Ma, N.; Huang, G.; Hiraku, Y.; Oikawa, S.; Murata, M. Promoter hypermethylation of Ras -related GTPase gene RRAD inactivates a tumor suppressor function in nasopharyngeal carcinoma. Cancer Lett. 2012, 323, 147. [Google Scholar] [CrossRef] [PubMed]
- Tseng, Y.H.; Vicent, D.; Zhu, J.; Niu, Y.; Adeyinka, A.; Moyers, J.S.; Watson, P.H.; Kahn, C.R. Regulation of growth and tumorigenicity of breast cancer cells by the low molecular weight GTPase Rad and nm23. Cancer Res. 2001, 61, 2071. [Google Scholar]
- Suzuki, M.; Shigematsu, H.; Shames, D.S.; Sunaga, N.; Takahashi, T.; Shivapurkar, N.; Iizasa, T.; Minna, J.D.; Fujisawa, T.; Gazdar, A.F. Methylation and Gene Silencing of the Ras-Related GTPase Gene in Lung and Breast Cancers. Ann. Surg. Oncol. 2007, 14, 1397. [Google Scholar] [CrossRef]
- Sova, P.; Feng, Q.; Geiss, G.; Wood, T.; Strauss, R.; Rudolf, V.; Lieber, A.; Kiviat, N. Discovery of novel methylation biomarkers in cervical carcinoma by global demethylation and microarray analysis. Cancer Epidemiol. Biomark. Prev. 2006, 15, 114. [Google Scholar] [CrossRef]
- Hsiao, B.-Y.; Chen, C.-C.; Hsieh, P.-C.; Chang, T.-K.; Yeh, Y.-C.; Wu, Y.-C.; Hsu, H.-S.; Wang, F.-F.; Chou, T.-Y. Rad is a p53 direct transcriptional target that inhibits cell migration and is frequently silenced in lung carcinoma cells. J. Mol. Med. 2011, 89, 481–492. [Google Scholar] [CrossRef]
- Young, T.; Mei, F.; Liu, J.; Bast, R.C., Jr.; Kurosky, A.; Cheng, X. Proteomics analysis of H-RAS-mediated oncogenic transformation in a genetically defined human ovarian cancer model. Oncogene 2011, 24, 6174–6184. [Google Scholar] [CrossRef][Green Version]
- Mei, F.C.; Young, T.W.; Liu, J.; Cheng, X. RAS-mediated epigenetic inactivation of OPCML in oncogenic transformation of human ovarian surface epithelial cells. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 2006, 20, 497. [Google Scholar] [CrossRef]
- Liu, J.; Yang, G.; Thompson-Lanza, J.A.; Glassman, A.; Hayes, K.; Patterson, A.; Marquez, R.T.; Auersperg, N.; Yu, Y.; Hahn, W.C.; et al. A Genetically Defined Model for Human Ovarian Cancer. Cancer Res. 2004, 64, 1655–1663. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.S.; Cho, J.Y.; Choi, H.S. Identification of Vibrio harveyi, Vibrio ichthyoenteri, and Photobacterium damselae isolated from olive flounder Paralichthys olivaceus in Korea by multiplex PCR developed using the rpoB gene. Fish. Sci. 2014, 80, 333–339. [Google Scholar] [CrossRef]
- Park, S.B.; Kwon, K.; Cha, I.S.; Jang, H.B.; Nho, S.W.; Fagutao, F.F.; Kim, Y.K.; Yu, J.E.; Jung, T.S. Development of a multiplex PCR assay to detect Edwardsiella tarda, Streptococcus parauberis, and Streptococcus iniae in olive flounder (Paralichthys olivaceus). J. Vet. Sci. 2014, 15, 163–166. [Google Scholar] [CrossRef] [PubMed]
- Bly, J.E.; Clem, L.W. Temperature and teleost immune functions. Fish Shellfish Immunol. 1992, 2, 159–171. [Google Scholar] [CrossRef]
- Zhe, J.; Xianling, F.; Qianhe, J.; Yulan, C.; Yan, G.; Xiaojing, Z.; Wang, L.; Zhang, Y.; Huang, W.; Fan, X.; et al. Aberrant Methylation of the Ras-related Associated with Diabetes Gene in Human Primary Esophageal Cancer. Anticancer Res. 2013, 33, 5199–5203. [Google Scholar]
- Szołtysek, K.; Janus, P.; Zając, G.; Stokowy, T.; Walaszczyk, A.; Widłak, W.; Wojtaś, B.; Gielniewski, B.; Cockell, S.; Perkins, N.D.; et al. RRAD, IL4I1, CDKN1A, and SERPINE1 genes are potentially co-regulated by NF-κB and p53 transcription factors in cells exposed to high doses of ionizing radiation. BMC Genom. 2018, 19, 813. [Google Scholar] [CrossRef]
- Liu, J.; Zhang, C.; Wu, R.; Lin, M.; Liang, Y.; Liu, J.; Wang, X.; Yang, B.; Feng, Z. RRAD inhibits the Warburg effect through negative regulation of the NF-κB signaling. Oncotarget 2015, 6, 14982–14992. [Google Scholar] [CrossRef][Green Version]
- Sommermann, T.G.; O’Neill, K.; Plas, D.R.; Cahir-McFarland, E. IKKβ and NF-κB transcription govern lymphoma cell survival through AKT-induced plasma membrane trafficking of GLUT1. Cancer Res. 2011, 71, 7291–7300. [Google Scholar] [CrossRef]
- Chen, Y.; Cao, B.; Zheng, W.; Sun, Y.; Xu, T. eIF3k inhibits NF-κB signaling by targeting MyD88 for ATG5-mediated autophagic degradation in teleost fish. J. Biol. Chem. 2022, 298, 101730. [Google Scholar] [CrossRef]
- Takai, Y.; Sasaki, T.; Matozaki, T. Small GTP-binding proteins. Physiol. Rev. 2001, 81, 153–208. [Google Scholar] [CrossRef]
- Bilan, P.J.; Moyers, J.S.; Kahn, C.R. The ras-related protein rad associates with the cytoskeleton in a non-lipid-dependent manner. Exp. Cell Res. 1998, 242, 391–400. [Google Scholar] [CrossRef] [PubMed]
- Ding, Y.; He, F.; Wen, H.; Li, J.; Qian, K.; Chi, M.; Ni, M.; Yin, X.; Bu, Y.; Zhao, Y.; et al. Polymorphism in exons CpG rich regions of the cyp17-II gene affecting its mRNA expression and reproductive endocrine levels in female Japanese flounder (Paralichthys olivaceus). Gen. Comp. Endocrinol. 2012, 179, 107–114. [Google Scholar] [CrossRef] [PubMed]
- Bhalla, M.; Heinzinger, L.R.; Morenikeji, O.B.; Marzullo, B.; Thomas, B.N.; Ghanem, E.N.B. Transcriptome profiling reveals CD73 and age-driven changes in neutrophil responses against Streptococcus pneumoniae. Infect. Immun. 2021, 89, e00258-21. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Li, Y.; Tan, X.; Liu, W.; He, X.; Pan, D.; Li, E.; Xu, L.; Long, L. Friend or Foe: Regulation, Downstream Effectors of RRAD in Cancer. Biomolecules 2023, 13, 477. [Google Scholar] [CrossRef]
- Sangrador-Vegas, A.; Smith, T.; Cairns, M. Cloning and characterization of a homologue of the alpha inhibitor of NF-κB in Rainbow trout (Oncorhynchus mykiss). Vet. Immunol. Immunopathol. 2005, 103, 1–7. [Google Scholar] [CrossRef]
- Wang, Y.; Wei, H.; Wang, X.; Du, L.; Zhang, A.; Zhou, H. Cellular activation, expression analysis and functional characterization of grass carp IκBα: Evidence for its involvement in fish NF-κB signaling pathway. Fish Shellfish Immunol. 2015, 42, 408–412. [Google Scholar] [CrossRef]
- Wang, R.; Huang, S.; Fu, X.; Huang, G.; Yan, X.; Yue, Z.; Chen, S.; Li, W.; Xu, A. The conserved ancient role of chordate PIAS as a multilevel repressor of the NF-κB pathway. Sci. Rep. 2017, 7, 17063. [Google Scholar] [CrossRef]
- Jin, P.; Lv, C.; Peng, S.; Cai, L.; Zhu, J.; Ma, F. Genome-wide organization, evolutionary diversification of the COMMD family genes of amphioxus (Branchiostoma belcheri) with the possible role in innate immunity. Fish Shellfish Immunol. 2018, 77, 31–39. [Google Scholar] [CrossRef]
- Vallabhapurapu, S.; Karin, M. Regulation and function of NF-κB transcription factors in the immune system. Annu. Rev. Immunol. 2009, 27, 693–733. [Google Scholar] [CrossRef]
- Baichwal, V.R.; Baeuerle, P.A. Apoptosis: Activate NF-κB or die? Curr. Biol. 1997, 7, R94–R96. [Google Scholar] [CrossRef]
- Boersma, B.; Jiskoot, W.; Lowe, P.; Bourquin, C. The interleukin-1 cytokine family members: Role in cancer pathogenesis and potential therapeutic applications in cancer immunotherapy. Cytokine Growth Factor Rev. 2021, 62, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Wangkahart, E.; Jumpalueang, S.; Ardprachan, S.; Phudkliang, J.; Sunthamala, P.; Pholchamat, S.; Qi, Z. Molecular Characterization and Expression Analysis of Novel Interleukin-1 Family Member (nIL-1Fm) Gene in Nile Tilapia (Oreochromis niloticus). J. Mar. Sci. Eng. 2022, 10, 1272. [Google Scholar] [CrossRef]
- Cao, Z.; Yang, X.; Li, T.; Liu, Z.; Li, P.; Zhou, Y.; Sun, Y. Molecular characterization and expression analysis of B-cell lymphoma-2 in Trachinotus ovatus and its role in apoptotic process. Front. Immunol. 2023, 14, 1129800. [Google Scholar] [CrossRef]
- Grimaldi, A.; Balestrieri, M.L.; D’Onofrio, N.; Di Domenico, G.; Nocera, C.; Lamberti, M.; Tonini, G.; Zoccoli, A.; Santini, D.; Caraglia, M.; et al. The synergistic effect of everolimus and chloroquine on endothelial cell number reduction is paralleled by increased apoptosis and reduced autophagy occurrence. PLoS ONE 2013, 8, e79658. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Beck, B.; Su, B.; Terhune, J.; Peatman, E. Early mucosal responses in blue catfish (Ictalurus furcatus) skin to Aeromonas hydrophila infection. Fish Shellfish Immunol. 2013, 34, 920–928. [Google Scholar] [CrossRef]
- Chen, H.; Cheng, Z.-Y.; Pan, Y.; Wang, Z.; Liu, Y.; Zhang, J.-Q. RASAL1 influences the proliferation and invasion of gastric cancer cells by regulating the RAS/ERK signaling pathway. Hum. Cell 2014, 27, 103–110. [Google Scholar] [CrossRef]
- Jones, P.A.; Baylin, S.B. The fundamental role of epigenetic events in cancer. Nat. Rev. Genet. 2002, 3, 415–428. [Google Scholar] [CrossRef]
- Wang, L.; Xu, X.; Zhang, Z.; Li, K.; Yang, Y.; Zheng, W.; Sun, H.; Chen, S. Transcriptome analysis and protein-protein interaction in resistant and susceptible families of Japanese flounder (Paralichthys olivaceus) to understand the mechanism against Edwardsiella tarda. Fish Shellfish Immunol. 2022, 123, 123. [Google Scholar] [CrossRef]
- Li, H.; Xu, W.; Zhu, Y.; Zhang, N.; Ma, J.; Sun, A.; Cui, Z.; Gao, F.; Wang, N.; Shao, C.; et al. Characterization and expression pattern of r-spondin1 in Cynoglossus semilaevis. J. Exp. Zool. Part B Mol. Dev. Evol. 2017, 328, 772–780. [Google Scholar] [CrossRef]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef]
- Zhu, Y.; Li, S.; Su, B.; Xue, T.; Cao, M.; Li, C. Genome-wide identification, characterization, and expression of the Toll-like receptors in Japanese flounder (Paralichthys olivaceus). Aquaculture 2021, 545, 737127. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhu, Y.; Yang, X.; Yang, Y.; Yan, X.; Li, C.; Chen, S. Identification and Functional Analysis of Ras-Related Associated with Diabetes Gene (rrad) in Edwardsiella piscicida-Resistant Individuals of Japanese Flounder (Paralichthys olivaceus). Int. J. Mol. Sci. 2024, 25, 10532. https://doi.org/10.3390/ijms251910532
Zhu Y, Yang X, Yang Y, Yan X, Li C, Chen S. Identification and Functional Analysis of Ras-Related Associated with Diabetes Gene (rrad) in Edwardsiella piscicida-Resistant Individuals of Japanese Flounder (Paralichthys olivaceus). International Journal of Molecular Sciences. 2024; 25(19):10532. https://doi.org/10.3390/ijms251910532
Chicago/Turabian StyleZhu, Ying, Xinsheng Yang, Yingming Yang, Xu Yan, Chao Li, and Songlin Chen. 2024. "Identification and Functional Analysis of Ras-Related Associated with Diabetes Gene (rrad) in Edwardsiella piscicida-Resistant Individuals of Japanese Flounder (Paralichthys olivaceus)" International Journal of Molecular Sciences 25, no. 19: 10532. https://doi.org/10.3390/ijms251910532
APA StyleZhu, Y., Yang, X., Yang, Y., Yan, X., Li, C., & Chen, S. (2024). Identification and Functional Analysis of Ras-Related Associated with Diabetes Gene (rrad) in Edwardsiella piscicida-Resistant Individuals of Japanese Flounder (Paralichthys olivaceus). International Journal of Molecular Sciences, 25(19), 10532. https://doi.org/10.3390/ijms251910532





