A GASA Protein Family Gene, CmGEG, Inhibits Petal Growth in Chrysanthemum
Abstract
1. Introduction
2. Results
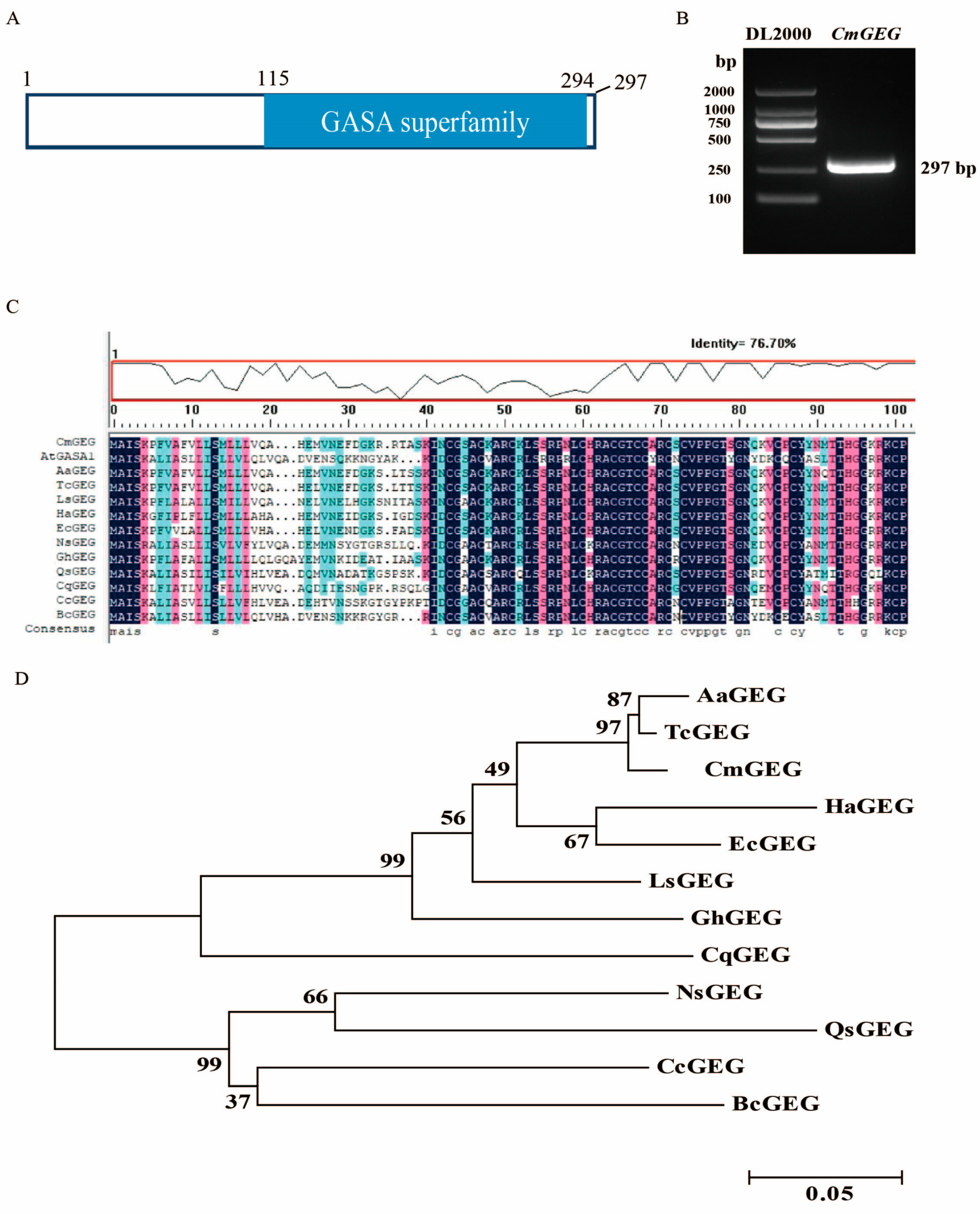
2.1. Cloning and Phylogenetic Analysis of CmGEG
2.2. Subcellular Localization of CmGEG
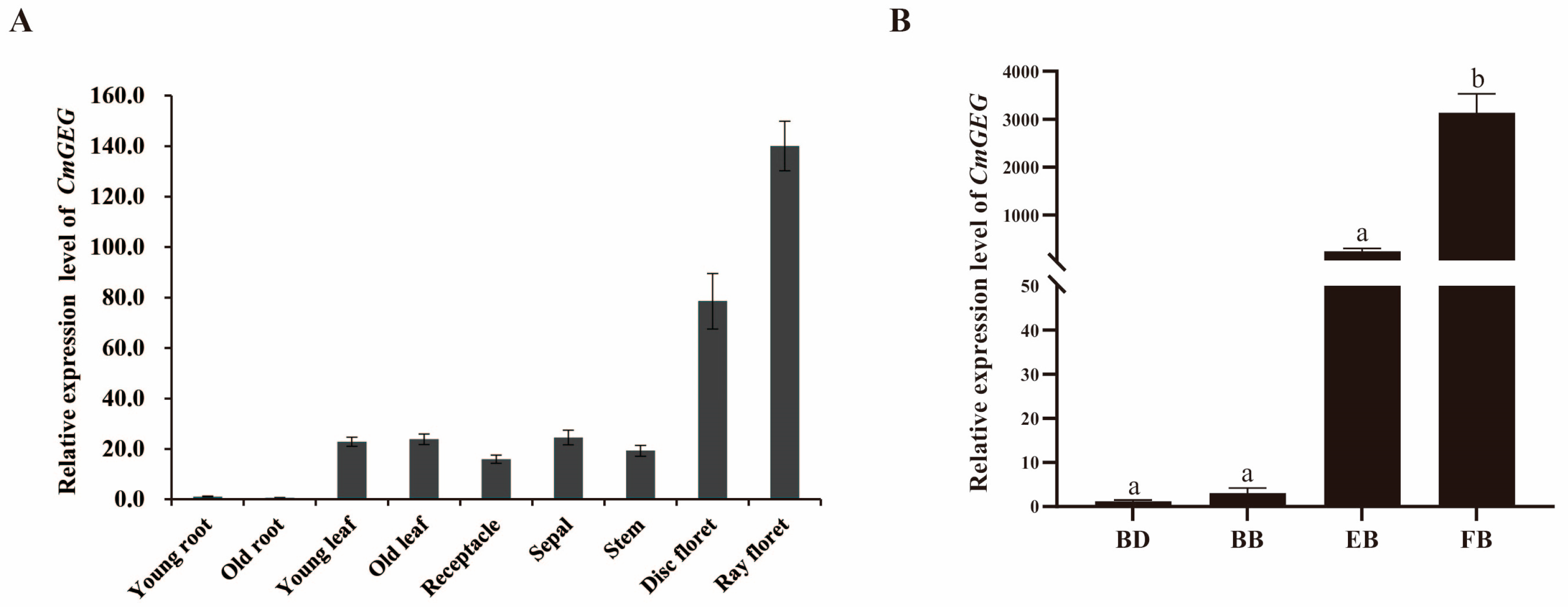
2.3. The Expression Pattern of CmGEG
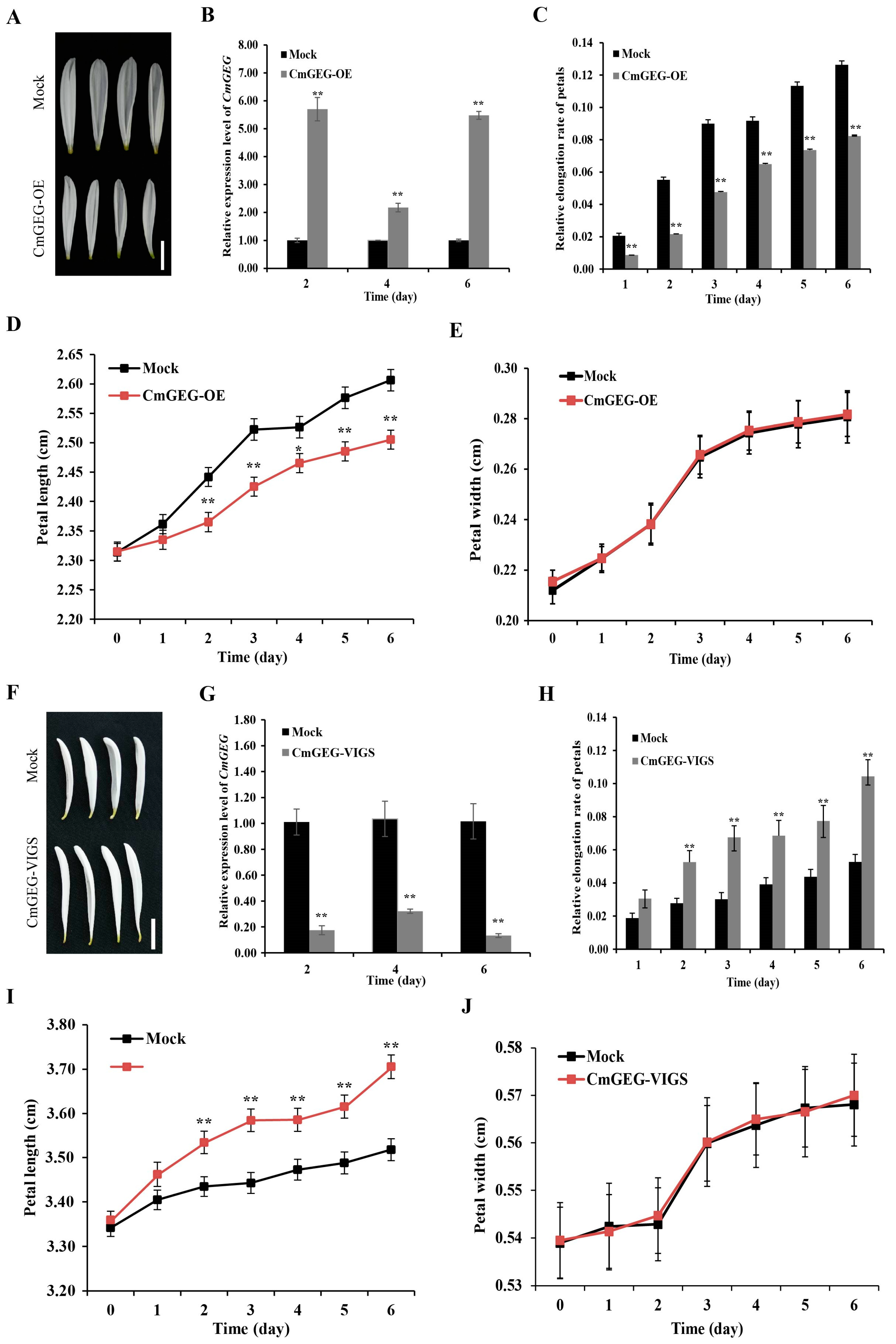
2.4. CmGEG Is Involved in Chrysanthemum Petal Elongation
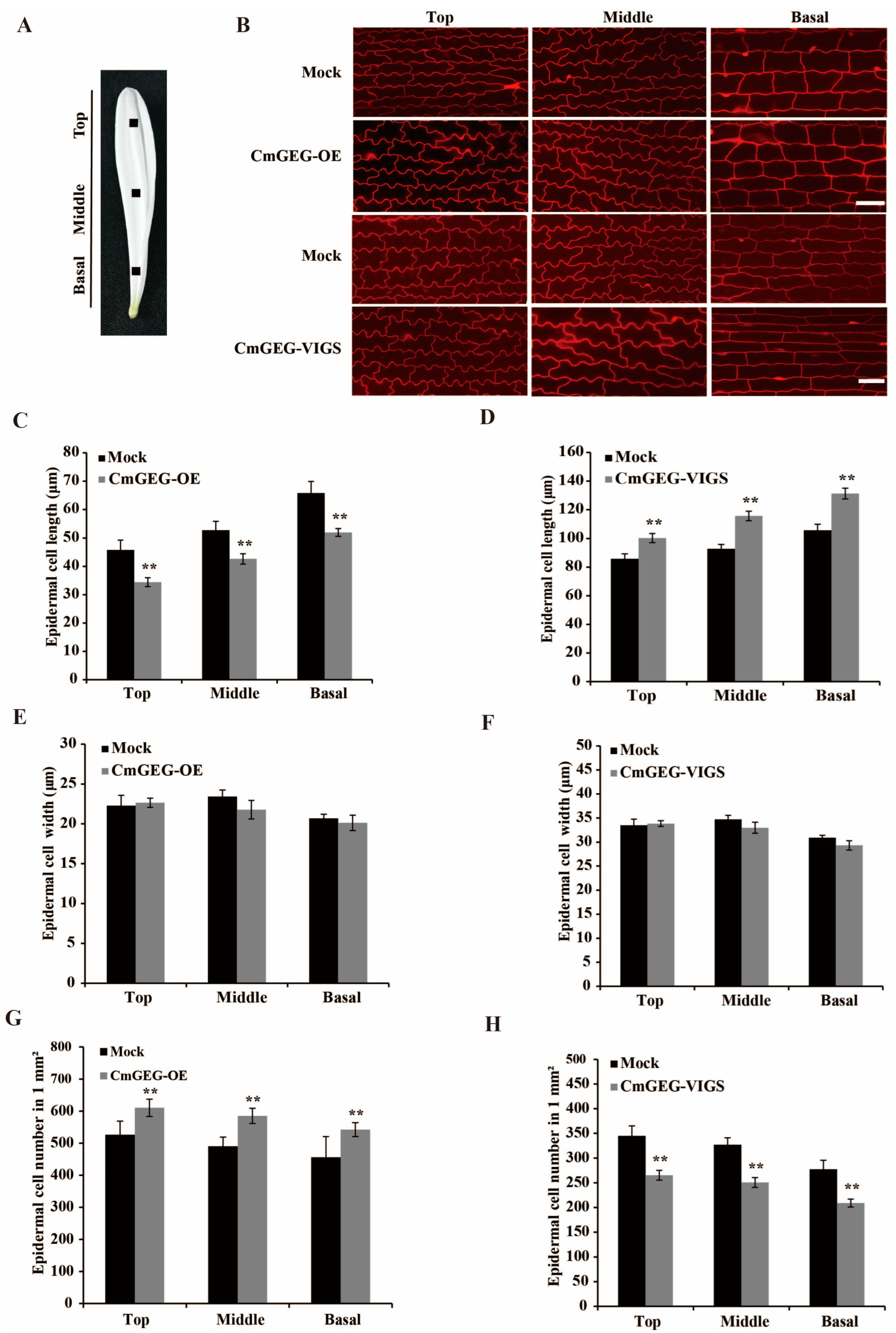
2.5. CmGEG Negatively Regulates Petal Size by Limiting Cell Expansion in Chrysanthemum
3. Discussion
3.1. The GASA Protein CmGEG Localize in the Cytoplasm and Cell Membrane
3.2. CmGEG Inhibited Petal Elongation Regulated by GA
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Cloning and Phylogenetic Analysis of CmGEG
4.3. Ectopic Expression of CmGEG in Arabidopsis
4.4. Subcellular Localization of CmGEG
4.5. Transient Transformation of the Petals
4.6. Measurement of the Petal Size and Cell Size
4.7. RNA Extraction and Quantitative RT-PCR (qRT-PCR)
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Amasino, R. Seasonal and developmental timing of flowering. Plant J. 2010, 61, 1001–1013. [Google Scholar] [CrossRef] [PubMed]
- Irish, V.F. The flowering of Arabidopsis flower development. Plant J. 2010, 61, 1014–1028. [Google Scholar] [CrossRef]
- Kieffer, M.; Davies, B. Developmental programmes in floral organ formation. Semin. Cell Dev. Biol. 2001, 12, 373–380. [Google Scholar] [CrossRef] [PubMed]
- Krizek, B.A.; Fletcher, J.C. Molecular mechanisms of flower development: An armchair guide. Nat. Rev. Genet. 2005, 6, 688–698. [Google Scholar] [CrossRef]
- Hermann, K.; Kuhlemeier, C. The genetic architecture of natural variation in flower morphology. Curr. Opin. Plant Biol. 2011, 14, 60–65. [Google Scholar] [CrossRef] [PubMed]
- Varaud, E.; Brioudes, F.; Szecsi, J.; Leroux, J.; Brown, S.; Perrot-Rechenmann, C.; Bendahmane, M. AUXIN RESPONSE FACTOR8 regulates Arabidopsis petal growth by interacting with the bHLH transcription factor BIGPETALp. Plant Cell 2011, 23, 973–983. [Google Scholar] [CrossRef] [PubMed]
- Alvarez-Buylla, E.R.; Benitez, M.; Corvera-Poire, A.; Chaos Cador, A.; de Folter, S.; Gamboa de Buen, A.; Garay-Arroyo, A.; Garcia-Ponce, B.; Jaimes-Miranda, F.; Perez-Ruiz, R.V.; et al. Flower development. Arab. Book 2010, 8, e0127. [Google Scholar] [CrossRef] [PubMed]
- Hepworth, J.; Lenhard, M. Regulation of plant lateral-organ growth by modulating cell number and size. Curr. Opin. Plant Biol. 2014, 17, 36–42. [Google Scholar] [CrossRef]
- Weiss, J.; Delgado-Benarroch, L.; Egea-Cortines, M. Genetic control of floral size and proportions. Int. J. Dev. Biol. 2005, 49, 513–525. [Google Scholar] [CrossRef]
- Jiang, Y.H.; Bao, L.; Jeong, S.Y.; Kim, S.K.; Xu, C.G.; Li, X.H.; Zhang, Q.F. XIAO is involved in the control of organ size by contributing to the regulation of signaling and homeostasis of brassinosteroids and cell cycling in rice. Plant J. 2012, 70, 398–408. [Google Scholar] [CrossRef]
- Lampugnani, E.R.; Kilinc, A.; Smyth, D.R. Auxin controls petal initiation in Arabidopsis. Development 2013, 140, 185–194. [Google Scholar] [CrossRef]
- Tabata, R.; Ikezaki, M.; Fujibe, T.; Aida, M.; Tian, C.E.; Ueno, Y.; Yamamoto, K.T.; Machida, Y.; Nakamura, K.; Ishiguro, S. Arabidopsis AUXIN RESPONSE FACTOR6 and 8 Regulate Jasmonic Acid Biosynthesis and Floral Organ Development via Repression of Class 1 KNOX Genes. Plant Cell Physiol. 2010, 51, 164–175. [Google Scholar] [CrossRef] [PubMed]
- Han, M.; Jin, X.; Yao, W.; Kong, L.; Huang, G.; Tao, Y.; Li, L.; Wang, X.; Wang, Y. A Mini Zinc-Finger Protein (MIF) from Gerbera hybrida Activates the GASA Protein Family Gene, GEG, to Inhibit Ray Petal Elongation. Front. Plant Sci. 2017, 8, 1649. [Google Scholar] [CrossRef] [PubMed]
- Huang, G.; Han, M.; Jian, L.; Chen, Y.; Sun, S.; Wang, X.; Wang, Y. An ETHYLENE INSENSITIVE3-LIKE1 Protein Directly Targets the GEG Promoter and Mediates Ethylene-Induced Ray Petal Elongation in Gerbera hybrida. Front. Plant Sci. 2019, 10, 1737. [Google Scholar] [CrossRef] [PubMed]
- Ma, N.; Xue, J.Q.; Li, Y.H.; Liu, X.J.; Dai, F.W.; Jia, W.S.; Luo, Y.B.; Gao, J.P. Rh-PIP2;1, a rose aquaporin gene, is involved in ethylene-regulated petal expansion. Plant Physiol. 2008, 148, 894–907. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Wu, Z.; Ma, N.; Gao, J. Regulation of the rose Rh-PIP2;1 promoter by hormones and abiotic stresses in Arabidopsis. Plant Cell Rep. 2009, 28, 185–196. [Google Scholar] [CrossRef]
- Wang, J.; Wang, H.; Ding, L.; Song, A.; Shen, F.; Jiang, J.; Chen, S.; Chen, F. Transcriptomic and hormone analyses reveal mechanisms underlying petal elongation in Chrysanthemum morifolium ‘Jinba’. Plant Mol. Biol. 2017, 93, 593–606. [Google Scholar] [CrossRef]
- Li, L.; Zhang, W.; Zhang, L.; Li, N.; Peng, J.; Wang, Y.; Zhong, C.; Yang, Y.; Sun, S.; Liang, S.; et al. Transcriptomic insights into antagonistic effects of gibberellin and abscisic acid on petal growth in Gerbera hybrida. Front. Plant Sci. 2015, 6, 168. [Google Scholar] [CrossRef]
- Huang, G.; Han, M.X.; Yao, W.; Wang, Y.Q. Transcriptome analysis reveals the regulation of brassinosteroids on petal growth in Gerbera hybrida. PeerJ 2017, 5, e3382. [Google Scholar] [CrossRef]
- Han, Y.; Zhang, C.; Yang, H.; Jiao, Y. Cytokinin pathway mediates APETALA1 function in the establishment of determinate floral meristems in Arabidopsis. Proc. Natl. Acad. Sci. USA 2014, 111, 6840–6845. [Google Scholar] [CrossRef]
- Qian, M.; Fan, Y.; Li, Y.; Liu, M.; Sun, W.; Duan, H.; Yu, M.; Chang, W.; Niu, Y.; Li, X.; et al. Genome-wide association study and transcriptome comparison reveal novel QTL and candidate genes that control petal size in rapeseed. J. Exp. Bot. 2021, 72, 3597–3610. [Google Scholar] [CrossRef]
- Peng, S.; Huang, S.; Liu, Z.; Feng, H. Mutation of ACX1, a Jasmonic Acid Biosynthetic Enzyme, Leads to Petal Degeneration in Chinese Cabbage (Brassica campestris ssp. pekinensis). Int. J. Mol. Sci. 2019, 20, 2310. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Kang, X.; Wei, L.; Zhang, D.; Lin, H. A gain-of-function mutation in Brassinosteroid-insensitive 2 alters Arabidopsis floral organ development by altering auxin levels. Plant Cell Rep. 2020, 39, 259–271. [Google Scholar] [CrossRef]
- Bartrina, I.; Otto, E.; Strnad, M.; Werner, T.; Schmulling, T. Cytokinin regulates the activity of reproductive meristems, flower organ size, ovule formation and thus seed yield in Arabidopsis thaliana. Plant Cell 2011, 23, 69–80. [Google Scholar] [CrossRef] [PubMed]
- Brioudes, F.; Joly, C.; Szecsi, J.; Varaud, E.; Leroux, J.; Bellvert, F.; Bertrand, C.; Bendahmane, M. Jasmonate controls late development stages of petal growth in Arabidopsis thaliana. Plant J. 2009, 60, 1070–1080. [Google Scholar] [CrossRef] [PubMed]
- Han, S.; Jiao, Z.; Niu, M.X.; Yu, X.; Huang, M.; Liu, C.; Wang, H.L.; Zhou, Y.; Mao, W.; Wang, X.; et al. Genome-Wide Comprehensive Analysis of the GASA Gene Family in Populus. Int. J. Mol. Sci. 2021, 22, 12336. [Google Scholar] [CrossRef] [PubMed]
- Herzog, M.; Dorne, A.M.; Grellet, F. GASA, a gibberellin-regulated gene family from Arabidopsis thaliana related to the tomato GAST1 gene. Plant Mol. Biol. 1995, 27, 743–752. [Google Scholar] [CrossRef]
- Roxrud, I.; Lid, S.E.; Fletcher, J.C.; Schmidt, E.D.; Opsahl-Sorteberg, H.G. GASA4, one of the 14-member Arabidopsis GASA family of small polypeptides, regulates flowering and seed development. Plant Cell Physiol. 2007, 48, 471–483. [Google Scholar] [CrossRef]
- Sun, S.L.; Wang, H.X.; Yu, H.M.; Zhong, C.M.; Zhang, X.X.; Peng, J.Z.; Wang, X.J. GASA14 regulates leaf expansion and abiotic stress resistance by modulating reactive oxygen species accumulation. J. Exp. Bot. 2013, 64, 1637–1647. [Google Scholar] [CrossRef]
- Zhang, S.C.; Wang, X.J. Expression pattern of GASA, downstream genes of DELLA, in Arabidopsis. Chin. Sci. Bull. 2008, 53, 3839–3846. [Google Scholar] [CrossRef]
- Zimmermann, R.; Sakai, H.; Hochholdinger, F. The Gibberellic Acid Stimulated-Like Gene Family in Maize and Its Role in Lateral Root Developmen. Plant Physiol. 2010, 152, 356–365. [Google Scholar] [CrossRef]
- Muhammad, I.; Li, W.Q.; Jing, X.Q.; Zhou, M.R.; Shalmani, A.; Ali, M.; Wei, X.Y.; Sharif, R.; Liu, W.T.; Chen, K.M. A systematic in silico prediction of gibberellic acid stimulated GASA family members: A novel small peptide contributes to floral architecture and transcriptomic changes induced by external stimuli in rice. J. Plant Physiol. 2019, 234, 117–132. [Google Scholar] [CrossRef]
- Nahirñak, V.; Rivarola, M.; Almasia, N.I.; Barón, M.P.B.; Hopp, H.E.; Vile, D.; Paniego, N.; Rovere, C.V. Snakin-1 affects reactive oxygen species and ascorbic acid levels and hormone balance in potato. PLoS ONE 2019, 14, e0214165. [Google Scholar] [CrossRef]
- Ben-Nissan, G.; Lee, J.Y.; Borohov, A.; Weiss, D. GIP, a Petunia hybrida GA-induced cysteine-rich protein: A possible role in shoot elongation and transition to flowering. Plant J. 2004, 37, 229–238. [Google Scholar] [CrossRef]
- Wu, T.L.; Zhong, Y.; Chen, M.; Wu, B.; Wang, T.; Jiang, B.; Zhong, G.Y. Analysis of CcGASA family members in Citrus clementina (Hort. ex Tan.) by a genome-wide approach. BMC Plant Biol. 2021, 21, 565. [Google Scholar] [CrossRef]
- Shi, L.; Gast, R.T.; Gopalraj, M.; Olszewski, N.E. Characterization of a shoot-specific, GA3- and ABA-regulated gene from tomato. Plant J. 1992, 2, 153–159. [Google Scholar] [CrossRef]
- Wang, L.; Wang, Z.; Xu, Y.Y.; Joo, S.H.; Kim, S.K.; Xue, Z.; Xu, Z.H.; Wang, Z.Y.; Chong, K. OsGSR1 is involved in crosstalk between gibberellins and brassinosteroids in rice. Plant J. 2009, 57, 498–510. [Google Scholar] [CrossRef] [PubMed]
- Zhong, C.; Xu, H.; Ye, S.; Wang, S.; Li, L.; Zhang, S.; Wang, X. Gibberellic Acid-Stimulated Arabidopsis6 Serves as an Integrator of Gibberellin, Abscisic Acid, and Glucose Signaling during Seed Germination in Arabidopsis. Plant Physiol. 2015, 169, 2288–2303. [Google Scholar]
- Moyano-Cañete, E.; Bellido, M.L.; García-Caparros, N.; Medina-Puche, L.; Amil-Ruiz, F.; González-Reyes, J.A.; Caballero, J.L.; Muñoz-Blanco, J.; Blanco-Portales, R. FaGAST2, a strawberry ripening-related gene, acts together with FaGAST1 to determine cell size of the fruit receptacl. Plant Cell Physiol. 2013, 54, 218–236. [Google Scholar] [CrossRef]
- Li, K.L.; Bai, X.; Li, Y.; Cai, H.; Ji, W.; Tang, L.L.; Wen, Y.D.; Zhu, Y.M. GsGASA1 mediated root growth inhibition in response to chronic cold stress is marked by the accumulation of DELLAs. J. Plant Physiol. 2011, 168, 2153–2160. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.C.; Wang, X.J. One new kind of phytohormonal signaling integrator: Up-and-coming GASA family genes. Plant Signal. Behav. 2017, 12, e1226453. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.C.; Yang, C.W.; Peng, J.Z.; Sun, S.L.; Wang, X.J. GASA5, a regulator of flowering time and stem growth in Arabidopsis thaliana. Plant Mol. Biol. 2009, 69, 745–759. [Google Scholar] [CrossRef]
- Furukawa, T.; Sakaguchi, N.; Shimada, H. Two OsGASR genes, rice GAST homologue genes that are abundant in proliferating tissues, show different expression patterns in developing panicles. Genes Genet. Syst. 2006, 81, 171–180. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Yue, M.X.; Zheng, X.K.; Gautam, M.; He, S.B.; Li, L.J. The Role of Promoter-Associated Histone Acetylation of Haem Oxygenase-1 (HO-1) and Giberellic Acid-Stimulated Like-1 (GSL-1) Genes in Heat-Induced Lateral Root Primordium Inhibition in Maize. Front. Plant Sci. 2018, 9, 1520. [Google Scholar] [CrossRef]
- Nahirñak, V.; Almasia, N.I.; Fernandez, P.V.; Hopp, H.E.; Estevez, J.M.; Carrari, F.; Vazquez-Rovere, C. Potato Snakin-1 Gene Silencing Affects Cell Division, Primary Metabolism, and Cell Wall Composition. Plant Physiol. 2012, 158, 252–263. [Google Scholar] [CrossRef]
- Kotilainen, M.; Helariutta, Y.; Mehto, M.; Pöllänen, E.; Albert, V.A.; Elomaa, P.; Teeri, T.H. GEG participates in the regulation of cell and organ shape during corolla and carpel development in Gerbera hybrida. Plant Cell 1999, 11, 1093–1104. [Google Scholar] [CrossRef]
- Peng, J.Z.; Lai, L.J.; Wang, X.J. PRGL: A cell wall proline-rich protein containning GASA domain in Gerbera hybrida. Sci. China Ser. C Life Sci. 2008, 51, 520–525. [Google Scholar] [CrossRef] [PubMed]
- Jiang, R.; Yuan, W.; Yao, W.; Jin, X.; Wang, X.; Wang, Y. A regulatory GhBPE–GhPRGL module maintains ray petal length in Gerbera hybrida. Mol. Hortic. 2022, 2, 9. [Google Scholar] [CrossRef]
- da Silva, J.A.T. Chrysanthemum: Advances in tissue culture, cryopreservation, postharvest technology, genetics and transgenic biotechnology. Biotechnol. Adv. 2003, 21, 715–766. [Google Scholar] [CrossRef]
- Ding, L.; Song, A.; Zhang, X.; Li, S.; Su, J.; Xia, W.; Zhao, K.; Zhao, W.; Guan, Y.; Fang, W.; et al. The core regulatory networks and hub genes regulating flower development in Chrysanthemum morifolium. Plant Mol. Biol. 2020, 103, 669–688. [Google Scholar] [CrossRef]
- Ding, L.; Zhao, K.; Zhang, X.; Song, A.; Su, J.; Hu, Y.; Zhao, W.; Jiang, J.; Chen, F. Comprehensive characterization of a floral mutant reveals the mechanism of hooked petal morphogenesis in Chrysanthemum morifolium. Plant Biotechnol. J. 2019, 17, 2325–2340. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.; Li, X.; Sun, M.; Zhang, T.; Pan, H.; Cheng, T.; Wang, J.; Zhang, Q. Identification and Characterization of CYC-Like Genes in Regulation of Ray Floret Development in Chrysanthemum morifolium. Front. Plant Sci. 2016, 7, 1633. [Google Scholar] [CrossRef]
- Yuan, C.Q.; Huang, D.; Yang, Y.; Sun, M.; Cheng, T.R.; Wang, J.; Pan, H.T.; Zhang, Q.X. CmCYC2-like transcription factors may interact with each other or bind to the promoter to regulate floral symmetry development in Chrysanthemum morifolium. Plant Mol. Biol. 2020, 103, 159–171. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Sun, M.; Pan, H.; Cheng, T.; Wang, J.; Zhang, Q. Two Cyc2CL transcripts (Cyc2CL-1 and Cyc2CL-2) may play key roles in the petal and stamen development of ray florets in chrysanthemum. BMC Plant Biol. 2021, 21, 105. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Sun, M.; Yuan, C.; Han, Y.; Zheng, T.; Cheng, T.; Wang, J.; Zhang, Q. Interactions between WUSCHEL- and CYC2-like Transcription Factors in Regulating the Development of Reproductive Organs in Chrysanthemum morifolium. Int. J. Mol. Sci. 2019, 20, 1276. [Google Scholar] [CrossRef]
- Narumi, T.; Aida, R.; Koyama, T.; Yamaguchi, H.; Sasaki, K.; Shikata, M.; Nakayama, M.; Ohme-Takagi, M.; Ohtsubo, N. Arabidopsis chimeric TCP3 repressor produces novel floral traits in Torenia fournieri and Chrysanthemum morifolium. Plant Biotechnol. 2011, 28, 131–140. [Google Scholar]
- Gao, Y.; Gao, Y.; Fan, M.; Yuan, L.; Wu, Z.; Zhang, Q. Overexpression of Chrysanthemum morifolium SVP gene delays blossoming and regulates inflorescence architecture in transgenic Arabidopsis. Can. J. Plant Sci. 2017, 97, 1130–1139. [Google Scholar]
- Gao, Y.H.; Gao, Y.K.; Wu, Z.P.; Bu, X.L.; Fan, M.; Zhang, Q.X. Characterization of TEMINAL FLOWER1 homologs CmTFL1c gene from Chrysanthemum morifolium. Plant Mol. Biol. 2019, 99, 587–601. [Google Scholar] [CrossRef]
- Cheng, P.L.; Liu, Y.N.; Yang, Y.M.; Chen, H.; Cheng, H.; Hu, Q.; Zhang, Z.X.; Gao, J.J.; Zhang, J.X.; Ding, L.; et al. CmBES1 is a regulator of boundary formation in chrysanthemum ray florets. Hortic. Res. 2020, 7, 129. [Google Scholar] [CrossRef]
- Yu, Z.Y.; Tian, C.; Guan, Y.X.; He, J.; Wang, Z.X.; Wang, L.K.; Lin, S.S.; Guan, Z.Y.; Fang, W.M.; Chen, S.M.; et al. Expression Analysis of TCP Transcription Factor Family in Autopolyploids of Chrysanthemum nankingense. Front. Plant Sci. 2022, 13, 860956. [Google Scholar] [CrossRef]
- Wang, J.J.; Guan, Y.X.; Ding, L.; Li, P.R.; Zhao, W.Q.; Jiang, J.F.; Chen, S.M.; Chen, F.D. The CmTCP20 gene regulates petal elongation growth in Chrysanthemum morifolium. Plant Sci. 2019, 280, 248–257. [Google Scholar] [CrossRef]
- Guan, Y.X.; Ding, L.; Jiang, J.F.; Jia, D.W.; Li, S.; Jin, L.; Zhao, W.Q.; Zhang, X.; Song, A.P.; Chen, S.M.; et al. The TIFY family protein CmJAZ1-like negatively regulates petal size via interaction with the bHLH transcription factor CmBPE2 in Chrysanthemum morifolium. Plant J. 2022, 112, 1489–1506. [Google Scholar] [CrossRef]
- Chen, B.; Sun, Y.; Tian, Z.; Fu, G.; Pei, X.; Pan, Z.; Nazir, M.F.; Song, S.; Li, H.; Wang, X.; et al. GhGASA10-1 promotes the cell elongation in fiber development through the phytohormones IAA-induced. BMC Plant Biol. 2021, 21, 448. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Liu, W.C.; Li, X.W.; Li, H.Y. Overexpression of GmGASA32 promoted soybean height by interacting with GmCDC25. Plant Signal. Behav. 2021, 16, 1855017. [Google Scholar] [CrossRef] [PubMed]
- Su, D.; Liu, K.D.; Yu, Z.S.; Li, Y.; Zhang, Y.X.; Zhu, Y.Q.; Wu, Y.; He, H.Y.; Zeng, X.D.; Chen, H.L.; et al. Genome-wide characterization of the tomato GASA family identifies SlGASA1 as a repressor of fruit ripening. Hortic. Res. 2023, 10, uhac222. [Google Scholar] [CrossRef] [PubMed]
- Wu, K.; Qu, Y.S.; Rong, H.; Han, X.; Tian, Y.T.; Xu, L.A. Identification and Expression Analysis of the Populus trichocarpa GASA-Gene Family. Int. J. Mol. Sci. 2022, 23, 1507. [Google Scholar] [CrossRef] [PubMed]
- Nahirnak, V.; Almasia, N.I.; Hopp, H.E.; Vazquez-Rovere, C. Snakin/GASA proteins: Involvement in hormone crosstalk and redox homeostasis. Plant Signal Behav. 2012, 7, 1004–1008. [Google Scholar] [CrossRef]
- Yang, M.Z.; Liu, C.Y.; Zhang, W.; Wu, J.; Zhong, Z.Q.; Yi, W.; Liu, H.; Leng, Y.; Sun, W.S.; Luan, A.P.; et al. Genome-Wide Identification and Characterization of Gibberellic Acid-Stimulated Arabidopsis Gene Family in Pineapple (Ananas comosus). Int. J. Mol. Sci. 2023, 24, 17063. [Google Scholar] [CrossRef] [PubMed]
- Bouteraa, M.T.; Ben Romdhane, W.; Baazaoui, N.; Alfaifi, M.Y.; Chouaibi, Y.; Ben Akacha, B.; Ben Hsouna, A.; Kacaniová, M.; Zeljkovic, S.C.; Garzoli, S.; et al. GASA Proteins: Review of Their Functions in Plant Environmental Stress Tolerance. Plants 2023, 12, 2045. [Google Scholar] [CrossRef]
- Fan, S.; Zhang, D.; Zhang, L.Z.; Gao, C.; Xin, M.Z.; Tahir, M.M.; Li, Y.M.; Ma, J.J.; Han, M.Y. Comprehensive analysis of GASA family members in the Malus domestica genome: Identification, characterization, and their expressions in response to apple flower induction. BMC Genom. 2017, 18, 827. [Google Scholar] [CrossRef]
- Wang, H.B.; Wei, T.; Wang, X.; Zhang, L.P.; Yang, M.L.; Chen, L.; Song, W.Q.; Wang, C.G.; Chen, C.B. Transcriptome Analyses from Mutant Salvia miltiorrhiza Reveals Important Roles for SmGASA4 during Plant Development. Int. J. Mol. Sci. 2018, 19, 2088. [Google Scholar] [CrossRef] [PubMed]
- Aubert, D.; Chevillard, M.; Dorne, A.M.; Arlaud, G.; Herzog, M. Expression patterns of GASA genes in Arabidopsis thaliana: The GASA4 gene is up-regulated by gibberellins in meristematic regions. Plant Mol. Biol. 1998, 36, 871–883. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Zhang, X.R.; Henriques, R.; Lin, S.S.; Niu, Q.W.; Chua, N.H. Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat. Protoc. 2006, 1, 641–646. [Google Scholar] [CrossRef] [PubMed]
- Wu, F.H.; Shen, S.C.; Lee, L.Y.; Lee, S.H.; Chan, M.T.; Lin, C.S. Tape-Arabidopsis Sandwich—A simpler Arabidopsis protoplast isolation method. Plant Methods 2009, 5, 16. [Google Scholar] [CrossRef]
- Gao, W.J.; Meng, Q.R.; Luo, H.; Chen, F.; Zhou, Y.W.; He, M. Transcriptional responses for biosynthesis of flavor volatiles in methyl jasmonate-treated Chrysanthemum indicum var. aromaticum leaves. Ind. Crop. Prod. 2020, 147, 112254. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
He, Z.; Jiang, R.; Wang, X.; Wang, Y. A GASA Protein Family Gene, CmGEG, Inhibits Petal Growth in Chrysanthemum. Int. J. Mol. Sci. 2024, 25, 3367. https://doi.org/10.3390/ijms25063367
He Z, Jiang R, Wang X, Wang Y. A GASA Protein Family Gene, CmGEG, Inhibits Petal Growth in Chrysanthemum. International Journal of Molecular Sciences. 2024; 25(6):3367. https://doi.org/10.3390/ijms25063367
Chicago/Turabian StyleHe, Ziying, Rui Jiang, Xiaojing Wang, and Yaqin Wang. 2024. "A GASA Protein Family Gene, CmGEG, Inhibits Petal Growth in Chrysanthemum" International Journal of Molecular Sciences 25, no. 6: 3367. https://doi.org/10.3390/ijms25063367
APA StyleHe, Z., Jiang, R., Wang, X., & Wang, Y. (2024). A GASA Protein Family Gene, CmGEG, Inhibits Petal Growth in Chrysanthemum. International Journal of Molecular Sciences, 25(6), 3367. https://doi.org/10.3390/ijms25063367



