Characterizing Common Factors Affecting Replication Initiation During H2O2 Exposure and Genetic Mutation-Induced Oxidative Stress in Escherichia coli
Abstract
:1. Introduction
2. Results
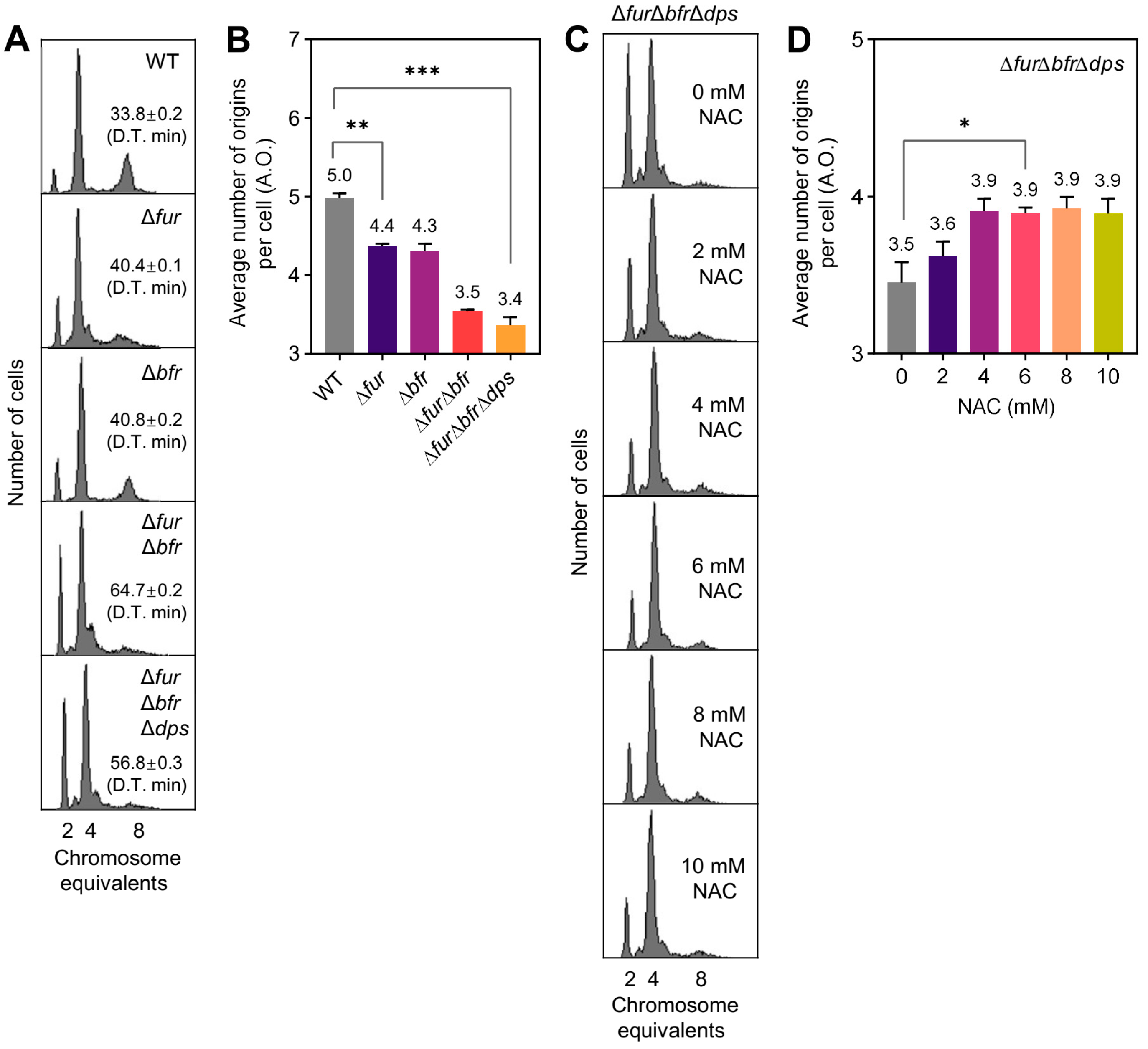
2.1. Defective Iron Regulation Mediates Delayed Replication Initiation and Can Be Reversed by the ROS Scavenger NAC
2.2. ATP Levels Associated with Delayed Replication Initiation Due to Oxidative Stress
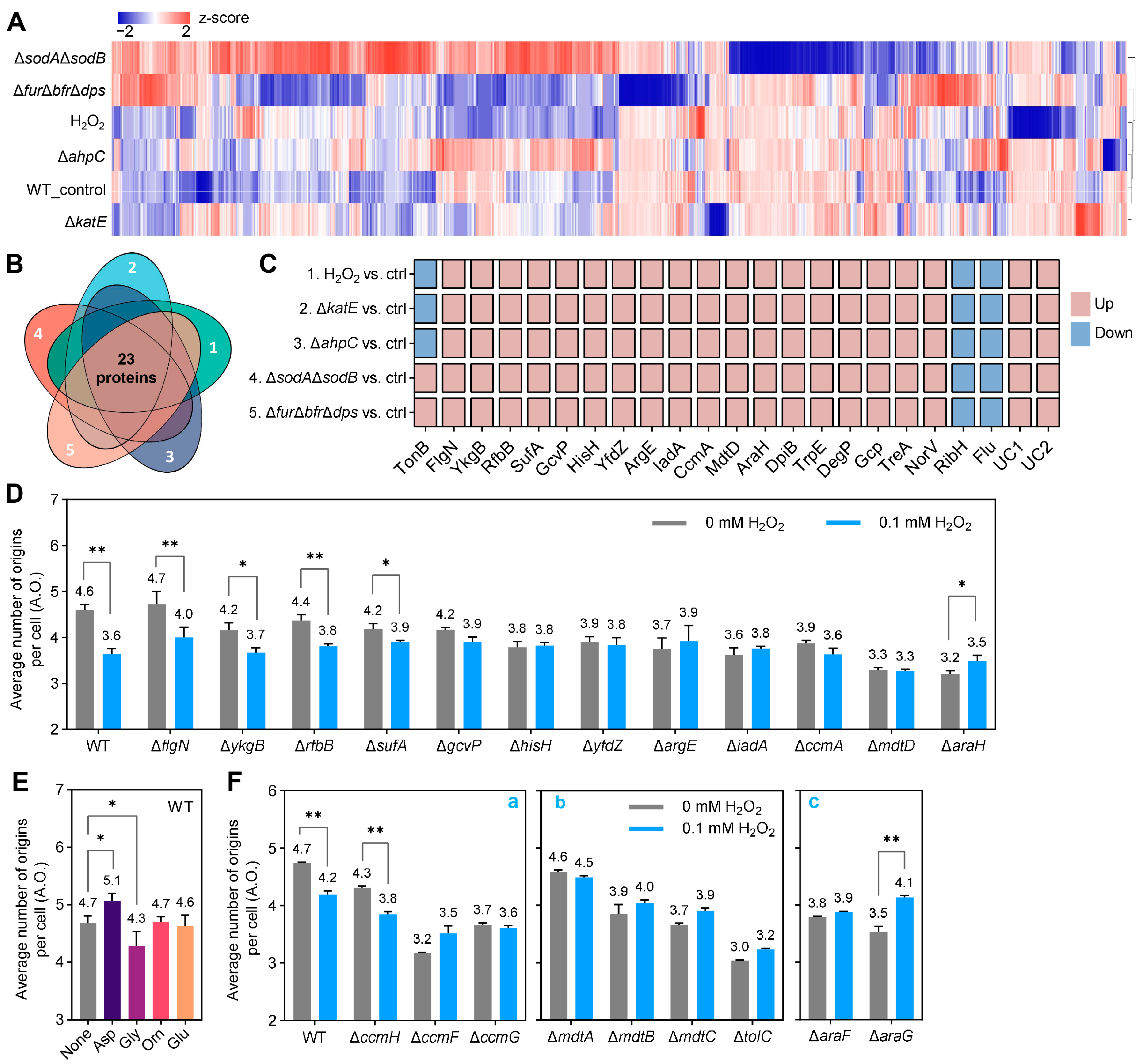
2.3. Proteomic Screening and Genetic Validation of Common Factors That Inhibit Replication Initiation Under Different Causes-Induced Oxidative Stresses
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains and Culture
4.2. Doubling Time Measurement
4.3. DNA Replication Patterns Determination
4.4. Proteomic Sample Preparation
4.5. Proteomic Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ROS | reactive oxygen species |
| VBNC | viable but not culturable |
| NAC | N-Acetylcysteine |
| CAA | Casamino acid |
| A.O. | average number of replication origins per cell |
| WT | wild-type |
References
- Brilli, M.; Fondi, M.; Fani, R.; Mengoni, A.; Ferri, L.; Bazzicalupo, M.; Biondi, E.G. The diversity and evolution of cell cycle regulation in alpha-proteobacteria: A comparative genomic analysis. BMC Syst. Biol. 2010, 4, 52. [Google Scholar] [CrossRef] [PubMed]
- Jonas, K. To divide or not to divide: Control of the bacterial cell cycle by environmental cues. Curr. Opin. Microbiol. 2014, 18, 54–60. [Google Scholar] [CrossRef] [PubMed]
- Wilmaerts, D.; Windels, E.M.; Verstraeten, N.; Michiels, J. General Mechanisms Leading to Persister Formation and Awakening. Trends Genet. 2019, 35, 401–411. [Google Scholar] [CrossRef]
- Qiao, J.; Du, D.; Wang, Y.; Xi, L.; Zhu, W.; Morigen. Uncovering the effects of non-lethal oxidative stress on replication initiation in Escherichia coli. Gene 2025, 933, 148992. [Google Scholar] [CrossRef] [PubMed]
- Helaine, S.; Cheverton, A.M.; Watson, K.G.; Faure, L.M.; Matthews, S.A.; Holden, D.W. Internalization of Salmonella by macrophages induces formation of nonreplicating persisters. Science 2014, 343, 204–208. [Google Scholar] [CrossRef]
- Liu, J.; Yang, L.; Kjellerup, B.V.; Xu, Z. Viable but nonculturable (VBNC) state, an underestimated and controversial microbial survival strategy. Trends Microbiol. 2023, 31, 1013–1023. [Google Scholar] [CrossRef]
- Lewis, K. Persister cells, dormancy and infectious disease. Nat. Rev. Microbiol. 2007, 5, 48–56. [Google Scholar] [CrossRef]
- Naghavi, M.; Vollset, S.E.; Ikuta, K.S.; Swetschinski, L.R.; Gray, A.P.; Wool, E.; Aguilar, G.R.; Mestrovic, T.; Smith, G.; Han, C.; et al. Global burden of bacterial antimicrobial resistance 1990–2021: A systematic analysis with forecasts to 2050. Lancet 2024, 404, 1199–1226. [Google Scholar] [CrossRef]
- Hoff, C.A.; Schmidt, S.S.; Hackert, B.J.; Worley, T.K.; Courcelle, J.; Courcelle, C.T. Events associated with DNA replication disruption are not observed in hydrogen peroxide-treated Escherichia coli. G3 Genes 2021, 11, jkab044. [Google Scholar] [CrossRef]
- Hutfilz, C.R.; Wang, N.E.; Hoff, C.A.; Lee, J.A.; Hackert, B.J.; Courcelle, J.; Courcelle, C.T. Manganese Is Required for the Rapid Recovery of DNA Synthesis following Oxidative Challenge in Escherichia coli. J. Bacteriol. 2019, 201, 24. [Google Scholar] [CrossRef]
- Wang, N.E.; Courcelle, E.J.; Coltman, S.M.; Spolek, R.L.; Courcelle, J.; Courcelle, C.T. Manganese transporters regulate the resumption of replication in hydrogen peroxide-stressed Escherichia coli. Biometals 2023, 36, 1361–1376. [Google Scholar] [CrossRef] [PubMed]
- Hantke, K. Regulation of ferric iron transport in Escherichia coli K12: Isolation of a constitutive mutant. Mol. Gen. Genet. 1981, 182, 288–292. [Google Scholar] [CrossRef] [PubMed]
- Andrews, S.C.; Harrison, P.M.; Guest, J.R. Cloning, sequencing, and mapping of the bacterioferritin gene (bfr) of Escherichia coli K-12. J. Bacteriol. 1989, 171, 3940–3947. [Google Scholar] [CrossRef] [PubMed]
- Ilari, A.; Ceci, P.; Ferrari, D.; Rossi, G.L.; Chiancone, E. Iron incorporation into Escherichia coli Dps gives rise to a ferritin-like microcrystalline core. J. Biol. Chem. 2002, 277, 37619–37623. [Google Scholar] [CrossRef]
- Chodavarapu, S.; Gomez, R.; Vicente, M.; Kaguni, J.M. Escherichia coli Dps interacts with DnaA protein to impede initiation: A model of adaptive mutation. Mol. Microbiol. 2008, 67, 1331–1346. [Google Scholar] [CrossRef]
- Colussi, C.; Albertini, M.C.; Coppola, S.; Rovidati, S.; Galli, F.; Ghibelli, L. H2O2-induced block of glycolysis as an active ADP-ribosylation reaction protecting cells from apoptosis. FASEB J. 2000, 14, 2266–2276. [Google Scholar] [CrossRef]
- Skarstad, K.; Katayama, T. Regulating DNA replication in bacteria. Cold Spring Harb. Perspect. Biol. 2013, 5, a012922. [Google Scholar] [CrossRef]
- Shi, Y.; Tang, M.; Sun, C.; Pan, Y.; Liu, L.; Long, Y.; Zheng, H. ATP Mimics pH-Dependent Dual Peroxidase-Catalase Activities Driving H2O2 Decomposition. CCS Chem. 2019, 1, 373–383. [Google Scholar] [CrossRef]
- Stauffer, L.T.; Plamann, M.D.; Stauffer, G.V. Cloning and characterization of the glycine-cleavage enzyme system of Escherichia coli. Gene 1986, 44, 219–226. [Google Scholar] [CrossRef]
- Klem, T.J.; Davisson, V.J. Imidazole glycerol phosphate synthase: The glutamine amidotransferase in histidine biosynthesis. Biochemistry 1993, 32, 5177–5186. [Google Scholar] [CrossRef]
- Kim, S.H.; Schneider, B.L.; Reitzer, L. Genetics and regulation of the major enzymes of alanine synthesis in Escherichia coli. J. Bacteriol. 2010, 192, 5304–5311. [Google Scholar] [CrossRef]
- Meinnel, T.; Schmitt, E.; Mechulam, Y.; Blanquet, S. Structural and biochemical characterization of the Escherichia coli argE gene product. J. Bacteriol. 1992, 174, 2323–2331. [Google Scholar] [CrossRef] [PubMed]
- Haley, E.E. Purification and properties of a beta-aspartyl peptidase from Escherichia coli. J. Biol. Chem. 1968, 243, 5748–5752. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Qimuge; Hao, J.; Yan, H.; Bach, T.; Fan, L.; Morigen. AspC-mediated aspartate metabolism coordinates the Escherichia coli cell cycle. PLoS ONE 2014, 9, e92229. [Google Scholar] [CrossRef] [PubMed]
- Verissimo, A.F.; Khalfaoui-Hassani, B.; Hwang, J.; Steimle, S.; Selamoglu, N.; Sanders, C.; Khatchikian, C.E.; Daldal, F. The thioreduction component CcmG confers efficiency and the heme ligation component CcmH ensures stereo-specificity during cytochrome c maturation. J. Biol. Chem. 2017, 292, 13154–13167. [Google Scholar] [CrossRef]
- Teelucksingh, T.; Thompson, L.K.; Zhu, S.; Kuehfuss, N.M.; Goetz, J.A.; Gilbert, S.E.; MacNair, C.R.; Geddes-McAlister, J.; Brown, E.D.; Cox, G. A genetic platform to investigate the functions of bacterial drug efflux pumps. Nat. Chem. Biol. 2022, 18, 1399–1409. [Google Scholar] [CrossRef]
- Scripture, J.B.; Voelker, C.; Miller, S.; O’Donnell, R.T.; Polgar, L.; Rade, J.; Horazdovsky, B.F.; Hogg, R.W. High-affinity L-arabinose transport operon. Nucleotide sequence and analysis of gene products. J. Mol. Biol. 1987, 197, 37–46. [Google Scholar] [CrossRef]
- Willis, L.; Huang, K.C. Sizing up the bacterial cell cycle. Nat. Rev. Microbiol. 2017, 15, 606–620. [Google Scholar] [CrossRef]
- Reyes-Lamothe, R.; Sherratt, D.J. The bacterial cell cycle, chromosome inheritance and cell growth. Nat. Rev. Microbiol. 2019, 17, 467–478. [Google Scholar] [CrossRef]
- Witz, G.; van Nimwegen, E.; Julou, T. Initiation of chromosome replication controls both division and replication cycles in E. coli through a double-adder mechanism. Elife 2019, 8, e48063. [Google Scholar] [CrossRef]
- Colin, A.; Micali, G.; Faure, L.; Cosentino Lagomarsino, M.; van Teeffelen, S. Two different cell-cycle processes determine the timing of cell division in Escherichia coli. Elife 2021, 10, e67495. [Google Scholar] [CrossRef]
- Si, F.; Le Treut, G.; Sauls, J.T.; Vadia, S.; Levin, P.A.; Jun, S. Mechanistic Origin of Cell-Size Control and Homeostasis in Bacteria. Curr. Biol. 2019, 29, 1760–1770.e1767. [Google Scholar] [CrossRef] [PubMed]
- Alekshun, M.N.; Levy, S.B. The mar regulon: Multiple resistance to antibiotics and other toxic chemicals. Trends Microbiol. 1999, 7, 410–413. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Wang, Y.; Wang, X.; Ou, D.; Ling, N.; Zhang, J.; Wu, Q.; Ye, Y. Role of the multiple efflux pump protein TolC on growth, morphology, and biofilm formation under nitric oxide stress in Cronobacter malonaticus. JDS Commun. 2021, 2, 98–103. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.J.; Wu, Y.C.; Kuo, C.J.; Hsuan, S.L.; Chen, T.H. TolC is important for bacterial survival and oxidative stress response in Salmonella enterica serovar Choleraesuis in an acidic environment. Vet. Microbiol. 2016, 193, 42–48. [Google Scholar] [CrossRef]
- Bogomolnaya, L.M.; Andrews, K.D.; Talamantes, M.; Maple, A.; Ragoza, Y.; Vazquez-Torres, A.; Andrews-Polymenis, H. The ABC-type efflux pump MacAB protects Salmonella enterica serovar typhimurium from oxidative stress. mBio 2013, 4, e00630-13. [Google Scholar] [CrossRef]
- Horemans, S.; Pitoulias, M.; Holland, A.; Pateau, E.; Lechaplais, C.; Ekaterina, D.; Perret, A.; Soultanas, P.; Janniere, L. Pyruvate kinase, a metabolic sensor powering glycolysis, drives the metabolic control of DNA replication. BMC Biol. 2022, 20, 87. [Google Scholar] [CrossRef]
- Murray, H.; Koh, A. Multiple regulatory systems coordinate DNA replication with cell growth in Bacillus subtilis. PLoS Genet. 2014, 10, e1004731. [Google Scholar] [CrossRef]
- Schmacht, M.; Lorenz, E.; Senz, M. Microbial production of glutathione. World J. Microbiol. Biotechnol. 2017, 33, 106. [Google Scholar] [CrossRef]
- Matilla, B.; Mauriz, J.L.; Culebras, J.M.; González-Gallego, J.; González, P. [Glycine: A cell-protecting anti-oxidant nutrient]. Nutr. Hosp. 2002, 17, 2–9. [Google Scholar]
- Coyle, J.T.; Puttfarcken, P. Oxidative stress, glutamate, and neurodegenerative disorders. Science 1993, 262, 689–695. [Google Scholar] [CrossRef] [PubMed]
- Torricelli, P.; Elia, A.C.; Magara, G.; Feriotto, G.; Forni, C.; Borromeo, I.; De Martino, A.; Tabolacci, C.; Mischiati, C.; Beninati, S. Reduction of oxidative stress and ornithine decarboxylase expression in a human prostate cancer cell line PC-3 by a combined treatment with α-tocopherol and naringenin. Amino Acids 2021, 53, 63–72. [Google Scholar] [CrossRef]
- Jenkins, C.M.; Yang, K.; Liu, G.; Moon, S.H.; Dilthey, B.G.; Gross, R.W. Cytochrome c is an oxidative stress-activated plasmalogenase that cleaves plasmenylcholine and plasmenylethanolamine at the sn-1 vinyl ether linkage. J. Biol. Chem. 2018, 293, 8693–8709. [Google Scholar] [CrossRef] [PubMed]
- Hendrickson, W.; Stoner, C.; Schleif, R. Characterization of the Escherichia coli araFGH and araJ promoters. J. Mol. Biol. 1990, 215, 497–510. [Google Scholar] [CrossRef]
- Hendrickson, W.; Flaherty, C.; Molz, L. Sequence elements in the Escherichia coli araFGH promoter. J. Bacteriol. 1992, 174, 6862–6871. [Google Scholar] [CrossRef]
- Zheng, D.; Constantinidou, C.; Hobman, J.L.; Minchin, S.D. Identification of the CRP regulon using in vitro and in vivo transcriptional profiling. Nucleic Acids Res. 2004, 32, 5874–5893. [Google Scholar] [CrossRef] [PubMed]
- Fic, E.; Bonarek, P.; Gorecki, A.; Kedracka-Krok, S.; Mikolajczak, J.; Polit, A.; Tworzydlo, M.; Dziedzicka-Wasylewska, M.; Wasylewski, Z. cAMP receptor protein from Escherichia coli as a model of signal transduction in proteins--a review. J. Mol. Microbiol. Biotechnol. 2009, 17, 1–11. [Google Scholar] [CrossRef]
- Zhu, W.; Chen, M.; Zhang, X.; Su, J.; Zhang, X.; Nong, Y.; Wang, B.; Guo, W.; Xue, Y.; Wang, D.; et al. Antibiotic tolerance due to restriction of cAMP-Crp regulation by mannitol, a non-glucose-family PTS carbon source. mSphere 2024, 9, e0077224. [Google Scholar] [CrossRef]
- Du, D.; Wang-Kan, X.; Neuberger, A.; van Veen, H.W.; Pos, K.M.; Piddock, L.J.V.; Luisi, B.F. Multidrug efflux pumps: Structure, function and regulation. Nat. Rev. Microbiol. 2018, 16, 523–539. [Google Scholar] [CrossRef]
- Fritz, G.; Megerle, J.A.; Westermayer, S.A.; Brick, D.; Heermann, R.; Jung, K.; Rädler, J.O.; Gerland, U. Single cell kinetics of phenotypic switching in the arabinose utilization system of E. coli. PLoS ONE 2014, 9, e89532. [Google Scholar] [CrossRef]
- Li, B.; Chen, X.; Yang, J.Y.; Gao, S.; Bai, F. Intracellular ATP concentration is a key regulator of bacterial cell fate. J. Bacteriol. 2024, 206, e0020824. [Google Scholar] [CrossRef] [PubMed]
- Benov, L. How superoxide radical damages the cell. Protoplasma 2001, 217, 33–36. [Google Scholar] [CrossRef]
- Sobota, J.M.; Gu, M.; Imlay, J.A. Intracellular hydrogen peroxide and superoxide poison 3-deoxy-D-arabinoheptulosonate 7-phosphate synthase, the first committed enzyme in the aromatic biosynthetic pathway of Escherichia coli. J. Bacteriol. 2014, 196, 1980–1991. [Google Scholar] [CrossRef] [PubMed]
- Imlay, J.A. Cellular defenses against superoxide and hydrogen peroxide. Annu. Rev. Biochem. 2008, 77, 755–776. [Google Scholar] [CrossRef]
- Datsenko, K.A.; Wanner, B.L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 2000, 97, 6640–6645. [Google Scholar] [CrossRef]
- Thomason, L.C.; Costantino, N.; Court, D.L. E. coli genome manipulation by P1 transduction. Curr. Protoc. Mol. Biol. 2007, 79, 1.17.1–1.17.8. [Google Scholar] [CrossRef]
- Morigen; Molina, F.; Skarstad, K. Deletion of the datA site does not affect once-per-cell-cycle timing but induces rifampin-resistant replication. J. Bacteriol. 2005, 187, 3913–3920. [Google Scholar] [CrossRef] [PubMed]
- Zhu, W.; Xi, L.; Qiao, J.; Du, D.; Wang, Y.; Morigen. Involvement of OxyR and Dps in the repression of replication initiation by DsrA small RNA in Escherichia coli. Gene 2023, 882, 147659. [Google Scholar] [CrossRef]
- Baba, T.; Ara, T.; Hasegawa, M.; Takai, Y.; Okumura, Y.; Baba, M.; Datsenko, K.A.; Tomita, M.; Wanner, B.L.; Mori, H. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: The Keio collection. Mol. Syst. Biol. 2006, 2, 2006.0008. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qiao, J.; Zhu, W.; Du, D.; Morigen, M. Characterizing Common Factors Affecting Replication Initiation During H2O2 Exposure and Genetic Mutation-Induced Oxidative Stress in Escherichia coli. Int. J. Mol. Sci. 2025, 26, 2968. https://doi.org/10.3390/ijms26072968
Qiao J, Zhu W, Du D, Morigen M. Characterizing Common Factors Affecting Replication Initiation During H2O2 Exposure and Genetic Mutation-Induced Oxidative Stress in Escherichia coli. International Journal of Molecular Sciences. 2025; 26(7):2968. https://doi.org/10.3390/ijms26072968
Chicago/Turabian StyleQiao, Jiaxin, Weiwei Zhu, Dongdong Du, and Morigen Morigen. 2025. "Characterizing Common Factors Affecting Replication Initiation During H2O2 Exposure and Genetic Mutation-Induced Oxidative Stress in Escherichia coli" International Journal of Molecular Sciences 26, no. 7: 2968. https://doi.org/10.3390/ijms26072968
APA StyleQiao, J., Zhu, W., Du, D., & Morigen, M. (2025). Characterizing Common Factors Affecting Replication Initiation During H2O2 Exposure and Genetic Mutation-Induced Oxidative Stress in Escherichia coli. International Journal of Molecular Sciences, 26(7), 2968. https://doi.org/10.3390/ijms26072968




