An Integrated Comprehensive Peptidomics and In Silico Analysis of Bioactive Peptide-Rich Milk Fermented by Three Autochthonous Cocci Strains
Abstract
1. Introduction
2. Results and Discussion
2.1. Whole-Genome Sequencing (WGS) and Genome Annotation of Bacterial Strains
2.2. Preparation of the Functional Fermented BP-Rich Lyophilised Dairy Product
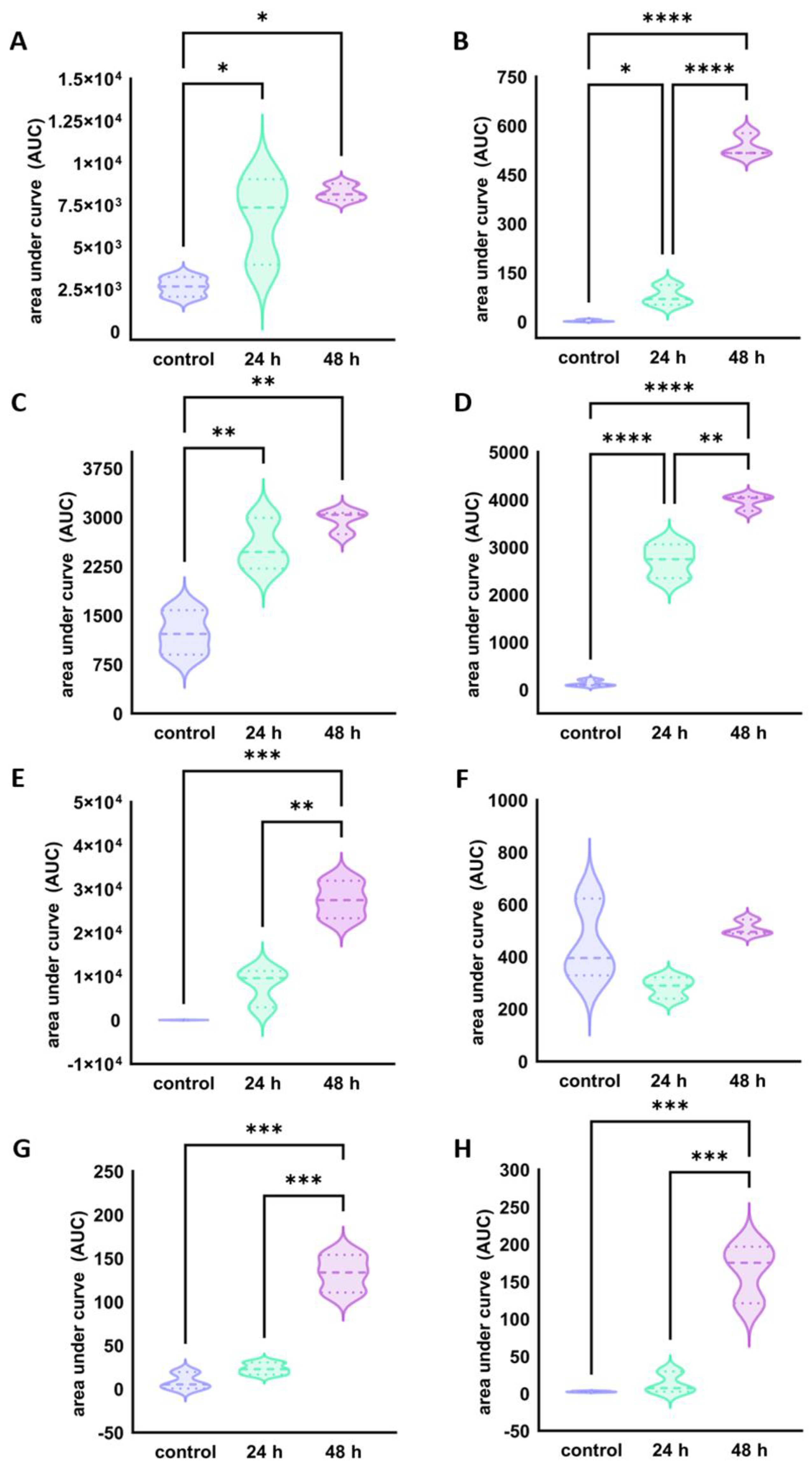
2.3. Peptidomic Profiling of the BP-Rich Lyophilised Dairy Product
| Precursor Protein | Peptide Sequence | FT | m/z [Da] | Biological Functions | Ref. No. | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| ACE | AO | AM | AT | IM | OP | AC | |||||
| α-s1-CN | 40VAPFPEVFGK49● | 48 h | 1090.61 | ||||||||
| κ-CN | 117ARHPHPHLSFM127●● * | 48 h | 1329.67 |  |  | [36,37] | |||||
| 45KYIPIQYVL53●● | both | 1136.70 |  | [38] | |||||||
| 45KYIPIQY51● | 48 h | 924.57 | |||||||||
| 54SRYPSYGLN62● | 48 h | 1056.53 | |||||||||
| 54SRYPSYGLNYY64● | 48 h | 1382.65 | |||||||||
| 63YYQQKPVALINN74● | both | 1450.80 | |||||||||
| 66QKPVALINNQFLPYPYYAKPA86● | 48 h | 2435.28 | |||||||||
| 74NQFLPYPYYAKPA86● | 48 h | 1571.83 | |||||||||
| 75QFLPYPYYAKPA86● | both | 1457.78 | |||||||||
| 87AVRSPAQILQWQVL100● | both | 1608.93 | |||||||||
| 117ARHPHPHLSF126● | 48 h | 1198.64 | |||||||||
| 63YYQQKPVALINNQFLPYPYYAKPA86● | 24 h | 2889.49 | |||||||||
| α-s2-CN | 209IQPKTKVIPYVRYL222●● | 24 h | 1718.04 |  |  | [39,40] | |||||
| 213TKVIPYVRYL222●● * | 24 h | 1251.75 |  | [41] | |||||||
| β-CN | 74VYPFPGPIPN83●● * | both | 1100.59 |  |  |  | [40,42,43] | ||||
| 208YQEPVLGPVRGPFPIIV224●● * | both | 1881.07 |  |  |  |  |  |  | [42,44,45,46,47,48] | ||
| 209QEPVLGPVRGPFPIIV224●● * | both | 1717.99 |  |  | [42,49] | ||||||
| 181SQSKVLPVPQKAVPYPQ197● * | both | 1865.04 |  |  | [40,50] | ||||||
| 198RDMPIQAF205● * | both | 977.51 |  | [44] | |||||||
| 207LYQEPVLGPVRGPFPIIV224● * | both | 1994.18 |  | [51] | |||||||
| 59ELQDKIHPF67● | 48 h | 1126.60 | |||||||||
| 62DKIHPFAQTQ71● | 48 h | 1184.62 | |||||||||
| 147NLHLPLPLLQ156● | 48 h | 1157.72 | |||||||||
| 185VLPVPQKAVPYPQ197● | 48 h | 1435.83 | |||||||||
| 210EPVLGPVRGPFPIIV224● | 48 h | 1589.97 | |||||||||
 ) indicates that the biological function listed in the column has been demonstrated for the peptide listed in the row.
) indicates that the biological function listed in the column has been demonstrated for the peptide listed in the row.2.4. In Silico Analysis of Peptides Released from Bovine Milk Fermented by Strains ZGBP5-51, ZGBP5-52 and ZGBP5-53
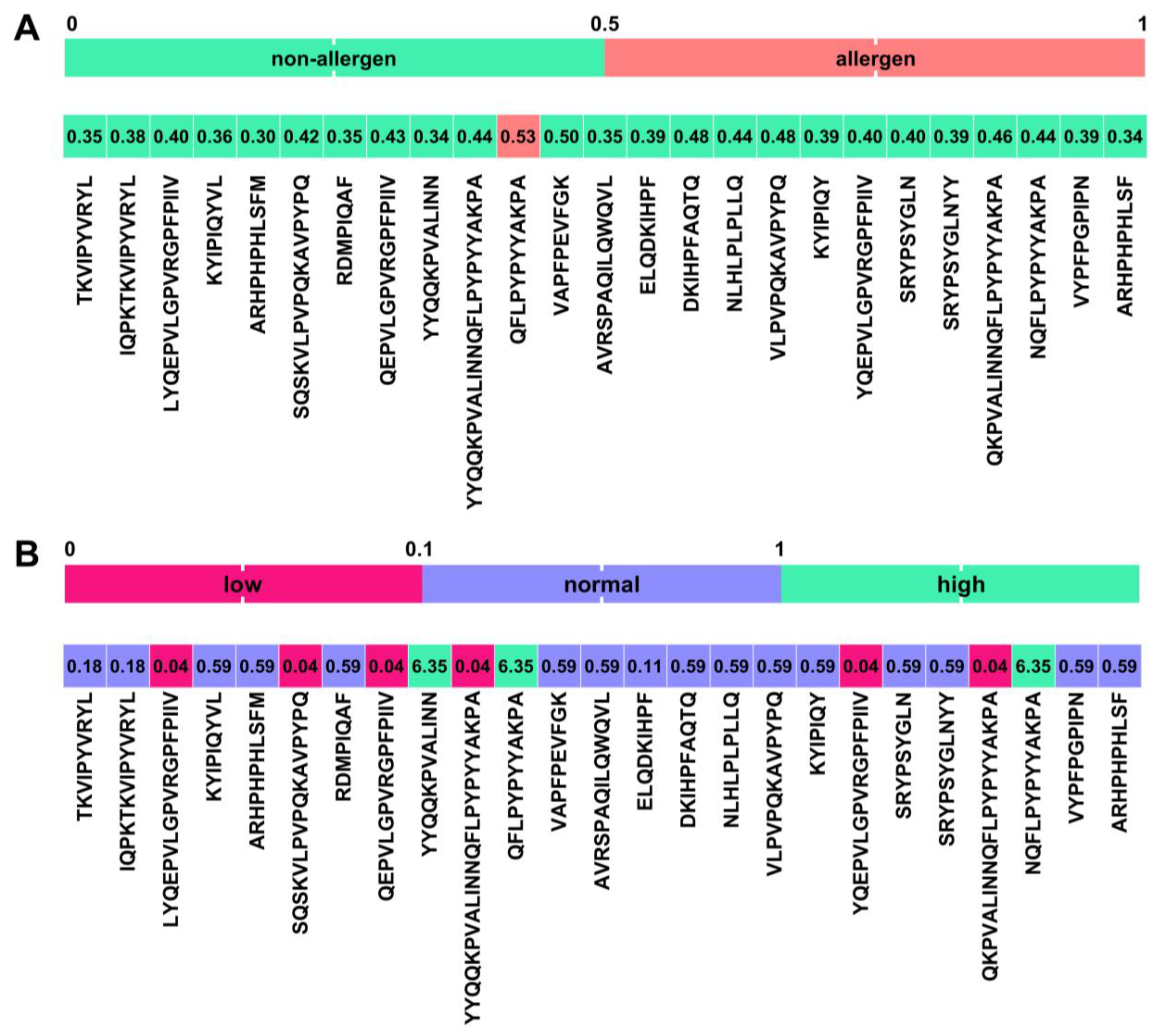
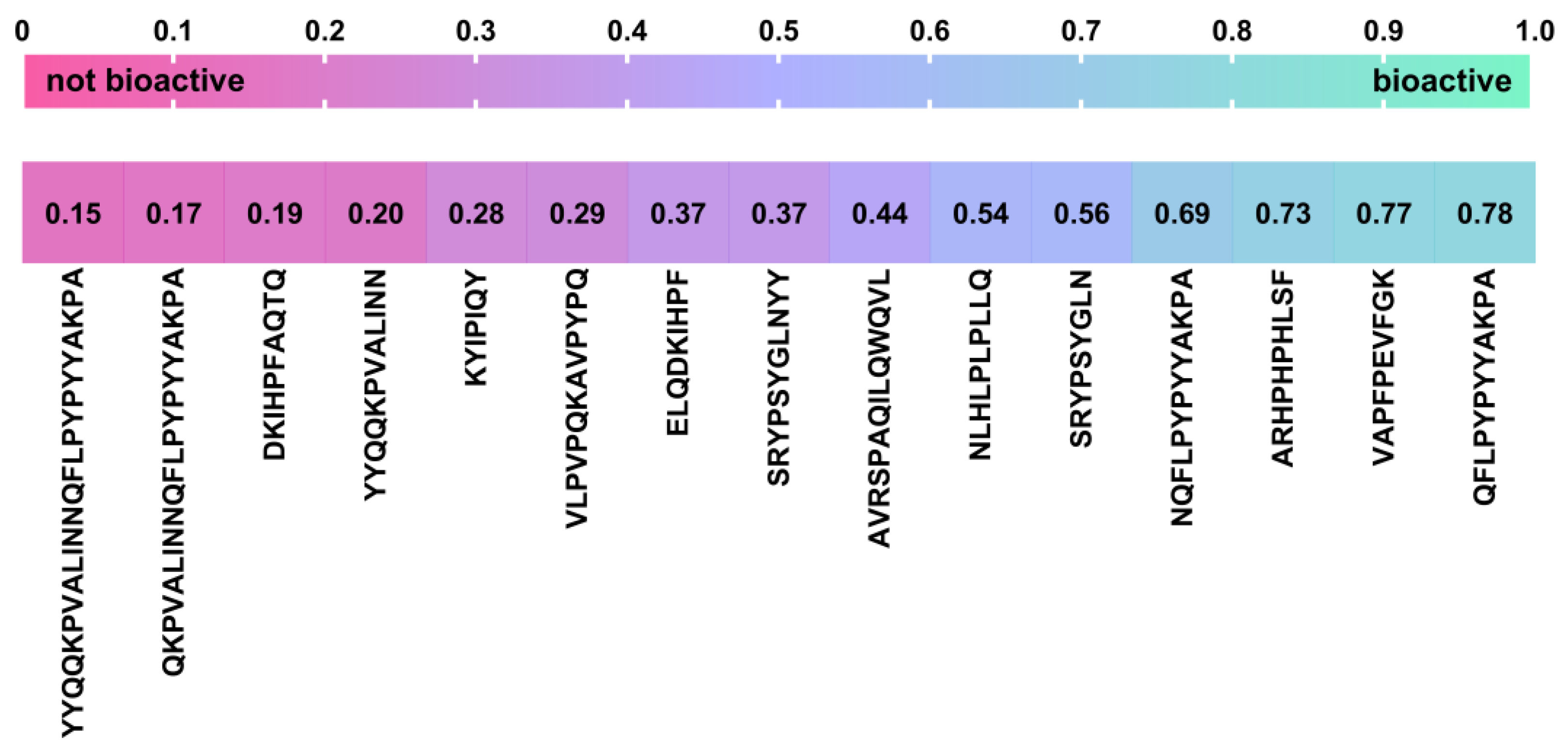
2.5. In Silico Identification of Potential Novel BPs Released during Microbial Fermentation of Bovine Milk by Strains ZGBP5-51, ZGBP5-52 and ZGBP5-53
3. Materials and Methods
3.1. Bacterial Strains and Cultivation Conditions
3.2. WGS and Genome Annotation of Bacterial Strains
3.3. Preparation of the Functional Fermented BP-Rich Lyophilised Dairy Product
3.4. Peptidomic Profiling of BP-Rich Lyophilised Dairy Product
3.4.1. Non-Targeted Peptide Analysis (NTA)
Preparation of the Samples
RP-Nano LC Workflow
MALDI-TOF/TOF MS
Peptide Identification
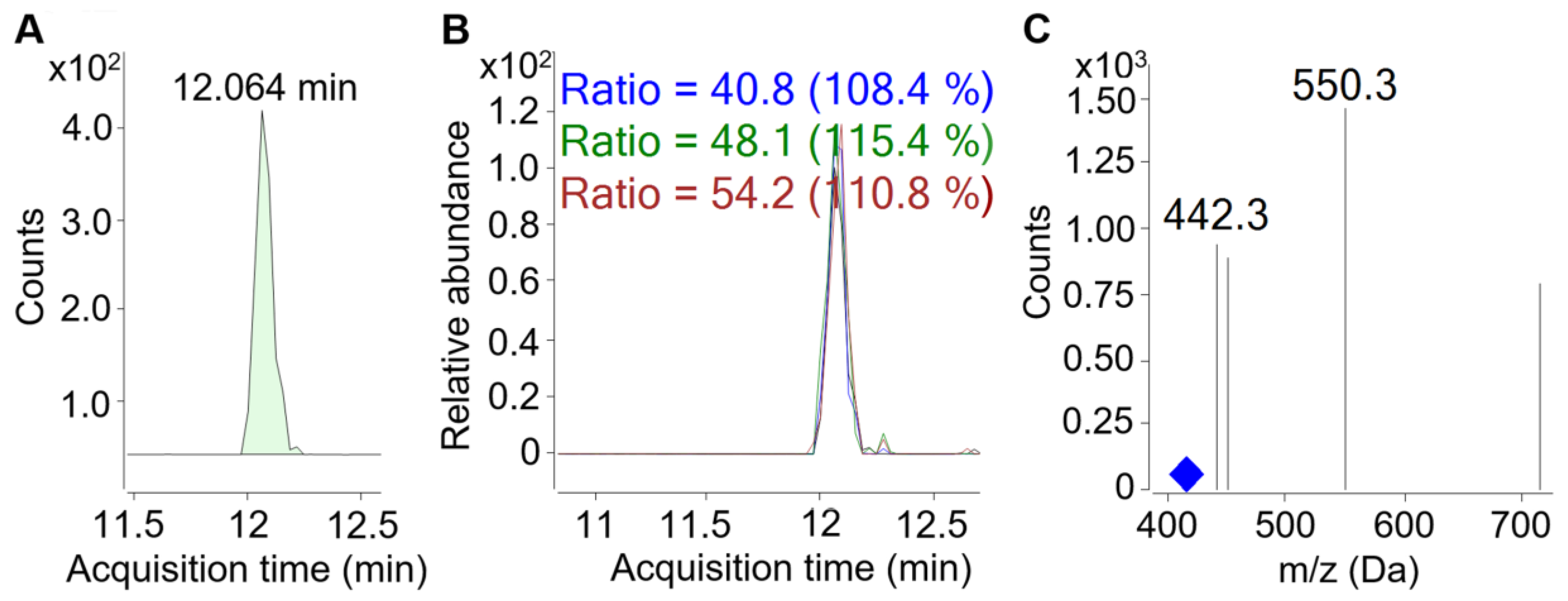
3.4.2. Targeted Peptide Analysis (TA) Using MS-Based MRM Method
3.5. In Silico Analyses
3.5.1. In Silico Analysis of Peptides Released from Bovine Milk Fermented by Strains ZGBP5-51, ZGBP5-52 and ZGBP5-53
3.5.2. In Silico Identification of Potential Novel BPs Released during the Fermentation of Bovine Milk by the Strains Lc. lactis subsp. lactis ZGBP5-51, E. faecium ZGBP5-52 and E. faecalis ZGBP5-53
3.6. Graphical Representation and Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Akbarian, M.; Khani, A.; Eghbalpour, S.; Uversky, V.N. Bioactive peptides: Synthesis, sources, applications, and proposed mechanisms of action. Int. J. Mol. Sci. 2022, 23, 1445. [Google Scholar] [CrossRef]
- Lemaire, M.; Ménard, O.; Cahu, A.; Nogret, I.; Briard-Bion, V.; Cudennec, B.; Cuinet, I.; Le Ruyet, P.; Baudry, C.; Dupont, D.; et al. Addition of dairy lipids and probiotic Lactobacillus fermentum in infant formulas modulates proteolysis and lipolysis with moderate consequences on gut physiology and metabolism in Yucatan piglets. Front. Nutr. 2021, 8, 615248. [Google Scholar] [CrossRef]
- Qiao, M.; Tu, M.; Wang, Z.; Mao, F.; Chen, H.; Qin, L.; Du, M. Identification and antithrombotic activity of peptides from blue mussel (Mytilus edulis) protein. Int. J. Mol. Sci. 2018, 19, 138. [Google Scholar] [CrossRef] [PubMed]
- Admassu, H.; Gasmalla, M.A.A.; Yang, R.; Zhao, W. Identification of bioactive peptides with α-amylase inhibitory potential from enzymatic protein hydrolysates of red seaweed (Porphyra spp). J. Agric. Food. Chem. 2018, 66, 4872–4882. [Google Scholar] [CrossRef]
- Daliri, E.B.; Oh, D.H.; Lee, B.H. Bioactive peptides. Foods 2017, 6, 32. [Google Scholar] [CrossRef] [PubMed]
- Sánchez, A.; Vázquez, A. Bioactive peptides: A review. Food Qual. Saf. 2017, 1, 29–46. [Google Scholar] [CrossRef]
- Tadesse, S.A.; Emire, S.A. Production and processing of antioxidant bioactive peptides: A driving force for the functional food market. Heliyon 2020, 6, e04765. [Google Scholar] [CrossRef] [PubMed]
- Hayes, M.; Stanton, C.; Fitzgerald, G.F.; Ross, R.P. Putting microbes to work: Dairy fermentation, cell factories and bioactive peptides. Part II: Bioactive peptide functions. Biotechnol. J. 2007, 2, 435–449. [Google Scholar] [CrossRef] [PubMed]
- Ali, S.M.; Salem, F.E.; Aboulwafa, M.M.; Shawky, R.M. Hypolipidemic activity of lactic acid bacteria: Adjunct therapy for potential probiotics. PLoS ONE 2022, 17, e0269953. [Google Scholar] [CrossRef] [PubMed]
- Raveschot, C.; Deracinois, B.; Bertrand, E.; Flahaut, C.; Frémont, M.; Drider, D.; Dhulster, P.; Cudennec, B.; Coutte, F. Integrated continuous bioprocess development for ACE-Inhibitory peptide production by Lactobacillus helveticus strains in membrane bioreactor. Front. Bioeng. Biotechnol. 2020, 8, 585815. [Google Scholar] [CrossRef]
- Almeida-Santos, A.C.; Novais, C.; Peixe, L.; Freitas, A.R. Enterococcus spp. as a producer and target of bacteriocins: A double-edged sword in the antimicrobial resistance crisis context. Antibiotics 2021, 10, 1215. [Google Scholar] [CrossRef]
- Dapkevicius, M.L.E.; Sgardioli, B.; Câmara, S.P.A.; Poeta, P.; Malcata, F.X. Current trends of enterococci in dairy products: A comprehensive review of their multiple roles. Foods 2021, 10, 821. [Google Scholar] [CrossRef]
- Novak, J.; Leboš Pavunc, A.; Butorac, K.; Banić, M.; Čuljak, N.; Rak, H.; Blažević, M.; Iveljić, A.-M.; Šušković, J.; Kos, B. Caseinolytic proteases of Lactobacillus and Lactococcus isolated from fermented dairy products. Mljekarstvo 2022, 72, 11–21. [Google Scholar] [CrossRef]
- Beganović, J.; Kos, B.; Leboš Pavunc, A.; Uroić, K.; Džidara, P.; Šušković, J. Proteolytic activity of probiotic strain Lactobacillus helveticus M92. Anaerobe 2013, 20, 58–64. [Google Scholar] [CrossRef] [PubMed]
- Van Tyne, D.; Gilmore, M.S. Friend turned foe: Evolution of enterococcal virulence and antibiotic resistance. Ann. Rev. Microbiol. 2014, 68, 337–356. [Google Scholar] [CrossRef] [PubMed]
- Golić, N.; Čadež, N.; Terzić-Vidojević, A.; Šuranská, H.; Beganović, J.; Lozo, J.; Kos, B.; Šušković, J.; Raspor, P.; Topisirović, L. Evaluation of lactic acid bacteria and yeast diversity in traditional white-pickled and fresh soft cheeses from mountain regions in Serbia and low laying regions in Croatia. Int. J. Food Microbiol. 2013, 166, 294–300. [Google Scholar] [CrossRef] [PubMed]
- Krawczyk, B.; Wityk, P.; Gałęcka, M.; Michalik, M. The many faces of Enterococcus spp.—Commensal, probiotic and opportunistic pathogen. Microorganisms 2021, 9, 1900. [Google Scholar] [CrossRef] [PubMed]
- García-Díez, J.; Saraiva, C. Use of starter cultures in foods from animal origin to improve their safety. Int. J. Environ. Res. Public Health 2021, 18, 2544. [Google Scholar] [CrossRef] [PubMed]
- Baccouri, O.; Boukerb, A.M.; Farhat, L.B.; Zébré, A.; Zimmermann, K.; Domann, E.; Cambronel, M.; Barreau, M.; Maillot, O.; Rincé, I.; et al. Probiotic potential and safety evaluation of Enterococcus faecalis OB14 and OB15, isolated from traditional Tunisian Testouri cheese and Rigouta, using physiological and genomic analysis. Front. Microbiol. 2019, 10, 881. [Google Scholar] [CrossRef] [PubMed]
- Franz, C.M.; Huch, M.; Abriouel, H.; Holzapfel, W.; Gálvez, A. Enterococci as probiotics and their implications in food safety. Int. J. Food Microbiol. 2011, 151, 125–140. [Google Scholar] [CrossRef]
- Allen, S.J.; Martinez, E.G.; Gregorio, G.V.; Dans, L.F. Probiotics for treating acute infectious diarrhoea. Cochrane Database Syst. Rev. 2010, 2010, CD003048. [Google Scholar] [CrossRef]
- Bhardwaj, A.; Malik, R.K.; Chauhan, P. Functional and safety aspects of enterococci in dairy foods. Indian J. Microbiol. 2008, 48, 317–325. [Google Scholar] [CrossRef]
- Giraffa, G. Functionality of enterococci in dairy products. Int. J. Food Microbiol. 2003, 88, 215–222. [Google Scholar] [CrossRef]
- Oumer, A.; Garde, S.; Gaya, P.; Medina, M.; Nuñez, M. The effects of cultivating lactic starter cultures with bacteriocin-producing lactic acid bacteria. J. Food Prot. 2001, 64, 81–86. [Google Scholar] [CrossRef]
- Centeno, J.A.; Menendez, S.; Hermida, M.; Rodríguez-Otero, J.L. Effects of the addition of Enterococcus faecalis in Cebreiro cheese manufacture. Int. J. Food Microbiol. 1999, 48, 97–111. [Google Scholar] [CrossRef]
- BCC Research: Global Markets Reports and Industry Analysis. Functional Foods and Beverages: Global Markets. (Report code FOD100B). Available online: https://www.bccresearch.com/market-research/food-and-beverage/functional-food-market.html (accessed on 12 September 2023).
- Kayaoglu, G.; Ørstavik, D. Virulence factors of Enterococcus faecalis: Relationship to endodontic disease. Crit. Rev. Oral Biol. Med. 2004, 15, 308–320. [Google Scholar] [CrossRef] [PubMed]
- Hollenbeck, B.L.; Rice, L.B. Intrinsic and acquired resistance mechanisms in Enterococcus. Virulence 2012, 3, 421–433. [Google Scholar] [CrossRef]
- Hellinger, R.; Sigurdsson, A.; Wu, W.; Romanova, E.V.; Li, L.; Sweedler, J.; Süssmuth, R.D.; Gruber, C.W. Peptidomics. Nat. Rev. Methods Primers 2023, 3, 25. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.P.; Dias, F.F.G.; Leite Nobrega de Moura Bell, J.M.; Barile, D. A complete workflow for discovering small bioactive peptides in foods by LC-MS/MS: A case study on almonds. Food Chem. 2022, 369, 130834. [Google Scholar] [CrossRef] [PubMed]
- Madsen, C.T.; Refsgaard, J.C.; Teufel, F.G.; Kjærulff, S.K.; Wang, Z.; Meng, G.; Jessen, C.; Heljo, P.; Jiang, Q.; Zhao, X.; et al. Combining mass spectrometry and machine learning to discover bioactive peptides. Nat. Commun. 2022, 13, 6235. [Google Scholar] [CrossRef]
- Daroit, D.J.; Brandelli, A. In vivo bioactivities of food protein-derived peptides-a current review. Curr. Opin. Food Sci. 2021, 39, 120–129. [Google Scholar] [CrossRef]
- Albenzio, M.; Santillo, A.; Caroprese, M.; Della Malva, A.; Marino, R. Bioactive peptides in animal food products. Foods 2017, 6, 35. [Google Scholar] [CrossRef] [PubMed]
- Schulze, B.; Jeon, Y.; Kaserzon, S.; Heffernan, A.; Dewapriya, P.; O’Brien, J.; Gomez Ramos, M.; Ghorbani Gorji, S.; Mueller, J.; Thomas, K.; et al. An assessment of quality assurance/quality control efforts in high resolution mass spectrometry non-target workflows for analysis of environmental samples. TrAC 2020, 133, 116063. [Google Scholar] [CrossRef]
- Thomas, C.E.; Sexton, W.; Benson, K.; Sutphen, R.; Koomen, J. Urine collection and processing for protein biomarker discovery and quantification. Cancer Epidemiol. Biomark. 2010, 19, 953–959. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, H.R.; Ahmed, A.S.; Miyata, T. Novel angiotensin-converting enzyme inhibitory peptides from caseins and whey proteins of goat milk. J. Adv. Res. 2017, 8, 63–71. [Google Scholar] [CrossRef] [PubMed]
- Tonolo, F.; Sandre, M.; Ferro, S.; Folda, A.; Scalcon, V.; Scutari, G.; Feller, E.; Marin, O.; Bindoli, A.; Rigobello, M.P. Milk-derived bioactive peptides protect against oxidative stress in a Caco-2 cell model. Food Funct. 2018, 9, 1245–1253. [Google Scholar] [CrossRef]
- Abdel-Hamid, M.; Romeih, E.; Gamba, R.R.; Nagai, E.; Suzuki, T.; Koyanagi, T.; Enomoto, T. The biological activity of fermented milk produced by Lactobacillus casei ATCC 393 during cold storage. Int. Dairy J. 2019, 91, 1–8. [Google Scholar] [CrossRef]
- Begunova, A.V.; Savinova, O.S.; Glazunova, O.A.; Moiseenko, K.V.; Rozhkova, I.V.; Fedorova, T.V. Development of antioxidant and antihypertensive properties during growth of Lactobacillus helveticus, Lactobacillus rhamnosus and Lactobacillus reuteri on cow’s milk: Fermentation and peptidomics study. Foods 2020, 10, 17. [Google Scholar] [CrossRef]
- Tonolo, F.; Fiorese, F.; Moretto, L.; Folda, A.; Scalcon, V.; Grinzato, A.; Ferro, S.; Arrigoni, G.; Bindoli, A.; Feller, E.; et al. Identification of new peptides from fermented milk showing antioxidant properties: Mechanism of action. Antioxidants 2020, 9, 117. [Google Scholar] [CrossRef]
- Alvarez-Ordóñez, A.; Begley, M.; Clifford, T.; Deasy, T.; Considine, K.; Hill, C. Structure-activity relationship of synthetic variants of the milk-derived antimicrobial peptide αs2-casein f(183–207). Appl. Environ. Microbiol. 2013, 79, 5179–5185. [Google Scholar] [CrossRef]
- Eisele, T.; Stressler, T.; Kranz, B.; Fischer, L. Bioactive peptides generated in an enzyme membrane reactor using Bacillus lentus alkaline peptidase. Eur. Food Res. Technol. 2013, 236, 483–490. [Google Scholar] [CrossRef]
- Boutrou, R.; Gaudichon, C.; Dupont, D.; Jardin, J.; Airinei, G.; Marsset-Baglieri, A.; Benamouzig, R.; Tomé, D.; Leonil, J. Sequential release of milk protein-derived bioactive peptides in the jejunum in healthy humans. Am. J. Clin. Nutr. 2013, 97, 1314–1323. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, N.; Akino, A.; Takano, T. Effect of the peptides derived from casein by an extracellular proteinase from Lactobacillus helveticus CP790. J. Dairy Sci. 1994, 77, 917–922. [Google Scholar] [CrossRef] [PubMed]
- Birkemo, G.A.; O’Sullivan, O.; Ross, R.P.; Hill, C. Antimicrobial activity of two peptides casecidin 15 and 17, found naturally in bovine colostrum. J. Appl. Microbiol. 2009, 106, 233–240. [Google Scholar] [CrossRef] [PubMed]
- Rojas-Ronquillo, R.; Cruz-Guerreroa, A.; Flores-Nájeraa, A.; Rodríguez-Serranoa, G.; Gómez-Ruiza, L.; Reyes-Grajedab, J.P.; Jiménez-Guzmána, J.; García-Garibay, M. Antithrombotic and angiotensin-converting enzyme inhibitory properties of peptides released from bovine casein by Lactobacillus casei Shirota. Int. Dairy J. 2012, 26, 147–154. [Google Scholar] [CrossRef]
- Sandré, C.; Gleizes, A.; Forestier, F.; Gorges-Kergot, R.; Chilmonczyk, S.; Léonil, J.; Moreau, M.-C.; Labarre, C. A peptide derived from bovine β-casein modulates functional properties of bone marrow-derived macrophages from germfree and human flora-associated mice. J. Nutr. 2001, 131, 2936–2942. [Google Scholar] [CrossRef] [PubMed]
- Sah, B.; Vasiljevic, T.; Mckechnie, S.; Donkor, O. Antioxidant peptides isolated from synbiotic yoghurt exhibit antiproliferative activities against HT-29 colon cancer cells. Int. Dairy J. 2016, 63, 99–106. [Google Scholar] [CrossRef]
- Lu, Y.; Govindasamy-Lucey, S.; Lucey, J.A. Angiotensin-I-converting enzyme-inhibitory peptides in commercial Wisconsin Cheddar cheeses of different ages. J Dairy Sci. 2016, 99, 41–52. [Google Scholar] [CrossRef]
- Elfahri, K. Anticarcinogenic Peptides Released from Milk Proteins by Lactobacillus Strains. Doctoral Dissertation, Victoria University, Victoria, Australia, 2018. [Google Scholar]
- Coste, M.; Rochet, V.; Léonil, J.; Mollé, D.; Bouhallab, S.; Tomé, D. Identification of C-terminal peptides of bovine beta-casein that enhance proliferation of rat lymphocytes. Immunol. Lett. 1992, 33, 41–46. [Google Scholar] [CrossRef]
- Fan, H.; Wang, J.; Liao, W.; Jiang, X.; Wu, J. Identification and characterization of gastrointestinal-resistant angiotensin-converting enzyme inhibitory peptides from egg white proteins. J. Agric. Food Chem. 2019, 67, 7147–7156. [Google Scholar] [CrossRef]
- Ong, L.; Shah, N.P. Release and identification of angiotensin-converting enzyme-inhibitory peptides as influenced by ripening temperatures and probiotic ad juncts in Cheddar cheeses. Food Sci. Technol. 2008, 41, 1555–1566. [Google Scholar] [CrossRef]
- Pihlanto-Leppälä, A.; Koskinen, P.; Piilola, K.; Tupasela, T.; Korhonen, H. Angiotensin I-converting enzyme inhibitory properties of whey protein digests: Concentration and characterization of active peptides. J. Dairy Res. 2000, 67, 53–64. [Google Scholar] [CrossRef] [PubMed]
- Sagardia, I.; Roa-Ureta, R.H.; Bald, C. A new QSAR model, for angiotensin I-converting enzyme inhibitory oligopeptides. Food Chem. 2013, 136, 1370–1376. [Google Scholar] [CrossRef]
- Caira, S.; Pinto, G.; Vitaglione, P.; Dal Piaz, F.; Ferranti, P.; Addeo, F. Identification of casein peptides in plasma of subjects after a cheese-enriched diet. Food Res. Int. 2016, 84, 108–112. [Google Scholar] [CrossRef]
- Snezhkina, A.V.; Kudryavtseva, A.V.; Kardymon, O.L.; Savvateeva, M.V.; Melnikova, N.V.; Krasnov, G.S.; Dmitriev, A.A. ROS generation and antioxidant defense systems in normal and malignant cells. Oxid. Med. Cell. Longev. 2019, 2019, 6175804. [Google Scholar] [CrossRef]
- Zhu, K.; Jia, H.; Sun, Y.; Dai, Y.; Zhang, C.; Guo, X.; Wang, T.; Zhu, L. Long-term phototransformation of microplastics under simulated sunlight irradiation in aquatic environments: Roles of reactive oxygen species. Water Res. 2020, 173, 115564. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Liu, A.; Hu, S.; Ares, I.; Martínez-Larrañaga, M.R.; Wang, X.; Martínez, M.; Anadón, A.; Martínez, M.A. Synthetic phenolic antioxidants: Metabolism, hazards and mechanism of action. Food Chem. 2021, 353, 129488. [Google Scholar] [CrossRef]
- Lafarga, T.; Sánchez-Zurano, A.; Villaró, S.; Morillas-España, A.; Acién, G. Industrial production of spirulina as a protein source for bioactive peptide generation. Trends Food Sci. Technol. 2021, 116, 176–185. [Google Scholar] [CrossRef]
- Samaei, S.P.; Ghorbani, M.; Tagliazucchi, D.; Martini, S.; Gotti, R.; Themelis, T.; Tesini, F.; Gianotti, A.; Gallina Toschi, T.; Babini, E. Functional, nutritional, antioxidant, sensory properties and comparative peptidomic profile of faba bean (Vicia faba, L.) seed protein hydrolysates and fortified apple juice. Food Chem. 2020, 330, 127120. [Google Scholar] [CrossRef]
- Sonklin, C.; Laohakunjit, N.; Kerdchoechuen, O. Assessment of antioxidant properties of membrane ultrafiltration peptides from mungbean meal protein hydrolysates. PeerJ 2018, 6, e5337. [Google Scholar] [CrossRef] [PubMed]
- Cheng, S.; Tu, M.; Liu, H.; Zhao, G.; Du, M. Food-derived antithrombotic peptides: Preparation, identification, and interactions with thrombin. Crit. Rev. Food Sci. Nutr. 2019, 59 (Suppl. S1), S81–S95. [Google Scholar] [CrossRef] [PubMed]
- Pavličević, M.; Marmiroli, N.; Maestri, E. Immunomodulatory peptides-A promising source for novel functional food production and drug discovery. Peptides 2022, 148, 170696. [Google Scholar] [CrossRef] [PubMed]
- Silano, M.; Di Benedetto, R.; Trecca, A.; Arrabito, G.; Leonardi, F.; De Vincenzi, M. A decapeptide from durum wheat prevents celiac peripheral blood lymphocytes from activation by gliadin peptides. Pediatr. Res. 2007, 61, 67–71. [Google Scholar] [CrossRef] [PubMed]
- Tyagi, A.; Daliri, E.B.; Kwami Ofosu, F.; Yeon, S.J.; Oh, D.H. Food-derived opioid peptides in human health: A review. Int. J. Mol. Sci. 2020, 21, 8825. [Google Scholar] [CrossRef]
- Abd-Talib, N.; Yaji, E.L.A.; Wahab, N.S.A.; Razali, N.; Len, K.Y.T.; Roslan, J.; Saari, N.; Pa’ee, K.F. Bioactive peptides and its alternative processes: A review. Biotechnol. Bioproc. E. 2022, 27, 306–335. [Google Scholar] [CrossRef]
- Liu, B.; Li, N.; Chen, F.; Zhang, J.; Sun, X.; Xu, L.; Fang, F. Review on the release mechanism and debittering technology of bitter peptides from protein hydrolysates. Compr. Rev. Food Sci. Food Saf. 2022, 21, 5153–5170. [Google Scholar] [CrossRef]
- Charoenkwan, P.; Nantasenamat, C.; Hasan, M.M.; Manavalan, B.; Shoombuatong, W. BERT4Bitter: A bidirectional encoder representations from transformers (BERT)-based model for improving the prediction of bitter peptides. Bioinformatics 2021, 37, 2556–2562. [Google Scholar] [CrossRef]
- Charoenkwan, P.; Nantasenamat, C.; Hasan, M.M.; Moni, M.A.; Lio’, P.; Shoombuatong, W. iBitter-Fuse: A novel sequence-based bitter peptide predictor by fusing multi-view features. Int. J. Mol. Sci. 2021, 22, 8958. [Google Scholar] [CrossRef]
- Sarabandi, K.; Mahoonak, A.S.; Hamishehkar, H.; Ghorbani, M.; Jafari, S.M. Protection of casein hydrolysates within nanoliposomes: Antioxidant and stability characterization. J. Food Eng. 2019, 251, 19–28. [Google Scholar] [CrossRef]
- Giroldi, M.; Grambusch, I.M.; Lehn Neutzling, D.; Volken de Souza, C.F. Encapsulation of dairy protein hydrolysates: Recent trends and future prospects. Dry. Technol. 2021, 39, 1513–1528. [Google Scholar] [CrossRef]
- Korhonen, H.; Pihlanto, A. Bioactive peptides: Production and functionality. Int. Dairy J. 2006, 16, 945–960. [Google Scholar] [CrossRef]
- Kyte, J.; Doolittle, R.F. A simple method for displaying the hydropathic character of a protein. J. Mol. Biol. 1982, 157, 105–132. [Google Scholar] [CrossRef] [PubMed]
- Cid, H.; Bunster, M.; Canales, M.; Gazitúa, F. Hydrophobicity and structural classes in proteins. Protein Eng. 1992, 5, 373–375. [Google Scholar] [CrossRef]
- Okella, H.; Okello, E.; Mtewa, A.G.; Ikiriza, H.; Kaggwa, B.; Aber, J.; Ndekezi, C.; Nkamwesiga, J.; Ajayi, C.O.; Mugeni, I.M.; et al. ADMET profiling and molecular docking of potential antimicrobial peptides previously isolated from African catfish, Clarias gariepinus. Front. Mol. Biosci. 2022, 9, 1039286. [Google Scholar] [CrossRef] [PubMed]
- Tunkel, A.R.; Rosser, S.W.; Hansen, E.J.; Scheld, W.M. Blood-brain barrier alterations in bacterial meningitis: Development of an in vitro model and observations on the effects of lipopolysaccharide. In Vitro Cell Dev. Biol. 1991, 27A, 113–120. [Google Scholar] [CrossRef]
- Di, L.; Breen, C.; Chambers, R.; Eckley, S.T.; Fricke, R.; Ghosh, A.; Harradine, P.; Kalvass, J.C.; Ho, S.; Lee, C.A.; et al. Industry perspective on contemporary protein-binding methodologies: Considerations for regulatory drug-drug interaction and related guidelines on highly bound drugs. J. Pharm. Sci. 2017, 106, 3442–3452. [Google Scholar] [CrossRef]
- Cao, H.; Liu, X.; Ulrih, N.P.; Sengupta, P.K.; Xiao, J. Plasma protein binding of dietary polyphenols to human serum albumin: A high performance affinity chromatography approach. Food Chem. 2019, 270, 257–263. [Google Scholar] [CrossRef]
- Lagorce, D.; Bouslama, L.; Becot, J.; Miteva, M.A.; Villoutreix, B.O. FAF-Drugs4: Free ADME-tox filtering computations for chemical biology and early stages drug discovery. Bioinformatics 2017, 33, 3658–3660. [Google Scholar] [CrossRef]
- Agyei, D.; Tsopmo, A.; Udenigwe, C.C. Bioinformatics and peptidomics approaches to the discovery and analysis of food-derived bioactive peptides. Anal. Bioanal. Chem. 2018, 410, 3463–3472. [Google Scholar] [CrossRef]
- Rodrigues, C.H.M.; Garg, A.; Keizer, D.; Pires, D.E.V.; Ascher, D.B. CSM-peptides: A computational approach to rapid identification of therapeutic peptides. Protein Sci. 2022, 31, e4442. [Google Scholar] [CrossRef]
- Mooney, C.; Haslam, N.J.; Pollastri, G.; Shields, D.C. Towards the improved discovery and design of functional peptides: Common features of diverse classes permit generalized prediction of bioactivity. PLoS ONE 2012, 7, e45012. [Google Scholar] [CrossRef]
- Popović, N.; Djokić, J.; Brdarić, E.; Dinić, M.; Terzić-Vidojević, A.; Golić, N.; Veljović, K. The Influence of heat-killed Enterococcus faecium BGPAS1-3 on the tight junction protein expression and immune function in differentiated Caco-2 Cells infected with Listeria monocytogenes ATCC 19111. Front. Microbiol. 2019, 10, 412. [Google Scholar] [CrossRef]
- Banić, M.; Uroić, K.; Leboš Pavunc, A.; Novak, J.; Zorić, K.; Durgo, K.; Petković, H.; Jamnik, P.; Kazazić, S.; Kazazić, S.; et al. Characterization of S-layer proteins of potential probiotic starter culture Lactobacillus brevis SF9B isolated from sauerkraut. LWT-Food Sci. Technol. 2018, 93, 257–267. [Google Scholar] [CrossRef]
- Del Fabbro, C.; Scalabrin, S.; Morgante, M.; Giorgi, F.M. An extensive evaluation of read trimming effects on Illumina NGS data analysis. PLoS ONE 2013, 8, e85024. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef]
- Olson, R.D.; Assaf, R.; Brettin, T.; Conrad, N.; Cucinell, C.; Davis, J.J.; Dempsey, D.M.; Dickerman, A.; Dietrich, E.M.; Kenyon, R.W.; et al. Introducing the Bacterial and Viral Bioinformatics Resource Center (BV-BRC): A resource combining PATRIC, IRD and ViPR. Nucleic Acids Res. 2023, 51, D678–D689. [Google Scholar] [CrossRef]
- Leboš Pavunc, A.; Turk, J.; Kos, B.; Beganović, J.; Frece, J.; Mahnet, S.; Kirin, S.; Šušković, J. Proizvodnja fermentiranih probiotičkih napitaka od permeata mlijeka obogaćenih retentatom sirutke i identifikacija prisutnih bakterija mliječne kiseline. Mljekarstvo 2009, 59, 11–19. [Google Scholar]
- Novak, J.; Butorac, K.; Leboš Pavunc, A.; Banić, M.; Butorac, A.; Lepur, A.; Oršolić, N.; Tonković, K.; Bendelja, K.; Čuljak, N.; et al. A lactic acid bacteria consortium impacted the content of casein-derived biopeptides in dried fresh cheese. Molecules 2021, 27, 160. [Google Scholar] [CrossRef]
- MacLean, B.; Tomazela, D.M.; Shulman, N.; Chambers, M.; Finney, G.L.; Frewen, B.; Kern, R.; Tabb, D.L.; Liebler, D.C.; MacCoss, M.J. Skyline: An open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 2010, 26, 966–968. [Google Scholar] [CrossRef]
- Nielsen, S.D.; Beverly, R.L.; Qu, Y.; Dallas, D.C. Milk bioactive peptide database: A comprehensive database of milk protein-derived bioactive peptides and novel visualization. Food Chem. 2017, 232, 673–682. [Google Scholar] [CrossRef]
- Qin, D.; Bo, W.; Zheng, X.; Hao, Y.; Li, B.; Zheng, J.; Liang, G. DFBP: A comprehensive database of food-derived bioactive peptides for peptidomics research. Bioinformatics 2022, 38, 3275–3280. [Google Scholar] [CrossRef] [PubMed]
- Minkiewicz, P.; Iwaniak, A.; Darewicz, M. BIOPEP-UWM Database of bioactive peptides: Current opportunities. Int. J. Mol. Sci. 2019, 20, 5978. [Google Scholar] [CrossRef]
- Xiong, G.; Wu, Z.; Yi, J.; Fu, L.; Yang, Z.; Hsieh, C.; Yin, M.; Zeng, X.; Wu, C.; Lu, A.; et al. ADMETlab 2.0: An integrated online platform for accurate and comprehensive predictions of ADMET properties. Nucleic Acids Res. 2021, 49, W5–W14. [Google Scholar] [CrossRef]





| AA Composition | Chemical Formula | Physiochemical Properties | ||||
|---|---|---|---|---|---|---|
| GRAVY | Net Charge (pH = 7) | pI | MW [Da] | ε [L/mol cm] | ||
| H-TKVIPYVRYL-OH | C61H98N14O14 | 0.34 | 2.00 | 10.04 | 1251.51 | 2560 |
| H-IQPKTKVIPYVRYL-OH | C83H136N20O19 | −0.08 | 3.00 | 10.34 | 1718.08 | 2560 |
| H-LYQEPVLGPVRGPFPIIV-OH | C97H152N22O23 | 0.67 | 0.00 | 7.00 | 1994.37 | 1280 |
| H-KYIPIQYVL-OH | C57H89N11O13 | 0.60 | 1.00 | 9.52 | 1136.38 | 2560 |
| H-ARHPHPHLSFM-OH | C60H88N20O13S | −0.71 | 1.30 | 11.05 | 1329.53 | 0 |
| H-SQSKVLPVPQKAVPYPQ-OH | C86H140N22O24 | −0.55 | 2.00 | 10.18 | 1866.16 | 1280 |
| H-RDMPIQAF-OH | C43H68N12O12S | −0.26 | 0.00 | 7.00 | 977.13 | 0 |
| H-QEPVLGPVRGPFPIIV-OH | C82H132N20O20 | 0.59 | 0.00 | 7.00 | 1718.04 | 0 |
| H-YYQQKPVALINN-OH | C67H103N17O19 | −0.65 | 1.00 | 9.52 | 1450.63 | 2560 |
| H-YYQQKPVALINNQFLPYPYYAKPA-OH | C141H201N31O35 | −0.57 | 2.00 | 9.57 | 2890.28 | 6400 |
| H-QFLPYPYYAKPA-OH | C74H100N14O17 | −0.49 | 1.00 | 9.39 | 1457.66 | 3840 |
| H-VAPFPEVFGK-OH | C54H79N11O13 | 0.48 | 0.00 | 7.00 | 1090.27 | 0 |
| H-AVRSPAQILQWQVL-OH | C74H121N21O19 | 0.41 | 1.00 | 11.05 | 1608.87 | 5690 |
| H-ELQDKIHPF-OH | C52H79N13O15 | −0.90 | −0.90 | 5.22 | 1126.26 | 0 |
| H-DKIHPFAQTQ-OH | C53H81N15O16 | −1.08 | 0.10 | 7.88 | 1184.29 | 0 |
| H-NLHLPLPLLQ-OH | C55H92N14O13 | 0.56 | 0.10 | 7.88 | 1157.40 | 0 |
| H-VLPVPQKAVPYPQ-OH | C69H110N16O17 | −0.03 | 1.00 | 9.72 | 1435.70 | 1280 |
| H-KYIPIQY-OH | C46H69N9O11 | −0.37 | 1.00 | 9.52 | 924.09 | 2560 |
| H-YQEPVLGPVRGPFPIIV-OH | C91H141N21O22 | 0.48 | 0.00 | 7.00 | 1881.21 | 1280 |
| H-SRYPSYGLN-OH | C47H69N13O15 | −1.16 | 1.00 | 9.58 | 1056.12 | 2560 |
| H-SRYPSYGLNYY-OH | C65H87N15O19 | −1.18 | 1.00 | 9.32 | 1382.47 | 5120 |
| H-QKPVALINNQFLPYPYYAKPA-OH | C118H175N27O29 | −0.36 | 2.00 | 9.78 | 2435.80 | 3840 |
| H-NQFLPYPYYAKPA-OH | C78H106N16O19 | −0.72 | 1.00 | 9.39 | 1571.76 | 3840 |
| H-VYPFPGPIPN-OH | C55H77N11O13 | −0.01 | 0.00 | 7.00 | 1100.26 | 1280 |
| H-ARHPHPHLSF-OH | C55H79N19O12 | −0.97 | 1.30 | 11.05 | 1198.33 | 0 |
| (A) 2D Structure and Physiochemical Properties of the Representative 213TKVIPYVRYL222 Peptide | ||||||||||||||||||||
| 2D structure | ||||||||||||||||||||
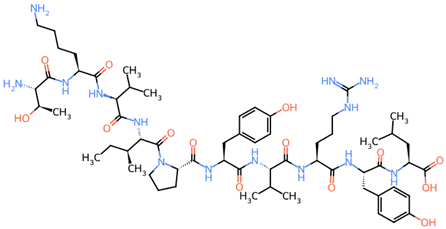 | ||||||||||||||||||||
| Radar chart of physicochemical properties | ||||||||||||||||||||
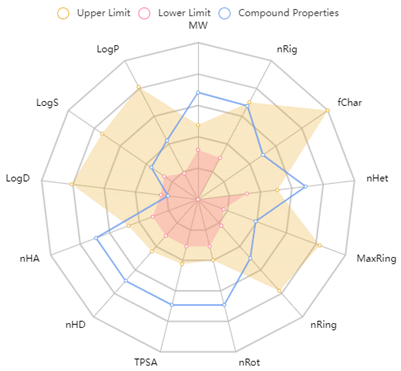 | MW | 1250.74 | ||||||||||||||||||
| nRig | 28 | |||||||||||||||||||
| fChar | 0 | |||||||||||||||||||
| nHet | 28 | |||||||||||||||||||
| MaxRing | 6 | |||||||||||||||||||
| nRing | 3 | |||||||||||||||||||
| nRot | 47 | |||||||||||||||||||
| TPSA | 465.04 | |||||||||||||||||||
| nHD | 20 | |||||||||||||||||||
| logD | 0.84 | |||||||||||||||||||
| logS | −3.06 | |||||||||||||||||||
| logP | 1.14 | |||||||||||||||||||
| (B) ADMET properties of peptides identified after fermentation of bovine milk by strains ZGBP5-51, ZGBP5-52 and ZGBP5-53. | ||||||||||||||||||||
| Peptide | PPB | VD | BBB penetration | Acute toxicity | Genotoxic carcinogenicity | Nongenotoxic carcinogenicity | Skin sensitisation | Aquatic toxicity | Nonbiodegradable rule | SureCheMBL Rule | ||||||||||
| Value [%] | Decision | Value | Decision | Value | Decision | Alerts | Decision | Alerts | Decision | Alerts | Decision | Alerts | Decision | Alerts | Decision | Alerts | Decision | Alerts | Decision | |
| 1 | 36.98 | ● | 0.55 | ● | 0.02 | ● | 0 | ● | 0 | ● | 0 | ● | 2 | ● | 0 | ● | 0 | ● | 0 | ● |
| 2 | 36.98 | ● | 0.43 | ● | 0.04 | ● | 0 | ● | 0 | ● | 0 | ● | 1 | ● | 0 | ● | 0 | ● | 0 | ● |
| 3 | 61.6 | ● | 0.62 | ● | 0.01 | ● | 0 | ● | 0 | ● | 0 | ● | 2 | ● | 0 | ● | 0 | ● | 0 | ● |
| 4 | 65.45 | ● | 0.55 | ● | 0.02 | ● | 0 | ● | 0 | ● | 0 | ● | 2 | ● | 0 | ● | 0 | ● | 0 | ● |
| 5 | 74.45 | ● | 0.64 | ● | 0.00 | ● | 0 | ● | 0 | ● | 0 | ● | 1 | ● | 0 | ● | 0 | ● | 0 | ● |
| 6 | 70.82 | ● | 0.63 | ● | 0.01 | ● | 0 | ● | 0 | ● | 0 | ● | 1 | ● | 0 | ● | 0 | ● | 0 | ● |
| 7 | 15.64 | ● | 0.53 | ● | 0.04 | ● | 0 | ● | 0 | ● | 0 | ● | 1 | ● | 0 | ● | 0 | ● | 0 | ● |
| 8 | 35.64 | ● | 0.46 | ● | 0.01 | ● | 0 | ● | 0 | ● | 0 | ● | 2 | ● | 0 | ● | 0 | ● | 0 | ● |
| 9 | 69.99 | ● | 0.48 | ● | 0.01 | ● | 0 | ● | 0 | ● | 0 | ● | 1 | ● | 0 | ● | 0 | ● | 0 | ● |
| 10 | 73.47 | ● | 0.64 | ● | 0.00 | ● | 0 | ● | 0 | ● | 0 | ● | 1 | ● | 0 | ● | 0 | ● | 0 | ● |
| 11 | 32.33 | ● | 0.59 | ● | 0.02 | ● | 0 | ● | 0 | ● | 0 | ● | 2 | ● | 0 | ● | 0 | ● | 0 | ● |
| 12 | 71.34 | ● | 0.62 | ● | 0.00 | ● | 0 | ● | 0 | ● | 0 | ● | 2 | ● | 0 | ● | 0 | ● | 0 | ● |
| 13 | 65.89 | ● | 0.58 | ● | 0.01 | ● | 0 | ● | 0 | ● | 0 | ● | 2 | ● | 0 | ● | 0 | ● | 0 | ● |
| 14 | 37.73 | ● | 0.50 | ● | 0.02 | ● | 0 | ● | 0 | ● | 0 | ● | 2 | ● | 0 | ● | 0 | ● | 0 | ● |
| 15 | 50.29 | ● | 0.50 | ● | 0.02 | ● | 0 | ● | 0 | ● | 0 | ● | 1 | ● | 0 | ● | 0 | ● | 0 | ● |
| 16 | 21.57 | ● | 0.61 | ● | 0.04 | ● | 0 | ● | 0 | ● | 0 | ● | 2 | ● | 0 | ● | 0 | ● | 0 | ● |
| 17 | 21.16 | ● | 0.47 | ● | 0.05 | ● | 0 | ● | 0 | ● | 0 | ● | 2 | ● | 0 | ● | 0 | ● | 0 | ● |
| 18 | 48.75 | ● | 0.53 | ● | 0.03 | ● | 0 | ● | 0 | ● | 0 | ● | 1 | ● | 0 | ● | 0 | ● | 0 | ● |
| 19 | 51.86 | ● | 0.63 | ● | 0.01 | ● | 0 | ● | 0 | ● | 0 | ● | 2 | ● | 0 | ● | 0 | ● | 0 | ● |
| 20 | 52.89 | ● | 0.50 | ● | 0.03 | ● | 0 | ● | 0 | ● | 0 | ● | 2 | ● | 0 | ● | 0 | ● | 0 | ● |
| 21 | 23.16 | ● | 0.52 | ● | 0.05 | ● | 0 | ● | 0 | ● | 0 | ● | 1 | ● | 0 | ● | 0 | ● | 0 | ● |
| 22 | 53.51 | ● | 0.61 | ● | 0.02 | ● | 0 | ● | 0 | ● | 0 | ● | 1 | ● | 0 | ● | 0 | ● | 0 | ● |
| 23 | 68.15 | ● | 0.59 | ● | 0.00 | ● | 0 | ● | 0 | ● | 0 | ● | 2 | ● | 0 | ● | 0 | ● | 0 | ● |
| 24 | 62.79 | ● | 0.60 | ● | 0.01 | ● | 0 | ● | 0 | ● | 0 | ● | 2 | ● | 0 | ● | 0 | ● | 0 | ● |
| 25 | 36.75 | ● | 0.41 | ● | 0.07 | ● | 0 | ● | 0 | ● | 0 | ● | 1 | ● | 0 | ● | 0 | ● | 0 | ● |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Banić, M.; Butorac, K.; Čuljak, N.; Butorac, A.; Novak, J.; Pavunc, A.L.; Rušanac, A.; Stanečić, Ž.; Lovrić, M.; Šušković, J.; et al. An Integrated Comprehensive Peptidomics and In Silico Analysis of Bioactive Peptide-Rich Milk Fermented by Three Autochthonous Cocci Strains. Int. J. Mol. Sci. 2024, 25, 2431. https://doi.org/10.3390/ijms25042431
Banić M, Butorac K, Čuljak N, Butorac A, Novak J, Pavunc AL, Rušanac A, Stanečić Ž, Lovrić M, Šušković J, et al. An Integrated Comprehensive Peptidomics and In Silico Analysis of Bioactive Peptide-Rich Milk Fermented by Three Autochthonous Cocci Strains. International Journal of Molecular Sciences. 2024; 25(4):2431. https://doi.org/10.3390/ijms25042431
Chicago/Turabian StyleBanić, Martina, Katarina Butorac, Nina Čuljak, Ana Butorac, Jasna Novak, Andreja Leboš Pavunc, Anamarija Rušanac, Željka Stanečić, Marija Lovrić, Jagoda Šušković, and et al. 2024. "An Integrated Comprehensive Peptidomics and In Silico Analysis of Bioactive Peptide-Rich Milk Fermented by Three Autochthonous Cocci Strains" International Journal of Molecular Sciences 25, no. 4: 2431. https://doi.org/10.3390/ijms25042431
APA StyleBanić, M., Butorac, K., Čuljak, N., Butorac, A., Novak, J., Pavunc, A. L., Rušanac, A., Stanečić, Ž., Lovrić, M., Šušković, J., & Kos, B. (2024). An Integrated Comprehensive Peptidomics and In Silico Analysis of Bioactive Peptide-Rich Milk Fermented by Three Autochthonous Cocci Strains. International Journal of Molecular Sciences, 25(4), 2431. https://doi.org/10.3390/ijms25042431







