RNA-Binding Protein-Mediated Alternative Splicing Regulates Abiotic Stress Responses in Plants
Abstract
:1. Introduction
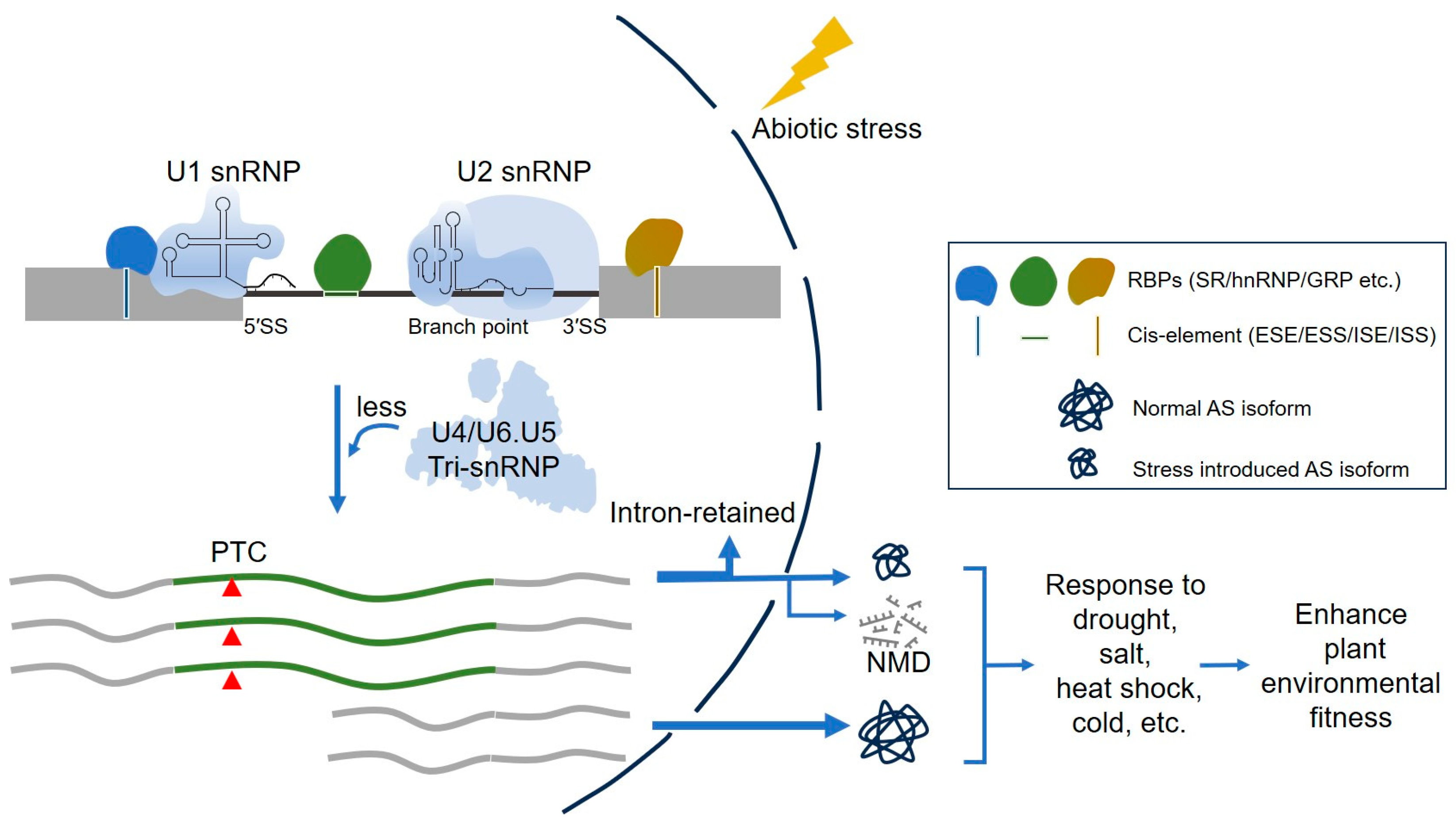
2. Pre-mRNA Splicing and Spliceosome Cycle
3. The Spliceosomal RBPs and Their RNA-Binding Elements in Plants
3.1. Splicing Regulatory Cis-Elements and Trans-Acting Factors in Splicing Regulation
3.2. The Spliceosomal RBPs in Plants
| RBPs | RBDs | Genes | Metazoan Homologues | Genes (H.s.) | Spliceosomal Complexes (H.s.) | References |
|---|---|---|---|---|---|---|
| SR proteins | ||||||
| SC35 | RRM_1 | AT5G64200 | SRSF2 | NP_003007 | A, B, B*, C | [33,75,76,77] |
| SR33/SCL33 | RRM_1 | AT1G55310 | SRSF10 | NP_006616 | B, B*, C | [75,76] |
| SCL30a | RRM_1 | AT3G13570 | ||||
| SCL30 | RRM_1 | AT3G55460 | ||||
| SCL28 | RRM_1 | AT5G18810 | ||||
| SR34 | RRM_1 | AT1G02840 | SRSF1/ASF/SF2 | NP_008855 | A, B, B*, C | [33,75,76,77] |
| SR34b | RRM_1 | AT4G02430 | ||||
| SR30 | RRM_1 | AT1G09140 | ||||
| SR34a | RRM_1 | AT3G49430 | ||||
| RSZp22/SRZ22 | RRM_1, zf-CCHC | AT4G31580 | SRSF7 | NP_001026854 | A, B (U1), B*, C | [33,75,76,77,78] |
| RSZp22a | RRM_1, zf-CCHC | AT2G24590 | ||||
| RSzp21/SRZ21 | RRM_1, zf-CCHC | AT1G23860 | ||||
| Tra/SFRS1 | RRM_1 | AT1G07350 | TRA2B | NP_004584 | A, B, B*, C | [33,75,76,77] |
| - | RRM_1 | AT4G35785 | TRA2A | NP_037425 | A, B (U1), C | [33,77,78] |
| SR-related proteins | ||||||
| SRRM1L | PWI | AT2G29210 | SRRM1 | NP_005830 | A, B (U1), B*, C | [33,75,76,77,78] |
| hnRNP family | ||||||
| RNPA/B_1 | RRM_1 | AT4G14300 | HNRNPA3 | NP_919223 | A, B, C | [33,75,76,77] |
| RNPA/B_2 | RRM_1 | AT2G33410 | ||||
| RNPA/B_3 | RRM_1 | AT5G55550 | ||||
| RNPA/B_4 | RRM_1 | AT4G26650 | ||||
| RNPA/B_5 | RRM_1 | AT5G47620 | ||||
| MSIL4 | RRM_1 | AT3G07810 | HNRNPA2B1 | NP_112533 | A, B (U1), C | [33,75,76,77,78] |
| RNPA/B_7 | RRM_1 | AT1G58470 | ||||
| RNPA/B_8a | RRM_1 | AT5G40490 | ||||
| RBGD3 | RRM_1 | AT3G13224 | ||||
| RNPA/B_8b | RRM_1 | AT1G17640 | ||||
| RNP_N1 | RRM_1 | AT3G13224 | ||||
| - | RRM_1 | AT5G46840 | ||||
| UBA2a | RRM_1, Nup35_RRM | AT3G56860 | ||||
| UBA2b | RRM_1 | AT2G41060 | ||||
| UBA2c | RRM_1 | AT3G15010 | ||||
| RBGD1 | RRM_1 | AT1G17640 | HNRPDL/DAP40a | NP_112740 | just identified in spliceosome | [79] |
| PEP/PEPPER | KH_1, KH_2 | AT4G26000 | HNRNPF | NP_004957 | B (U1), C | [75,76,77,78] |
| hnRNP-H/RNPH/F_1 | RRM_1 | AT5G66010 | ||||
| hnRNP-H/RNPH/F_2 | RRM_1 | AT3G20890 | ||||
| hnRNP-G1 | RRM_1, zf-CCHC | AT5G04280 | RBMX/hnRNP G | NP_002130 | A, B (U1), B*, C | [33,75,76,77,78] |
| RZ-1A/hnRNP-G2 | RRM_1, zf-CCHC | AT3G26420 | ||||
| RZ-1B/hnRNP-G3 | RRM_1, zf-CCHC | AT1G60650 | ||||
| PTB3 | RRM_1, RRM_5 | AT1G43190 | PTBP1/hnRNP I | NP_787041 | just identified in spliceosome | [79] |
| PTB1 | RRM_1, RRM_5 | AT3G01150 | ||||
| PTB2 | RRM_1, RRM_5 | AT5G53180 | ||||
| LIF2/hnRNP-R1 | RRM_1 | AT4G00830 | HNRNPR | NP_005817 | A, B (U1), B*, C | [33,75,76,77,78] |
| GRP7 | RRM_1 | AT2G21660 | CIRBP | NP_001271 | just identified in spliceosome | [79] |
| GRP8 | RRM_1 | AT4G39260 | ||||
3.3. Plant RBP Binding Elements in Pre-mRNA
| RBPs | Protein Family | RNA Elements | Target Determination Methods | PPI-Spliceosome Components | AS Events | References |
|---|---|---|---|---|---|---|
| SCL33 | SR protein | GAAG repeat | splicing reporter approach; EMSA | U1-70K | A3SS | [80,85] |
| SC35 | SR protein | AGAAGA | matriX motifs (XX motif) method | U1-70K | IR | [85,86] |
| SR45 | SR protein | GGNGG | RIP-seq | U1-70K; U2AF35b | IR; A5SS; A3SS | [74,81,85,87] |
| GAA/GA repeat (5′SS) CUU/UC repeat (3′SS) | ||||||
| SRRM1L | SR-related protein | CU-rich | RIP-seq; RNA-seq; EMSA | U1-70K | IR | [70] |
| PTB1/2 | hnRNP-like | pyrimidine motifs (UC-rich) | splicing reporter analysis; EMSA | U2AF65 (Speculation based on conservative mechanisms) | ES | [26,88] |
| GRP7 | hnRNP-like | GUUUC (U-rich) | iCLIP | U1-70K | A5SS; A3SS; IR | [72,83,84,89] |
| RZ-1C | glycine-rich protein | U-rich; GA-rich | eCLIP/SELEX | SR proteins | co-transcriptional splicing | [82,90] |
| GRP20 | glycine-rich protein | purine-rich (GA-rich) | mRNA seq; EMSA | PRP18 (U5 snRNP) | ES (for micro- and small exon) | [91] |
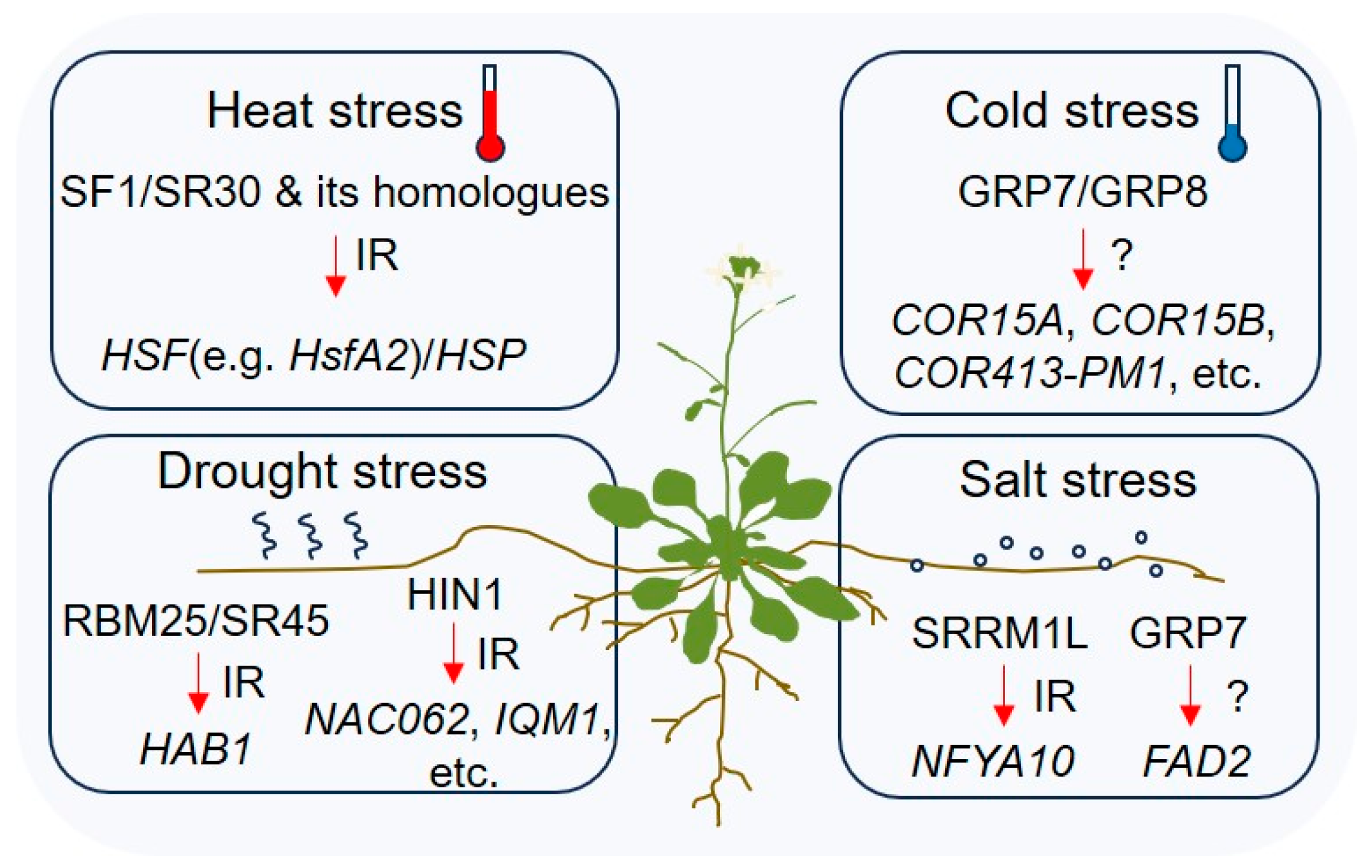
4. RBP-Mediated Alternative Splicing in the Regulation of Plant Abiotic Stresses
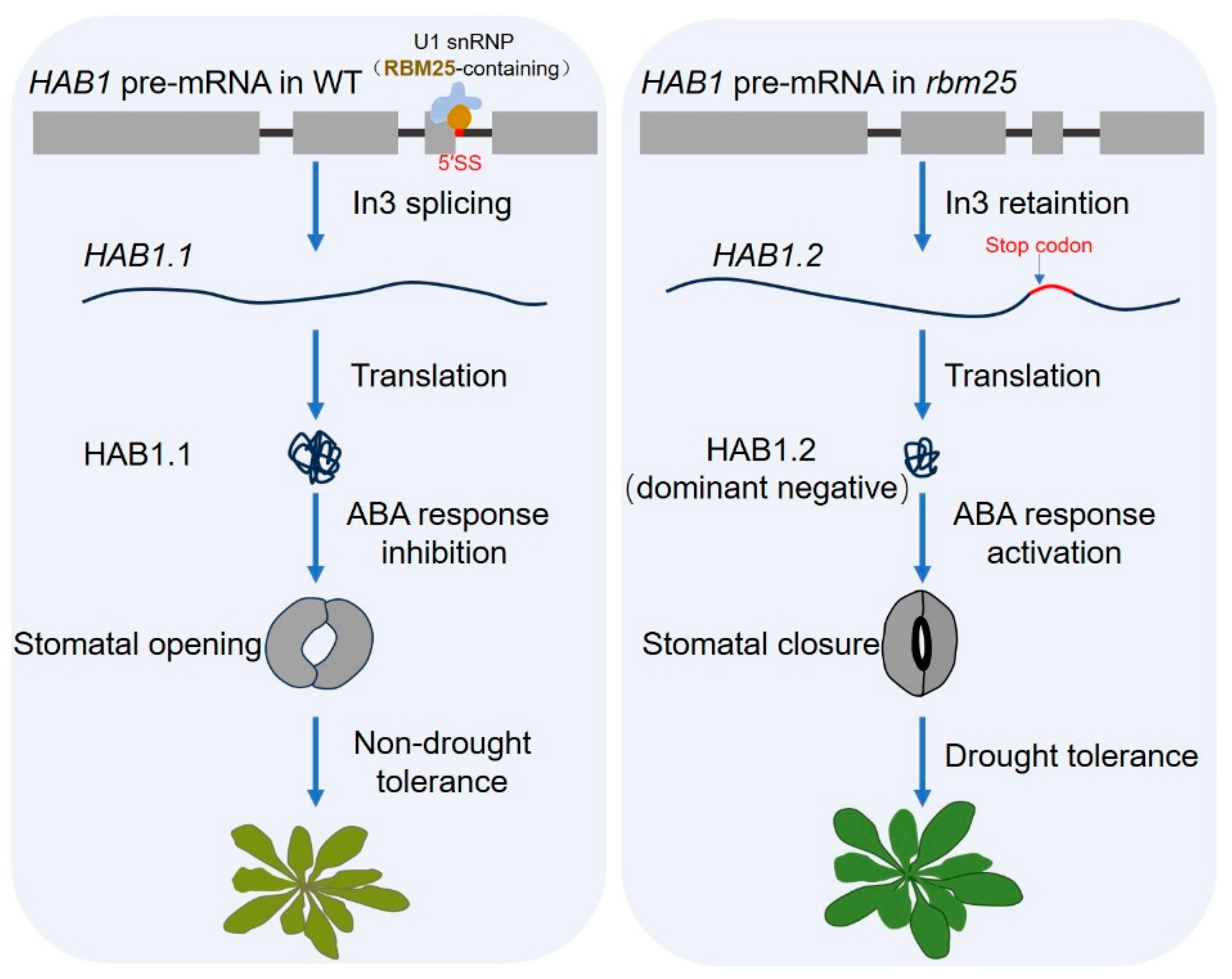
4.1. Drought Stress Response
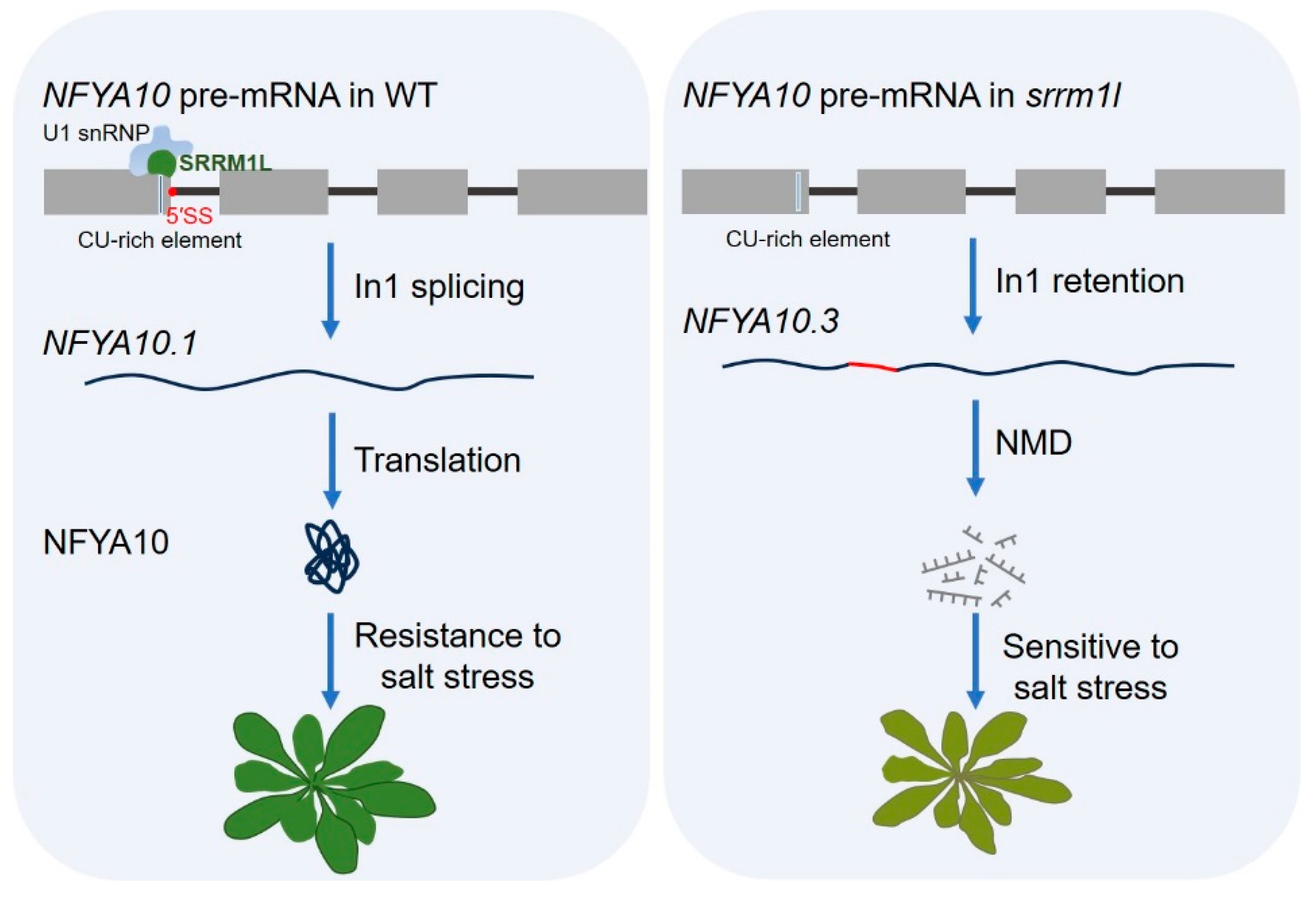
4.2. Salt Stress Response
4.3. Heat Shock Response
4.4. Cold Stress Response
5. Conclusions and Prospections
Author Contributions
Funding
Conflicts of Interest
References
- Shang, X.; Cao, Y.; Ma, L. Alternative splicing in plant genes: A means of regulating the environmental fitness of plants. Int. J. Mol. Sci. 2017, 18, 432. [Google Scholar] [CrossRef] [PubMed]
- Laloum, T.; Martín, G.; Duque, P. Alternative splicing control of abiotic stress responses. Trends Plant Sci. 2018, 23, 140–150. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Jia, Z.; Pu, Q.; Tian, Y.; Zhu, F.; Liu, Y. ABA mediates plant development and abiotic stress via alternative splicing. Int. J. Mol. Sci. 2022, 23, 3796. [Google Scholar] [CrossRef]
- Pomeranz Krummel, D.A.; Oubridge, C.; Leung, A.K.; Li, J.; Nagai, K. Crystal structure of human spliceosomal U1 snRNP at 5.5 Å resolution. Nature 2009, 458, 475–480. [Google Scholar] [CrossRef] [PubMed]
- Valcarcel, J.; Gaur, R.K.; Singh, R.; Green, M.R. Interaction of U2AF65 RS region with pre-mRNA branch point and promotion of base pairing with U2 snRNA. Science 1994, 273, 1706–1709. [Google Scholar] [CrossRef] [PubMed]
- Gozani, O.; Potashkin, J.; Reed, R. A potential role for U2AF-SAP155 interactions in recruiting U2 snRNP to the branch site. Mol. Cell Biol. 1998, 18, 4752–4760. [Google Scholar] [CrossRef] [PubMed]
- Wan, R.; Bai, R.; Zhan, X.; Shi, Y. How is precursor messenger RNA spliced by the spliceosome? Annu. Rev. Biochem. 2020, 89, 333–358. [Google Scholar] [CrossRef]
- Bradley, R.K.; Anczuków, O. RNA splicing dysregulation and the hallmarks of cancer. Nat. Rev. Cancer 2023, 23, 135–155. [Google Scholar] [CrossRef] [PubMed]
- Dong, C.; He, F.; Berkowitz, O.; Liu, J.; Cao, P.; Tang, M.; Shi, H.; Wang, W.; Li, Q.; Shen, Z.; et al. Alternative splicing plays a critical role in maintaining mineral nutrient homeostasis in rice (Oryza sativa). Plant Cell 2018, 30, 2267–2285. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Han, Y.; Liu, H.; Wang, X.; Sun, J.; Zhao, B.; Li, W.; Tian, J.; Liang, Y.; Yan, J.; et al. Genome-wide association analyses reveal the importance of alternative splicing in diversifying gene function and regulating phenotypic variation in maize. Plant Cell 2018, 30, 1404–1423. [Google Scholar] [CrossRef] [PubMed]
- Martín, G.; Márquez, Y.; Mantica, F.; Duque, P.; Irimia, M. Alternative splicing landscapes in Arabidopsis thaliana across tissues and stress conditions highlight major functional differences with animals. Genome Biol. 2021, 22, 35. [Google Scholar] [CrossRef]
- Feng, J.; Li, J.; Gao, Z.; Lu, Y.; Yu, J.; Zheng, Q.; Yan, S.; Zhang, W.; He, H.; Ma, L.; et al. SKIP confers osmotic tolerance during salt stress by controlling alternative gene splicing in Arabidopsis. Mol. Plant 2015, 8, 1038–1052. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Xia, C.; Feng, J.; Yang, D.; Wu, F.; Cao, Y.; Li, L.; Ma, L. The SNW domain of SKIP is required for its integration into the spliceosome and its interaction with the Paf1 complex in Arabidopsis. Mol. Plant 2016, 9, 1040–1050. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Yang, J.; Shang, X.; Lv, W.; Xia, C.; Wang, C.; Feng, J.; Cao, Y.; He, H.; Ma, L. SKIP regulates environmental fitness and floral transition by forming two distinct complexes in Arabidopsis. New Phytol. 2019, 224, 321–335. [Google Scholar] [CrossRef] [PubMed]
- Kim, E.; Magen, A.; Ast, G. Different levels of alternative splicing among eukaryotes. Nucleic Acids Res. 2007, 35, 125–131. [Google Scholar] [CrossRef] [PubMed]
- Jia, J.; Long, Y.; Zhang, H.; Li, Z.; Liu, Z.; Zhao, Y.; Lu, D.; Jin, X.; Deng, X.; Xia, R.; et al. Post-transcriptional splicing of nascent RNA contributes to widespread intron retention in plants. Nat. Plants 2020, 6, 780–788. [Google Scholar] [CrossRef]
- Syed, N.H.; Kalyna, M.; Marquez, Y.; Barta, A.; Brown, J.W.S. Alternative splicing in plants—Coming of age. Trends Plant Sci. 2012, 17, 616–623. [Google Scholar] [CrossRef] [PubMed]
- Boothby, T.C.; Zipper, R.S.; van der Weele, C.M.; Wolniak, S.M. Removal of retained introns regulates translation in the rapidly developing gametophyte of Marsilea vestita. Dev. Cell 2013, 24, 517–529. [Google Scholar] [CrossRef] [PubMed]
- Ule, J.; Blencowe, B.J. Alternative splicing regulatory networks: Functions, mechanisms, and evolution. Mol. Cell 2019, 76, 329–345. [Google Scholar] [CrossRef] [PubMed]
- Tao, Y.; Zhang, Q.; Wang, H.; Yang, X.; Mu, H. Alternative splicing and related RNA binding proteins in human health and disease. Signal Transduct. Target. Ther. 2024, 9, 26. [Google Scholar] [CrossRef] [PubMed]
- Lunde, B.M.; Moore, C.; Varani, G. RNA-binding proteins: Modular design for efficient function. Nat. Rev. Mol. Cell Biol. 2007, 8, 479–490. [Google Scholar] [CrossRef]
- Howard, J.M.; Sanford, J.R. The RNAissance family: SR proteins as multifaceted regulators of gene expression. Wiley Interdiscip. Rev. RNA 2015, 6, 93–110. [Google Scholar] [CrossRef]
- Geuens, T.; Bouhy, D.; Timmerman, V. The hnRNP family: Insights into their role in health and disease. Hum. Genet. 2016, 135, 851–867. [Google Scholar] [CrossRef] [PubMed]
- He, S.; Valkov, E.; Cheloufi, S.; Murn, J. The nexus between RNA-binding proteins and their effectors. Nat. Rev. Genet. 2023, 24, 276–294. [Google Scholar] [CrossRef]
- Lorković, Z.J.; Barta, A. Genome analysis: RNA recognition motif (RRM) and K homology (KH) domain RNA-binding proteins from the flowering plant Arabidopsis thaliana. Nucleic Acids Res. 2002, 30, 623–635. [Google Scholar] [CrossRef] [PubMed]
- Burgardt, R.; Lambert, D.; Heuwieser, C.; Sack, M.; Wagner, G.; Weinberg, Z.; Wachter, A. Positioning of pyrimidine motifs around cassette exons defines their PTB-dependent splicing in Arabidopsis. Plant J. 2024, 118, 2202–2218. [Google Scholar] [CrossRef] [PubMed]
- Köster, T.; Meyer, K. Plant ribonomics: Proteins in search of RNA partners. Trends Plant Sci. 2018, 23, 352–365. [Google Scholar] [CrossRef]
- Xue, R.; Mo, R.; Cui, D.; Cheng, W.; Wang, H.; Qin, J.; Liu, Z. Alternative splicing in the regulatory circuit of plant temperature response. Int. J. Mol. Sci. 2023, 24, 3878. [Google Scholar] [CrossRef] [PubMed]
- Wahl, M.C.; Will, C.L.; Lührmann, R. The spliceosome: Design principles of a dynamic RNP machine. Cell 2009, 136, 701–718. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, M.E.; Charenton, C.; Nagai, K. RNA splicing by the spliceosome. Annu. Rev. Biochem. 2020, 89, 359–388. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, S.H.; Silva, J.; Burstein, D.; Pupko, T.; Eyras, E.; Ast, G. Large-scale comparative analysis of splicing signals and their corresponding splicing factors in eukaryotes. Genome Res. 2008, 18, 88–103. [Google Scholar] [CrossRef]
- Shi, Y. Mechanistic insights into precursor messenger RNA splicing by the spliceosome. Nat. Rev. Mol. Cell Biol. 2017, 18, 655–670. [Google Scholar] [CrossRef] [PubMed]
- Behzadnia, N.; Goals, M.M.; Hartmuth, K.; Sander, B.; Kastner, B.; Deckert, J.; Dube, P.; Will, C.L.; Urlaub, H.; Stark, H.; et al. Composition and three-dimensional EM structure of double affinity-purified, human prespliceosomal A complexes. EMBO J. 2007, 26, 1737–1748. [Google Scholar] [CrossRef]
- Zhang, X.; Zhan, X.; Bian, T.; Yang, F.; Li, P.; Lu, Y.; Xing, Z.; Fan, R.; Zhang, Q.C.; Shi, Y. Structural insights into branch site proofreading by human spliceosome. Nat. Struct. Mol. Biol. 2024, 31, 835–845. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.H.; Galej, W.P.; Bai, X.C.; Savva, C.G.; Newman, A.J.; Scheres, S.H.; Nagai, K. The architecture of the spliceosomal U4/U6.U5 tri-snRNP. Nature 2015, 523, 47–52. [Google Scholar] [CrossRef]
- Wan, R.; Yan, C.; Bai, R.; Wang, L.; Huang, M.; Wong, C.C.; Shi, Y. The 3.8 Å structure of the U4/U6.U5 tri-snRNP: Insights into spliceosome assembly and catalysis. Science 2016, 351, 466–475. [Google Scholar] [CrossRef]
- Agafonov, D.E.; Kastner, B.; Dybkov, O.; Hofele, R.V.; Liu, W.T.; Urlaub, H.; Lührmann, R.; Stark, H. Molecular architecture of the human U4/U6.U5 tri-snRNP. Science 2016, 351, 1416–1420. [Google Scholar] [CrossRef] [PubMed]
- Yan, C.; Wan, R.; Bai, R.; Huang, G.; Shi, Y. Structure of a yeast activated spliceosome at 3.5 Å resolution. Science 2016, 353, 904–911. [Google Scholar] [CrossRef] [PubMed]
- Townsend, C.; Leelaram, M.N.; Agafonov, D.E.; Dybkov, O.; Will, C.L.; Bertram, K.; Urlaub, H.; Kastner, B.; Stark, H.; Lührmann, R. Mechanism of protein-guided folding of the active site U2/U6 RNA during spliceosome activation. Science 2020, 370, eabc3753. [Google Scholar] [CrossRef] [PubMed]
- Bai, R.; Wan, R.; Yan, C.; Jia, Q.; Lei, J.; Shi, Y. Mechanism of spliceosome remodeling by the ATPase/helicase Prp2 and its coactivator Spp2. Science 2020, 371, eabe8863. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Bian, T.; Zhan, X.; Chen, Z.; Xing, Z.; Larsen, N.A.; Zhang, X.; Shi, Y. Mechanisms of the RNA helicases DDX42 and DDX46 in human U2 snRNP assembly. Nat. Commun. 2023, 14, 897. [Google Scholar] [CrossRef]
- Wan, R.; Yan, C.; Bai, R.; Huang, G.; Shi, Y. Structure of a yeast catalytic step I spliceosome at 3.4 Å resolution. Science 2016, 353, 895–904. [Google Scholar] [CrossRef] [PubMed]
- Yan, C.; Wan, R.; Bai, R.; Huang, G.; Shi, Y. Structure of a yeast step II catalytically activated spliceosome. Science 2017, 355, 149–155. [Google Scholar] [CrossRef] [PubMed]
- Zhan, X.; Lu, Y.; Zhang, X.; Yan, C.; Shi, Y. Mechanism of exon ligation by human spliceosome. Mol. Cell 2022, 82, 2769–2778. [Google Scholar] [CrossRef] [PubMed]
- Fu, X.D.; Ares, M., Jr. Context-dependent control of alternative splicing by RNA-binding proteins. Nat. Rev. Genet. 2014, 15, 689–701. [Google Scholar] [CrossRef]
- Lim, L.P.; Burge, C.B. A computational analysis of sequence features involved in recognition of short introns. Proc. Natl. Acad. Sci. USA 2001, 98, 11193–11198. [Google Scholar] [CrossRef]
- Wang, Z.; Burge, C.B. Splicing regulation: From a parts list of regulatory elements to an integrated splicing code. RNA 2008, 14, 802–813. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Rolish, M.E.; Yeo, G.; Tung, V.; Mawson, M.; Burge, C.B. Systematic identification and analysis of exonic splicing silencers. Cell 2004, 119, 831–845. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Ma, M.; Xiao, X.; Wang, Z. Intronic splicing enhancers, cognate splicing factors and context-dependent regulation rules. Nat. Struct. Mol. Biol. 2012, 19, 1044–1052. [Google Scholar] [CrossRef]
- Wang, Y.; Xiao, X.; Zhang, J.; Choudhury, R.; Robertson, A.; Li, K.; Ma, M.; Burge, C.B.; Wang, Z. A complex network of factors with overlapping affinities represses splicing through intronic elements. Nat. Struct. Mol. Biol. 2013, 20, 36–45. [Google Scholar] [CrossRef]
- Hui, J.; Hung, L.H.; Heiner, M.; Schreiner, S.; Neumüller, N.; Reither, G.; Haas, S.A.; Bindereif, A. Intronic CA-repeat and CA-rich elements: A new class of regulators of mammalian alternative splicing. EMBO J. 2005, 24, 1988–1998. [Google Scholar] [CrossRef] [PubMed]
- Shen, H.; Green, M.R. RS domains contact splicing signals and promote splicing by a common mechanism in yeast through humans. Genes Dev. 2006, 20, 1755–1765. [Google Scholar] [CrossRef] [PubMed]
- Shen, H.; Kan, J.L.; Green, M.R. Arginine-serine-rich domains bound at splicing enhancers contact the branchpoint to promote prespliceosome assembly. Mol. Cell 2004, 13, 367–376. [Google Scholar] [CrossRef]
- Pozzoli, U.; Sironi, M. Silencers regulate both constitutive and alternative splicing events in mammals. Cell Mol. Life Sci. 2005, 62, 1579–1604. [Google Scholar] [CrossRef]
- Izquierdo, J.M.; Majos, N.; Bonnal, S.; Martinez, C.; Castelo, R.; Guigo, R.; Bilbao, D.; Valcarcel, J. Regulation of fas alternative splicing by antagonistic effects of TIA-1 and PTB on exon definition. Mol. Cell 2005, 19, 475–484. [Google Scholar] [CrossRef]
- Sharma, S.; Falick, A.M.; Black, D.L. Polypyrimidine tract binding protein blocks the 5′ splice site-dependent assembly of U2AF and the prespliceosomal E complex. Mol. Cell 2005, 19, 485–496. [Google Scholar] [CrossRef] [PubMed]
- Nasim, F.U.; Hutchison, S.; Cordeau, M.; Chabot, B. High-affinity hnRNP A1 binding sites and duplex-forming inverted repeats have similar effects on 5′ splice site selection in support of a common looping out and repression mechanism. RNA 2002, 8, 1078–1089. [Google Scholar] [CrossRef]
- Koncz, C.; Dejong, F.; Villacorta, N.; Szakonyi, D.; Koncz, Z. The spliceosome-activating complex: Molecular mechanisms underlying the function of a pleiotropic regulator. Front. Plant Sci. 2012, 3, 9. [Google Scholar] [CrossRef]
- Siomi, H.; Matunis, M.J.; Michael, W.M.; Dreyfuss, G. The pre-mRNA binding K protein contains a novel evolutionarily conserved motif. Nucleic Acids Res. 1993, 21, 1193–1198. [Google Scholar] [CrossRef]
- Ryter, J.M.; Schultz, S.C. Molecular basis of double-stranded RNA-protein interactions: Structure of a dsRNA-binding domain complexed with dsRNA. EMBO J. 1998, 17, 7505–7513. [Google Scholar] [CrossRef]
- Corley, M.; Burns, M.C.; Yeo, G.W. How RNA-binding proteins interact with RNA: Molecules and mechanisms. Mol. Cell 2020, 78, 9–29. [Google Scholar] [CrossRef] [PubMed]
- Oubridge, C.; Ito, N.; Evans, P.R.; Teo, C.H.; Nagai, K. Crystal structure at 1.92 Å resolution of the RNA-binding domain of the U1A spliceosomal protein complexed with an RNA hairpin. Nature 1994, 372, 432–438. [Google Scholar] [CrossRef]
- Cléry, A.; Blatter, M.; Allain, F.H. RNA recognition motifs: Boring? Not quite. Curr. Opin. Struct. Biol. 2008, 18, 290–298. [Google Scholar] [CrossRef] [PubMed]
- Auweter, S.D.; Oberstrass, F.C.; Allain, F.H. Sequence-specific binding of single-stranded RNA: Is there a code for recognition? Nucleic Acids Res. 2006, 34, 4943–4959. [Google Scholar] [CrossRef] [PubMed]
- Valverde, R.; Edwards, L.; Regan, L. Structure and function of KH domains. FEBS J. 2008, 275, 2712–2726. [Google Scholar] [CrossRef]
- Font, J.; Mackay, J.P. Beyond DNA: Zinc finger domains as RNA-binding modules. Methods Mol. Biol. 2010, 649, 479–491. [Google Scholar]
- Hall, T.M. Multiple modes of RNA recognition by zinc finger proteins. Curr. Opin. Struc Biol. 2005, 15, 367–373. [Google Scholar] [CrossRef] [PubMed]
- Szymczyna, B.R.; Bowman, J.; McCracken, S.; Pineda-Lucena, A.; Lu, Y.; Cox, B.; Lambermon, M.; Graveley, B.R.; Arrowsmith, C.H.; Blencowe, B.J. Structure and function of the PWI motif: A novel nucleic acid-binding domain that facilitates pre-mRNA processing. Genes Dev. 2003, 17, 461–475. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Ji, H.; Yuan, B.; Wang, S.; Su, C.; Yao, B.; Zhao, H.; Li, X. ABA signalling is fine-tuned by antagonistic HAB1 variants. Nat. Commun. 2015, 6, 8138. [Google Scholar] [CrossRef] [PubMed]
- Sun, Q.; Sun, Y.; Liu, X.; Li, M.; Li, Q.; Xiao, J.; Xu, P.; Zhang, S.; Ding, X. Regulation of plant resistance to salt stress by the SnRK1-dependent splicing factor SRRM1L. New Phytol. 2024, 242, 2093–2114. [Google Scholar] [CrossRef] [PubMed]
- Reddy, A.S.; Shad Ali, G. Plant serine/arginine-rich proteins: Roles in precursor messenger RNA splicing, plant development, and stress responses. Wiley Interdiscip. Rev. RNA 2011, 2, 875–889. [Google Scholar] [CrossRef] [PubMed]
- Streitner, C.; Köster, T.; Simpson, C.G.; Shaw, P.; Danisman, S.; Brown, J.W.; Staiger, D. An hnRNP-like RNA-binding protein affects alternative splicing by in vivo interaction with transcripts in Arabidopsis thaliana. Nucleic Acids Res. 2012, 40, 11240–11255. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Cheng, K.; Li, J.; Deng, Z.; Zhang, C.; Zhu, H. Roles of plant glycine-rich RNA-binding proteins in development and stress responses. Int. J. Mol. Sci. 2021, 22, 5849. [Google Scholar] [CrossRef] [PubMed]
- Day, I.S.; Golovkin, M.; Palusa, S.G.; Link, A.; Ali, G.S.; Thomas, J.; Richardson, D.N.; Reddy, A.S. Interactions of SR45, an SR-like protein, with spliceosomal proteins and an intronic sequence: Insights into regulated splicing. Plant J. 2012, 71, 936–947. [Google Scholar] [CrossRef] [PubMed]
- Deckert, J.; Hartmuth, K.; Boehringer, D.; Behzadnia, N.; Will, C.L.; Kastner, B.; Stark, H.; Urlaub, H.; Lührmann, R. Protein composition and electron microscopy structure of affinity-purified human spliceosomal B complexes isolated under physiological conditions. Mol. Cell Biol. 2006, 26, 5528–5543. [Google Scholar] [CrossRef] [PubMed]
- Bessonov, S.; Anokhina, M.; Krasauskas, A.; Golas, M.M.; Sander, B.; Will, C.L.; Urlaub, H.; Stark, H.; Lührmann, R. Characterization of purified human Bact spliceosomal complexes reveals compositional and morphological changes during spliceosome activation and first step catalysis. RNA 2010, 16, 2384–2403. [Google Scholar] [CrossRef]
- Bessonov, S.; Anokhina, M.; Will, C.L.; Urlaub, H.; Lührmann, R. Isolation of an active step I spliceosome and composition of its RNP core. Nature 2008, 452, 846–850. [Google Scholar] [CrossRef] [PubMed]
- Makarova, O.V.; Makarov, E.M.; Urlaub, H.; Will, C.L.; Gentzel, M.; Wilm, M.; Lührmann, R. A subset of human 35S U5 proteins, including Prp19, function prior to catalytic step 1 of splicing. EMBO J. 2004, 23, 2381–2391. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Licklider, L.J.; Gygi, S.P.; Reed, R. Comprehensive proteomic analysis of the human spliceosome. Nature 2002, 419, 182–185. [Google Scholar] [CrossRef] [PubMed]
- Thomas, J.; Palusa, S.G.; Prasad, K.V.; Ali, G.S.; Surabhi, G.K.; Ben-Hur, A.; Abdel-Ghany, S.E.; Reddy, A.S. Identification of an intronic splicing regulatory element involved in auto-regulation of alternative splicing of SCL33 pre-mRNA. Plant J. 2012, 72, 935–946. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.N.; Shi, Y.; Powers, J.J.; Gowda, N.B.; Zhang, C.; Ibrahim, H.M.M.; Ball, H.B.; Chen, S.L.; Lu, H.; Mount, S.M. Transcriptome analyses reveal SR45 to be a neutral splicing regulator and a suppressor of innate immunity in Arabidopsis thaliana. BMC Genom. 2017, 18, 772. [Google Scholar] [CrossRef] [PubMed]
- Zhu, D.; Mao, F.; Tian, Y.; Lin, X.; Gu, L.; Gu, H.; Qu, L.J.; Wu, Y.; Wu, Z. The features and regulation of co-transcriptional splicing in Arabidopsis. Mol. Plant 2020, 13, 278–294. [Google Scholar] [CrossRef] [PubMed]
- Meyer, K.; Köster, T.; Nolte, C.; Weinholdt, C.; Lewinski, M.; Grosse, I.; Staiger, D. Adaptation of iCLIP to plants determines the binding landscape of the clock-regulated RNA-binding protein AtGRP7. Genome Biol. 2017, 18, 204. [Google Scholar] [CrossRef] [PubMed]
- Lewinski, M.; Steffen, A.; Kachariya, N.; Elgner, M.; Schmal, C.; Messini, N.; Köster, T.; Reichel, M.; Sattler, M.; Zarnack, K.; et al. Arabidopsis thaliana GLYCINE RICH RNA-BINDING PROTEIN 7 interaction with its iCLIP target LHCB1.1 correlates with changes in RNA stability and circadian oscillation. Plant J. 2024, 118, 203–224. [Google Scholar] [CrossRef] [PubMed]
- Golovkin, M.; Reddy, A.S. An SC35-like protein and a novel serine/arginine-rich protein interact with Arabidopsis U1-70K protein. J. Biol. Chem. 1999, 274, 36428–36438. [Google Scholar] [CrossRef] [PubMed]
- Yan, Q.; Xia, X.; Sun, Z.; Fang, Y. Depletion of Arabidopsis SC35 and SC35-like serine/arginine-rich proteins affects the transcription and splicing of a subset of genes. PLoS Genet. 2017, 13, e1006663. [Google Scholar] [CrossRef]
- Xing, D.; Wang, Y.; Hamilton, M.; Ben-Hur, A.; Reddy, A.S. Transcriptome-wide identification of RNA targets of Arabidopsis SERINE/ARGININE-RICH45 uncovers the unexpected roles of this RNA binding protein in RNA processing. Plant Cell 2015, 27, 3294–3308. [Google Scholar] [CrossRef] [PubMed]
- Rühl, C.; Stauffer, E.; Kahles, A.; Wagner, G.; Drechsel, G.; Rätsch, G.; Wachter, A. Polypyrimidine tract binding protein homologs from Arabidopsis are key regulators of alternative splicing with implications in fundamental developmental processes. Plant Cell 2012, 24, 4360–4375. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Yang, T.; Wang, B.; Lin, Q.; Zhu, S.; Li, C.; Ma, Y.; Tang, J.; Xing, J.; Li, X.; et al. RALF1-FERONIA complex affects splicing dynamics to modulate stress responses and growth in plants. Sci. Adv. 2020, 6, 1622. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.; Zhu, D.; Lin, X.; Miao, J.; Gu, L.; Deng, X.; Yang, Q.; Sun, K.; Zhu, D.; Cao, X.; et al. RNA binding proteins RZ-1B and RZ-1C play critical roles in regulating pre-mRNA splicing and gene expression during development in Arabidopsis. Plant Cell 2016, 28, 55–73. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Ma, X.; Hu, Y.; Feng, G.; Guo, C.; Zhang, X.; Ma, H. Regulation of micro- and small-exon retention and other splicing processes by GRP20 for flower development. Nat. Plants 2024, 10, 66–85. [Google Scholar] [CrossRef] [PubMed]
- Cheng, C.; Wang, Z.; Yuan, B.; Li, X. RBM25 mediates abiotic responses in plants. Front. Plant Sci. 2017, 8, 292. [Google Scholar] [CrossRef]
- Carvalho, R.F.; Carvalho, S.D.; Duque, P. The plant-specific SR45 protein negatively regulates glucose and ABA signaling during early seedling development in Arabidopsis. Plant Physiol. 2010, 154, 772–783. [Google Scholar] [CrossRef]
- Chong, G.L.; Foo, M.H.; Lin, W.D.; Wong, M.M.; Verslues, P.E. Highly ABA-Induced 1 (HAI1)-Interacting protein HIN1 and drought acclimation-enhanced splicing efficiency at intron retention sites. Proc. Natl. Acad. Sci. USA 2019, 116, 22376–22385. [Google Scholar] [CrossRef] [PubMed]
- Alhabsi, A.; Butt, H.; Kirschner, G.K.; Blilou, I.; Mahfouz, M.M. SCR106 splicing factor modulates abiotic stress responses by maintaining RNA splicing in rice. J. Exp. Bot. 2024, 75, 802–818. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.S.; Jung, H.J.; Lee, H.J.; Kim, K.A.; Goh, C.H.; Woo, Y.; Oh, S.H.; Han, Y.S.; Kang, H. Glycine-rich RNA-binding protein7 affects abiotic stress responses by regulating stomata opening and closing in Arabidopsis thaliana. Plant J. 2008, 55, 455–466. [Google Scholar] [CrossRef]
- Sugio, A.; Dreos, R.; Aparicio, F.; Maule, A.J. The cytosolic protein response as a subcomponent of the wider heat shock response in Arabidopsis. Plant Cell 2009, 21, 642–665. [Google Scholar] [CrossRef]
- Liu, J.; Sun, N.; Liu, M.; Liu, J.; Du, B.; Wang, X.; Qi, X. An autoregulatory loop controlling Arabidopsis HsfA2 expression: Role of heat shock-induced alternative splicing. Plant Physiol. 2013, 162, 512–521. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Q.; Zhou, Y.; Liu, Z.; Zhang, L.; Song, G.; Guo, Z.; Wang, W.; Qu, X.; Zhu, Y.; Yang, D. An alternatively spliced heat shock transcription factor, OsHSFA2dI, functions in the heat stress-induced unfolded protein response in rice. Plant Biol. 2015, 17, 419–429. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Mesihovic, A.; Jiménez-Gómez, J.M.; Röth, S.; Gebhardt, P.; Bublak, D.; Bovy, A.; Scharf, K.D.; Schleiff, E.; Fragkostefanakis, S. Natural variation in HsfA2 pre-mRNA splicing is associated with changes in thermotolerance during tomato domestication. New Phytol. 2020, 225, 1297–1310. [Google Scholar] [CrossRef] [PubMed]
- Ling, Y.; Mahfouz, M.M.; Zhou, S. Pre-mRNA alternative splicing as a modulator for heat stress response in plants. Trends Plant Sci. 2021, 26, 1153–1170. [Google Scholar] [CrossRef] [PubMed]
- Palusa, S.G.; Reddy, A.S. Differential recruitment of splice variants from SR pre-mRNAs to polysomes during development and in response to stresses. Plant Cell Physiol. 2015, 56, 421–427. [Google Scholar] [CrossRef]
- Yang, C.; Luo, A.; Lu, H.P.; Davis, S.J.; Liu, J.X. Diurnal regulation of alternative splicing associated with thermotolerance in rice by two glycine-rich RNA-binding proteins. Sci. Bull. 2024, 69, 59–71. [Google Scholar] [CrossRef] [PubMed]
- Jang, Y.H.; Park, H.Y.; Lee, K.C.; Thu, M.P.; Kim, S.K.; Suh, M.C.; Kang, H.; Kim, J.K. A homolog of splicing factor SF1 is essential for development and is involved in the alternative splicing of pre-mRNA in Arabidopsis thaliana. Plant J. 2014, 78, 591–603. [Google Scholar] [CrossRef]
- Lee, K.C.; Jang, Y.H.; Kim, S.K.; Park, H.Y.; Thu, M.P.; Lee, J.H.; Kim, J.K. RRM domain of Arabidopsis splicing factor SF1 is important for pre-mRNA splicing of a specific set of genes. Plant Cell Rep. 2017, 36, 1083–1095. [Google Scholar] [CrossRef] [PubMed]
- Lu, S.; Gao, C.; Wang, Y.; He, Y.; Du, J.; Chen, M.; Zhao, H.; Fang, H.; Wang, B.; Cao, Y. Phylogenetic analysis of the plant U2 snRNP auxiliary factor large subunit a gene family in response to developmental cues and environmental stimuli. Front. Plant Sci. 2021, 12, 739671. [Google Scholar] [CrossRef]
- Ling, Y.; Serrano, N.; Gao, G.; Atia, M.; Mokhtar, M.; Woo, Y.H.; Bazin, J.; Veluchamy, A.; Benhamed, M.; Crespi, M.; et al. Thermopriming triggers splicing memory in Arabidopsis. J. Exp. Bot. 2018, 69, 2659–2675. [Google Scholar] [CrossRef]
- Zhong, Y.; Luo, Y.; Sun, J.; Qin, X.; Gan, P.; Zhou, Z.; Qian, Y.; Zhao, R.; Zhao, Z.; Cai, W.; et al. Pan-transcriptomic analysis reveals alternative splicing control of cold tolerance in rice. Plant Cell 2024, 36, 2117–2139. [Google Scholar] [CrossRef]
- Butt, H.; Bazin, J.; Prasad, K.V.S.K.; Awad, N.; Crespi, M.; Reddy, A.S.N.; Mahfouz, M.M. The rice serine/arginine splicing factor RS33 regulates pre-mRNA splicing during abiotic stress responses. Cells 2022, 11, 1796. [Google Scholar] [CrossRef] [PubMed]
- Guo, L.; Wang, S.; Jiao, X.; Ye, X.; Deng, D.; Liu, H.; Li, Y.; Van de Peer, Y.; Wu, W. Convergent and/or parallel evolution of RNA-binding proteins in angiosperms after polyploidization. New Phytol. 2024, 242, 1377–1393. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.H.; Kwak, K.J.; Kim, M.K.; Park, S.J.; Yang, K.Y.; Kang, H. Expression of Arabidopsis glycine-rich RNA-binding protein AtGRP2 or AtGRP7 improves grain yield of rice (Oryza sativa) under drought stress conditions. Plant Sci. 2014, 214, 106–112. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guo, Y.; Shang, X.; Ma, L.; Cao, Y. RNA-Binding Protein-Mediated Alternative Splicing Regulates Abiotic Stress Responses in Plants. Int. J. Mol. Sci. 2024, 25, 10548. https://doi.org/10.3390/ijms251910548
Guo Y, Shang X, Ma L, Cao Y. RNA-Binding Protein-Mediated Alternative Splicing Regulates Abiotic Stress Responses in Plants. International Journal of Molecular Sciences. 2024; 25(19):10548. https://doi.org/10.3390/ijms251910548
Chicago/Turabian StyleGuo, Ying, Xudong Shang, Ligeng Ma, and Ying Cao. 2024. "RNA-Binding Protein-Mediated Alternative Splicing Regulates Abiotic Stress Responses in Plants" International Journal of Molecular Sciences 25, no. 19: 10548. https://doi.org/10.3390/ijms251910548





