Hfq-Antisense RNA I Binding Regulates RNase E-Dependent RNA Stability and ColE1 Plasmid Copy Number
Abstract
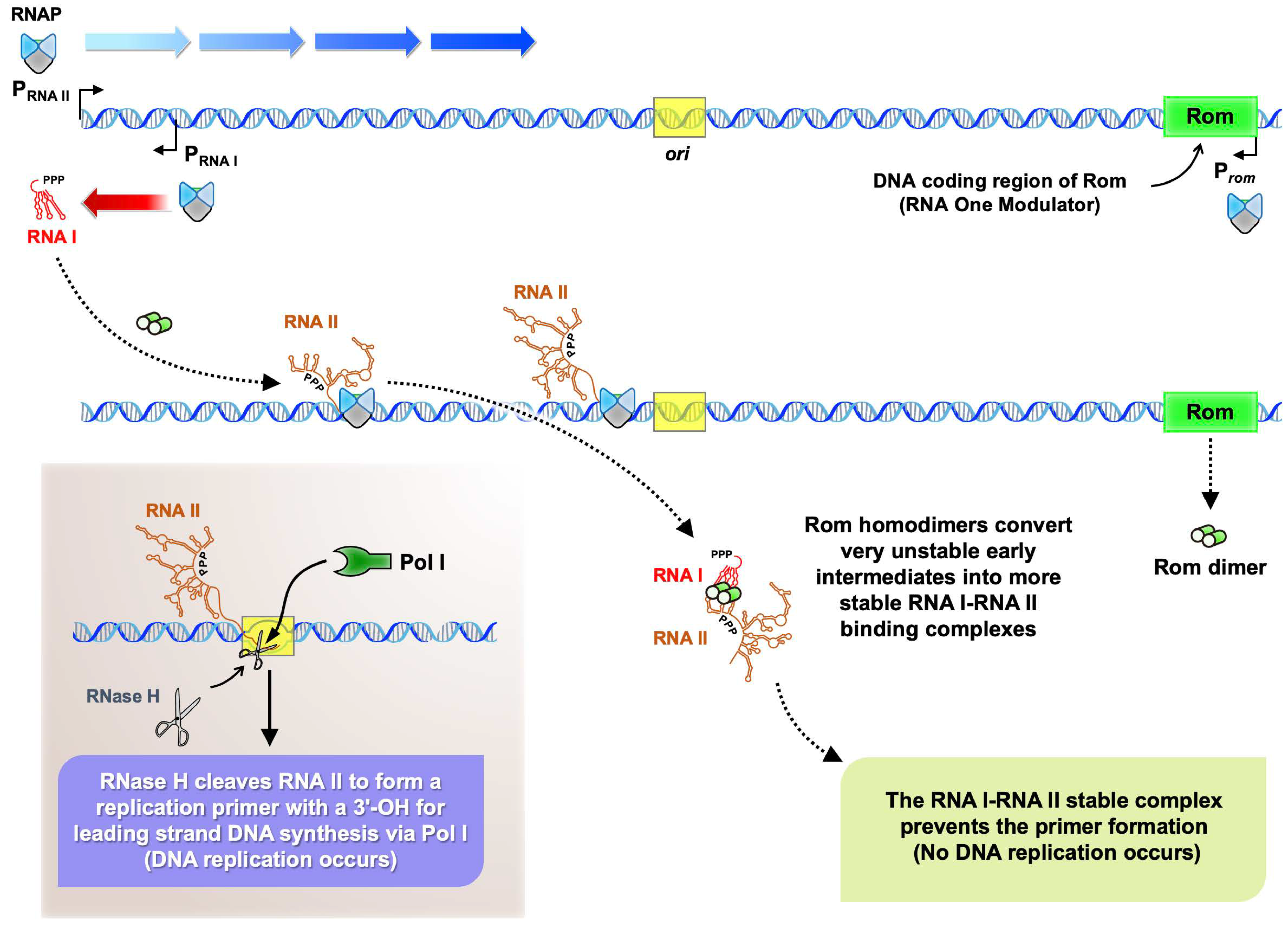
:1. Introduction
2. Results
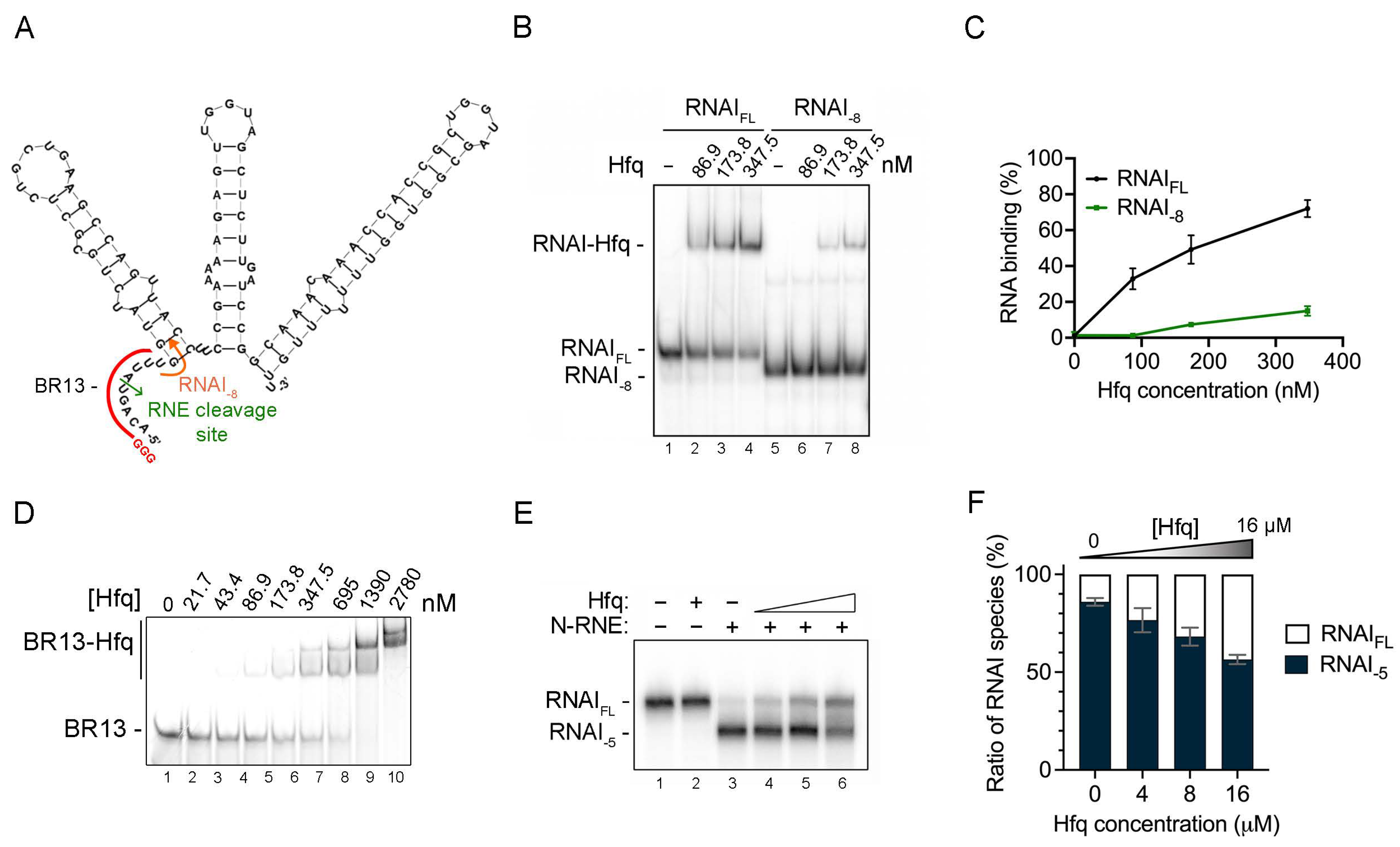
2.1. Deletion of the RNA I 5′ End Single-Stranded Region Significantly Reduces Hfq–RNA I Binding, and High-Affinity Hfq Binding Limits RNase E Cleavage
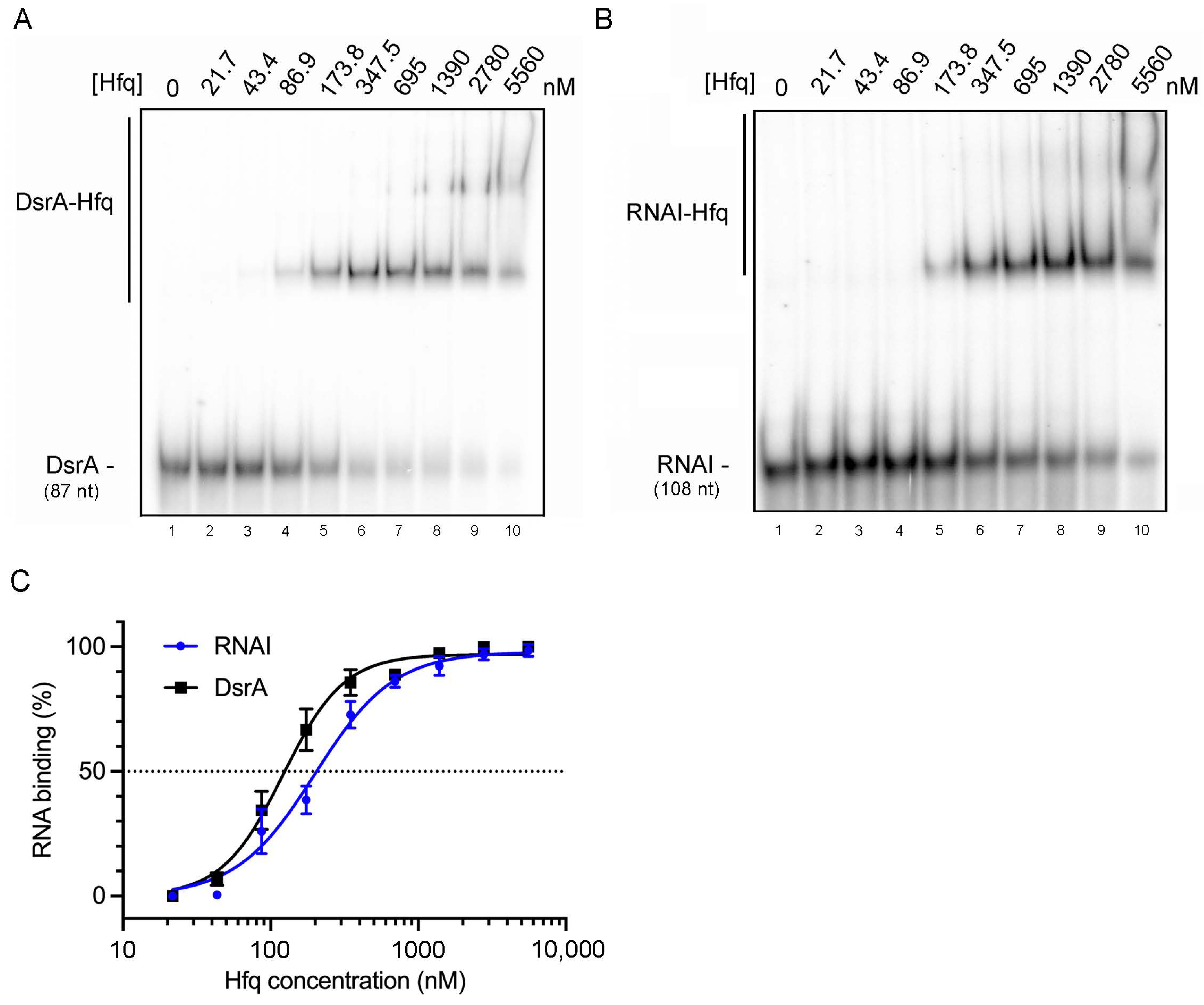
2.2. RNA I Displays a Sub-Micromolar Binding Affinity for Hfq Protein In Vitro
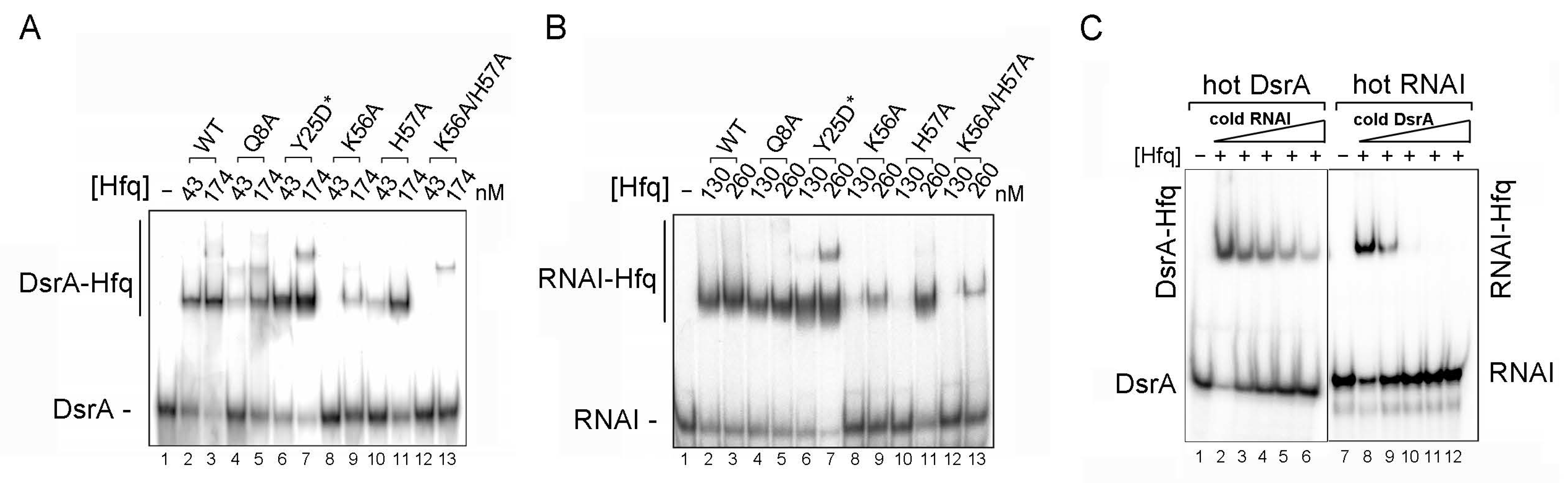
2.3. Proximal Face Mutants of Hfq Decrease Both DsrA and RNA I Binding, and These Two RNAs Compete with Each Other for Hfq Binding
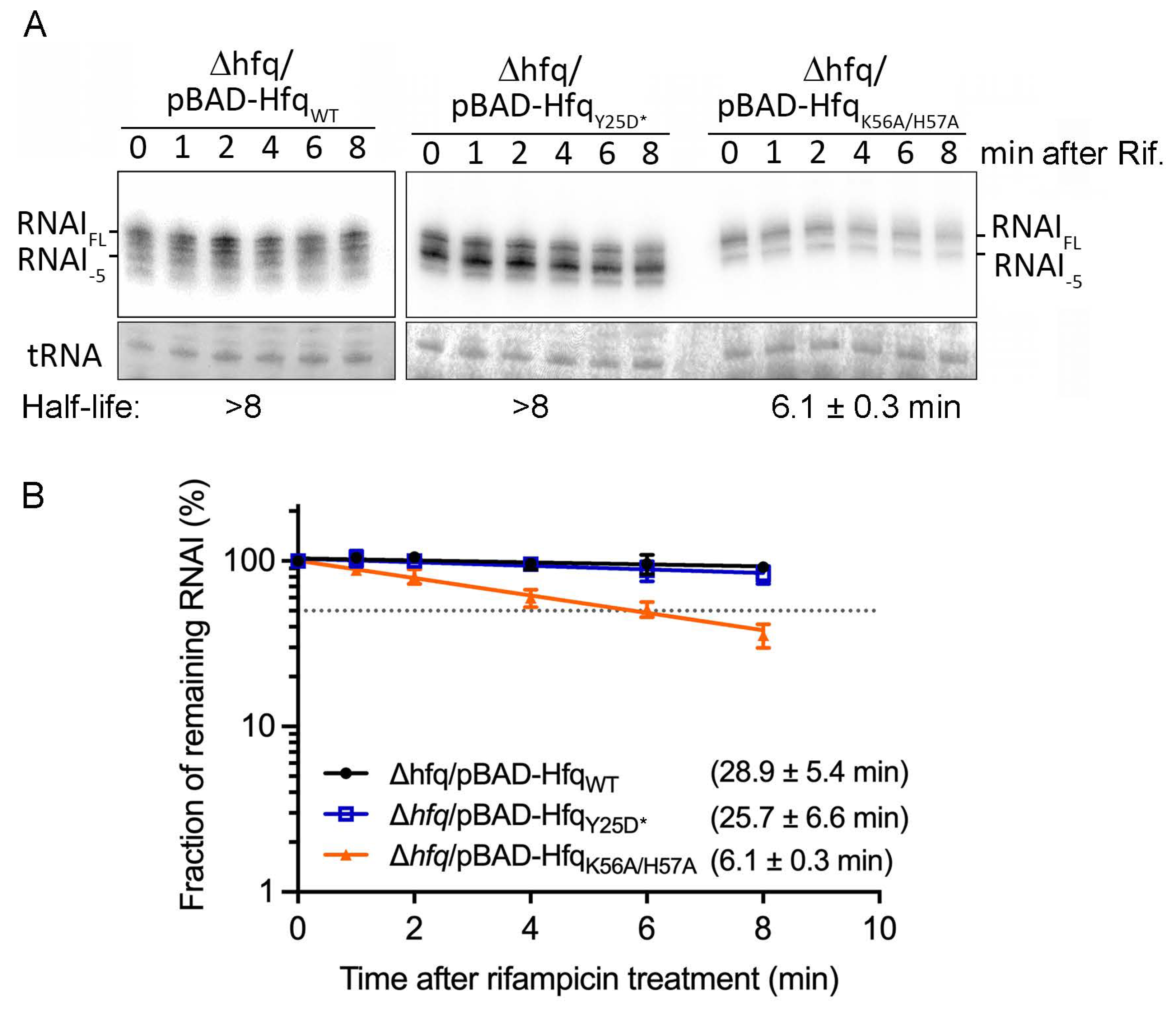
2.4. Hfq protects RNA I from RNase E-Mediated Degradation Predominantly through Its Proximal but Not Distal Face in E. coli
2.5. Hfq Controls ColE1 Plasmid Copy Number by Regulating RNA I Stability
3. Discussion
3.1. Hfq Might Have Other Binding Sites in Addition to the 5′ end Single-Stranded AU-Rich Region of RNA I
3.2. Hfq Binding Face for RNA I
3.3. How Mutations of Hfq Binding Surfaces Affect RNA I Stability
3.4. Possible Competition between Plasmid-Borne Antisense RNA and Chromosome-Borne sRNAs
4. Materials and Methods
4.1. Bacterial Strains and Plasmid Construction
4.2. Protein Expression and Purification
4.3. RNA Preparation, N-RNE In Vitro Cleavage, In Vitro Protein–RNA Gel-Shift Assay, and RNA–Protein Competition Assay
4.4. RNA Half-Life Determination, Total RNA Extraction, and Northern Blot Analysis
4.5. Plasmid Copy Number Determination
4.6. DNA Isolation and Quantitative PCR (qPCR)
4.7. Quantification and Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Waters, L.S.; Storz, G. Regulatory RNAs in bacteria. Cell 2009, 136, 615–628. [Google Scholar] [CrossRef] [PubMed]
- Santiago-Frangos, A.; Woodson, S.A. Hfq chaperone brings speed dating to bacterial sRNA. Wiley Interdiscip. Rev. RNA 2018, 9, e1475. [Google Scholar] [CrossRef] [PubMed]
- Gottesman, S.; Storz, G. Bacterial small RNA regulators: Versatile roles and rapidly evolving variations. Cold Spring Harb. Perspect. Biol. 2011, 3, a003798. [Google Scholar] [CrossRef] [PubMed]
- Kawamoto, H.; Koide, Y.; Morita, T.; Aiba, H. Base-pairing requirement for RNA silencing by a bacterial small RNA and acceleration of duplex formation by Hfq. Mol. Microbiol. 2006, 61, 1013–1022. [Google Scholar] [CrossRef] [PubMed]
- Møller, T.; Franch, T.; Udesen, C.; Gerdes, K.; Valentin-Hansen, P. Spot 42 RNA mediates discoordinate expression of the E. coli galactose operon. Genes Dev. 2002, 16, 1696–1706. [Google Scholar] [CrossRef] [PubMed]
- Papenfort, K.; Vanderpool, C.K. Target activation by regulatory RNAs in bacteria. FEMS Microbiol. Rev. 2015, 39, 362–378. [Google Scholar] [CrossRef] [PubMed]
- De Lay, N.; Schu, D.J.; Gottesman, S. Bacterial small RNA-based negative regulation: Hfq and its accomplices. J. Biol. Chem. 2013, 288, 7996–8003. [Google Scholar] [CrossRef]
- Melamed, S.; Peer, A.; Faigenbaum-Romm, R.; Gatt, Y.E.; Reiss, N.; Bar, A.; Altuvia, Y.; Argaman, L.; Margalit, H. Global Mapping of Small RNA-Target Interactions in Bacteria. Mol. Cell 2016, 63, 884–897. [Google Scholar] [CrossRef]
- Han, K.; Tjaden, B.; Lory, S. GRIL-seq provides a method for identifying direct targets of bacterial small regulatory RNA by in vivo proximity ligation. Nat. Microbiol. 2016, 2, 16239. [Google Scholar] [CrossRef]
- Iosub, I.A.; van Nues, R.W.; McKellar, S.W.; Nieken, K.J.; Marchioretto, M.; Sy, B.; Tree, J.J.; Viero, G.; Granneman, S. Hfq CLASH uncovers sRNA-target interaction networks linked to nutrient availability adaptation. eLife 2020, 9, e54655. [Google Scholar] [CrossRef]
- Pearl Mizrahi, S.; Elbaz, N.; Argaman, L.; Altuvia, Y.; Katsowich, N.; Socol, Y.; Bar, A.; Rosenshine, I.; Margalit, H. The impact of Hfq-mediated sRNA-mRNA interactome on the virulence of enteropathogenic Escherichia coli. Sci. Adv. 2021, 7, eabi8228. [Google Scholar] [CrossRef] [PubMed]
- Franze de Fernandez, M.T.; Eoyang, L.; August, J.T. Factor fraction required for the synthesis of bacteriophage Qbeta-RNA. Nature 1968, 219, 588–590. [Google Scholar] [CrossRef]
- de Fernandez, M.T.F.; Hayward, W.S.; August, J.T. Bacterial proteins required for replication of phage Qβ ribonucleic acid: Purification and properties of host factor I, a ribonucleic acid-binding protein. J. Biol. Chem. 1972, 247, 824–831. [Google Scholar] [CrossRef]
- Brennan, R.G.; Link, T.M. Hfq structure, function and ligand binding. Curr. Opin. Microbiol. 2007, 10, 125–133. [Google Scholar] [CrossRef]
- Schumacher, M.A.; Pearson, R.F.; Moller, T.; Valentin-Hansen, P.; Brennan, R.G. Structures of the pleiotropic translational regulator Hfq and an Hfq-RNA complex: A bacterial Sm-like protein. EMBO J. 2002, 21, 3546–3556. [Google Scholar] [CrossRef] [PubMed]
- Link, T.M.; Valentin-Hansen, P.; Brennan, R.G. Structure of Escherichia coli Hfq bound to polyriboadenylate RNA. Proc. Natl. Acad. Sci. USA 2009, 106, 19292–19297. [Google Scholar] [CrossRef] [PubMed]
- Panja, S.; Schu, D.J.; Woodson, S.A. Conserved arginines on the rim of Hfq catalyze base pair formation and exchange. Nucleic Acids Res. 2013, 41, 7536–7546. [Google Scholar] [CrossRef]
- Fortas, E.; Piccirilli, F.; Malabirade, A.; Militello, V.; Trepout, S.; Marco, S.; Taghbalout, A.; Arluison, V. New insight into the structure and function of Hfq C-terminus. Biosci. Rep. 2015, 35, e00190. [Google Scholar] [CrossRef] [PubMed]
- Santiago-Frangos, A.; Kavita, K.; Schu, D.J.; Gottesman, S.; Woodson, S.A. C-terminal domain of the RNA chaperone Hfq drives sRNA competition and release of target RNA. Proc. Natl. Acad. Sci. USA 2016, 113, E6089–E6096. [Google Scholar] [CrossRef]
- Santiago-Frangos, A.; Jeliazkov, J.R.; Gray, J.J.; Woodson, S.A. Acidic C-terminal domains autoregulate the RNA chaperone Hfq. eLife 2017, 6, e27049. [Google Scholar] [CrossRef]
- Moll, I.; Afonyushkin, T.; Vytvytska, O.; Kaberdin, V.R.; Blasi, U. Coincident Hfq binding and RNase E cleavage sites on mRNA and small regulatory RNAs. RNA 2003, 9, 1308–1314. [Google Scholar] [CrossRef] [PubMed]
- Masse, E.; Escorcia, F.E.; Gottesman, S. Coupled degradation of a small regulatory RNA and its mRNA targets in Escherichia coli. Genes Dev. 2003, 17, 2374–2383. [Google Scholar] [CrossRef] [PubMed]
- Sittka, A.; Lucchini, S.; Papenfort, K.; Sharma, C.M.; Rolle, K.; Binnewies, T.T.; Hinton, J.C.; Vogel, J. Deep sequencing analysis of small noncoding RNA and mRNA targets of the global post-transcriptional regulator. Hfq. PLoS Genet. 2008, 4, e1000163. [Google Scholar] [CrossRef] [PubMed]
- Bilusic, I.; Popitsch, N.; Rescheneder, P.; Schroeder, R.; Lybecker, M. Revisiting the coding potential of the E. coli genome through Hfq co-immunoprecipitation. RNA Biol. 2014, 11, 641–654. [Google Scholar] [CrossRef] [PubMed]
- Lorenz, C.; Gesell, T.; Zimmermann, B.; Schoeberl, U.; Bilusic, I.; Rajkowitsch, L.; Waldsich, C.; von Haeseler, A.; Schroeder, R. Genomic SELEX for Hfq-binding RNAs identifies genomic aptamers predominantly in antisense transcripts. Nucleic Acids Res. 2010, 38, 3794–3808. [Google Scholar] [CrossRef]
- Opdyke, J.A.; Kang, J.G.; Storz, G. GadY, a small-RNA regulator of acid response genes in Escherichia coli. J. Bacteriol. 2004, 186, 6698–6705. [Google Scholar] [CrossRef]
- Ross, J.A.; Ellis, M.J.; Hossain, S.; Haniford, D.B. Hfq restructures RNA-IN and RNA-OUT and facilitates antisense pairing in the Tn10/IS10 system. RNA 2013, 19, 670–684. [Google Scholar] [CrossRef] [PubMed]
- Brantl, S. Antisense-RNA regulation and RNA interference. Biochim. Et Biophys. Acta (BBA)-Gene Struct. Expr. 2002, 1575, 15–25. [Google Scholar] [CrossRef]
- Tomizawa, J.; Itoh, T.; Selzer, G.; Som, T. Inhibition of ColE1 RNA primer formation by a plasmid-specified small RNA. Proc. Natl. Acad. Sci. USA 1981, 78, 1421–1425. [Google Scholar] [CrossRef]
- Stougaard, P.; Molin, S.; Nordstrom, K. RNAs involved in copy-number control and incompatibility of plasmid R1. Proc. Natl. Acad. Sci. USA 1981, 78, 6008–6012. [Google Scholar] [CrossRef]
- Morita, M.; Oka, A. The structure of a transcriptional unit on colicin E1 plasmid. Eur. J. Biochem. 1979, 97, 435–443. [Google Scholar] [CrossRef] [PubMed]
- Cesareni, G.; Helmer-Citterich, M.; Castagnoli, L. Control of ColE1 plasmid replication by antisense RNA. Trends Genet. TIG 1991, 7, 230–235. [Google Scholar] [CrossRef] [PubMed]
- Itoh, T.; Tomizawa, J. Formation of an RNA primer for initiation of replication of ColE1 DNA by ribonuclease H. Proc. Natl. Acad. Sci. USA 1980, 77, 2450–2454. [Google Scholar] [CrossRef] [PubMed]
- Itoh, T.; Tomizawa, J. Initiation of replication of plasmid ColE1 DNA by RNA polymerase, ribonuclease H, and DNA polymerase I. In Cold Spring Harbor Symposia on Quantitative Biology; Cold Spring Harbor Laboratory Press: Woodbury, NY, USA, 1979; pp. 409–417. [Google Scholar]
- Masukata, H.; Tomizawa, J. Control of primer formation for ColE1 plasmid replication: Conformational change of the primer transcript. Cell 1986, 44, 125–136. [Google Scholar] [CrossRef] [PubMed]
- Masukata, H.; Tomizawa, J.-I. A mechanism of formation of a persistent hybrid between elongating RNA and template DNA. Cell 1990, 62, 331–338. [Google Scholar] [CrossRef] [PubMed]
- Masukata, H.; Tomizawa, J.-I. Effects of point mutations on formation and structure of the RNA primer for ColE1 DNA replication. Cell 1984, 36, 513–522. [Google Scholar] [CrossRef] [PubMed]
- Tomizawa, J. Control of ColE1 plasmid replication: Binding of RNA I to RNA II and inhibition of primer formation. Cell 1986, 47, 89–97. [Google Scholar] [CrossRef] [PubMed]
- Tomizawa, J. Control of ColE1 plasmid replication: The process of binding of RNA I to the primer transcript. Cell 1984, 38, 861–870. [Google Scholar] [CrossRef]
- Tomizawa, J. Control of ColE1 plasmid replication. Interaction of Rom protein with an unstable complex formed by RNA I and RNA II. J. Mol. Biol. 1990, 212, 695–708. [Google Scholar] [CrossRef]
- Lin-Chao, S.; Cohen, S.N. The rate of processing and degradation of antisense RNAI regulates the replication of ColE1-type plasmids in vivo. Cell 1991, 65, 1233–1242. [Google Scholar] [CrossRef]
- Xu, F.; Lin-Chao, S.; Cohen, S.N. The Escherichia coli pcnB gene promotes adenylylation of antisense RNAI of ColE1-type plasmids in vivo and degradation of RNAI decay intermediates. Proc. Natl. Acad. Sci. USA 1993, 90, 6756–6760. [Google Scholar] [CrossRef]
- Tomcsanyi, T.; Apirion, D. Processing enzyme ribonuclease E specifically cleaves RNA I. An inhibitor of primer formation in plasmid DNA synthesis. J. Mol. Biol. 1985, 185, 713–720. [Google Scholar] [CrossRef] [PubMed]
- McDowall, K.J.; Kaberdin, V.R.; Wu, S.W.; Cohen, S.N.; Lin-Chao, S. Site-specific RNase E cleavage of oligonucleotides and inhibition by stem-loops. Nature 1995, 374, 287–290. [Google Scholar] [CrossRef]
- Cech, G.M.; Pakula, B.; Kamrowska, D.; Wegrzyn, G.; Arluison, V.; Szalewska-Palasz, A. Hfq protein deficiency in Escherichia coli affects ColE1-like but not lambda plasmid DNA replication. Plasmid 2014, 73, 10–15. [Google Scholar] [CrossRef] [PubMed]
- Wien, F.; Kubiak, K.; Turbant, F.; Mosca, K.; Węgrzyn, G.; Arluison, V. Synchrotron Radiation Circular Dichroism, a New Tool to Probe Interactions between Nucleic Acids Involved in the Control of ColE1-Type Plasmid Replication. Appl. Sci. 2022, 12, 2639. [Google Scholar] [CrossRef]
- Ikeda, Y.; Yagi, M.; Morita, T.; Aiba, H. Hfq binding at RhlB-recognition region of RNase E is crucial for the rapid degradation of target mRNAs mediated by sRNAs in Escherichia coli. Mol. Microbiol. 2011, 79, 419–432. [Google Scholar] [CrossRef] [PubMed]
- Lalaouna, D.; Masse, E. The spectrum of activity of the small RNA DsrA: Not so narrow after all. Curr. Genet. 2016, 62, 261–264. [Google Scholar] [CrossRef] [PubMed]
- Brescia, C.C.; Mikulecky, P.J.; Feig, A.L.; Sledjeski, D.D. Identification of the Hfq-binding site on DsrA RNA: Hfq binds without altering DsrA secondary structure. RNA 2003, 9, 33–43. [Google Scholar] [CrossRef] [PubMed]
- Zhang, A.; Schu, D.J.; Tjaden, B.C.; Storz, G.; Gottesman, S. Mutations in interaction surfaces differentially impact E. coli Hfq association with small RNAs and their mRNA targets. J. Mol. Biol. 2013, 425, 3678–3697. [Google Scholar] [CrossRef]
- Lease, R.A.; Woodson, S.A. Cycling of the Sm-like protein Hfq on the DsrA small regulatory RNA. J. Mol. Biol. 2004, 344, 1211–1223. [Google Scholar] [CrossRef]
- Mikulecky, P.J.; Kaw, M.K.; Brescia, C.C.; Takach, J.C.; Sledjeski, D.D.; Feig, A.L. Escherichia coli Hfq has distinct interaction surfaces for DsrA, rpoS and poly(A) RNAs. Nat. Struct. Mol. Biol. 2004, 11, 1206–1214. [Google Scholar] [CrossRef] [PubMed]
- Olejniczak, M. Despite similar binding to the Hfq protein regulatory RNAs widely differ in their competition performance. Biochemistry 2011, 50, 4427–4440. [Google Scholar] [CrossRef] [PubMed]
- Lin-Chao, S.; Bremer, H. Activities of the RNAI and RNAII promoters of plasmid pBR322. J. Bacteriol. 1987, 169, 1217–1222. [Google Scholar] [CrossRef] [PubMed]
- Ali Azam, T.; Iwata, A.; Nishimura, A.; Ueda, S.; Ishihama, A. Growth phase-dependent variation in protein composition of the Escherichia coli nucleoid. J. Bacteriol. 1999, 181, 6361–6370. [Google Scholar] [CrossRef] [PubMed]
- Kaberdin, V.R.; Chao, Y.H.; Lin-Chao, S. RNase E cleaves at multiple sites in bubble regions of RNA I stem loops yielding products that dissociate differentially from the enzyme. J. Biol. Chem. 1996, 271, 13103–13109. [Google Scholar] [CrossRef] [PubMed]
- Richards, J.; Belasco, J.G. Obstacles to Scanning by RNase E Govern Bacterial mRNA Lifetimes by Hindering Access to Distal Cleavage Sites. Mol. Cell 2019, 74, 284–295.e285. [Google Scholar] [CrossRef] [PubMed]
- He, L.; Soderbom, F.; Wagner, E.G.; Binnie, U.; Binns, N.; Masters, M. PcnB is required for the rapid degradation of RNAI, the antisense RNA that controls the copy number of ColE1-related plasmids. Mol. Microbiol. 1993, 9, 1131–1142. [Google Scholar] [CrossRef] [PubMed]
- Mohanty, B.K.; Maples, V.F.; Kushner, S.R. The Sm-like protein Hfq regulates polyadenylation dependent mRNA decay in Escherichia coli. Mol. Microbiol. 2004, 54, 905–920. [Google Scholar] [CrossRef] [PubMed]
- Hajnsdorf, E.; Regnier, P. Host factor Hfq of Escherichia coli stimulates elongation of poly(A) tails by poly(A) polymerase I. Proc. Natl. Acad. Sci. USA 2000, 97, 1501–1505. [Google Scholar] [CrossRef]
- Hussein, R.; Lim, H.N. Disruption of small RNA signaling caused by competition for Hfq. Proc. Natl. Acad. Sci. USA 2011, 108, 1110–1115. [Google Scholar] [CrossRef]
- Moon, K.; Gottesman, S. Competition among Hfq-binding small RNAs in Escherichia coli. Mol. Microbiol. 2011, 82, 1545–1562. [Google Scholar] [CrossRef]
- Małecka, E.M.; Strozecka, J.; Sobanska, D.; Olejniczak, M. Structure of bacterial regulatory RNAs determines their performance in competition for the chaperone protein Hfq. Biochemistry 2015, 54, 1157–1170. [Google Scholar] [CrossRef] [PubMed]
- Kwiatkowska, J.; Wroblewska, Z.; Johnson, K.A.; Olejniczak, M. The binding of Class II sRNA MgrR to two different sites on matchmaker protein Hfq enables efficient competition for Hfq and annealing to regulated mRNAs. RNA 2018, 24, 1761–1784. [Google Scholar] [CrossRef] [PubMed]
- Wagner, E.G. Cycling of RNAs on Hfq. RNA Biol. 2013, 10, 619–626. [Google Scholar] [CrossRef] [PubMed]
- Guyer, M.S.; Reed, R.R.; Steitz, J.A.; Low, K.B. Identification of a sex-factor-affinity site in E. coli as gamma delta. In Cold Spring Harbor Symposia on Quantitative Biology; Cold Spring Harbor Laboratory Press: Woodbury, NY, USA, 1981; Volume 45, pp. 135–140. [Google Scholar] [CrossRef]
- Lopez, P.J.; Marchand, I.; Joyce, S.A.; Dreyfus, M. The C-terminal half of RNase E, which organizes the Escherichia coli degradosome, participates in mRNA degradation but not rRNA processing in vivo. Mol. Microbiol. 1999, 33, 188–199. [Google Scholar] [CrossRef]
- Baba, T.; Ara, T.; Hasegawa, M.; Takai, Y.; Okumura, Y.; Baba, M.; Datsenko, K.A.; Tomita, M.; Wanner, B.L.; Mori, H. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: The Keio collection. Mol. Syst. Biol. 2006, 2, 2006-0008. [Google Scholar] [CrossRef]
- Thomason, L.C.; Costantino, N.; Court, D.L. E. coli genome manipulation by P1 transduction. Curr. Protoc. Mol. Biol. 2007, 79, 1–17. [Google Scholar] [CrossRef]
- Liu, H.; Naismith, J.H. An efficient one-step site-directed deletion, insertion, single and multiple-site plasmid mutagenesis protocol. BMC Biotechnol. 2008, 8, 91. [Google Scholar] [CrossRef]
- Singh, D.; Chang, S.J.; Lin, P.H.; Averina, O.V.; Kaberdin, V.R.; Lin-Chao, S. Regulation of ribonuclease E activity by the L4 ribosomal protein of Escherichia coli. Proc. Natl. Acad. Sci. USA 2009, 106, 864–869. [Google Scholar] [CrossRef]
- Hellman, L.M.; Fried, M.G. Electrophoretic mobility shift assay (EMSA) for detecting protein–nucleic acid interactions. Nat. Protoc. 2007, 2, 1849–1861. [Google Scholar] [CrossRef]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef] [PubMed]
- Bustin, S.A.; Benes, V.; Garson, J.A.; Hellemans, J.; Huggett, J.; Kubista, M.; Mueller, R.; Nolan, T.; Pfaffl, M.W.; Shipley, G.L. The MIQE Guidelines: M inimum I nformation for Publication of Q uantitative Real-Time PCR Experiments. Clin. Chem. 2009, 55, 4611–4622. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]


| Wildtype | Proximal Face Mutants | Distal Face Mutant | ||||
|---|---|---|---|---|---|---|
| HfqWT | Q8A | K56A | H57A | K56/H57A | Y25D* | |
| DsrA | 100% | 46.9 ± 4.0% b | 24.4 ± 4.3% a | 66.5 ± 17.0% d | 13.2 ± 6.9% b | 162.7 ± 13.7% c |
| RNA I | 100% | 86.5 ± 3.6% b | 36.1 ± 5.1% b | 72.7 ± 5.0% c | 13.8 ± 8.6% b | 125.2 ± 22.4% d |
| Primer Name | Primer Sequence | Purpose |
|---|---|---|
| Primer 1 | 5′-CCG CTC GAG AAT GGC TAA GGG GCA AT-3′ | pBAD-His-Hfq construction |
| Primer 2 | 5′-CCG GAA TTC TTA TTC GGT TTC TTC GCT GTC CTG TTG-3′ | pBAD-His-Hfq construction |
| Primer 3 | 5′-GGA ATT CCA TAT GGC TAA GGG GCA ATC TTT ACA-3′ | pBAD-Hfq construction |
| Primer 4 | 5′-GGG CAA TCT TTA GCA GAT CCG TTC CTG AAC GCA CTG CG-3′ | Hfq Q8A mutagenesis |
| Primer 5 | 5′-GTT CAG GAA CGG ATC TGC TAA AGA TTG CCC CTT AG-3′ | Hfq Q8A mutagenesis |
| Primer 6 | 5′-GTT TCT ATT GAT TTG GTG AAT GGT ATT AAG CTG CAA GGG C-3′ | Hfq Y25D* mutagenesis |
| Primer 7 | 5′-CCA TTC ACC AAA TCA ATA GAA ACT GGA ACA CG-3′ | Hfq Y25D* mutagenesis |
| Primer 8 | 5′-GGT TTA CGC GCA CGC GAT TTC TAC TGT TGT CC-3′ | Hfq K56A mutagenesis |
| Primer 9 | 5′-GAA ATC GCG TGC GCG TAA ACC ATC TGG CTG AC-3′ | Hfq K56A mutagenesis |
| Primer 10 | 5′-CAG CCA GAT GGT TTA CAA GGC CGC GAT TTC TAC TGT TGT CC-3′ | Hfq H57A mutagenesis |
| Primer 11 | 5′-GGA CAA CAG TAG AAA TCG CGG CCT TGT AAA CCA TCT GGC TG-3′ | Hfq H57A mutagenesis |
| Primer 12 | 5′-GGT TTA CGC GGC CGC GAT TTC TAC TGT TGT CC-3′ | Hfq K56A/H57A mutagenesis |
| Primer 13 | 5′-GAA ATC GCG GCC GCG TAA ACC ATC TGG CTG AC-3′ | Hfq K56A/H57A mutagenesis |
| Primer 14 | 5′-TAA TAC GAC TCA CTA TAG GGA CAG TAT TTG GTA TCT GCG C-3′ | RNA IFL T7 template |
| Primer 15 | 5′-ACA AAA AAA CCA CCG CTA CC-3′ | RNA IFL T7 template |
| Primer 16 | 5′-TAA TAC GAC TCA CTA TAG GGT GGT ATC TGC GCT CTG C-3′ | RNA I-8 T7 template |
| Primer 17 | 5′-TAA TAC GAC TCA CTA TAG GGA ACA CAT CAG ATT TCC TGG-3′ | DsrA T7 template |
| Primer 18 | 5′-AAA TCC CGA CCC TGA GG-3′ | DsrA T7 template |
| Primer 19 | 5′-GTA ACT GGC TTC AGC AGA GCG CAG ATA CC-3′ | RNA I oligo probe |
| Primer 20 | CGG TTC TAT GGT GGT TGA | qPCR primer for mreB |
| Primer 21 | GCG CAC AGA AGA GGA GTA | qPCR primer for mreB |
| Primer 22 | TTC CGG CTG GCT GGT TTA TT | qPCR primer for ampA |
| Primer 23 | TGA CTC CCC GTC GTG TAG AT | qPCR primer for ampA |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, W.-S.; Lin-Chao, S. Hfq-Antisense RNA I Binding Regulates RNase E-Dependent RNA Stability and ColE1 Plasmid Copy Number. Int. J. Mol. Sci. 2024, 25, 3955. https://doi.org/10.3390/ijms25073955
Wang W-S, Lin-Chao S. Hfq-Antisense RNA I Binding Regulates RNase E-Dependent RNA Stability and ColE1 Plasmid Copy Number. International Journal of Molecular Sciences. 2024; 25(7):3955. https://doi.org/10.3390/ijms25073955
Chicago/Turabian StyleWang, Wei-Syuan, and Sue Lin-Chao. 2024. "Hfq-Antisense RNA I Binding Regulates RNase E-Dependent RNA Stability and ColE1 Plasmid Copy Number" International Journal of Molecular Sciences 25, no. 7: 3955. https://doi.org/10.3390/ijms25073955






