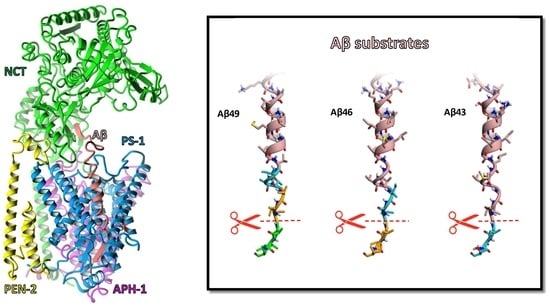
Conformational Changes and Unfolding of β-Amyloid Substrates in the Active Site of γ-Secretase
Abstract
:1. Introduction
2. Results and Discussion
3. Methods and Materials
3.1. Preparation of the Structure
3.2. Molecular Dynamics (MD) Simulations
3.3. Steered Molecular Dynamics (SMD) Simulations
3.4. Statistical Procedure and the Box-Plot
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Hardy, J.; Selkoe, D.J. The amyloid hypothesis of Alzheimer’s disease: Progress and problems on the road to therapeutics. Science 2002, 297, 353–356. [Google Scholar] [CrossRef]
- Pensalfini, A.; Albay, R., 3rd; Rasool, S.; Wu, J.W.; Hatami, A.; Arai, H.; Margol, L.; Milton, S.; Poon, W.W.; Corrada, M.M.; et al. Intracellular amyloid and the neuronal origin of Alzheimer neuritic plaques. Neurobiol. Dis. 2014, 71, 53–61. [Google Scholar] [CrossRef] [PubMed]
- Steiner, H.; Fukumori, A.; Tagami, S.; Okochi, M. Making the final cut: Pathogenic amyloid-beta peptide generation by gamma-secretase. Cell Stress 2018, 2, 292–310. [Google Scholar] [CrossRef]
- Jagust, W. Imaging the evolution and pathophysiology of Alzheimer disease. Nat. Rev. Neurosci. 2018, 19, 687–700. [Google Scholar] [CrossRef]
- Haass, C.; Selkoe, D.J. Soluble protein oligomers in neurodegeneration: Lessons from the Alzheimer’s amyloid beta-peptide. Nat. Rev. Mol. Cell Biol. 2007, 8, 101–112. [Google Scholar] [CrossRef]
- Yang, G.; Zhou, R.; Guo, X.; Yan, C.; Lei, J.; Shi, Y. Structural basis of gamma-secretase inhibition and modulation by small molecule drugs. Cell 2021, 184, 521–533.e4. [Google Scholar] [CrossRef] [PubMed]
- Feng, M.; Santhanam, R.K.; Xing, H.; Zhou, M.; Jia, H. Inhibition of gamma-secretase/Notch pathway as a potential therapy for reversing cancer drug resistance. Biochem. Pharmacol. 2024, 220, 115991. [Google Scholar] [CrossRef]
- Lichtenthaler, S.F. Alpha-secretase cleavage of the amyloid precursor protein: Proteolysis regulated by signaling pathways and protein trafficking. Curr. Alzheimer Res. 2012, 9, 165–177. [Google Scholar] [CrossRef] [PubMed]
- Lichtenthaler, S.F.; Haass, C.; Steiner, H. Regulated intramembrane proteolysis—Lessons from amyloid precursor protein processing. J. Neurochem. 2011, 117, 779–796. [Google Scholar] [CrossRef]
- Orzel, U.; Jakowiecki, J.; Mlynarczyk, K.; Filipek, S. The Role of Cholesterol in Amyloidogenic Substrate Binding to the gamma-Secretase Complex. Biomolecules 2021, 11, 935. [Google Scholar] [CrossRef]
- Sanders, C.R. How gamma-secretase hits a moving target. eLife 2016, 5, e20043. [Google Scholar] [CrossRef]
- Bolduc, D.M.; Montagna, D.R.; Seghers, M.C.; Wolfe, M.S.; Selkoe, D.J. The amyloid-beta forming tripeptide cleavage mechanism of gamma-secretase. eLife 2016, 5, e17578. [Google Scholar] [CrossRef]
- Koch, M.; Enzlein, T.; Chen, S.Y.; Petit, D.; Lismont, S.; Zacharias, M.; Hopf, C.; Chavez-Gutierrez, L. APP substrate ectodomain defines amyloid-beta peptide length by restraining gamma-secretase processivity and facilitating product release. EMBO J. 2023, 42, e114372. [Google Scholar] [CrossRef]
- Suzuki, R.; Takahashi, H.; Yoshida, C.; Hidaka, M.; Ogawa, T.; Futai, E. Specific Mutations near the Amyloid Precursor Protein Cleavage Site Increase gamma-Secretase Sensitivity and Modulate Amyloid-beta Production. Int. J. Mol. Sci. 2023, 24, 3970. [Google Scholar] [CrossRef]
- Pauli, T.M.; Julius, A.; Costa, F.; Eschrig, S.; Moosmuller, J.; Fischer, L.; Schanzenbach, C.; Schmidt, F.C.; Ortner, M.; Langosch, D. Interaction of Substrates with gamma-Secretase at the Level of Individual Transmembrane Helices-A Methodological Approach. Int. J. Mol. Sci. 2023, 24, 14396. [Google Scholar] [CrossRef]
- Watanabe, H.; Yoshida, C.; Hidaka, M.; Ogawa, T.; Tomita, T.; Futai, E. Specific Mutations in Aph1 Cause gamma-Secretase Activation. Int. J. Mol. Sci. 2022, 23, 507. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Ma, G.; Yan, X.; Li, X.; Wang, E.; Xu, X.X.; Zhao, J.B.; Ma, X.; Zeng, J. Microbial infection promotes amyloid pathology in a mouse model of Alzheimer’s disease via modulating gamma-secretase. Mol. Psychiatry 2024. [Google Scholar] [CrossRef] [PubMed]
- Zhou, R.; Yang, G.; Guo, X.; Zhou, Q.; Lei, J.; Shi, Y. Recognition of the amyloid precursor protein by human gamma-secretase. Science 2019, 363, eaaw0930. [Google Scholar] [CrossRef] [PubMed]
- Hitzenberger, M.; Zacharias, M. Structural Modeling of gamma-Secretase Abeta n Complex Formation and Substrate Processing. ACS Chem. Neurosci. 2019, 10, 1826–1840. [Google Scholar] [CrossRef]
- Bai, X.C.; Rajendra, E.; Yang, G.; Shi, Y.; Scheres, S.H. Sampling the conformational space of the catalytic subunit of human gamma-secretase. eLife 2015, 4, e11182. [Google Scholar] [CrossRef] [PubMed]
- Kukar, T.L.; Ladd, T.B.; Robertson, P.; Pintchovski, S.A.; Moore, B.; Bann, M.A.; Ren, Z.; Jansen-West, K.; Malphrus, K.; Eggert, S.; et al. Lysine 624 of the amyloid precursor protein (APP) is a critical determinant of amyloid beta peptide length: Support for a sequential model of gamma-secretase intramembrane proteolysis and regulation by the amyloid beta precursor protein (APP) juxtamembrane region. J. Biol. Chem. 2011, 286, 39804–39812. [Google Scholar] [CrossRef] [PubMed]
- Bhattarai, A.; Devkota, S.; Bhattarai, S.; Wolfe, M.S.; Miao, Y. Mechanisms of gamma-Secretase Activation and Substrate Processing. ACS Cent. Sci. 2020, 6, 969–983. [Google Scholar] [CrossRef] [PubMed]
- Bhattarai, A.; Devkota, S.; Do, H.N.; Wang, J.; Bhattarai, S.; Wolfe, M.S.; Miao, Y. Mechanism of Tripeptide Trimming of Amyloid beta-Peptide 49 by gamma-Secretase. J. Am. Chem. Soc. 2022, 144, 6215–6226. [Google Scholar] [CrossRef] [PubMed]
- Chan, R.B.; Oliveira, T.G.; Cortes, E.P.; Honig, L.S.; Duff, K.E.; Small, S.A.; Wenk, M.R.; Shui, G.; Di Paolo, G. Comparative lipidomic analysis of mouse and human brain with Alzheimer disease. J. Biol. Chem. 2012, 287, 2678–2688. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Ye, S.; Tang, Y.; Tong, W.; Sun, S. Brain cholesterol homeostasis and its association with neurodegenerative diseases. Neurochem. Int. 2023, 171, 105635. [Google Scholar] [CrossRef] [PubMed]
- Aguayo-Ortiz, R.; Straub, J.E.; Dominguez, L. Influence of membrane lipid composition on the structure and activity of gamma-secretase. Phys. Chem. Chem. Phys. 2018, 20, 27294–27304. [Google Scholar] [CrossRef] [PubMed]
- Takami, M.; Nagashima, Y.; Sano, Y.; Ishihara, S.; Morishima-Kawashima, M.; Funamoto, S.; Ihara, Y. gamma-Secretase: Successive tripeptide and tetrapeptide release from the transmembrane domain of beta-carboxyl terminal fragment. J. Neurosci. 2009, 29, 13042–13052. [Google Scholar] [CrossRef]
- Yang, G.; Zhou, R.; Zhou, Q.; Guo, X.; Yan, C.; Ke, M.; Lei, J.; Shi, Y. Structural basis of Notch recognition by human gamma-secretase. Nature 2019, 565, 192–197. [Google Scholar] [CrossRef]
- Janssen, J.C.; Beck, J.A.; Campbell, T.A.; Dickinson, A.; Fox, N.C.; Harvey, R.J.; Houlden, H.; Rossor, M.N.; Collinge, J. Early onset familial Alzheimer’s disease: Mutation frequency in 31 families. Neurology 2003, 60, 235–239. [Google Scholar] [CrossRef]
- Sun, L.; Zhou, R.; Yang, G.; Shi, Y. Analysis of 138 pathogenic mutations in presenilin-1 on the in vitro production of Abeta42 and Abeta40 peptides by gamma-secretase. Proc. Natl. Acad. Sci. USA 2017, 114, E476–E485. [Google Scholar] [CrossRef]
- Gomez-Isla, T.; Wasco, W.; Pettingell, W.P.; Gurubhagavatula, S.; Schmidt, S.D.; Jondro, P.D.; McNamara, M.; Rodes, L.A.; DiBlasi, T.; Growdon, W.B.; et al. A novel presenilin-1 mutation: Increased beta-amyloid and neurofibrillary changes. Ann. Neurol. 1997, 41, 809–813. [Google Scholar] [CrossRef]
- Jiao, B.; Liu, H.; Guo, L.; Xiao, X.; Liao, X.; Zhou, Y.; Weng, L.; Zhou, L.; Wang, X.; Jiang, Y.; et al. The role of genetics in neurodegenerative dementia: A large cohort study in South China. NPJ Genom. Med. 2021, 6, 69. [Google Scholar] [CrossRef]
- Wallon, D.; Rousseau, S.; Rovelet-Lecrux, A.; Quillard-Muraine, M.; Guyant-Marechal, L.; Martinaud, O.; Pariente, J.; Puel, M.; Rollin-Sillaire, A.; Pasquier, F.; et al. The French series of autosomal dominant early onset Alzheimer’s disease cases: Mutation spectrum and cerebrospinal fluid biomarkers. J. Alzheimers Dis. 2012, 30, 847–856. [Google Scholar] [CrossRef]
- Borroni, B.; Pilotto, A.; Bonvicini, C.; Archetti, S.; Alberici, A.; Lupi, A.; Gennarelli, M.; Padovani, A. Atypical presentation of a novel Presenilin 1 R377W mutation: Sporadic, late-onset Alzheimer disease with epilepsy and frontotemporal atrophy. Neurol. Sci. 2012, 33, 375–378. [Google Scholar] [CrossRef]
- Orzel, U.; Pasznik, P.; Miszta, P.; Lorkowski, M.; Niewieczerzal, S.; Jakowiecki, J.; Filipek, S. GS-SMD server for steered molecular dynamics of peptide substrates in the active site of the gamma-secretase complex. Nucleic Acids Res. 2023, 51, W251–W262. [Google Scholar] [CrossRef]
- Guzman-Ocampo, D.C.; Aguayo-Ortiz, R.; Velasco-Bolom, J.L.; Gupta, P.L.; Roitberg, A.E.; Dominguez, L. Elucidating the Protonation State of the gamma-Secretase Catalytic Dyad. ACS Chem. Neurosci. 2023, 14, 261–269. [Google Scholar] [CrossRef]
- Lee, J.; Cheng, X.; Swails, J.M.; Yeom, M.S.; Eastman, P.K.; Lemkul, J.A.; Wei, S.; Buckner, J.; Jeong, J.C.; Qi, Y.; et al. CHARMM-GUI Input Generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM Simulations Using the CHARMM36 Additive Force Field. J. Chem. Theory Comput. 2016, 12, 405–413. [Google Scholar] [CrossRef] [PubMed]
- Case, D.A.; Cheatham, T.E.; Darden, T.; Gohlke, H.; Luo, R.; Merz, K.M.; Onufriev, A.; Simmerling, C.; Wang, B.; Woods, R.J. The Amber biomolecular simulation programs. J. Comput. Chem. 2005, 26, 1668–1688. [Google Scholar] [CrossRef] [PubMed]
- Klauda, J.B.; Venable, R.M.; Freites, J.A.; O’Connor, J.W.; Tobias, D.J.; Mondragon-Ramirez, C.; Vorobyov, I.; MacKerell, A.D., Jr.; Pastor, R.W. Update of the CHARMM all-atom additive force field for lipids: Validation on six lipid types. J. Phys. Chem. B 2010, 114, 7830–7843. [Google Scholar] [CrossRef] [PubMed]
- Krautler, V.; Van Gunsteren, W.F.; Hunenberger, P.H. A fast SHAKE: Algorithm to solve distance constraint equations for small molecules in molecular dynamics simulations. J. Comput. Chem. 2001, 22, 501–508. [Google Scholar] [CrossRef]
- Essmann, U.; Perera, L.; Berkowitz, M.L.; Darden, T.; Lee, H.; Pedersen, L.G. A Smooth Particle Mesh Ewald Method. J. Chem. Phys. 1995, 103, 8577–8593. [Google Scholar] [CrossRef]
- Lazaridis, T. Effective energy function for proteins in lipid membranes. Proteins 2003, 52, 176–192. [Google Scholar] [CrossRef]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. Model. 1996, 14, 33–38. [Google Scholar] [CrossRef] [PubMed]
- Land, H.; Humble, M.S. YASARA: A Tool to Obtain Structural Guidance in Biocatalytic Investigations. Methods Mol. Biol. 2018, 1685, 43–67. [Google Scholar] [CrossRef] [PubMed]
- The PyMOL Molecular Graphics System, Version 2.5.0. Schrödinger, LLC.
- Hunter, J.D. Matplotlib: A 2D Graphics Environment. Comput. Sci. Eng. 2007, 9, 90–95. [Google Scholar] [CrossRef]










| Mutation | Biological Effect | Ref. |
|---|---|---|
| R269G | Early-onset AD; decreased protease activity with APP; increased Aβ42/Aβ40 ratio. | [30] |
| R269H | Early-onset AD; increased Aβ (42 + 43)/(37 + 38 + 40) ratio in cells; decreased GS activity. | [29,31,32] |
| R377M | Early-onset AD; uncertain significance, but in silico algorithm predicted it is deleterious. | [29] |
| R377W | In vitro, decreased Aβ42 production and abrogated Aβ40 production; nearly abolishes protease activity with APP. | [30,33,34] |
| K380R | Unknown, but in silico predictions suggest damaging effect. | [32] |
| Del377-381 | Loss of Notch1 and APP-C83 cleavage. | [18,28] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jakowiecki, J.; Orzeł, U.; Miszta, P.; Młynarczyk, K.; Filipek, S. Conformational Changes and Unfolding of β-Amyloid Substrates in the Active Site of γ-Secretase. Int. J. Mol. Sci. 2024, 25, 2564. https://doi.org/10.3390/ijms25052564
Jakowiecki J, Orzeł U, Miszta P, Młynarczyk K, Filipek S. Conformational Changes and Unfolding of β-Amyloid Substrates in the Active Site of γ-Secretase. International Journal of Molecular Sciences. 2024; 25(5):2564. https://doi.org/10.3390/ijms25052564
Chicago/Turabian StyleJakowiecki, Jakub, Urszula Orzeł, Przemysław Miszta, Krzysztof Młynarczyk, and Sławomir Filipek. 2024. "Conformational Changes and Unfolding of β-Amyloid Substrates in the Active Site of γ-Secretase" International Journal of Molecular Sciences 25, no. 5: 2564. https://doi.org/10.3390/ijms25052564