Codon Bias of the DDR1 Gene and Transcription Factor EHF in Multiple Species
Abstract
1. Introduction
2. Results
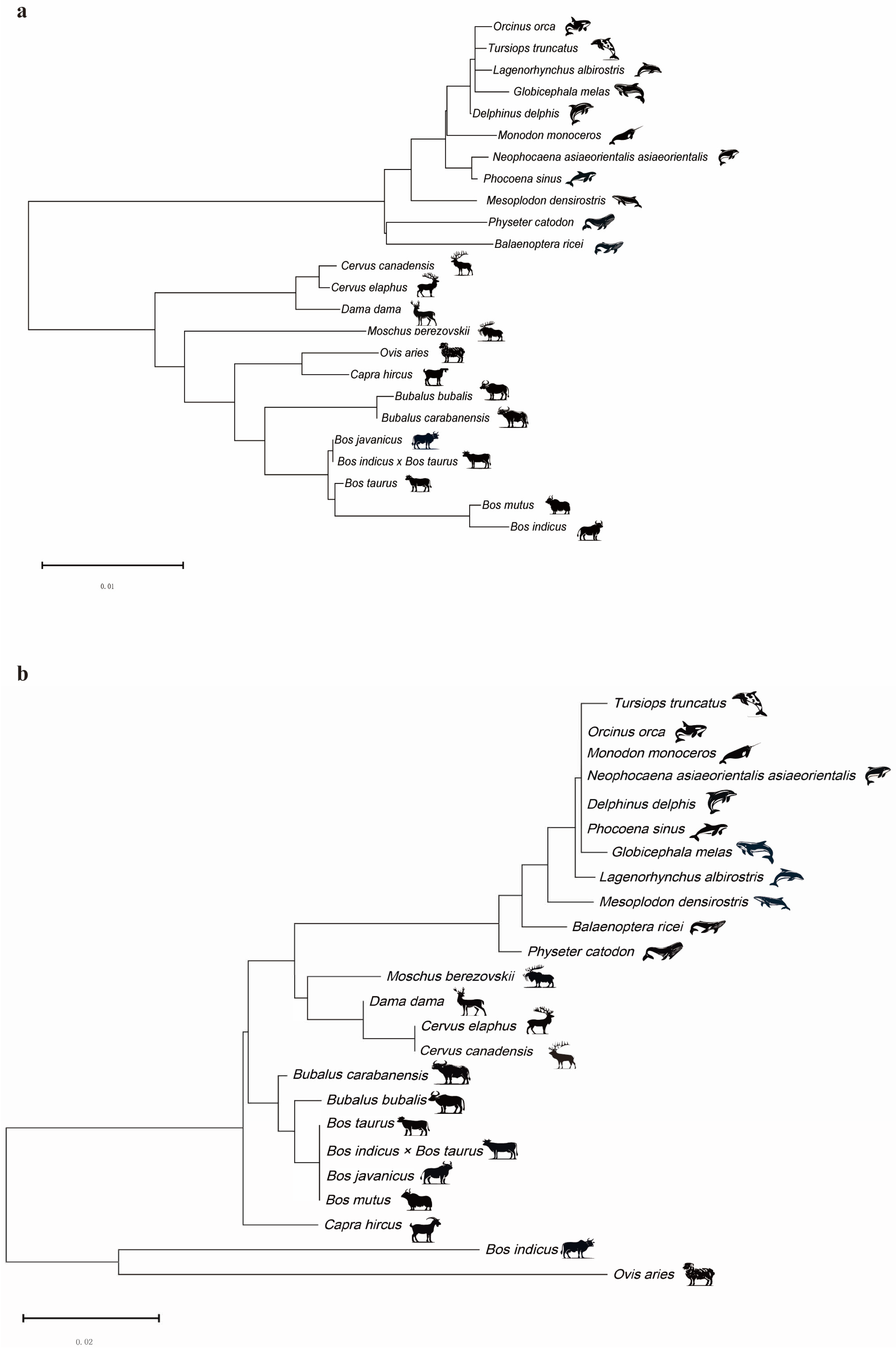
2.1. Construction of DDR1 Gene and EHF Phylogenetic Trees
2.2. Codon Usage Patterns for the DDR1 Gene and EHF
2.3. Relative Synonymous Codon Usage (RSCU) Values Analysis and Determination of Putative Optimal Codons for DDR1 and EHF across 24 Species
2.4. Hierarchical Clustering Analysis of RSCU for DDR1 and EHF
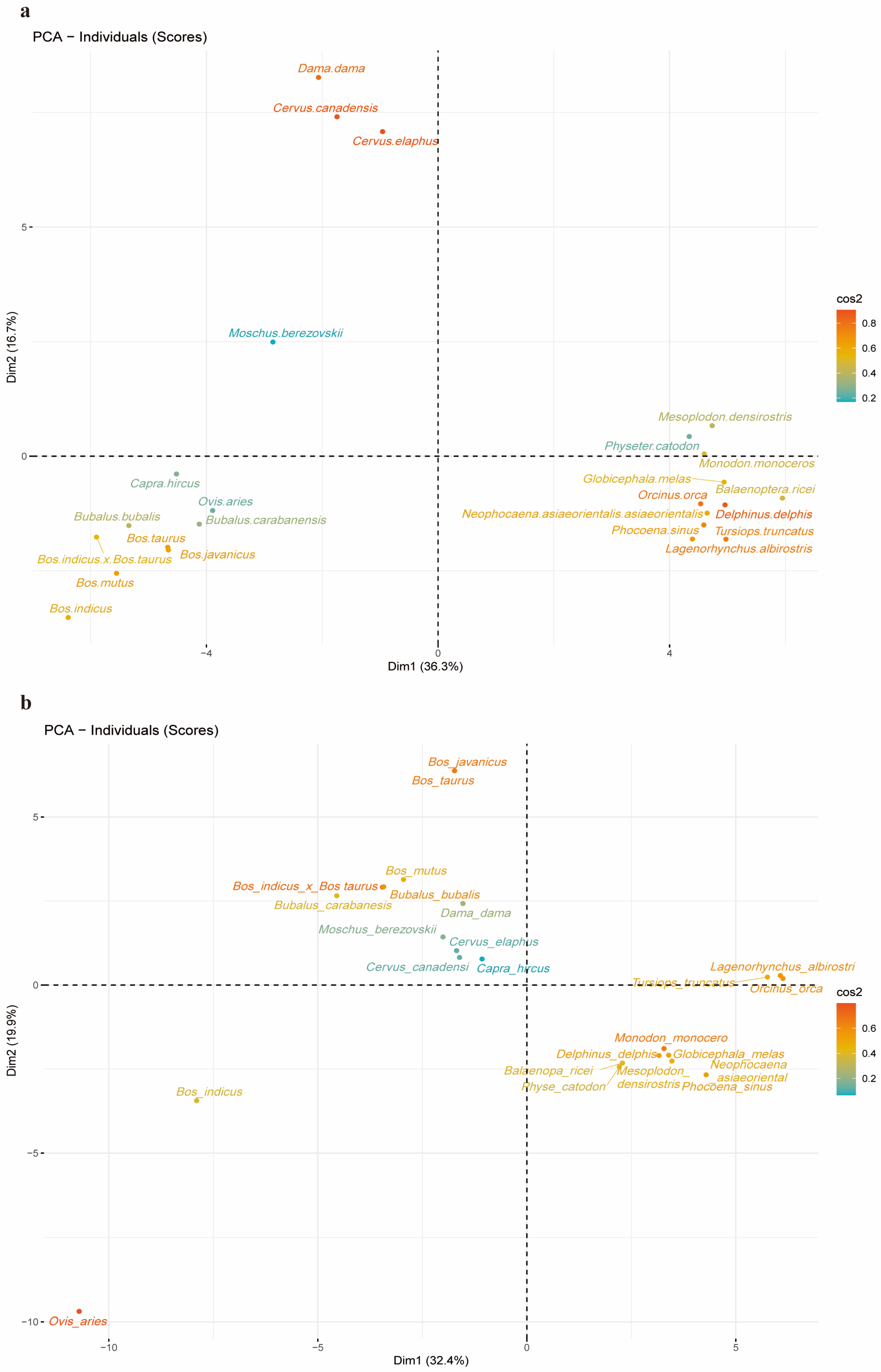
2.5. PCA Analysis of 24 Species and Codons Separately Using RSCU Values
2.6. Correlation Analysis of Codon Usage Preference between DDR1 Gene and EHF
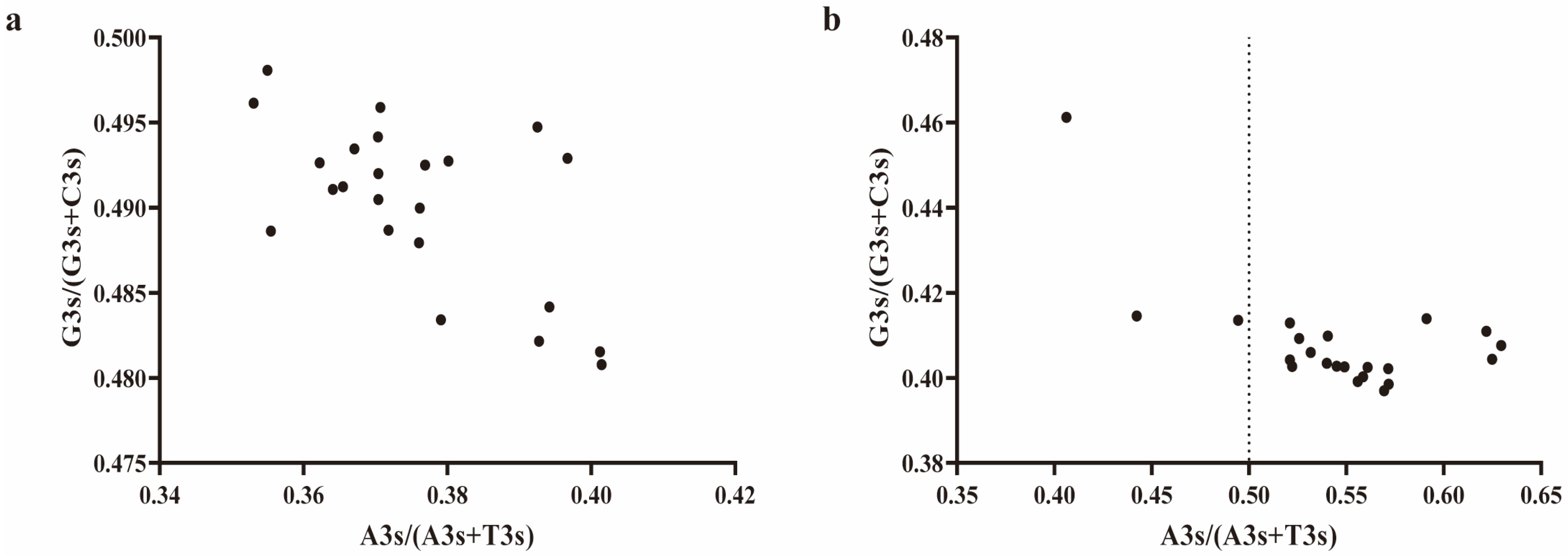
2.7. Analysis of DDR1 Gene and EHF Third Codon Bias
2.8. Multidimensional Clustering Analysis of CAI for DDR1–EHF Based on K-Means
3. Discussion
4. Materials and Methods
4.1. DDR1 Gene Data and EHF TF Data Collection
4.2. Phylogenetic Trees and Hierarchical Cluster Analysis
4.3. Parametric Statistical Methods for Codons
4.4. Codon Correlation and Third Codon Analysis
4.5. PCA and K-Means Cluster Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Do, D.N.; Dudemaine, P.L.; Li, R.; Ibeagha-Awemu, E.M. Co-Expression Network and Pathway Analyses Reveal Important Modules of miRNAs Regulating Milk Yield and Component Traits. Int. J. Mol. Sci. 2017, 18, 1560. [Google Scholar] [CrossRef] [PubMed]
- Barker, K.T.; Martindale, J.E.; Mitchell, P.J.; Kamalati, T.; Page, M.J.; Phippard, D.J.; Dale, T.C.; Gusterson, B.A.; Crompton, M.R. Expression patterns of the novel receptor-like tyrosine kinase, DDR, in human breast tumours. Oncogene 1995, 10, 569–575. [Google Scholar]
- Faraci-Orf, E.; McFadden, C.; Vogel, W.F. DDR1 signaling is essential to sustain Stat5 function during lactogenesis. J. Cell. Biochem. 2006, 97, 109–121. [Google Scholar] [CrossRef]
- Rauner, G.; Jin, D.X.; Miller, D.H.; Gierahn, T.M.; Li, C.M.; Sokol, E.S.; Feng, Y.X.; Mathis, R.A.; Love, J.C.; Gupta, P.B.; et al. Breast tissue regeneration is driven by cell-matrix interactions coordinating multi-lineage stem cell differentiation through DDR1. Nat. Commun. 2021, 12, 7116. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Hu, Y.; Pan, Y.; Xiong, Y.; Zhang, Y.; Han, M.; Dong, K.; Song, J.; Liang, H.; Ding, Z.; et al. DDR1 promotes hepatocellular carcinoma metastasis through recruiting PSD4 to ARF6. Oncogene 2022, 41, 1821–1834. [Google Scholar] [CrossRef] [PubMed]
- Ngai, D.; Mohabeer, A.L.; Mao, A.; Lino, M.; Bendeck, M.P. Stiffness-responsive feedback autoregulation of DDR1 expression is mediated by a DDR1-YAP/TAZ axis. Matrix Biol. 2022, 110, 129–140. [Google Scholar] [CrossRef]
- Sun, X.; Wu, B.; Chiang, H.C.; Deng, H.; Zhang, X.; Xiong, W.; Liu, J.; Rozeboom, A.M.; Harris, B.T.; Blommaert, E.; et al. Tumour DDR1 promotes collagen fibre alignment to instigate immune exclusion. Nature 2021, 599, 673–678. [Google Scholar] [CrossRef]
- Shenoy, G.P.; Pal, R.; Purwarga Matada, G.S.; Singh, E.; Raghavendra, N.M.; Dhiwar, P.S. Discoidin domain receptor inhibitors as anticancer agents: A systematic review on recent development of DDRs inhibitors, their resistance and structure activity relationship. Bioorg. Chem. 2023, 130, 106215. [Google Scholar] [CrossRef]
- Vogel, W.F.; Aszódi, A.; Alves, F.; Pawson, T. Discoidin domain receptor 1 tyrosine kinase has an essential role in mammary gland development. Mol. Cell. Biol. 2001, 21, 2906–2917. [Google Scholar] [CrossRef]
- Toscani, A.M.; Aguilera, P.; Coluccio Leskow, F. Discoidin domain receptor 1 regulates ErbB2/ErbB3 signaling in mammary epithelial cells. FEBS Lett. 2022, 596, 2795–2807. [Google Scholar] [CrossRef]
- Shen, W.K.; Chen, S.Y.; Gan, Z.Q.; Zhang, Y.Z.; Yue, T.; Chen, M.M.; Xue, Y.; Hu, H.; Guo, A.Y. AnimalTFDB 4.0: A comprehensive animal transcription factor database updated with variation and expression annotations. Nucleic Acids Res. 2023, 51, D39–D45. [Google Scholar] [CrossRef] [PubMed]
- Biswas, S.K.; Banerjee, S.; Baker, G.W.; Kuo, C.Y.; Chowdhury, I. The Mammary Gland: Basic Structure and Molecular Signaling during Development. Int. J. Mol. Sci. 2022, 23, 3883. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.; Zhang, Q.; Hu, F.F.; Liu, C.J.; Guo, A.Y. A comprehensive survey for human transcription factors on expression, regulation, interaction, phenotype and cancer survival. Brief Bioinform. 2021, 22, bbab002. [Google Scholar] [CrossRef] [PubMed]
- Nitta, K.R.; Jolma, A.; Yin, Y.; Morgunova, E.; Kivioja, T.; Akhtar, J.; Hens, K.; Toivonen, J.; Deplancke, B.; Furlong, E.E.; et al. Conservation of transcription factor binding specificities across 600 million years of bilateria evolution. eLife 2015, 4, e04837. [Google Scholar] [CrossRef] [PubMed]
- Barrera, L.A.; Vedenko, A.; Kurland, J.V.; Rogers, J.M.; Gisselbrecht, S.S.; Rossin, E.J.; Woodard, J.; Mariani, L.; Kock, K.H.; Inukai, S.; et al. Survey of variation in human transcription factors reveals prevalent DNA binding changes. Science 2016, 351, 1450–1454. [Google Scholar] [CrossRef]
- Xie, G.Y.; Xia, M.; Miao, Y.R.; Luo, M.; Zhang, Q.; Guo, A.Y. FFLtool: A web server for transcription factor and miRNA feed forward loop analysis in human. Bioinformatics 2020, 36, 2605–2607. [Google Scholar] [CrossRef]
- Zhang, Q.; Liu, W.; Zhang, H.M.; Xie, G.Y.; Miao, Y.R.; Xia, M.; Guo, A.Y. hTFtarget: A Comprehensive Database for Regulations of Human Transcription Factors and Their Targets. Genom. Proteom. Bioinform. 2020, 18, 120–128. [Google Scholar] [CrossRef]
- Bochert, M.A.; Kleinbaum, L.A.; Sun, L.Y.; Burton, F.H. Molecular cloning and expression of Ehf, a new member of the ets transcription factor/oncoprotein gene family. Biochem. Biophys. Res. Commun. 1998, 246, 176–181. [Google Scholar] [CrossRef]
- Li, W.; Okabe, A.; Usui, G.; Fukuyo, M.; Matsusaka, K.; Rahmutulla, B.; Mano, Y.; Hoshii, T.; Funata, S.; Hiura, N.; et al. Activation of EHF via STAT3 phosphorylation by LMP2A in Epstein-Barr virus-positive gastric cancer. Cancer Sci. 2021, 112, 3349–3362. [Google Scholar] [CrossRef]
- Brenne, K.; Nymoen, D.A.; Hetland, T.E.; Trope, C.G.; Davidson, B. Expression of the ETS transcription factor EHF in serous ovarian carcinoma effusions is a marker of poor survival. Hum. Pathol. 2012, 43, 496–505. [Google Scholar] [CrossRef]
- Wang, L.; Xing, J.; Cheng, R.; Shao, Y.; Li, P.; Zhu, S.; Zhang, S. Abnormal Localization and Tumor Suppressor Function of Epithelial Tissue-Specific Transcription Factor ESE3 in Esophageal Squamous Cell Carcinoma. PLoS ONE 2015, 10, e0126319. [Google Scholar] [CrossRef]
- Taniue, K.; Oda, T.; Hayashi, T.; Okuno, M.; Akiyama, T. A member of the ETS family, EHF, and the ATPase RUVBL1 inhibit p53-mediated apoptosis. EMBO Rep. 2011, 12, 682–689. [Google Scholar] [CrossRef] [PubMed]
- Kleinbaum, L.A.; Duggan, C.; Ferreira, E.; Coffey, G.P.; Butticè, G.; Burton, F.H. Human chromosomal localization, tissue/tumor expression, and regulatory function of the ETS family gene EHF. Biochem. Biophys. Res. Commun. 1999, 264, 119–126. [Google Scholar] [CrossRef]
- Galang, C.K.; Muller, W.J.; Foos, G.; Oshima, R.G.; Hauser, C.A. Changes in the expression of many Ets family transcription factors and of potential target genes in normal mammary tissue and tumors. J. Biol. Chem. 2004, 279, 11281–11292. [Google Scholar] [CrossRef]
- He, J.; Pan, Y.; Hu, J.; Albarracin, C.; Wu, Y.; Dai, J.L. Profile of Ets gene expression in human breast carcinoma. Cancer Biol. Ther. 2007, 6, 76–82. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Novoa, E.M.; Pavon-Eternod, M.; Pan, T.; Ribas de Pouplana, L. A role for tRNA modifications in genome structure and codon usage. Cell 2012, 149, 202–213. [Google Scholar] [CrossRef]
- Quax, T.E.; Claassens, N.J.; Söll, D.; van der Oost, J. Codon Bias as a Means to Fine-Tune Gene Expression. Mol. Cell 2015, 59, 149–161. [Google Scholar] [CrossRef] [PubMed]
- Parvathy, S.T.; Udayasuriyan, V.; Bhadana, V. Codon usage bias. Mol. Biol. Rep. 2022, 49, 539–565. [Google Scholar] [CrossRef] [PubMed]
- Ma, Q.P.; Li, C.; Wang, J.; Wang, Y.; Ding, Z.T. Analysis of synonymous codon usage in FAD7 genes from different plant species. Genet. Mol. Res. 2015, 14, 1414–1422. [Google Scholar] [CrossRef]
- Liu, Y. A code within the genetic code: Codon usage regulates co-translational protein folding. Cell Commun. Signal. 2020, 18, 145. [Google Scholar] [CrossRef]
- Pedersen, A.K.; Wiuf, C.; Christiansen, F.B. A codon-based model designed to describe lentiviral evolution. Mol. Biol. Evol. 1998, 15, 1069–1081. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Mazumdar, P.; Binti Othman, R.; Mebus, K.; Ramakrishnan, N.; Ann Harikrishna, J. Codon usage and codon pair patterns in non-grass monocot genomes. Ann. Bot. 2017, 120, 893–909. [Google Scholar] [CrossRef] [PubMed]
- Sharp, P.M.; Li, W.H. An evolutionary perspective on synonymous codon usage in unicellular organisms. J. Mol. Evol. 1986, 24, 28–38. [Google Scholar] [CrossRef]
- Gu, W.; Zhou, T.; Ma, J.; Sun, X.; Lu, Z. The relationship between synonymous codon usage and protein structure in Escherichia coli and Homo sapiens. Biosystems 2004, 73, 89–97. [Google Scholar] [CrossRef]
- Moriyama, E.N.; Powell, J.R. Codon usage bias and tRNA abundance in Drosophila. J. Mol. Evol. 1997, 45, 514–523. [Google Scholar] [CrossRef]
- Moriyama, E.N.; Powell, J.R. Gene length and codon usage bias in Drosophila melanogaster, Saccharomyces cerevisiae and Escherichia coli. Nucleic Acids Res. 1998, 26, 3188–3193. [Google Scholar] [CrossRef]
- Gu, W.; Zhou, T.; Ma, J.; Sun, X.; Lu, Z. Folding type specific secondary structure propensities of synonymous codons. IEEE Trans. Nanobiosci. 2003, 2, 150–157. [Google Scholar]
- Romero, H.; Zavala, A.; Musto, H. Codon usage in Chlamydia trachomatis is the result of strand-specific mutational biases and a complex pattern of selective forces. Nucleic Acids Res. 2000, 28, 2084–2090. [Google Scholar] [CrossRef]
- Lee, S.; Weon, S.; Lee, S.; Kang, C. Relative codon adaptation index, a sensitive measure of codon usage bias. Evol. Bioinform. Online 2010, 6, 47–55. [Google Scholar] [CrossRef]
- Wright, F. The ‘effective number of codons’ used in a gene. Gene 1990, 87, 23–29. [Google Scholar] [CrossRef]
- Ikemura, T. Correlation between the abundance of Escherichia coli transfer RNAs and the occurrence of the respective codons in its protein genes. J. Mol. Biol. 1981, 146, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, K.L.; Yang, I.C. Incorporating PCA and fuzzy-ART techniques into achieve organism classification based on codon usage consideration. Comput. Biol. Med. 2008, 38, 886–893. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Lu, Y.; Song, Y.; Jing, L. Analysis of codon usage bias of WRKY transcription factors in Helianthus annuus. BMC Genom. Data 2022, 23, 46. [Google Scholar] [CrossRef]
- Srihari, J.; Sakthi, A.; Balakrishnan, N.; Duraialagaraja, S.; Varatharajalu, U. Study of Expression of Indigenous Bt cry2AX1 Gene in T 3 Progeny of Cotton and its Efficacy Against Helicoverpa armigera (Hubner). Braz. Arch. Biol. Technol. 2020, 63, 2020. [Google Scholar]
- Batard, Y.; Hehn, A.; Nedelkina, S.; Schalk, M.; Pallett, K.; Schaller, H.; Werck-Reichhart, D. Increasing expression of P450 and P450-reductase proteins from monocots in heterologous systems. Arch. Biochem. Biophys. 2000, 379, 161–169. [Google Scholar] [CrossRef]
- Xu, Y.; Liu, K.; Han, Y.; Xing, Y.; Zhang, Y.; Yang, Q.; Zhou, M. Codon usage bias regulates gene expression and protein conformation in yeast expression system P. pastoris. Microb. Cell Fact. 2021, 20, 91. [Google Scholar] [CrossRef] [PubMed]
- Peden, J.F. Analysis of Codon Usage. Ph.D. Thesis, University of Nottingham, Nottingham, UK, 1999. [Google Scholar]
- Bazzini, A.A.; Del Viso, F.; Moreno-Mateos, M.A.; Johnstone, T.G.; Vejnar, C.E.; Qin, Y.; Yao, J.; Khokha, M.K.; Giraldez, A.J. Codon identity regulates mRNA stability and translation efficiency during the maternal-to-zygotic transition. EMBO J. 2016, 35, 2087–2103. [Google Scholar] [CrossRef]
- Radrizzani, S.; Kudla, G.; Izsvák, Z.; Hurst, L.D. Selection on synonymous sites: The unwanted transcript hypothesis. Nat. Rev. Genet. 2024, 25, 431–448. [Google Scholar] [CrossRef] [PubMed]
- Xing, K.; He, X. Reassessing the “duon” hypothesis of protein evolution. Mol. Biol. Evol. 2015, 32, 1056–1062. [Google Scholar] [CrossRef]
- Chen, L.; Liu, T.; Yang, D.; Nong, X.; Xie, Y.; Fu, Y.; Wu, X.; Huang, X.; Gu, X.; Wang, S.; et al. Analysis of codon usage patterns in Taenia pisiformis through annotated transcriptome data. Biochem. Biophys. Res. Commun. 2013, 430, 1344–1348. [Google Scholar] [CrossRef]
- Dhindsa, R.S.; Copeland, B.R.; Mustoe, A.M.; Goldstein, D.B. Natural Selection Shapes Codon Usage in the Human Genome. Am. J. Hum. Genet. 2020, 107, 83–95. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Yang, Q.; Zhao, F. Synonymous but Not Silent: The Codon Usage Code for Gene Expression and Protein Folding. Annu. Rev. Biochem. 2021, 90, 375–401. [Google Scholar] [CrossRef] [PubMed]
- Saul, A.; Battistutta, D. Codon usage in Plasmodium falciparum. Mol. Biochem. Parasitol. 1988, 27, 35–42. [Google Scholar] [CrossRef]
- Xiong, B.; Wang, T.; Huang, S.; Liao, L.; Wang, X.; Deng, H.; Zhang, M.; He, J.; Sun, G.; He, S.; et al. Analysis of Codon Usage Bias in Xyloglucan Endotransglycosylase (XET) Genes. Int. J. Mol. Sci. 2023, 24, 6108. [Google Scholar] [CrossRef] [PubMed]
- Duret, L. Evolution of synonymous codon usage in metazoans. Curr. Opin. Genet. Dev. 2002, 12, 640–649. [Google Scholar] [CrossRef] [PubMed]
- Nasrullah, I.; Butt, A.M.; Tahir, S.; Idrees, M.; Tong, Y. Genomic analysis of codon usage shows influence of mutation pressure, natural selection, and host features on Marburg virus evolution. BMC Evol. Biol. 2015, 15, 174. [Google Scholar] [CrossRef]
- Vicario, S.; Moriyama, E.N.; Powell, J.R. Codon usage in twelve species of Drosophila. BMC Evol. Biol. 2007, 7, 226. [Google Scholar] [CrossRef]
- Tao, P.; Dai, L.; Luo, M.; Tang, F.; Tien, P.; Pan, Z. Analysis of synonymous codon usage in classical swine fever virus. Virus Genes 2009, 38, 104–112. [Google Scholar] [CrossRef]
- Wang, H.; Liu, S.; Zhang, B.; Wei, W. Analysis of Synonymous Codon Usage Bias of Zika Virus and Its Adaption to the Hosts. PLoS ONE 2016, 11, e0166260. [Google Scholar] [CrossRef]
- Wang, D.; Yang, B. Analysis of codon usage bias of thioredoxin in apicomplexan protozoa. Parasit. Vectors 2023, 16, 431. [Google Scholar] [CrossRef]
- Ouyang, T.; Zhong, J.; Chai, Z.; Wang, J.; Zhang, M.; Wu, Z.; Xin, J. Codon Usage Bias and Cluster Analysis of the MMP-2 and MMP-9 Genes in Seven Mammals. Genet. Res. 2022, 2022, 2823356. [Google Scholar] [CrossRef]
- Chakraborty, S.; Uddin, A.; Choudhury, M.N. Factors affecting the codon usage bias of SRY gene across mammals. Gene 2017, 630, 13–20. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, S.; Nag, D.; Mazumder, T.H.; Uddin, A. Codon usage pattern and prediction of gene expression level in Bungarus species. Gene 2017, 604, 48–60. [Google Scholar] [CrossRef] [PubMed]
- Somaratne, Y.; Guan, D.L.; Wang, W.Q.; Zhao, L.; Xu, S.Q. The Complete Chloroplast Genomes of Two Lespedeza Species: Insights into Codon Usage Bias, RNA Editing Sites, and Phylogenetic Relationships in Desmodieae (Fabaceae: Papilionoideae). Plants 2019, 9, 51. [Google Scholar] [CrossRef] [PubMed]
- Bu, Y.; Wu, X.; Sun, N.; Man, Y.; Jing, Y. Codon usage bias predicts the functional MYB10 gene in Populus. J. Plant Physiol. 2021, 265, 153491. [Google Scholar] [CrossRef]
- Crick, F.H.; Barnett, L.; Brenner, S.; Watts-Tobin, R.J. General nature of the genetic code for proteins. Nature 1961, 192, 1227–1232. [Google Scholar] [CrossRef]
- Nirenberg, M. Historical review: Deciphering the genetic code—A personal account. Trends Biochem. Sci. 2004, 29, 46–54. [Google Scholar] [CrossRef]
- Matthaei, H.; Nirenberg, M.W. The dependence of cell-free protein synthesis in E. coli upon RNA prepared from ribosomes. Biochem. Biophys. Res. Commun. 1961, 4, 404–408. [Google Scholar] [CrossRef]
- Lambert, S.A.; Jolma, A.; Campitelli, L.F.; Das, P.K.; Yin, Y.; Albu, M.; Chen, X.; Taipale, J.; Hughes, T.R.; Weirauch, M.T. The Human Transcription Factors. Cell 2018, 172, 650–665. [Google Scholar] [CrossRef]
- Weidemüller, P.; Kholmatov, M.; Petsalaki, E.; Zaugg, J.B. Transcription factors: Bridge between cell signaling and gene regulation. Proteomics 2021, 21, e2000034. [Google Scholar] [CrossRef]
- Kwon, K.C.; Chan, H.T.; León, I.R.; Williams-Carrier, R.; Barkan, A.; Daniell, H. Codon Optimization to Enhance Expression Yields Insights into Chloroplast Translation. Plant Physiol. 2016, 172, 62–77. [Google Scholar] [CrossRef] [PubMed]
- Baeza, M.; Sepulveda, D.; Cifuentes, V.; Alcaíno, J. Codon usage bias in yeasts and its correlation with gene expression, growth temperature, and protein structure. Front. Microbiol. 2024, 15, 1414422. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.-J.; Zhou, J.; Li, Z.-F.; Wang, L.; Gu, X.; Zhong, Y. Comparative Analysis of Codon Usage Patterns Among Mitochondrion, Chloroplast and Nuclear Genes in Triticum aestivum L. J. Integr. Plant Biol. 2007, 49, 246–254. [Google Scholar] [CrossRef]
- Sueoka, N. Intrastrand parity rules of DNA base composition and usage biases of synonymous codons. J. Mol. Evol. 1995, 40, 318–325. [Google Scholar] [CrossRef]
- Sueoka, N. Translation-coupled violation of Parity Rule 2 in human genes is not the cause of heterogeneity of the DNA G+C content of third codon position. Gene 1999, 238, 53–58. [Google Scholar] [CrossRef]
- Kassambara, A.; Mundt, F. Extract and Visualize the Results of Multivariate Data Analyses [R Package Factoextra Version 1.0.7]; R Foundation: Indianapolis, IN, USA, 2020. [Google Scholar]
- Husson, F.; Josse, J.; Le, S.; Mazet, J. Multivariate exploratory data analysis and data mining. Cran 2020, 1, 1–130. [Google Scholar]
- Ikotun, A.M.; Ezugwu, A.E. Boosting k-means clustering with symbiotic organisms search for automatic clustering problems. PLoS ONE 2022, 17, e0272861. [Google Scholar] [CrossRef]



| Species | T3s/% | C3s/% | A3s/% | G3s/% | CAI | CBI | Fop | Nc | GC3s/% | GC/% | Gravy | Aromo |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Bos taurus | 17.23 | 47.39 | 11.21 | 44.48 | 0.197 | 0.071 | 0.442 | 41.49 | 75.90 | 63.30 | −0.23 | 0.09 |
| Bos indicus × Bos taurus | 17.49 | 46.58 | 11.30 | 45.61 | 0.199 | 0.066 | 0.441 | 41.59 | 75.70 | 62.90 | −0.22 | 0.09 |
| Bos javanicus | 17.23 | 47.39 | 11.21 | 44.48 | 0.198 | 0.072 | 0.443 | 41.67 | 75.90 | 63.30 | −0.22 | 0.09 |
| Bos indicus | 16.73 | 47.94 | 11.22 | 44.39 | 0.198 | 0.082 | 0.448 | 41.72 | 76.30 | 63.40 | −0.20 | 0.09 |
| Bos mutus | 17.04 | 47.72 | 11.02 | 44.43 | 0.199 | 0.082 | 0.449 | 41.78 | 76.20 | 63.30 | −0.20 | 0.09 |
| Bubalus carabanensis | 16.82 | 47.85 | 11.27 | 44.44 | 0.202 | 0.083 | 0.449 | 41.80 | 76.20 | 63.40 | −0.22 | 0.09 |
| Bubalus bubalis | 17.08 | 47.13 | 11.23 | 45.81 | 0.205 | 0.081 | 0.450 | 41.82 | 76.10 | 63.00 | −0.21 | 0.09 |
| Capra hircus | 17.21 | 47.40 | 10.41 | 46.00 | 0.207 | 0.087 | 0.453 | 41.85 | 76.60 | 63.30 | −0.22 | 0.09 |
| Ovis aries | 17.98 | 46.59 | 10.59 | 45.8 | 0.208 | 0.088 | 0.454 | 41.94 | 75.80 | 63.00 | −0.21 | 0.09 |
| Cervus elaphus | 18.77 | 46.28 | 11.11 | 44.23 | 0.204 | 0.083 | 0.450 | 41.96 | 74.60 | 62.90 | −0.22 | 0.09 |
| Dama dama | 18.41 | 46.87 | 11.24 | 43.86 | 0.205 | 0.087 | 0.452 | 42.00 | 74.90 | 63.00 | −0.22 | 0.09 |
| Moschus berezovskii | 18.12 | 46.68 | 10.92 | 44.48 | 0.203 | 0.074 | 0.444 | 42.00 | 75.30 | 63.20 | −0.21 | 0.09 |
| Cervus canadensis | 18.90 | 46.15 | 10.82 | 44.53 | 0.204 | 0.083 | 0.450 | 42.01 | 74.70 | 63.00 | −0.22 | 0.09 |
| Globicephala melas | 18.35 | 47.23 | 10.12 | 45.13 | 0.210 | 0.094 | 0.456 | 42.02 | 75.90 | 63.20 | −0.27 | 0.09 |
| Phocoena sinus | 18.28 | 46.74 | 10.53 | 45.13 | 0.214 | 0.101 | 0.460 | 42.02 | 75.60 | 63.20 | −0.22 | 0.09 |
| Mesoplodon densirostris | 18.54 | 46.08 | 11.37 | 44.76 | 0.208 | 0.083 | 0.449 | 42.14 | 74.70 | 62.90 | −0.22 | 0.09 |
| Lagenorhynchus albirostri | 18.12 | 46.68 | 10.66 | 45.21 | 0.210 | 0.097 | 0.457 | 42.16 | 75.60 | 63.10 | −0.22 | 0.09 |
| Delphinus delphis | 18.38 | 46.41 | 10.66 | 45.21 | 0.210 | 0.091 | 0.454 | 42.19 | 75.40 | 63.00 | −0.22 | 0.09 |
| Tursiops truncatus | 18.12 | 46.81 | 10.66 | 45.06 | 0.211 | 0.095 | 0.456 | 42.19 | 75.60 | 63.10 | −0.22 | 0.09 |
| Neophocaena asiaeoriental asiaeoriental | 18.54 | 46.48 | 10.53 | 45.13 | 0.214 | 0.101 | 0.460 | 42.21 | 75.30 | 63.10 | −0.22 | 0.09 |
| Balaenoptera ricei | 19.35 | 45.75 | 10.56 | 45.05 | 0.206 | 0.076 | 0.445 | 42.32 | 74.60 | 62.80 | −0.23 | 0.09 |
| Monodon monoceros | 18.64 | 46.28 | 11.24 | 44.46 | 0.211 | 0.096 | 0.456 | 42.52 | 74.70 | 62.80 | −0.22 | 0.09 |
| Orcinus orca | 18.38 | 46.28 | 10.81 | 45.21 | 0.209 | 0.090 | 0.452 | 42.57 | 75.30 | 63.00 | −0.22 | 0.09 |
| Physeter catodon | 19.17 | 45.63 | 10.55 | 45.28 | 0.208 | 0.089 | 0.452 | 42.66 | 74.80 | 63.00 | −0.22 | 0.09 |
| Species | T3s/% | C3s/% | A3s/% | G3s/% | CAI | CBI | Fop | Nc | GC3s/% | GC% | Gravy | Aromo |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Bos taurus | 21.38 | 46.54 | 30.92 | 32.87 | 0.277 | 0.215 | 0.554 | 57.52 | 59.90 | 49.10 | −0.95 | 0.10 |
| Bos indicus × Bos taurus | 22.67 | 49.78 | 25.73 | 34.02 | 0.292 | 0.226 | 0.56 | 54.01 | 63.10 | 50.70 | −0.81 | 0.11 |
| Bos javanicus | 21.38 | 46.54 | 30.92 | 32.87 | 0.277 | 0.215 | 0.554 | 57.52 | 59.90 | 49.10 | −0.95 | 0.10 |
| Bos indicus | 27.37 | 47.37 | 21.71 | 33.54 | 0.304 | 0.187 | 0.536 | 50.46 | 61.70 | 50.80 | −0.47 | 0.10 |
| Bos mutus | 21.95 | 49.76 | 28.04 | 33.52 | 0.287 | 0.206 | 0.55 | 53.46 | 62.30 | 50.40 | −0.87 | 0.12 |
| Bubalus carabanensis | 25.57 | 47.03 | 25.00 | 33.16 | 0.276 | 0.199 | 0.542 | 54.68 | 60.90 | 50.00 | −0.64 | 0.12 |
| Bubalus bubalis | 24.22 | 48.88 | 26.34 | 33.16 | 0.29 | 0.204 | 0.548 | 54.06 | 61.60 | 50.10 | −0.79 | 0.11 |
| Capra hircus | 23.21 | 48.66 | 27.80 | 32.81 | 0.284 | 0.218 | 0.555 | 55.03 | 61.20 | 50.20 | −0.80 | 0.11 |
| Ovis aries | 34.60 | 37.26 | 23.67 | 31.90 | 0.249 | 0.108 | 0.483 | 54.81 | 53.60 | 49.40 | −0.28 | 0.12 |
| Cervus elaphus | 23.21 | 49.11 | 25.73 | 34.02 | 0.274 | 0.187 | 0.537 | 53.93 | 62.60 | 50.80 | −0.79 | 0.11 |
| Dama dama | 22.64 | 50.00 | 24.74 | 33.71 | 0.284 | 0.242 | 0.567 | 53.56 | 63.60 | 51.00 | −0.68 | 0.12 |
| Moschus berezovskii | 21.88 | 50.00 | 25.73 | 34.72 | 0.288 | 0.23 | 0.562 | 53.42 | 63.70 | 51.20 | −0.78 | 0.11 |
| Cervus canadensis | 23.21 | 49.11 | 25.24 | 34.54 | 0.274 | 0.187 | 0.537 | 54.31 | 63.00 | 50.90 | −0.79 | 0.11 |
| Globicephala melas | 21.17 | 50.45 | 26.79 | 33.67 | 0.272 | 0.198 | 0.544 | 52.53 | 63.30 | 50.90 | −0.77 | 0.11 |
| Phocoena sinus | 21.17 | 49.55 | 28.23 | 33.33 | 0.269 | 0.185 | 0.537 | 53.24 | 62.30 | 50.60 | −0.76 | 0.11 |
| Mesoplodon densirostris | 22.97 | 49.55 | 26.96 | 33.51 | 0.272 | 0.175 | 0.532 | 52.32 | 62.10 | 50.20 | −0.80 | 0.12 |
| Lagenorhynchus albirostri | 19.23 | 48.08 | 32.03 | 32.64 | 0.251 | 0.175 | 0.532 | 56.16 | 60.70 | 49.10 | −0.92 | 0.10 |
| Delphinus delphis | 21.72 | 50.23 | 26.44 | 33.85 | 0.271 | 0.202 | 0.546 | 53.08 | 63.20 | 50.70 | −0.78 | 0.11 |
| Tursiops truncatus | 19.87 | 46.79 | 32.68 | 32.64 | 0.253 | 0.175 | 0.532 | 57.44 | 59.70 | 48.80 | −0.92 | 0.10 |
| Neophocaena asiaeoriental asiaeoriental | 21.17 | 49.55 | 28.23 | 33.33 | 0.269 | 0.185 | 0.537 | 53.24 | 62.30 | 50.60 | −0.76 | 0.11 |
| Balaenoptera ricei | 20.27 | 51.35 | 27.05 | 34.02 | 0.266 | 0.198 | 0.544 | 53.15 | 64.10 | 50.80 | −0.80 | 0.11 |
| Monodon monoceros | 20.72 | 50.90 | 27.40 | 33.51 | 0.272 | 0.199 | 0.544 | 53.05 | 63.30 | 50.70 | −0.76 | 0.11 |
| Orcinus orca | 19.23 | 47.44 | 32.68 | 32.64 | 0.253 | 0.175 | 0.532 | 56.58 | 60.20 | 49.00 | −0.92 | 0.10 |
| Physeter catodon | 21.62 | 50.45 | 27.05 | 33.51 | 0.27 | 0.198 | 0.544 | 52.62 | 63.00 | 50.60 | −0.79 | 0.11 |
| Amino Acid | Codon | Number | RSCU | Amino Acid | Codon | Number | RSCU |
|---|---|---|---|---|---|---|---|
| Phe | UUU | 318 | 0.77 | Ala | GCU | 253 | 0.60 |
| UUC * | 506 | 1.23 | GCC * | 1005 | 2.38 | ||
| Leu | UUA | 105 | 0.24 | GCA | 177 | 0.42 | |
| UUG | 155 | 0.36 | GCG | 252 | 0.60 | ||
| CUU | 182 | 0.42 | Tyr | UAU | 281 | 0.87 | |
| CUC * | 664 | 1.52 | UAC * | 366 | 1.13 | ||
| CUA | 141 | 0.33 | His | CAU | 150 | 0.58 | |
| CUG * | 1368 | 3.14 | CAC * | 369 | 1.42 | ||
| Ile | AUU | 74 | 0.36 | Gln | CAA | 74 | 0.19 |
| AUC * | 522 | 2.52 | CAG * | 682 | 1.81 | ||
| AUA | 26 | 0.13 | Asn | AAU | 212 | 0.66 | |
| Val | GUU | 118 | 0.32 | AAC * | 427 | 1.34 | |
| GUC | 301 | 0.83 | Lys | AAA | 45 | 0.16 | |
| GUA | 75 | 0.21 | AAG * | 513 | 1.84 | ||
| GUG * | 958 | 2.63 | Asp | GAU | 475 | 0.79 | |
| Ser | UCU | 157 | 0.75 | GAC * | 726 | 1.21 | |
| UCC * | 289 | 1.38 | Glu | GAA | 150 | 0.25 | |
| UCA | 77 | 0.37 | GAG * | 1049 | 1.75 | ||
| UCG | 79 | 0.38 | Cys | UGU | 127 | 0.62 | |
| AGU | 148 | 0.71 | UGC * | 282 | 1.38 | ||
| AGC * | 505 | 2.41 | Arg | CGU | 95 | 0.35 | |
| Pro | CCU | 423 | 0.89 | CGC * | 428 | 1.59 | |
| CCC * | 864 | 1.82 | CGA | 127 | 0.47 | ||
| CCA | 343 | 0.72 | CGG * | 630 | 2.33 | ||
| CCG | 268 | 0.56 | AGA | 62 | 0.23 | ||
| Thr | ACU | 124 | 0.60 | AGG * | 277 | 1.03 | |
| ACC * | 443 | 2.15 | Gly | GGU | 151 | 0.30 | |
| ACA | 126 | 0.61 | GGC * | 825 | 1.64 | ||
| ACG | 133 | 0.65 | GGA | 268 | 0.53 | ||
| GGG * | 767 | 1.53 |
| Amino Acid | Codon | Number | RSCU | Amino Acid | Codon | Number | RSCU |
|---|---|---|---|---|---|---|---|
| Phe | UUU | 87 | 0.64 | Ala | GCU | 37 | 0.54 |
| UUC * | 164 | 1.36 | GCC * | 116 | 1.76 | ||
| Leu | UUA | 71 | 0.70 | GCA | 63 | 0.97 | |
| UUG | 74 | 0.73 | GCG | 49 | 0.72 | ||
| CUU | 66 | 0.61 | Tyr | UAU | 110 | 0.83 | |
| CUC * | 187 | 1.74 | UAC * | 156 | 1.17 | ||
| CUA | 33 | 0.32 | His | CAU | 81 | 0.63 | |
| CUG * | 201 | 1.91 | CAC * | 165 | 1.37 | ||
| Ile | AUU | 85 | 0.89 | Gln | CAA | 67 | 0.35 |
| AUC * | 197 | 2.07 | CAG * | 327 | 1.65 | ||
| AUA | 3 | 0.03 | Asn | AAU | 189 | 0.85 | |
| Val | GUU | 23 | 0.36 | AAC * | 253 | 1.15 | |
| GUC * | 56 | 1.08 | Lys | AAA * | 242 | 1.06 | |
| GUA * | 67 | 1.27 | AAG | 214 | 0.94 | ||
| GUG * | 69 | 1.29 | Asp | GAU | 73 | 0.46 | |
| Ser | UCU | 32 | 0.36 | GAC * | 243 | 1.54 | |
| UCC * | 109 | 1.24 | Glu | GAA * | 276 | 1.24 | |
| UCA | 35 | 0.41 | GAG | 168 | 0.76 | ||
| UCG | 10 | 0.12 | Cys | UGU | 3 | 0.04 | |
| AGU | 81 | 0.91 | UGC * | 107 | 1.96 | ||
| AGC* | 259 | 2.95 | Arg | CGU | 34 | 0.62 | |
| Pro | CCU* | 95 | 1.35 | CGC | 33 | 0.59 | |
| CCC | 66 | 0.96 | CGA * | 61 | 1.10 | ||
| CCA* | 64 | 1.00 | CGG * | 68 | 1.36 | ||
| CCG | 45 | 0.69 | AGA | 77 | 1.47 | ||
| Thr | ACU | 74 | 0.66 | AGG * | 48 | 0.88 | |
| ACC* | 211 | 1.90 | Gly | GGU | 70 | 0.70 | |
| ACA | 110 | 1.00 | GGC | 101 | 0.99 | ||
| ACG | 51 | 0.44 | GGA | 95 | 0.94 | ||
| GGG * | 137 | 1.38 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Z.; Li, W.; Wang, Z.; Ma, S.; Zheng, F.; Liu, H.; Zhang, X.; Ding, Y.; Yin, Z.; Zheng, X. Codon Bias of the DDR1 Gene and Transcription Factor EHF in Multiple Species. Int. J. Mol. Sci. 2024, 25, 10696. https://doi.org/10.3390/ijms251910696
Zhang Z, Li W, Wang Z, Ma S, Zheng F, Liu H, Zhang X, Ding Y, Yin Z, Zheng X. Codon Bias of the DDR1 Gene and Transcription Factor EHF in Multiple Species. International Journal of Molecular Sciences. 2024; 25(19):10696. https://doi.org/10.3390/ijms251910696
Chicago/Turabian StyleZhang, Zhiyong, Wenxi Li, Ziyang Wang, Shuya Ma, Fangyuan Zheng, Hongyu Liu, Xiaodong Zhang, Yueyun Ding, Zongjun Yin, and Xianrui Zheng. 2024. "Codon Bias of the DDR1 Gene and Transcription Factor EHF in Multiple Species" International Journal of Molecular Sciences 25, no. 19: 10696. https://doi.org/10.3390/ijms251910696
APA StyleZhang, Z., Li, W., Wang, Z., Ma, S., Zheng, F., Liu, H., Zhang, X., Ding, Y., Yin, Z., & Zheng, X. (2024). Codon Bias of the DDR1 Gene and Transcription Factor EHF in Multiple Species. International Journal of Molecular Sciences, 25(19), 10696. https://doi.org/10.3390/ijms251910696






