Natural Allelic Variations of Bch10G006400 Controlling Seed Size in Chieh-qua (Benincasa hispida Cogn. var. Chieh-qua How)
Abstract
:1. Introduction
2. Results
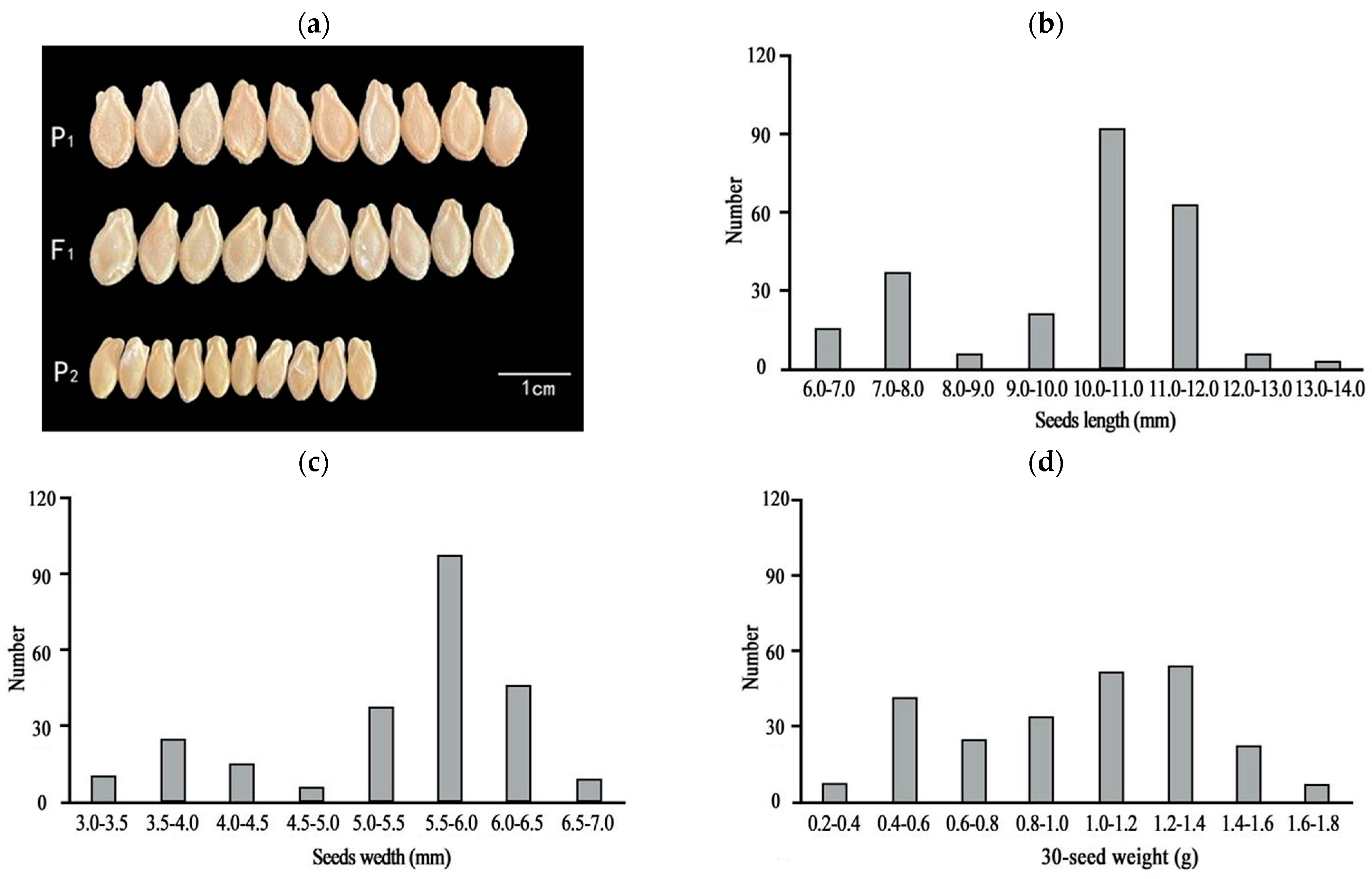
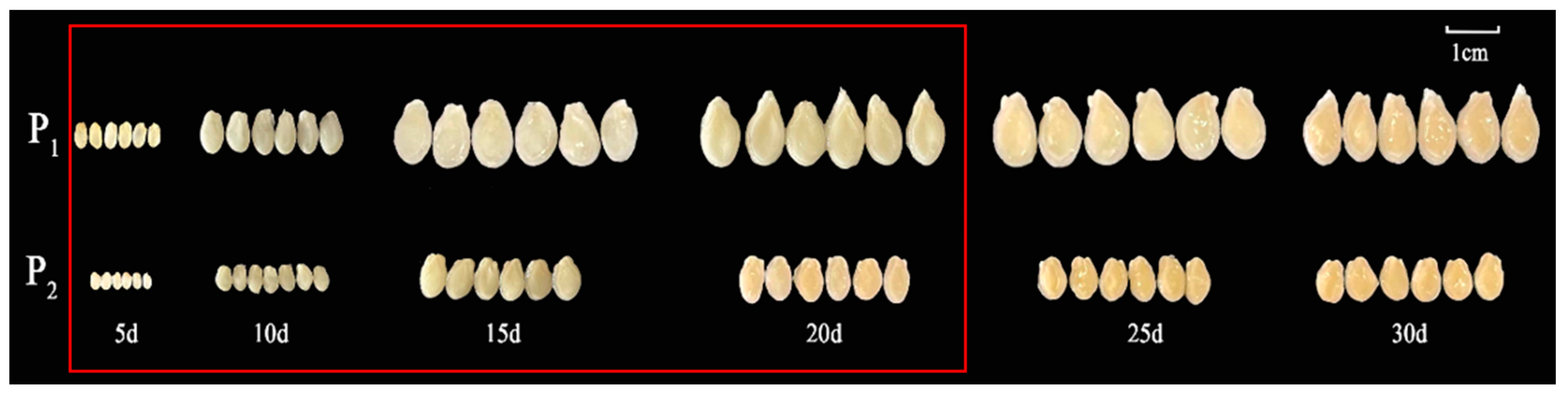
2.1. Inheritance and Phenotypic Characterization of Seed Size in Chieh-qua
2.2. Preliminary Mapping of the Seed Size Gene
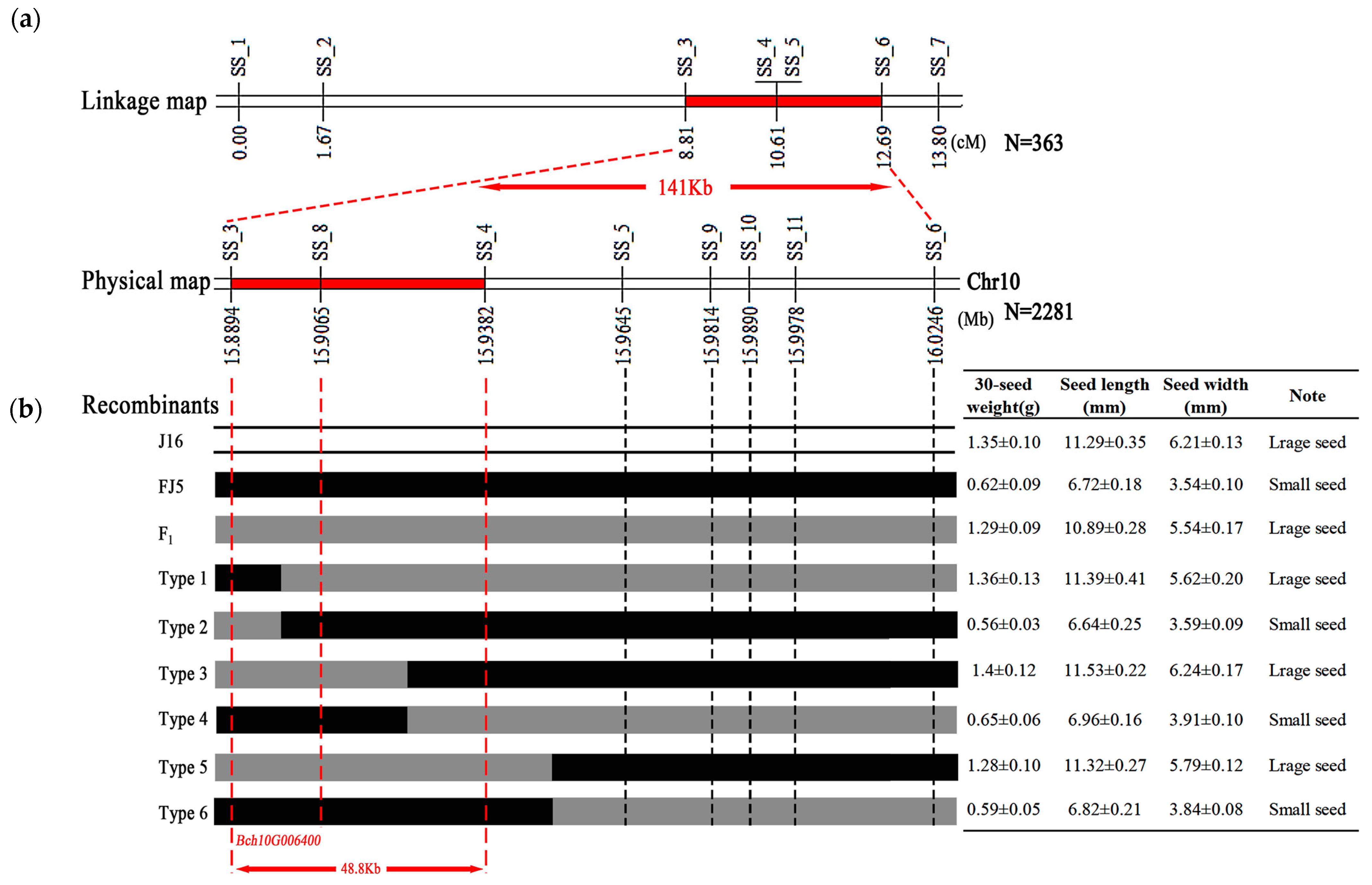
2.3. Fine Mapping of the Seed Size Gene
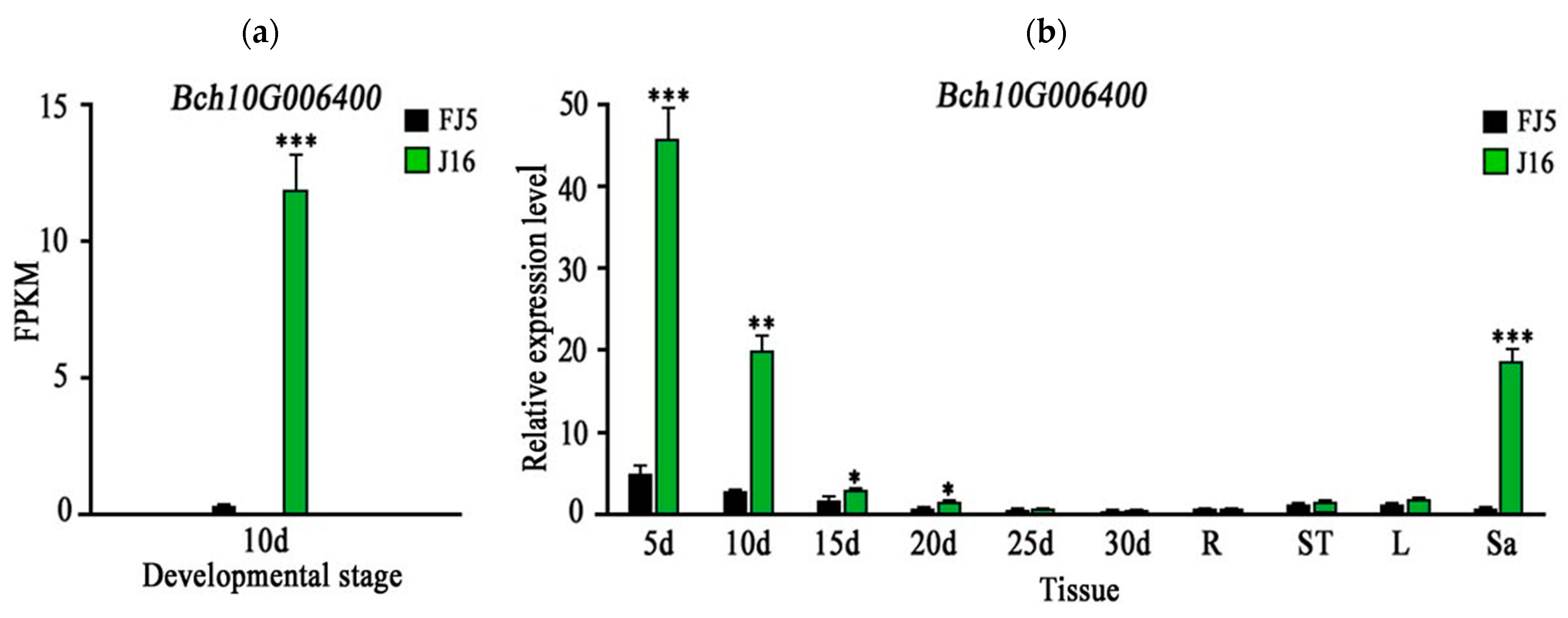
2.4. RNA-Seq and qRT-PCR Analysis of Candidate Genes
2.5. Sequence Characterization of the Candidate Genes
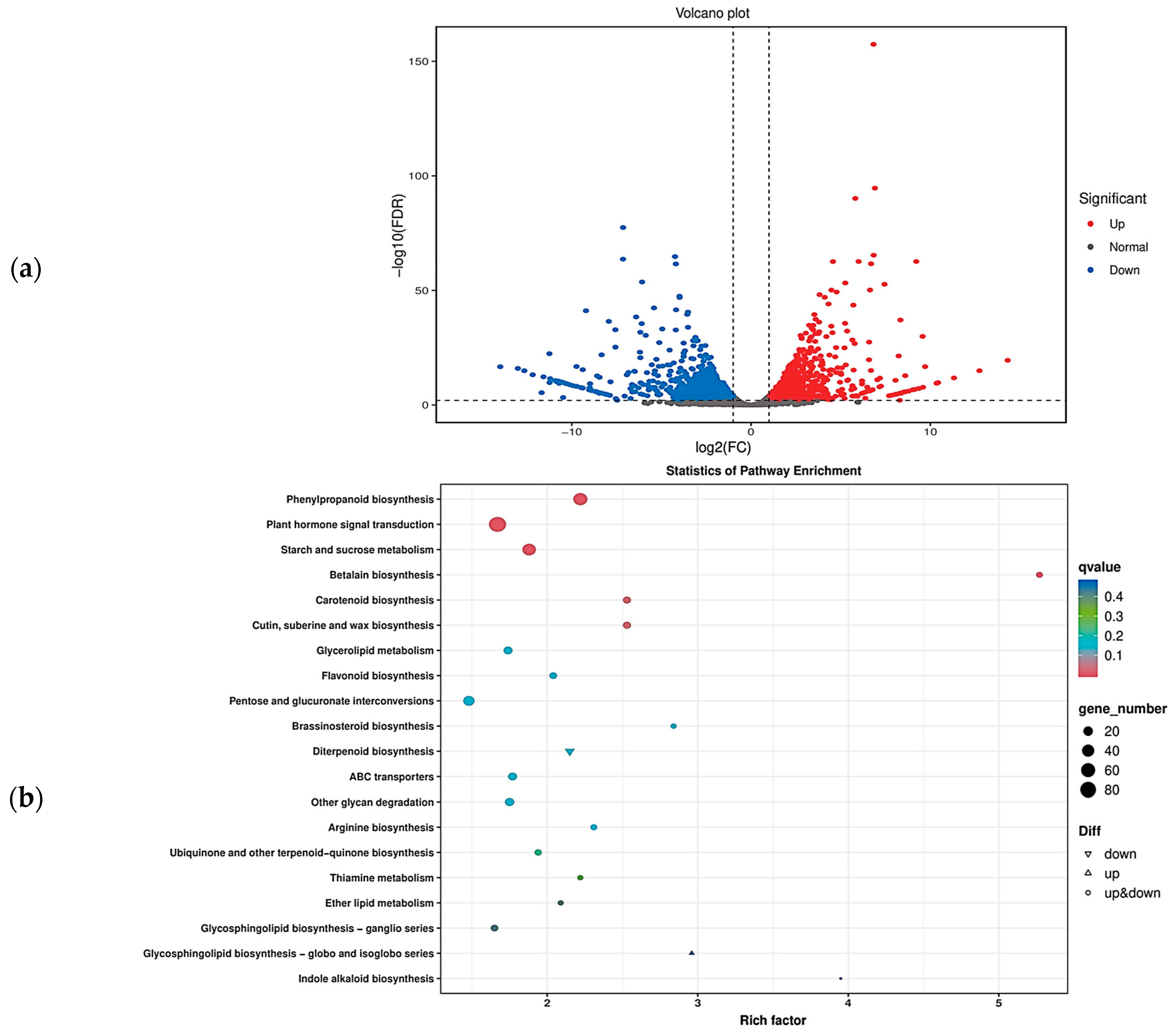
2.6. Transcriptome Analysis of the Seed Size of Chieh-qua
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Phenotyping
4.2. BSA-Seq Analysis
4.3. Linkage Mapping Analysis
4.4. Fine Mapping Analysis
4.5. RNA-Seq Analysis
4.6. qRT-PCR Analysis
4.7. Candidate Gene Cloning and Sequence Comparison
4.8. Development of the dCAPS Maker
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lin, J.Y.; Xie, W.P.; Tan, X.; Qiao, Y.C.; Xie, L.F. A New Early-maturing Chieh-qua Cultivar ‘Guanhua 5’. Acta Hortic. Sin. 2013, 40, 1857–1858. [Google Scholar]
- Zhang, J.; Zhang, X.; Liu, X.; Pai, Q.; Wang, Y.; Wu, X. Molecular Network for Regulation of Seed Size in Plants. Int. J. Mol. Sci. 2023, 24, 10666. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zheng, L.; Corke, F.; Smith, C.; Bevan, M.W. Control of final seed and organ size by the DA1 gene family in Arabidopsis thaliana. Genes Dev. 2008, 22, 1331–1336. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Xu, R.; Li, Y. Molecular Networks of Seed Size Control in Plants. Annu. Rev. Plant Biol. 2019, 70, 435–463. [Google Scholar] [CrossRef] [PubMed]
- Shi, C.; Ren, Y.; Liu, L.; Wang, F.; Zhang, H.; Tian, P.; Pan, T.; Wang, Y.; Jing, R.; Liu, T.; et al. Ubiquitin Specific Protease 15 Has an Important Role in Regulating Grain Width and Size in Rice. Plant Physiol. 2019, 180, 381–391. [Google Scholar] [CrossRef] [PubMed]
- Haughn, G.; Chaudhury, A. Genetic analysis of seed coat development in Arabidopsis. Trends Plant Sci. 2005, 10, 472–477. [Google Scholar] [CrossRef]
- Cao, J.; Li, G.; Qu, D.; Li, X.; Wang, Y. Into the Seed: Auxin Controls Seed Development and Grain Yield. Int. J. Mol. Sci. 2020, 21, 1662. [Google Scholar] [CrossRef]
- Li, N.; Li, Y. Signaling pathways of seed size control in plants. Curr. Opin. Plant Biol. 2016, 33, 23–32. [Google Scholar] [CrossRef]
- Gasparis, S.; Miłoszewski, M.M. Genetic Basis of Grain Size and Weight in Rice, Wheat, and Barley. Int. J. Mol. Sci. 2023, 24, 16921. [Google Scholar] [CrossRef]
- Li, N.; Shang, J.; Wang, J.; Zhou, D.; Li, N.; Ma, S. Fine mapping and discovery of candidate genes for seed size in watermelon by genome survey sequencing. Sci. Rep. 2018, 8, 17843. [Google Scholar] [CrossRef]
- Gong, C.; Zhao, S.; Yang, D.; Lu, X.; Anees, M.; He, N.; Zhu, H.; Zhao, Y.; Liu, W. Genome-wide association analysis provides molecular insights into the natural variation of watermelon seed size. Hortic. Res. 2022, 19, uhab074. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; He, J.; Zhao, Y.; Wu, T.; Zhou, X.; Ding, Y.; Kong, L.; Wang, X.; Wang, Y.; Li, J.; et al. EAR1 Negatively Regulates ABA Signaling by Enhancing 2C Protein Phosphatase Activity. Plant Cell. 2018, 30, 815–834. [Google Scholar] [CrossRef]
- Yang, Y.; Qin, Y.; Xie, C.; Zhao, F.; Zhao, J.; Liu, D.; Chen, S.; Fuglsang, A.T.; Palmgren, M.G.; Schumaker, K.S.; et al. The Arabidopsis chaperone J3 regulates the plasma membrane H+-ATPase through interaction with the PKS5 kinase. Plant Cell. 2010, 22, 1313–1332. [Google Scholar] [CrossRef]
- Zhou, X.; Hao, H.; Zhang, Y.; Bai, Y.; Zhu, W.; Qin, Y.; Yuan, F.; Zhao, F.; Wang, M.; Hu, J.; et al. SOS2-LIKE PROTEIN KINASE5, an SNF1-RELATED PROTEIN KINASE3-Type Protein Kinase, Is Important for Abscisic Acid Responses in Arabidopsis through Phosphorylation of ABSCISIC ACID-INSENSITIVE5. Plant Physiol. 2015, 168, 659–676. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wang, J.; Guo, S.; Tian, S.; Zhang, J.; Ren, Y.; Li, M.; Gong, G.; Zhang, H.; Xu, Y. CRISPR/Cas9-mediated mutagenesis of ClBG1 decreased seed size and promoted seed germination in watermelon. Hortic. Res. 2021, 8, 70. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wang, C.; Han, H.; Luo, Y.; Wang, Z.; Yan, C.; Xu, W.; Qu, S. Construction of a High-Density Genetic Map and Analysis of Seed-Related Traits Using Specific Length Amplified Fragment Sequencing for Cucurbita maxima. Front. Plant Sci. 2020, 10, 1782. [Google Scholar] [CrossRef] [PubMed]
- Mi, C.X. Phenotypes Study and GWAS Analysis Based on Spectral and Imaging Information of Pumpkin Seeds; Zhejiang University: Hangzhou, China, 2021. [Google Scholar]
- Li, J.F. Analysis of Genetic Characteristics of Melon Seed Traits; Heilongjiang Bayi Agricultural University: Daqing, China, 2022. [Google Scholar]
- Chen, L.L.; Qin, Z.W.; Zhou, X.Y.; Xin, M.; Wu, T. Molecular marker and genetic analysis of cucumber seed length. Chin. Sci. Bull. 2012, 28, 165–170. [Google Scholar]
- Min, W.; Han, M.; Sheng-Ping, Z.; Shu-Lin, L.; Shao-Yun, D.; Ye, W.; Xing-Fang, G. Inheritance analysis and QTL mapping of cucumber seed size. Acta Hortic. Sin. 2014, 41, 63–72. [Google Scholar]
- Guo, Y.; Gao, M.; Liang, X.; Xu, M.; Liu, X.; Zhang, Y.; Liu, X.; Liu, J.; Gao, Y.; Qu, S.; et al. Quantitative Trait Loci for Seed Size Variation in Cucurbits—A Review. Front. Plant Sci. 2020, 11, 304. [Google Scholar] [CrossRef]
- Yang, W.; Wang, P.; Liu, T.; Nong, L.; Cheng, Z.; Su, L.; Bai, W.; Deng, Y.; Chen, Z.; Liu, Z. Fine mapping of the major gene BhHLS1 controlling seed size in wax gourd (Benincasa hispida). Front. Plant Sci. 2023, 14, 1266796. [Google Scholar] [CrossRef]
- Abe, A.; Kosugi, S.; Yoshida, K.; Natsume, S.; Takagi, H.; Kanzaki, H.; Matsumura, H.; Yoshida, K.; Mitsuoka, C.; Tamiru, M.; et al. Genome sequencing reveals agronomically important loci in rice using MutMap. Nat. Biotechnol. 2012, 30, 174–178. [Google Scholar] [CrossRef] [PubMed]
- Takagi, H.; Abe, A.; Yoshida, K.; Kosugi, S.; Natsume, S.; Mitsuoka, C.; Uemura, A.; Utsushi, H.; Tamiru, M.; Takuno, S.; et al. QTL-seq: Rapid mapping of quantitative trait loci in rice by whole genome resequencing of DNA from two bulked populations. Plant J. Cell Mol. Biol. 2013, 74, 174–183. [Google Scholar] [CrossRef]
- Zhang, L.; Li, D.Q. MORC2 regulates DNA damage response through a PARP1-dependent pathway. Nucleic Acids Res. 2019, 47, 8502–8520. [Google Scholar] [CrossRef]
- Xu, X.; Wei, C.; Liu, Q.; Qu, W.; Qi, X.; Xu, Q.; Chen, X. The major-effect quantitative trait locus Fnl7.1 encodes a late embryogenesis abundant protein associated with fruit neck length in cucumber. Plant Biotechnol. J. 2020, 18, 1598–1609. [Google Scholar] [CrossRef] [PubMed]
- Gebremeskel, H.; Umer, M.J.; Hongju, Z.; Li, B.; Shengjie, Z.; Yuan, P.; Xuqiang, L.; Nan, H.; Wenge, L. Genetic mapping and molecular characterization of the delayed green gene dg in watermelon (Citrullus lanatus). Front. Plant Sci. 2023, 14, 1152644. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Liu, Z.; Cheng, Z.; Gou, J.; Chen, J.; Yu, W.; Wang, P. Identification and Application of BhAPRR2 Controlling Peel Colour in Wax Gourd (Benincasa hispida). Front. Plant Sci. 2021, 12, 716772. [Google Scholar] [CrossRef] [PubMed]
- Jin, G.; Xu, M.; Zou, M.; Duan, S. The Processing, Gene Regulation, Biological Functions, and Clinical Relevance of N4-Acetylcytidine on RNA: A Systematic Review. Mol. Ther. Nucleic Acids 2020, 20, 13–24. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Wang, X.; Pan, Q.; Li, P.; Liu, Y.; Lu, X.; Zhong, W.; Li, M.; Han, L.; Li, J.; et al. QTG-Seq Accelerates QTL Fine Mapping through QTL Partitioning and Whole-Genome Sequencing of Bulked Segregant Samples. Mol. Plant 2019, 12, 426–437. [Google Scholar] [CrossRef]
- Oh, T.I.; Lee, Y.M.; Lim, B.O.; Lim, J.H. Inhibition of NAT10 Suppresses Melanogenesis and Melanoma Growth by Attenuating Microphthalmia-Associated Transcription Factor (MITF) Expression. Int. J. Mol. Sci. 2017, 18, 1924. [Google Scholar] [CrossRef]
- Roman, G.; Ecker, J.R. Genetic analysis of a seedling stress response to ethylene in Arabidopsis. Philos. Trans. R Soc. Lond B Biol. Sci. 1995, 350, 75–81. [Google Scholar] [PubMed]
- Liu, N.N. Molecular Mechanism of HOOKLESS1 Regulating Flowering Time of Arabidopsis thaliana; Nanjing Normal University: Nanjing, China, 2022. [Google Scholar]
- Liao, C.J.; Lai, Z.; Lee, S.; Yun, D.J.; Mengiste, T. Arabidopsis HOOKLESS1 Regulates Responses to Pathogens and Abscisic Acid through Interaction with MED18 and Acetylation of WRKY33 and ABI5 Chromatin. Plant Cell 2016, 28, 1662–1681. [Google Scholar] [CrossRef]
- Jin, H.; Zhu, Z. HOOKLESS1 is a positive regulator in Arabidopsis thermomorphogenesis. Sci. China Life Sci. 2019, 62, 423–425. [Google Scholar] [CrossRef] [PubMed]
- Jin, H.; Lin, J.; Zhu, Z. PIF4 and HOOKLESS1 Impinge on Common Transcriptome and Isoform Regulation in Thermomorphogenesis. Plant Commun. 2020, 1, 100034. [Google Scholar] [CrossRef] [PubMed]
- Chang, K.N.; Zhong, S.; Weirauch, M.T.; Hon, G.; Pelizzola, M.; Li, H.; Huang, S.S.; Schmitz, R.J.; Urich, M.A.; Kuo, D.; et al. Temporal transcriptional response to ethylene gas drives growth hormone cross-regulation in Arabidopsis. eLife 2020, 2, e00675. [Google Scholar] [CrossRef]
- Neubauer, M.; Innes, R.W. Loss of the Acetyltransferase NAA50 Induces Endoplasmic Reticulum Stress and Immune Responses and Suppresses Growth. Plant Physiol. 2020, 183, 1838–1854. [Google Scholar] [CrossRef] [PubMed]
- Armbruster, L.; Linster, E.; Boyer, J.B.; Brünje, A.; Eirich, J.; Stephan, I.; Bienvenut, W.V.; Weidenhausen, J.; Meinnel, T.; Hell, R.; et al. NAA50 Is an Enzymatically Active Nα-Acetyltransferase That Is Crucial for Development and Regulation of Stress Responses. Plant Physiol. 2020, 183, 1502–1516. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Sun, J.; Wang, R.; Zhang, Z.; Liu, N.; Jin, H.; Zhong, B.; Zhu, Z. The origin, evolution and functional divergence of HOOKLESS1 in plants. Commun. Biol. 2023, 6, 460. [Google Scholar] [CrossRef] [PubMed]
- Porebski, S.; Bailey, L.G.; Baum, B.R. Modification of a ctab dna extraction protocol for plants containing high polysaccharide and polyphenol components. Plant Mol. Biol. Rep. 1997, 15, 8–15. [Google Scholar]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef]
- Hill, J.T.; Demarest, B.L.; Bisgrove, B.W.; Gorsi, B.; Su, Y.C.; Yost, H.J. MMAPPR: Mutation mapping analysis pipeline for pooled RNA-seq. Genome Res. 2013, 23, 687–697. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.C.; Mendell, J.T.; Salzberg, S.L. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef] [PubMed]
- Florea, L.; Song, L.; Salzberg, S.L. Thousands of exon skipping events differentiate among splicing patterns in sixteen human tissues. F1000Research 2013, 2, 188. [Google Scholar] [CrossRef] [PubMed]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene ontology: Tool for the unification of biology. Gene Ontol. Consort. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef] [PubMed]
- Eddy, S.R. Profile hidden Markov models. Bioinformatics 1998, 14, 755–763. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Bateman, A.; Clements, J.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Heger, A.; Hetherington, K.; Holm, L.; Mistry, J.; et al. Pfam: The protein families database. Nucleic Acids Res. 2014, 42, D222–D230. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Williams, B.A.; Pertea, G.; Mortazavi, A.; Kwan, G.; van Baren, M.J.; Salzberg, S.L.; Wold, B.J.; Pachter, L. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 2010, 28, 511–515. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]


| Generation | Seed Length (mm) | Seed Width (mm) | 30-Seed Weight (g) |
|---|---|---|---|
| P1 | 11.44 ± 0.27 a | 5.97 ± 0.14 a | 1.34 ± 0.10 a |
| P2 | 6.50 ± 0.24 b | 3.43 ± 0.15 b | 0.62 ± 0.07 b |
| F1 | 11.22 ± 0.81 a | 5.83 ± 0.16 a | 1.32 ± 0.14 a |
| Sample ID | SNP_314 | SNP_400 | InDel_551 | Length (mm) | Width (mm) |
|---|---|---|---|---|---|
| D2-2-2 | G | G | AGA | 11.49 ± 0.36 | 6.46 ± 0.56 |
| D4-1-2 | G | G | AGA | 12.11 ± 0.57 | 6.13 ± 0.39 |
| 0401 | G | G | AGA | 11.2 ± 0.28 | 5.91 ± 0.48 |
| Tai 13 | G | G | AGA | 11.26 ± 0.58 | 6.37 ± 0.40 |
| 17075 | C | G | AGA | 6.81 ± 0.44 | 3.72 ± 0.29 |
| 17063 | C | G | AGA | 4.98 ± 0.49 | 2.91 ± 0.21 |
| 17011 | C | G | AGA | 6.60 ± 0.84 | 3.55 ± 0.18 |
| 19FJ1-1 | G | T | AGA | 6.31 ± 0.79 | 3.44 ± 0.46 |
| 17097 | G | T | AGA | 5.86 ± 0.29 | 3.11 ± 0.28 |
| 19FJ28-2 | G | G | - | 5.69 ± 0.46 | 3.19 ± 0.37 |
| 17045 | G | G | - | 5.66 ± 0.41 | 3.43 ± 0.23 |
| 1714 | G | G | - | 5.68 ± 0.54 | 3.20 ± 0.37 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gao, Y.; Peng, J.; Qiao, Y.; Wang, G. Natural Allelic Variations of Bch10G006400 Controlling Seed Size in Chieh-qua (Benincasa hispida Cogn. var. Chieh-qua How). Int. J. Mol. Sci. 2024, 25, 4236. https://doi.org/10.3390/ijms25084236
Gao Y, Peng J, Qiao Y, Wang G. Natural Allelic Variations of Bch10G006400 Controlling Seed Size in Chieh-qua (Benincasa hispida Cogn. var. Chieh-qua How). International Journal of Molecular Sciences. 2024; 25(8):4236. https://doi.org/10.3390/ijms25084236
Chicago/Turabian StyleGao, Yin, Jiazhu Peng, Yanchun Qiao, and Guoping Wang. 2024. "Natural Allelic Variations of Bch10G006400 Controlling Seed Size in Chieh-qua (Benincasa hispida Cogn. var. Chieh-qua How)" International Journal of Molecular Sciences 25, no. 8: 4236. https://doi.org/10.3390/ijms25084236





