Genome-Wide Identification, Characterization, and Expression Analysis of Four Subgroup Members of the GH13 Family in Wheat (Triticum aestivum L.)
Abstract
:1. Introduction
2. Results
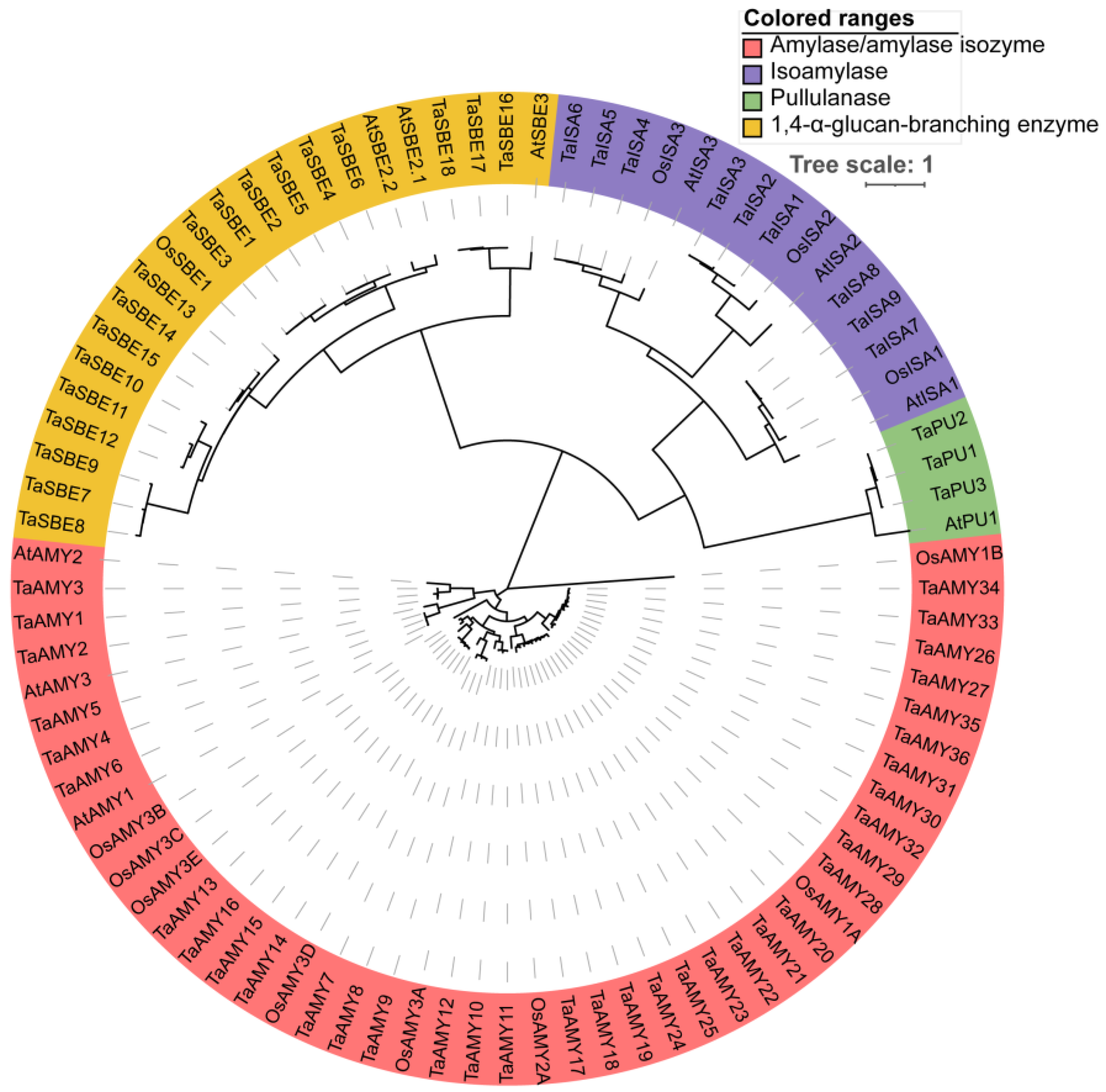
2.1. Identification and Phylogenetic Analysis of GH13 Family Members in Wheat Genome
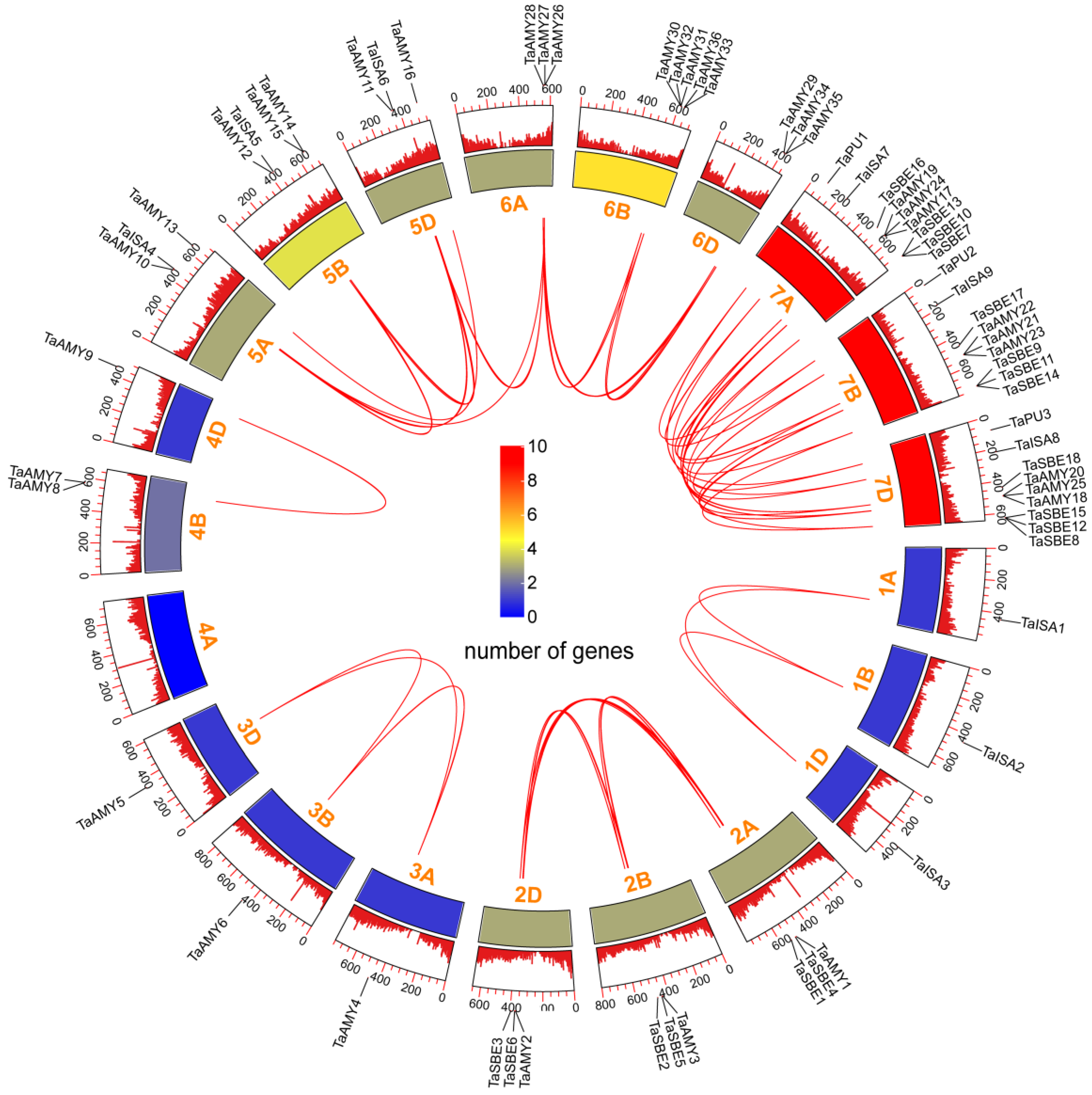
2.2. Chromosomal Locations, Synteny Analysis, and Duplication Events of TaGH13 Family
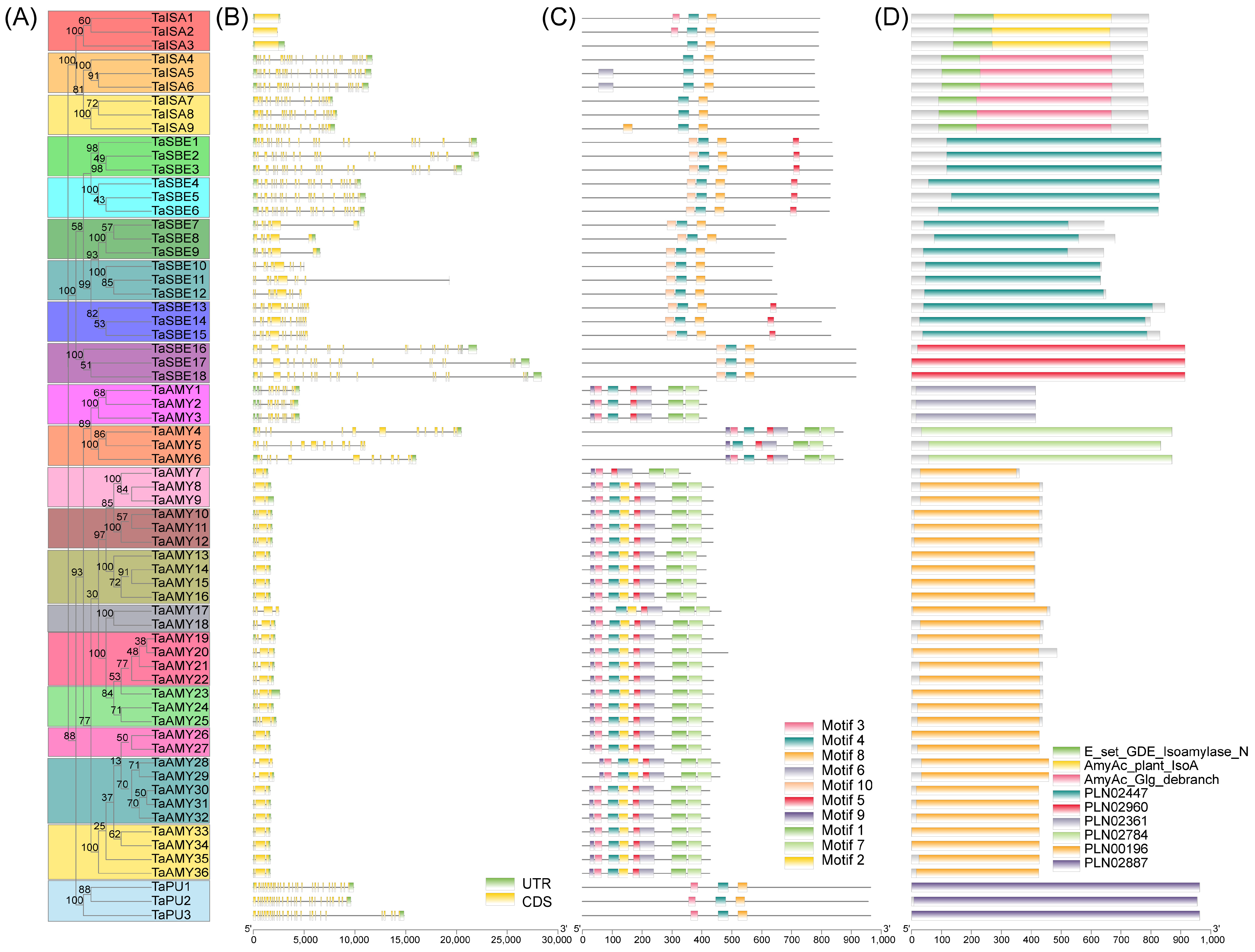
2.3. Gene Structure, Motif, and Domain Composition of the TaGH13 Family
2.4. Prediction of Cis-Acting Elements and Transcription Factor (TF) Binding Sites in TaGH13 Promoters
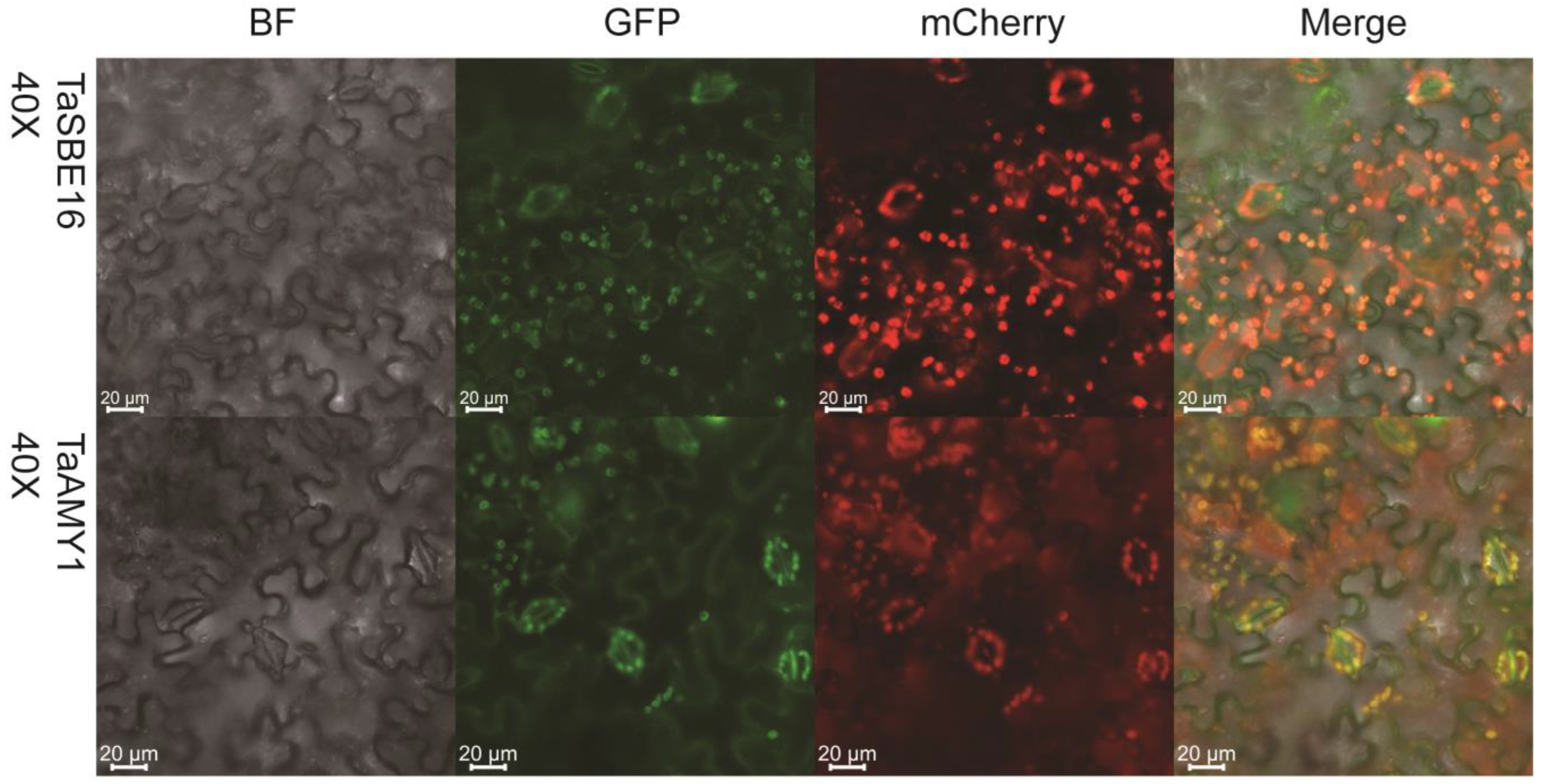
2.5. Subcellular Localization of TaGH13 Family Proteins
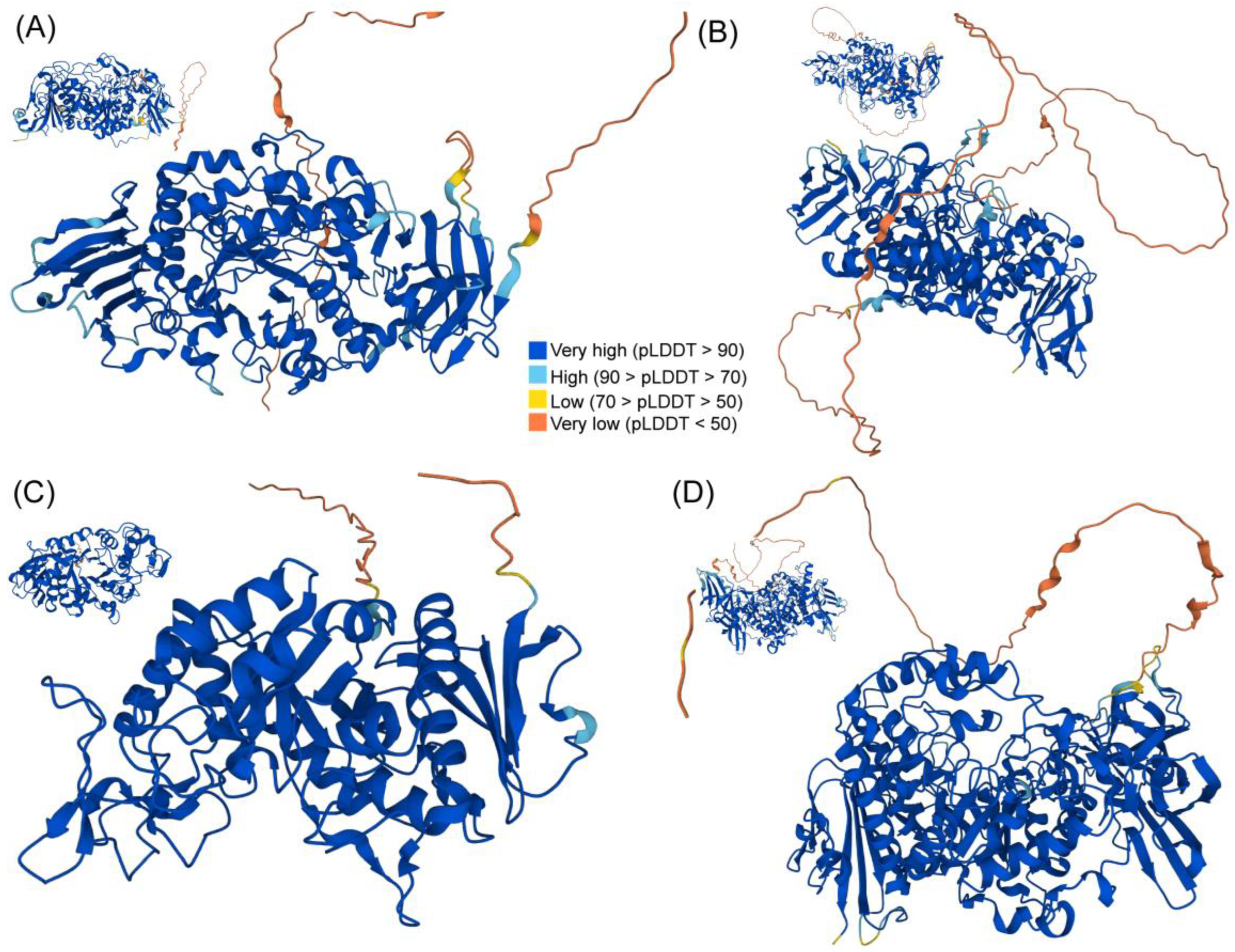
2.6. Three-Dimensional (3D) Structure Analysis of TaGH13 Proteins
2.7. Analysis of GH13 Expression Patterns Using Online Data
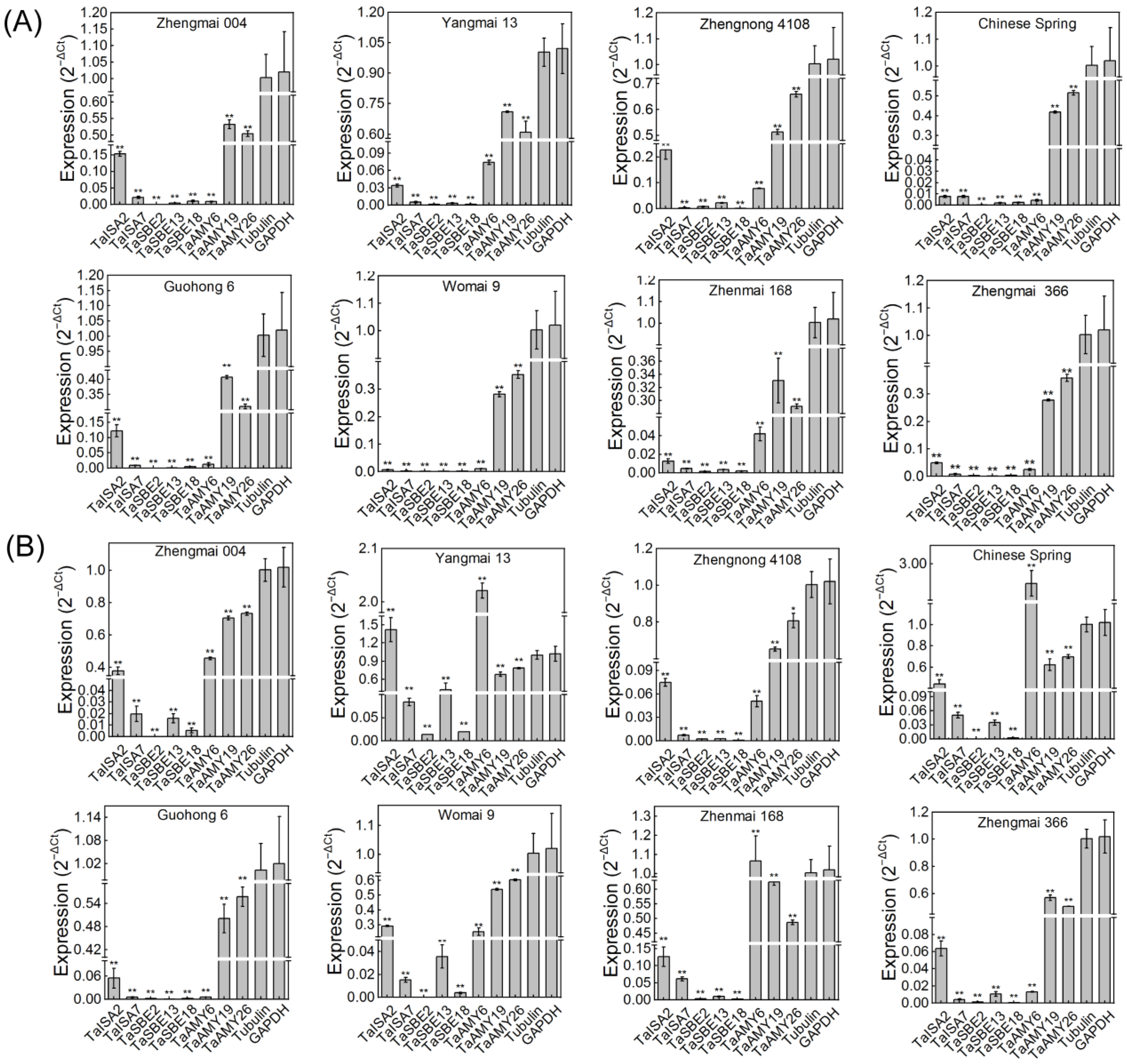
2.8. Expression of GH13 Members during Germination in Different Varieties of Wheat
3. Discussion
4. Materials and Methods
4.1. Identification and Characteristics of GH13 Genes in Wheat
4.2. Chromosomal Location, Synteny Analysis, Phylogenetic Relationships, and Gene Duplication Analysis of TaGH13 Genes
4.3. Sequence Analysis, Motif Analysis, Cis-Acting Elements Analysis, and 3D Structure Analysis of TaGH13 Members
4.4. Subcellular Localization Analysis
4.5. Expression Analysis of TaGH13 Genes Using In Silico Methods and qRT-PCR
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kilian, B.; Martin, W.; Salamini, F. Genetic diversity, evolution and domestication of wheat and barley in the Fertile Crescent. In Evolution in Action: Case Studies in Adaptive Radiation, Speciation and the Origin of Biodiversity; Springer: Berlin/Heidelberg, Germany, 2010; pp. 137–166. [Google Scholar]
- Bertoft, E. Understanding starch structure: Recent progress. Agronomy 2017, 7, 56. [Google Scholar] [CrossRef]
- Bornscheuer, U.; Buchholz, K.; Seibel, J. Enzymatic degradation of (ligno) cellulose. Angew. Chem. Int. Ed. 2014, 53, 10876–10893. [Google Scholar] [CrossRef]
- Jespersen, H.M.; Ann MacGregor, E.; Henrissat, B.; Sierks, M.R.; Svensson, B. Starch-and glycogen-debranching and branching enzymes: Prediction of structural features of the catalytic (β/α) 8-barrel domain and evolutionary relationship to other amylolytic enzymes. J. Protein Chem. 1993, 12, 791–805. [Google Scholar] [CrossRef]
- Kumar, V. Identification of the sequence motif of glycoside hydrolase 13 family members. Bioinformation 2011, 6, 61. [Google Scholar] [CrossRef]
- Machovič, M.; Janeček, Š. Domain evolution in the GH13 pullulanase subfamily with focus on the carbohydrate-binding module family 48. Biologia 2008, 63, 1057–1068. [Google Scholar] [CrossRef]
- Drula, E.; Garron, M.-L.; Dogan, S.; Lombard, V.; Henrissat, B.; Terrapon, N. The carbohydrate-active enzyme database: Functions and literature. Nucleic Acids Res. 2022, 50, D571–D577. [Google Scholar] [CrossRef]
- Janecek, S.t. α-Amylase family: Molecular biology and evolution. Prog. Biophys. Mol. Biol. 1997, 67, 67–97. [Google Scholar] [CrossRef] [PubMed]
- Pujadas, G.; Palau, J. Evolution of α-amylases: Architectural features and key residues in the stabilization of the (β/α) 8 scaffold. Mol. Biol. Evol. 2001, 18, 38–54. [Google Scholar] [CrossRef] [PubMed]
- Van Der Maarel, M.J.; Van der Veen, B.; Uitdehaag, J.C.; Leemhuis, H.; Dijkhuizen, L. Properties and applications of starch-converting enzymes of the α-amylase family. J. Biotechnol. 2002, 94, 137–155. [Google Scholar] [CrossRef] [PubMed]
- Janecek, S.; Svensson, B.; Henrissat, B. Domain evolution in the α-amylase family. J. Mol. Evol. 1997, 45, 322–331. [Google Scholar] [CrossRef] [PubMed]
- MacGregor, E.A.; Janeček, Š.; Svensson, B. Relationship of sequence and structure to specificity in the α-amylase family of enzymes. Biochim. Biophys. Acta (BBA)-Protein Struct. Mol. Enzymol. 2001, 1546, 1–20. [Google Scholar] [CrossRef]
- Kadziola, A.; Abe, J.-i.; Svensson, B.; Haser, R. Crystal and molecular structure of barley α-amylase. J. Mol. Biol. 1994, 239, 104–121. [Google Scholar] [CrossRef]
- Katsuya, Y.; Mezaki, Y.; Kubota, M.; Matsuura, Y. Three-dimensional structure of Pseudomonas isoamylase at 2.2 Å resolution. J. Mol. Biol. 1998, 281, 885–897. [Google Scholar] [CrossRef] [PubMed]
- Mikami, B.; Iwamoto, H.; Malle, D.; Yoon, H.-J.; Demirkan-Sarikaya, E.; Mezaki, Y.; Katsuya, Y. Crystal structure of pullulanase: Evidence for parallel binding of oligosaccharides in the active site. J. Mol. Biol. 2006, 359, 690–707. [Google Scholar] [CrossRef] [PubMed]
- Uitdehaag, J.; Mosi, R.; Kalk, K.H.; van der Veen, B.A.; Dijkhuizen, L.; Withers, S.G.; Dijkstra, B.W. X-ray structures along the reaction pathway of cyclodextrin glycosyltransferase elucidate catalysis in the α-amylase family. Nat. Struct. Biol. 1999, 6, 432–436. [Google Scholar] [CrossRef] [PubMed]
- Fincher, G.B. Molecular and cellular biology associated with endosperm mobilization in germinating cereal grains. Annu. Rev. Plant Biol. 1989, 40, 305–346. [Google Scholar] [CrossRef]
- Li, Q.; Yang, A. Comparative studies on seed germination of two rice genotypes with different tolerances to low temperature. Environ. Exp. Bot. 2020, 179, 104216. [Google Scholar] [CrossRef]
- Streb, S.; Delatte, T.; Umhang, M.; Eicke, S.; Schorderet, M.; Reinhardt, D.; Zeeman, S.C. Starch granule biosynthesis in Arabidopsis is abolished by removal of all debranching enzymes but restored by the subsequent removal of an endoamylase. Plant Cell 2008, 20, 3448–3466. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Xue, L.; Sun, J.; Zuo, J. The Arabidopsis BE1 gene, encoding a putative glycoside hydrolase localized in plastids, plays crucial roles during embryogenesis and carbohydrate metabolism. J. Integr. Plant Biol. 2010, 52, 273–288. [Google Scholar] [CrossRef]
- Xia, H.; Yandeau-Nelson, M.; Thompson, D.B.; Guiltinan, M.J. Deficiency of maize starch-branching enzyme i results in altered starch fine structure, decreased digestibility and reduced coleoptile growth during germination. BMC Plant Biol. 2011, 11, 95. [Google Scholar] [CrossRef]
- Li, Q.-F.; Zhang, G.-Y.; Dong, Z.-W.; Yu, H.-X.; Gu, M.-H.; Sun, S.S.; Liu, Q.-Q. Characterization of expression of the OsPUL gene encoding a pullulanase-type debranching enzyme during seed development and germination in rice. Plant Physiol. Biochem. 2009, 47, 351–358. [Google Scholar] [CrossRef] [PubMed]
- James, M.G.; Robertson, D.S.; Myers, A.M. Characterization of the maize gene sugary1, a determinant of starch composition in kernels. Plant Cell 1995, 7, 417–429. [Google Scholar] [PubMed]
- Rahman, A.; Wong, K.-s.; Jane, J.-l.; Myers, A.M.; James, M.G. Characterization of SU1 isoamylase, a determinant of storage starch structure in maize. Plant Physiol. 1998, 117, 425–435. [Google Scholar] [CrossRef]
- Qi, W.; Liu, J.; Yu, T.; Huang, S.; Song, R.; Qiao, Z. Ae1/Sbe1 maize-derived high amylose improves gut barrier function and ameliorates type II diabetes in high-fat diet-fed mice by increasing Akkermansia. Front. Nutr. 2022, 9, 999020. [Google Scholar] [CrossRef]
- Luo, J.; Ahmed, R.; Kosar-Hashemi, B.; Larroque, O.; Butardo, V.M.; Tanner, G.J.; Colgrave, M.L.; Upadhyaya, N.M.; Tetlow, I.J.; Emes, M.J. The different effects of starch synthase IIa mutations or variation on endosperm amylose content of barley, wheat and rice are determined by the distribution of starch synthase I and starch branching enzyme IIb between the starch granule and amyloplast stroma. Theor. Appl. Genet. 2015, 128, 1407–1419. [Google Scholar]
- Thitisaksakul, M.; Jiménez, R.C.; Arias, M.C.; Beckles, D.M. Effects of environmental factors on cereal starch biosynthesis and composition. J. Cereal Sci. 2012, 56, 67–80. [Google Scholar] [CrossRef]
- Tanamachi, K.; Miyazaki, M.; Matsuo, K.; Suriyasak, C.; Tamada, A.; Matsuyama, K.; Iwaya-Inoue, M.; Ishibashi, Y. Differential responses to high temperature during maturation in heat-stress-tolerant cultivars of Japonica rice. Plant Prod. Sci. 2016, 19, 300–308. [Google Scholar] [CrossRef]
- Thalmann, M.; Santelia, D. Starch as a determinant of plant fitness under abiotic stress. New Phytol. 2017, 214, 943–951. [Google Scholar] [CrossRef]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME Suite: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, W202–W208. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Force, A.; Lynch, M.; Pickett, F.B.; Amores, A.; Yan, Y.-l.; Postlethwait, J. Preservation of duplicate genes by complementary, degenerative mutations. Genetics 1999, 151, 1531–1545. [Google Scholar] [CrossRef]
- Acevedo, E.; Silva, P.; Silva, H. Wheat growth and physiology. Bread Wheat Improv. Prod. 2002, 30, 39–70. [Google Scholar]
- Stam, M.R.; Danchin, E.G.; Rancurel, C.; Coutinho, P.M.; Henrissat, B. Dividing the large glycoside hydrolase family 13 into subfamilies: Towards improved functional annotations of α-amylase-related proteins. Protein Eng. Des. Sel. 2006, 19, 555–562. [Google Scholar] [CrossRef]
- Kubo, A.; Fujita, N.; Harada, K.; Matsuda, T.; Satoh, H.; Nakamura, Y. The starch-debranching enzymes isoamylase and pullulanase are both involved in amylopectin biosynthesis in rice endosperm. Plant Physiol. 1999, 121, 399–410. [Google Scholar] [CrossRef]
- Yamasaki, Y.; Nakashima, S.; Konno, H. Pullulanase from rice endosperm. Acta Biochim. Pol. 2008, 55, 507–510. [Google Scholar] [CrossRef]
- Facon, M.; Lin, Q.; Azzaz, A.M.; Hennen-Bierwagen, T.A.; Myers, A.M.; Putaux, J.-L.; Roussel, X.; D’Hulst, C.; Wattebled, F. Distinct functional properties of isoamylase-type starch debranching enzymes in monocot and dicot leaves. Plant Physiol. 2013, 163, 1363–1375. [Google Scholar] [CrossRef] [PubMed]
- Streb, S.; Zeeman, S.C. Starch metabolism in Arabidopsis. Arab. Book/Am. Soc. Plant Biol. 2012, 10, e0160. [Google Scholar] [CrossRef] [PubMed]
- Kurotani, A.; Tokmakov, A.A.; Sato, K.-I.; Stefanov, V.E.; Yamada, Y.; Sakurai, T. Localization-specific distributions of protein pI in human proteome are governed by local pH and membrane charge. BMC Mol. Cell Biol. 2019, 20, 36. [Google Scholar] [CrossRef] [PubMed]
- Tokmakov, A.A.; Kurotani, A.; Sato, K.-I. Protein pI and intracellular localization. Front. Mol. Biosci. 2021, 8, 775736. [Google Scholar] [CrossRef]
- Peltier, J.-B.; Friso, G.; Kalume, D.E.; Roepstorff, P.; Nilsson, F.; Adamska, I.; van Wijka, K.J. Proteomics of the chloroplast: Systematic identification and targeting analysis of lumenal and peripheral thylakoid proteins. Plant Cell 2000, 12, 319–341. [Google Scholar] [CrossRef] [PubMed]
- Kitajima, A.; Asatsuma, S.; Okada, H.; Hamada, Y.; Kaneko, K.; Nanjo, Y.; Kawagoe, Y.; Toyooka, K.; Matsuoka, K.; Takeuchi, M. The rice α-amylase glycoprotein is targeted from the Golgi apparatus through the secretory pathway to the plastids. Plant Cell 2009, 21, 2844–2858. [Google Scholar] [CrossRef]
- Seung, D.; Thalmann, M.; Sparla, F.; Abou Hachem, M.; Lee, S.K.; Issakidis-Bourguet, E.; Svensson, B.; Zeeman, S.C.; Santelia, D. Arabidopsis thaliana AMY3 is a unique redox-regulated chloroplastic α-amylase. J. Biol. Chem. 2013, 288, 33620–33633. [Google Scholar] [CrossRef]
- Doyle, E.A.; Lane, A.M.; Sides, J.M.; Mudgett, M.B.; Monroe, J.D. An α-amylase (At4g25000) in Arabidopsis leaves is secreted and induced by biotic and abiotic stress. Plant Cell Environ. 2007, 30, 388–398. [Google Scholar] [CrossRef]
- Blanc, G.; Hokamp, K.; Wolfe, K.H. A recent polyploidy superimposed on older large-scale duplications in the Arabidopsis genome. Genome. Res. 2003, 13, 137–144. [Google Scholar] [CrossRef]
- Bowers, J.E.; Chapman, B.A.; Rong, J.; Paterson, A.H. Unravelling angiosperm genome evolution by phylogenetic analysis of chromosomal duplication events. Nature 2003, 422, 433–438. [Google Scholar] [CrossRef] [PubMed]
- Paterson, A.; Bowers, J.; Chapman, B. Ancient polyploidization predating divergence of the cereals, and its consequences for comparative genomics. Proc. Natl. Acad. Sci. USA 2004, 101, 9903–9908. [Google Scholar] [CrossRef] [PubMed]
- Panchy, N.; Lehti-Shiu, M.; Shiu, S.-H. Evolution of gene duplication in plants. Plant Physiol. 2016, 171, 2294–2316. [Google Scholar] [CrossRef]
- Qiao, X.; Li, Q.; Yin, H.; Qi, K.; Li, L.; Wang, R.; Zhang, S.; Paterson, A.H. Gene duplication and evolution in recurring polyploidization–diploidization cycles in plants. Genome Biol. 2019, 20, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Consortium, T.I.W.G.S.; Appels, R.; Eversole, K.; Stein, N.; Feuillet, C.; Keller, B.; Rogers, J.; Pozniak, C.J.; Choulet, F.; Distelfeld, A.; et al. Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 2018, 361, eaar7191. [Google Scholar] [CrossRef]
- El Baidouri, M.; Murat, F.; Veyssiere, M.; Molinier, M.; Flores, R.; Burlot, L.; Alaux, M.; Quesneville, H.; Pont, C.; Salse, J. Reconciling the evolutionary origin of bread wheat (Triticum aestivum). New Phytol. 2017, 213, 1477–1486. [Google Scholar] [CrossRef] [PubMed]
- Consortium, I.W.G.S.; Mayer, K.F.; Rogers, J.; Doležel, J.; Pozniak, C.; Eversole, K.; Feuillet, C.; Gill, B.; Friebe, B.; Lukaszewski, A.J. A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 2014, 345, 1251788. [Google Scholar]
- Damaris, R.N.; Lin, Z.; Yang, P.; He, D. The rice alpha-amylase, conserved regulator of seed maturation and germination. Int. J. Mol. Sci. 2019, 20, 450. [Google Scholar] [CrossRef]
- Yang, T.; Li, H.; Li, L.; Wei, W.; Huang, Y.; Xiong, F.; Wei, M. Genome-wide characterization and expression analysis of α-amylase and β-amylase genes underlying drought tolerance in cassava. BMC Genom. 2023, 24, 190. [Google Scholar] [CrossRef]
- Nakamura, Y.; Utsumi, Y.; Sawada, T.; Aihara, S.; Utsumi, C.; Yoshida, M.; Kitamura, S. Characterization of the reactions of starch branching enzymes from rice endosperm. Plant Cell Physiol. 2010, 51, 776–794. [Google Scholar] [CrossRef]
- Xu, X.; Fang, J.; Wang, W.; Guo, J.; Chen, P.; Cheng, J.; Shen, Z. Expression of a bacterial α-amylase gene in transgenic rice seeds. Transgenic Res. 2008, 17, 645–650. [Google Scholar] [CrossRef]
- Lin, Q.; Facon, M.; Putaux, J.L.; Dinges, J.R.; Wattebled, F.; d’Hulst, C.; Hennen-Bierwagen, T.A.; Myers, A.M. Function of isoamylase-type starch debranching enzymes ISA 1 and ISA 2 in the Z ea mays leaf. New Phytol. 2013, 200, 1009–1021. [Google Scholar] [CrossRef] [PubMed]
- Boyer, L.; Roussel, X.; Courseaux, A.; Ndjindji, O.M.; Lancelon-Pin, C.; Putaux, J.L.; Tetlow, I.J.; Emes, M.J.; Pontoire, B.; D’hulst, C. Expression of Escherichia coli glycogen branching enzyme in an Arabidopsis mutant devoid of endogenous starch branching enzymes induces the synthesis of starch-like polyglucans. Plant Cell Environ. 2016, 39, 1432–1447. [Google Scholar] [CrossRef] [PubMed]
- Ranjhan, S.; Karrer, E.E.; Rodriguez, R.L. Localizing a-amylase gene expression in germinated rice grains. Plant Cell Physiol. 1992, 33, 73–79. [Google Scholar]
- Asatsuma, S.; Sawada, C.; Itoh, K.; Okito, M.; Kitajima, A.; Mitsui, T. Involvement of α-amylase I-1 in starch degradation in rice chloroplasts. Plant Cell Physiol. 2005, 46, 858–869. [Google Scholar] [CrossRef]
- Fujita, N.; Toyosawa, Y.; Utsumi, Y.; Higuchi, T.; Hanashiro, I.; Ikegami, A.; Akuzawa, S.; Yoshida, M.; Mori, A.; Inomata, K. Characterization of pullulanase (PUL)-deficient mutants of rice (Oryza sativa L.) and the function of PUL on starch biosynthesis in the developing rice endosperm. J. Exp. Bot. 2009, 60, 1009–1023. [Google Scholar] [CrossRef] [PubMed]
- Orzechowski, S. Starch metabolism in leaves. Acta Biochim. Pol. 2008, 55, 435–445. [Google Scholar] [CrossRef]
- Rushton, P.J.; Somssich, I.E.; Ringler, P.; Shen, Q.J. WRKY transcription factors. Trends Plant Sci. 2010, 15, 247–258. [Google Scholar] [CrossRef]
- Swaminathan, K.; Peterson, K.; Jack, T. The plant B3 superfamily. Trends Plant Sci. 2008, 13, 647–655. [Google Scholar] [CrossRef]
- Dubos, C.; Stracke, R.; Grotewold, E.; Weisshaar, B.; Martin, C.; Lepiniec, L. MYB transcription factors in Arabidopsis. Trends Plant Sci. 2010, 15, 573–581. [Google Scholar] [CrossRef]
- Kiełbowicz-Matuk, A. Involvement of plant C2H2-type zinc finger transcription factors in stress responses. Plant Sci. 2012, 185, 78–85. [Google Scholar] [CrossRef] [PubMed]
- Feng, K.; Hou, X.-L.; Xing, G.-M.; Liu, J.-X.; Duan, A.-Q.; Xu, Z.-S.; Li, M.-Y.; Zhuang, J.; Xiong, A.-S. Advances in AP2/ERF super-family transcription factors in plant. Crit. Rev. Biotechnol. 2020, 40, 750–776. [Google Scholar] [CrossRef] [PubMed]
- Pandey, R.; Agarwal, R.; Jeevaratnam, K.; Sharma, G. Osmotic stress-induced alterations in rice (Oryza sativa L.) and recovery on stress release. Plant Growth Regul. 2004, 42, 79–87. [Google Scholar] [CrossRef]
- Xiao, Y.; Huang, X.; Shen, Y.; Huang, Z. A novel wheat α-amylase inhibitor gene, tahps, significantly improves the salt and drought tolerance of transgenic arabidopsis. Physiol. Plant. 2013, 148, 273–283. [Google Scholar] [CrossRef] [PubMed]
- Gilding, E.K.; Frere, C.H.; Cruickshank, A.; Rada, A.K.; Prentis, P.J.; Mudge, A.M.; Mace, E.S.; Jordan, D.R.; Godwin, I.D. Allelic variation at a single gene increases food value in a drought-tolerant staple cereal. Nat. Commun. 2013, 4, 1483. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; ur Rehman, S.; Yamaguchi, H.; Hitachi, K.; Tsuchida, K.; Yamaguchi, T.; Sunohara, Y.; Matsumoto, H.; Komatsu, S. Proteomic analysis of the effect of plant-derived smoke on soybean during recovery from flooding stress. J. Proteom. 2018, 181, 238–248. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.S.; Hwang, W.H.; Jeong, J.H.; Ahn, S.H.; Baek, J.S.; Jeong, H.Y.; Park, H.K.; Ku, B.I.; Yun, J.T.; Lee, G.H. Analysis of the distribution of assimilation products and the characteristics of transcriptomes in rice by submergence during the ripening stage. BMC Genom. 2019, 20, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Zhu, T.; Wang, L.; Rimbert, H.; Rodriguez, J.C.; Deal, K.R.; De Oliveira, R.; Choulet, F.; Keeble-Gagnère, G.; Tibbits, J.; Rogers, J. Optical maps refine the bread wheat Triticum aestivum cv. Chinese Spring genome assembly. The. Plant. J. 2021, 107, 303–314. [Google Scholar] [CrossRef] [PubMed]
- Cheng, C.; Liu, F.; Sun, X.; Wang, B.; Liu, J.; Ni, X.; Hu, C.; Deng, G.; Tong, Z.; Zhang, Y. Genome-wide identification of FAD gene family and their contributions to the temperature stresses and mutualistic and parasitic fungi colonization responses in banana. Int. J. Biol. Macromol. 2022, 204, 661–676. [Google Scholar] [CrossRef] [PubMed]
- Choudhuri, S. Bioinformatics for Beginners: Genes, Genomes, Molecular Evolution, Databases and Analytical Tools; Elsevier: Amsterdam, The Netherlands, 2014. [Google Scholar]
- Wang, J.; Chitsaz, F.; Derbyshire, M.K.; Gonzales, N.R.; Gwadz, M.; Lu, S.; Marchler, G.H.; Song, J.S.; Thanki, N.; Yamashita, R.A. The conserved domain database in 2023. Nucleic Acids Res. 2023, 51, D384–D388. [Google Scholar] [CrossRef] [PubMed]
- Paysan-Lafosse, T.; Blum, M.; Chuguransky, S.; Grego, T.; Pinto, B.L.; Salazar, G.A.; Bileschi, M.L.; Bork, P.; Bridge, A.; Colwell, L. InterPro in 2022. Nucleic Acids Res. 2023, 51, D418–D427. [Google Scholar] [CrossRef]
- Artimo, P.; Jonnalagedda, M.; Arnold, K.; Baratin, D.; Csardi, G.; de Castro, E.; Duvaud, S.; Flegel, V.; Fortier, A.; Gasteiger, E.; et al. ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res. 2012, 40, W597–W603. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; DeBarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.-h.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef]
- Xu, B.; Yang, Z. pamlX: A Graphical User Interface for PAML. Mol. Biol. Evol. 2013, 30, 2723–2724. [Google Scholar] [CrossRef]
- Tyagi, S.; Verma, P.C.; Singh, K.; Upadhyay, S.K. Molecular characterization of ascorbate peroxidase (APX) and APX-related (APX-R) genes in Triticum aestivum L. Genomics 2020, 112, 4208–4223. [Google Scholar] [CrossRef]
- Rombauts, S.; Déhais, P.; Van Montagu, M.; Rouzé, P. PlantCARE, a plant cis-acting regulatory element database. Nucleic Acids Res. 1999, 27, 295–296. [Google Scholar] [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Zidek, A.; Potapenko, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Lai, Y.; Lv, L.; Ji, M.; Han, K.; Yan, D.; Lu, Y.; Peng, J.; Rao, S.; Yan, F. Fasciclin-like arabinogalactan gene family in Nicotiana benthamiana: Genome-wide identification, classification and expression in response to pathogens. BMC Plant Biol. 2020, 20, 305. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.; Shim, C.; Lee, J.; Wangchuk, C. Hot Water Treatment as Seed Disinfection Techniques for Organic and Eco-Friendly Environmental Agricultural Crop Cultivation. Agriculture 2022, 12, 1081. [Google Scholar] [CrossRef]
- Vergara-Irigaray, M.; Fookes, M.C.; Thomson, N.R.; Tang, C.M. RNA-seq analysis of the influence of anaerobiosis and FNR on Shigella flexneri. BMC Genom. 2014, 15, 438. [Google Scholar] [CrossRef]


| Name | Gene ID | Number of Amino Acids | Molecular Weight | Theoretical pI | Instability Index | Aliphatic Index | Grand Average Hydropathicity |
|---|---|---|---|---|---|---|---|
| TaISA1 | TraesCS1A02G247100 | 793 | 85,711.73 | 6.31 | 45.6 | 86.09 | −0.03 |
| TaISA2 | TraesCS1B02G257700 | 788 | 85,511.58 | 6.46 | 47.11 | 87.74 | −0.024 |
| TaISA3 | TraesCS1D02G246300 | 789 | 85,662.71 | 6.75 | 45.42 | 85.65 | −0.056 |
| TaISA4 | TraesCS5A02G248700 | 775 | 86,074.49 | 6.1 | 35.38 | 74.12 | −0.428 |
| TaISA5 | TraesCS5B02G246400 | 776 | 86,254.8 | 6.13 | 35.72 | 73.53 | −0.432 |
| TaISA6 | TraesCS5D02G255800 | 776 | 86,226.7 | 6.07 | 35.56 | 73.16 | −0.439 |
| TaISA7 | TraesCS7A02G251400 | 790 | 88,718.4 | 5.72 | 40.26 | 69.59 | −0.343 |
| TaISA8 | TraesCS7D02G249500 | 791 | 88,822.5 | 5.65 | 39.91 | 69.53 | −0.341 |
| TaISA9 | TraesCS7B02G139700 | 790 | 88,715.53 | 5.84 | 41.48 | 69.97 | −0.333 |
| TaSBE1 | TraesCS2A02G310300 | 834 | 94,039.27 | 5.71 | 39.16 | 65.32 | −0.511 |
| TaSBE2 | TraesCS2B02G327300 | 836 | 94,367.65 | 5.56 | 38.15 | 66.67 | −0.499 |
| TaSBE3 | TraesCS2D02G308600 | 836 | 94,433.73 | 5.63 | 39.65 | 65.41 | −0.511 |
| TaSBE4 | TraesCS2A02G293400 | 828 | 93,628.97 | 5.4 | 38.9 | 70.53 | −0.481 |
| TaSBE5 | TraesCS2B02G309500 | 828 | 93,662.12 | 5.46 | 37.7 | 71.12 | −0.475 |
| TaSBE6 | TraesCS2D02G290800 | 825 | 93,184.56 | 5.49 | 37.3 | 71.14 | −0.46 |
| TaSBE7 | TraesCS7A02G549300 | 645 | 72,979.6 | 7.16 | 37.76 | 74.7 | −0.465 |
| TaSBE8 | TraesCS7D02G535600 | 680 | 76,951.02 | 6.61 | 37.44 | 76.59 | −0.432 |
| TaSBE9 | TraesCS7B02G472300 | 642 | 72,473.91 | 6.59 | 36.62 | 74.3 | −0.474 |
| TaSBE10 | TraesCS7A02G549200 | 635 | 71,927.42 | 6.27 | 34.31 | 69.09 | −0.414 |
| TaSBE11 | TraesCS7B02G472400 | 632 | 71,376.67 | 6.27 | 35.71 | 70.68 | −0.369 |
| TaSBE12 | TraesCS7D02G535500 | 650 | 73,492.25 | 6.55 | 35.56 | 71.12 | −0.35 |
| TaSBE13 | TraesCS7A02G549100 | 846 | 95,331.37 | 6.43 | 34 | 64.81 | −0.501 |
| TaSBE14 | TraesCS7B02G472500 | 798 | 90,203.54 | 6.11 | 31.37 | 66.74 | −0.479 |
| TaSBE15 | TraesCS7D02G535400 | 830 | 93,577.25 | 6.27 | 33.07 | 64.76 | −0.522 |
| TaSBE16 | TraesCS7A02G336400 | 914 | 105,036.2 | 6.23 | 39.21 | 70.51 | −0.547 |
| TaSBE17 | TraesCS7B02G248000 | 914 | 104,905.1 | 6.24 | 38.97 | 70.62 | −0.536 |
| TaSBE18 | TraesCS7D02G344000 | 914 | 104,889 | 6.19 | 39.29 | 70.73 | −0.537 |
| TaAMY1 | TraesCS2A02G289800 | 415 | 47,254.95 | 5.44 | 47.36 | 70.02 | −0.52 |
| TaAMY2 | TraesCS2D02G287800 | 415 | 47,111.72 | 5.44 | 47.59 | 70.96 | −0.504 |
| TaAMY3 | TraesCS2B02G306400 | 415 | 47,225.89 | 5.44 | 48 | 70.96 | −0.518 |
| TaAMY4 | TraesCS3A02G248000 | 871 | 97,498.62 | 5.48 | 38.15 | 75.36 | −0.452 |
| TaAMY5 | TraesCS3D02G248000 | 833 | 93,182.72 | 5.62 | 36.57 | 74.95 | −0.46 |
| TaAMY6 | TraesCS3B02G276700 | 871 | 97228.37 | 5.63 | 38.5 | 74.35 | −0.459 |
| TaAMY7 | TraesCS4B02G285600 | 361 | 40,278.79 | 6.22 | 33.29 | 85.07 | −0.194 |
| TaAMY8 | TraesCS4B02G285400 | 438 | 48,330.78 | 6.24 | 34.77 | 81.74 | −0.184 |
| TaAMY9 | TraesCS4D02G284400 | 437 | 48,337.83 | 6.75 | 36.28 | 82.17 | −0.194 |
| TaAMY10 | TraesCS5A02G238100 | 436 | 47,610 | 5.95 | 29.27 | 81.67 | −0.189 |
| TaAMY11 | TraesCS5D02G245000 | 436 | 47,656.04 | 6.27 | 29.49 | 80.32 | −0.228 |
| TaAMY12 | TraesCS5B02G236600 | 436 | 47,570.97 | 6.08 | 28.92 | 81.88 | −0.204 |
| TaAMY13 | TraesCS5A02G464500 | 413 | 45,370.5 | 8.01 | 26.06 | 78.64 | −0.296 |
| TaAMY14 | TraesCS5B02G476000 | 413 | 45,458.6 | 8.63 | 26.66 | 79.35 | −0.298 |
| TaAMY15 | TraesCS5B02G475700 | 413 | 45,349.51 | 8.62 | 25.91 | 80.29 | −0.283 |
| TaAMY16 | TraesCS5D02G477100 | 413 | 45,461.68 | 8.47 | 26.08 | 80.07 | −0.303 |
| TaAMY17 | TraesCS7A02G384000 | 463 | 50,860.8 | 6.14 | 27.1 | 80.54 | −0.158 |
| TaAMY18 | TraesCS7D02G380500 | 440 | 48,343.83 | 5.47 | 22.65 | 83.64 | −0.137 |
| TaAMY19 | TraesCS7A02G383200 | 437 | 47,584.79 | 5.69 | 20.54 | 78.65 | −0.15 |
| TaAMY20 | TraesCS7D02G379700 | 486 | 53,511.52 | 5.66 | 22.18 | 75.72 | −0.203 |
| TaAMY21 | TraesCS7B02G286100 | 438 | 47,717.9 | 5.64 | 20.54 | 79.16 | −0.152 |
| TaAMY22 | TraesCS7B02G286000 | 438 | 47,809.05 | 5.66 | 20.2 | 80.25 | −0.155 |
| TaAMY23 | TraesCS7B02G286700 | 439 | 47,989.28 | 5.48 | 20.79 | 80.73 | −0.144 |
| TaAMY24 | TraesCS7A02G383900 | 437 | 47,788.09 | 5.6 | 20.08 | 78.19 | −0.161 |
| TaAMY25 | TraesCS7D02G380400 | 437 | 47,877.09 | 5.69 | 20.6 | 77.76 | −0.182 |
| TaAMY26 | TraesCS6A02G334200 | 427 | 47,378.47 | 5.83 | 22.1 | 81.1 | −0.306 |
| TaAMY27 | TraesCS6A02G334100 | 427 | 47,249.36 | 6 | 19.86 | 81.33 | −0.281 |
| TaAMY28 | TraesCS6A02G319300 | 459 | 50,862.44 | 5.97 | 23.11 | 80.15 | −0.323 |
| TaAMY29 | TraesCS6D02G298500 | 459 | 50,789.34 | 5.89 | 22.45 | 81 | −0.302 |
| TaAMY30 | TraesCS6B02G349500 | 425 | 47,165.24 | 5.84 | 20.07 | 81.27 | −0.3 |
| TaAMY31 | TraesCS6B02G349800 | 425 | 47,209.29 | 5.84 | 20.18 | 81.51 | −0.302 |
| TaAMY32 | TraesCS6B02G349700 | 425 | 47,209.29 | 5.84 | 20.07 | 81.27 | −0.307 |
| TaAMY33 | TraesCS6B02G364900 | 427 | 47,332.49 | 6.01 | 22.08 | 81.36 | −0.301 |
| TaAMY34 | TraesCS6D02G313300 | 427 | 47,346.51 | 5.92 | 23.27 | 81.57 | −0.295 |
| TaAMY35 | TraesCS6D02G313500 | 427 | 47,322.52 | 6.1 | 20.52 | 79.06 | −0.294 |
| TaAMY36 | TraesCS6B02G364800 | 425 | 47,093.21 | 5.92 | 20.59 | 81.27 | −0.291 |
| TaPU1 | TraesCS7A02G133500 | 963 | 106,019.4 | 5.57 | 34.03 | 79.41 | −0.275 |
| TaPU2 | TraesCS7B02G034600 | 955 | 104,967 | 5.44 | 32.5 | 80.69 | −0.263 |
| TaPU3 | TraesCS7D02G133100 | 963 | 105,770.9 | 5.31 | 33.16 | 79.29 | −0.266 |
| Gene ID | Name |
|---|---|
| At4g25000 | AtAMY1 |
| At1g76130 | AtAMY2 |
| At1g69830 | AtAMY3 |
| At2g36390 | AtSBE2.1 |
| At5g03650 | AtSBE2.2 |
| At3g20440 | AtSBE3 |
| At5g04360 | AtPU1 |
| At2g39930 | AtISA1 |
| At1g03310 | AtISA2 |
| At4g09020 | AtISA3 |
| Os02g0765600 | OsAMY1A |
| Os01g0357400 | OsAMY1B |
| Os06g0713800 | OsAMY2A |
| Os09g0457400 | OsAMY3A |
| Os09g0457600 | OsAMY3B |
| Os09g0457800 | OsAMY3C |
| Os08g0473900 | OsAMY3D |
| Os08g0473600 | OsAMY3E |
| Os08g0520900 | OsISA1 |
| Os05g0393700 | OsISA2 |
| Os09g0469400 | OsISA3 |
| Os06g0726400 | OsSBE1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yin, Y.; Cui, D.; Sun, H.; Guan, P.; Zhang, H.; Chi, Q.; Jiao, Z. Genome-Wide Identification, Characterization, and Expression Analysis of Four Subgroup Members of the GH13 Family in Wheat (Triticum aestivum L.). Int. J. Mol. Sci. 2024, 25, 3399. https://doi.org/10.3390/ijms25063399
Yin Y, Cui D, Sun H, Guan P, Zhang H, Chi Q, Jiao Z. Genome-Wide Identification, Characterization, and Expression Analysis of Four Subgroup Members of the GH13 Family in Wheat (Triticum aestivum L.). International Journal of Molecular Sciences. 2024; 25(6):3399. https://doi.org/10.3390/ijms25063399
Chicago/Turabian StyleYin, Yue, Dongjie Cui, Hao Sun, Panfeng Guan, Hanfeng Zhang, Qing Chi, and Zhen Jiao. 2024. "Genome-Wide Identification, Characterization, and Expression Analysis of Four Subgroup Members of the GH13 Family in Wheat (Triticum aestivum L.)" International Journal of Molecular Sciences 25, no. 6: 3399. https://doi.org/10.3390/ijms25063399






