Abstract
During a study of Botryosphaeriaceae species associated with grapevine trunk diseases in the Czech Republic, a collection of 22 Botryosphaeriaceae-like strains were isolated from four cultivars (Blaufränkisch, Pálava, Pinot Noir, and Welschriesling) in four distinct vineyards. Based on morphology and DNA sequence data (ITS, tub2, and tef), four species were identified: Botryosphaeria dothidea, Diplodia mutila, D. seriata, and Neofusicoccum parvum. These species are reported for the first time from grapevine in the Czech Republic. Relationships between vascular lesions and particular species were highlighted in this study. Diplodia seriata was the most frequently isolated species, present in all four sampled cultivars, while D. mutila was the least frequent, present only in ‘Pálava’. The cultivar Pinot Noir was the most tolerant host for Botryosphaeriaceae fungi.
1. Introduction
Grapevine (Vitis vinifera L.) is one of the Czech Republic’s most valuable fruit crops. In 2021, registered vineyards covered an area of 16,360 hectares, producing 90,060 tonnes annually, with an estimated market value of $77,162,000 USD [1]. During the last few decades, an increased incidence of grapevine trunk diseases (GTDs) has been reported in grape-producing countries worldwide [2,3], with estimated economical loses exceeding 1 billion dollars annually [4].
The Botryosphaeriaceae family comprises a diverse group of cosmopolitan fungi, responsible for dieback and canker diseases in various woody hosts, including grapevines [5]. More than 26 different Botryosphaeriaceae species have been associated with Botryosphaeria dieback of grapevine [6]. External symptoms of Botryosphaeria dieback on grapevine include leaf spots, leaf wilting, fruit rots, perennial cankers, cordon dieback, and sudden plant mortality, while internal wood symptoms manifest as wedge-shaped necroses and dark lines beneath the bark [7].
Plants are usually infected by fungal spores that colonize the plants through winter pruning wounds. Besides infection through pruning wounds, the presence of latent infections caused by Botryosphaeriaceae fungi has been well documented in nurseries during the grapevine propagation process [8,9,10,11]. It was confirmed that Botryosphaeriaceae fungi can live within their host as endophytes or latent pathogens that become pathogenic when their hosts are exposed to stress conditions [12,13].
Due to a lack of studies, very little is known about the incidence of Botryosphaeriaceae pathogens in Czech vineyards. Thus, the aim of this study was to provide a comprehensive overview of the Botryosphaeriaceae fungi responsible for Botryosphaeria dieback in the Czech Republic.
2. Materials and Methods
2.1. Collection and Isolation
Plant material displaying symptoms of dieback (Figure 1) and asymptomatic material, in the case of a young 3-year-old vineyard, were collected from four commercial vineyards located in the South Moravia region of the Czech Republic with the permission of landowner (Table 1). The field observation and sampling were performed in July 2019. In total, 40 grapevines (ten plants per vineyard) were sampled and immediately transported to the laboratory of Mendeleum–Institute of Genetics, Mendel University, the Czech Republic, for further processing. Trunks and arms were debarked using a sterile scalpel and cut longitudinally and transversely to identify the type and location of internal wood necrosis. Bark-less wood tissues were subjected to surface sterilization. From each tissue, wood fragments, approx. 1 cm3, were cut and surface sterilized with 1% sodium hypochlorite for ten minutes and then rinsed three times with sterile distilled water, following protocols previously described [14]. The disinfected wood fragments were cut into small chips of 5 × 2 mm and aseptically transferred onto Petri dishes (five chips per plate) containing potato dextrose agar (PDA, HiMedia, Mumbai, India) supplemented with 0.5 g/L streptomycin sulfate (Sigma–Aldrich, St. Louis, MO, USA). The plates were incubated at 25 °C in the dark for four weeks, and fungal growth was checked every two days. Newly developed mycelia were immediately transferred to new PDA plates and purified using hyphal tip isolation [15]. All fungal isolates were deposited in MEND-F, Fungal Culture Collection of Mendeleum, Mendel University in Brno, the Czech Republic.
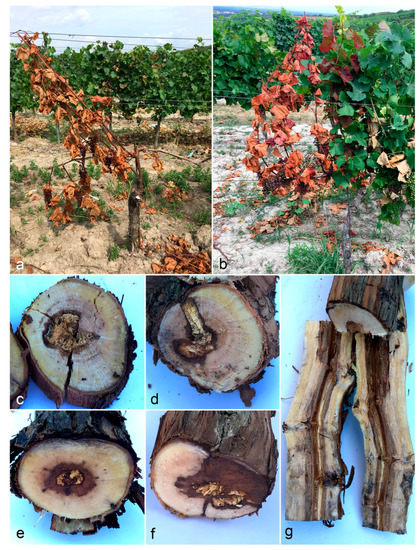

Figure 1.
Typical symptoms of sampled plants. (a,b) Apoplexy. (c–g) Internal wood necroses.

Table 1.
Sampled localities and sampling characterization.
2.2. Morphology
Botryosphaeriaceae-like isolates were selected according to the keys provided in the study by Phillips et al. [5]. Culture characteristics were determined on PDA incubated for 7 days at 25 °C in the dark. Water agar plates (WA, HiMedia, Mumbai, India) with double autoclaved pine needles were incubated for 1–3 weeks at 25 °C with exposure to near-UV light to induce sporulation.
2.3. DNA Extraction and Amplification
Genomic DNA was extracted from 7-day-old mycelium grown on PDA at 25 °C in darkness using a NucleoSpin DNA extraction kit (Macherey-Nagel, Düren, Germany) following the manufacturer’s protocol. To confirm the identity of the fungal species, fragments of three genes were amplified: internal transcribed spacer region (ITS), beta-tubulin (tub2), and translation elongation factor 1-alpha (tef). PCR was performed utilizing G2 Flexi DNA polymerase (Promega, Madison, USA), and the primers are listed in Table 2, following protocols previously described [16,17]. Resulting products were purified using NucleoSpin Gel and PCR Clean-up Kit (Macherey-Nagel, Düren, Germany) following the manufacturer’s protocol. Subsequently, the purified products were sequenced from both ends using the Sanger method at Eurofins Genomics (Ebersberg, Germany).

Table 2.
Primers used for PCR amplification and sequencing.
2.4. Phylogenetic Analyses
To identify the isolates, newly generated DNA sequences, together with those retrieved from GenBank, were subjected to phylogenetic analyses (Table 3). The dataset of each gene was aligned separately using the MAFFT v. 7 employing the European Bioinformatics Institute platform (EMBL-EBI, https://www.ebi.ac.uk, accessed on 1 February 2023) [22]. Obtained alignment was manually checked and edited when necessary, using Geneious Prime® (v.2023.0.1., Biomatters Ltd., Auckland, New Zeland). Concatenated dataset was built in Sequence Matrix v.1.8 [23], and the missing information sites were denoted by a question mark. The combined (ITS, tub2, and tef) dataset was subjected to Maximum Likelihood (ML) analyses. Phylogenetic trees were constructed using IQ-TREE 2 [24], running 1000 bootstrap replicates. The best model for ML analyses was selected according to the Akaike Information Criterion (AIC). Bayesian analyses (BI) employed MrBayes v. 3.2.7 [25,26]. The BI analyses included four parallel runs of 50 M generations starting from a random tree topology, every 1000 generations were sampled, and the first 25% of the trees were discarded as the ‘burn-in’. The most suitable substitution model was determined separately for each locus using jModelTest v. 2.1.7 [27]. Trees were visualized in iTOL v. 6.7 [28] and edited in Adobe Illustrator CC 2019. The resulting trees of both methods shared a similar topology; thus, we decided to present ML trees with support values of both methods–bootstrap (BS) and posterior probabilities (PP) labelled at the nodes. Values below 0.85 (PP) and 75% (BS) support are not shown or indicated with a hyphen. The alignments and corresponding trees are available on Figshare (10.6084/m9.figshare.22837472).

Table 3.
Fungal species and barcodes used in phylogenetic analyses.
3. Results
3.1. Fungal Isolation
In total, 204 isolates were obtained from the 40 sampled plants. A preliminary morphological characterization revealed 22 isolates that displayed morphological and growth characteristics consistent with the Botryosphaeriaceae family.
3.2. Phylogenetic Analyses
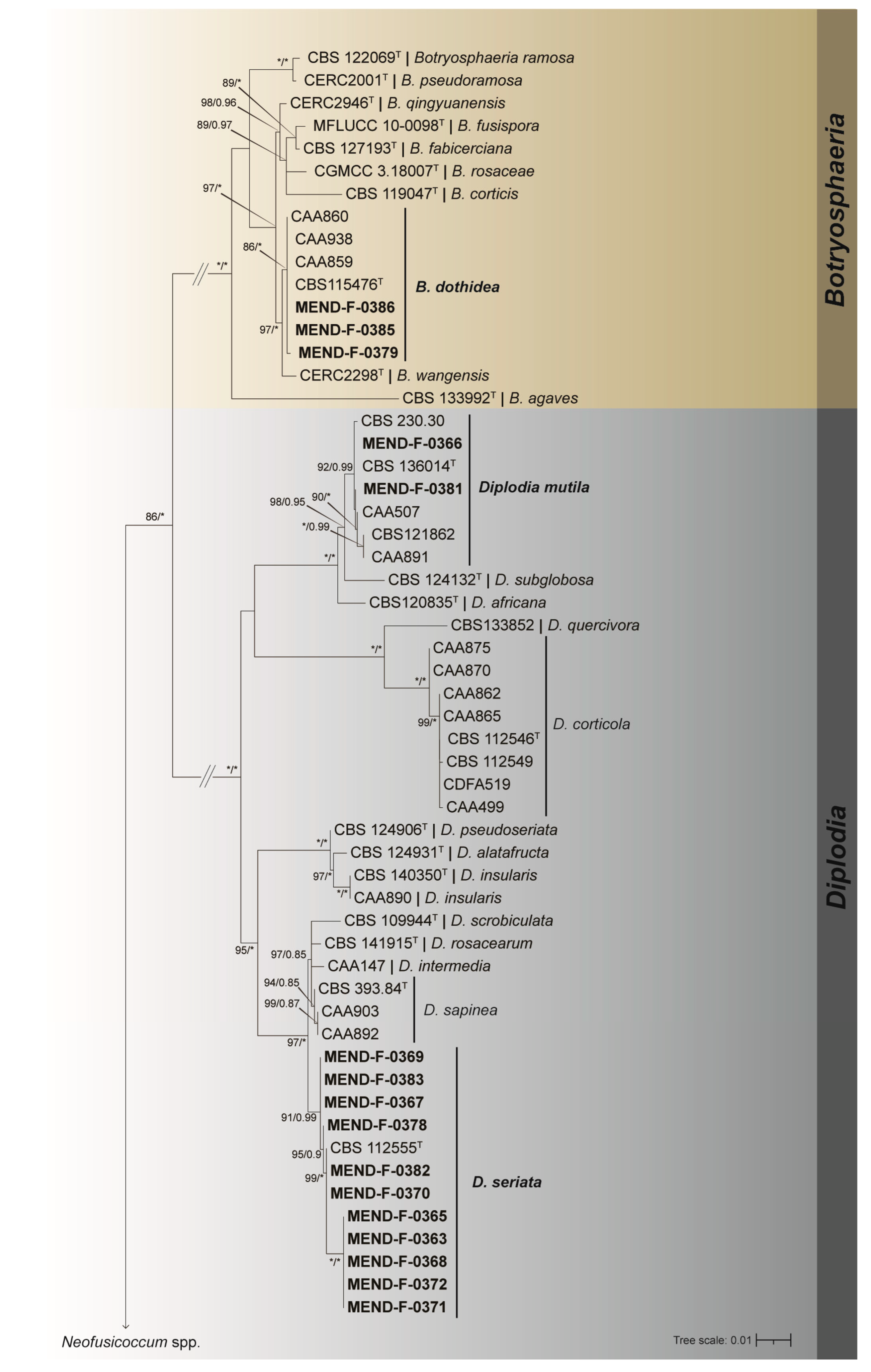
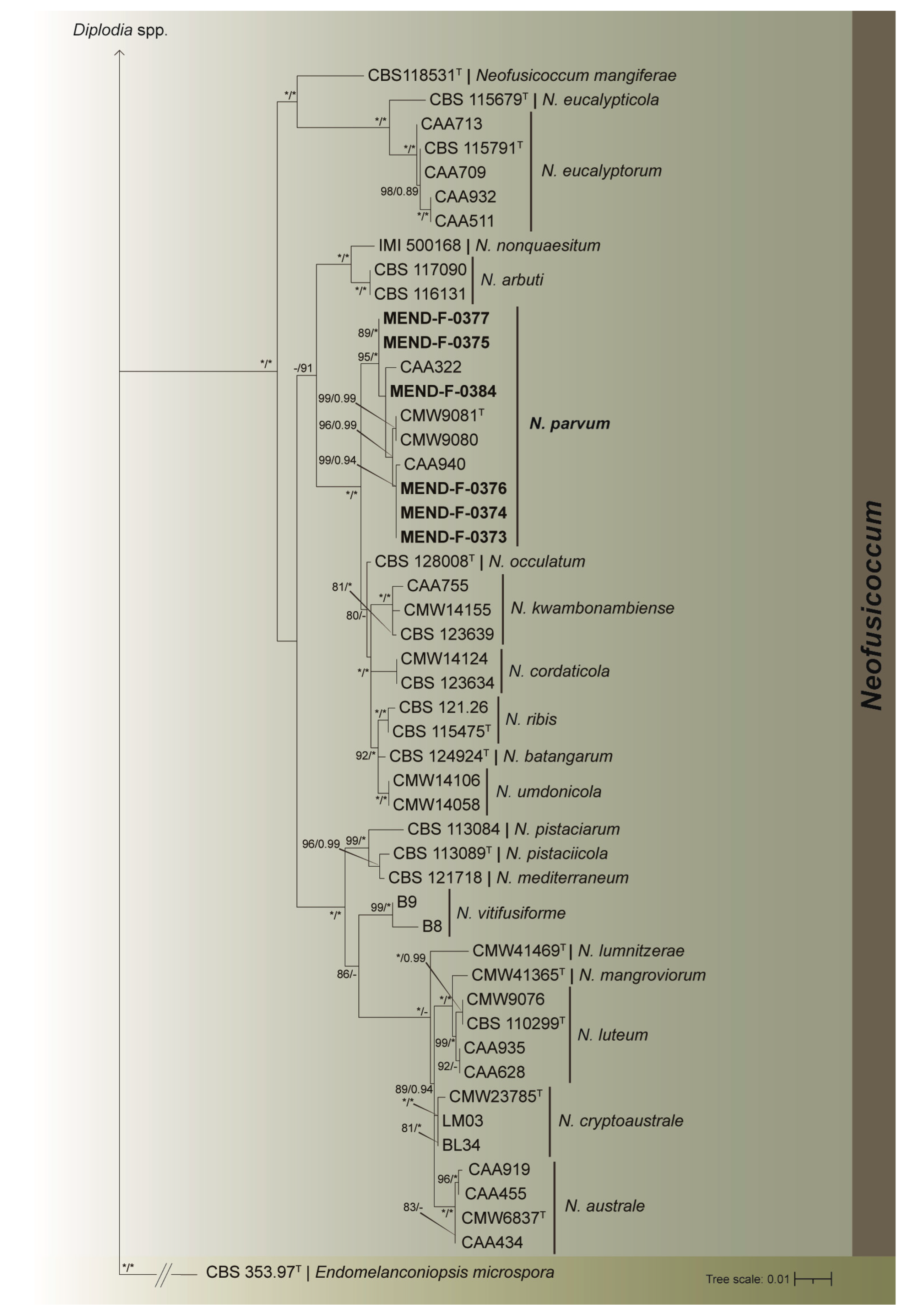
Molecular identification was performed on the 22 representative isolates, and their identity confirmed employing three-gene based (ITS, tub2, tef) phylogenetic analyses. The dataset consisted of sequences from 106 isolates (Table 3), including the outgroup Endomelanconiopsis microspora (CBS 353.97T). The combined dataset contained a total of 1259 characters, including alignment gaps. Among these characters, 822 were conserved, 351 provided informative data for parsimony analysis, and 86 were unique. Detailed results for each individual gene dataset, along with the corresponding models used, can be found in Table 4. The ML/BI analyses (Figure 2 and Figure 3) placed 11 isolates in group with the type strain of D. seriata (CBS 112555) with strong support of 91/0.99 (BP/pp); six isolates formed a fully supported clade with the type strain (CMW 9081) and three other Neofusicoccum parvum strains; three isolates were placed in group with the type strain (CBS 115476) and three other strains of Botryosphaeria dothidea with robust 97/1.0 (BP/pp) support; finally, two isolates were displayed in a well-supported clade 98/0.95 (BP/pp) with the type strain (CBS 121862) and three other strains of D. mutila.

Table 4.
Detailed characteristics of phylogeny datasets.
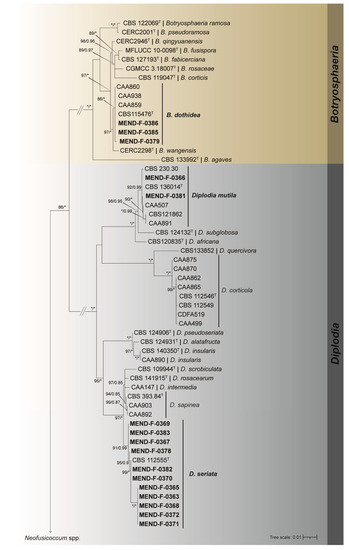
Figure 2.
Maximum likelihood tree generated from the combined (ITS, tef, and tub2) Botryosphaeriaceae dataset. Support values of both methods–bootstrap (BS) and posterior probabilities (pp) labelled at the nodes. Values below 75% (BS) and 0.85 (pp) support are not shown or indicated with a hyphen. Asterisk represents full support. Strains obtained in this study are highlighted in bold. T indicates ex-type strain. The tree continues in Figure 3.
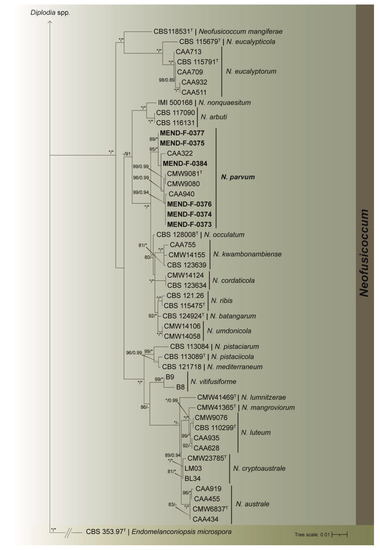
Figure 3.
Maximum likelihood tree generated from the combined (ITS, tef, and tub2) Botryosphaeriaceae dataset. Support values of both methods–bootstrap (BS) and posterior probabilities (pp) labelled at the nodes. Values below 75% (BS) and 0.85 (pp) support are not shown or indicated with a hyphen. Asterisk represents full support. Strains obtained in this study are highlighted in bold. T indicates ex-type strain. Endomelanconiopsis microspora strain CBS 353.97T served as an outgroup.
3.3. Species Diversity in Different Grapevine Varieties and Wood necrosis
Diplodia seriata was the most frequently isolated species (11 isolates), present in all four sampled varieties, followed by N. parvum (n = 6) isolated from both red varieties, B. dothidea (n = 3) detected only in cf. Pinot Noir, and D. mutila (n = 2) detected only in cf. Pálava.
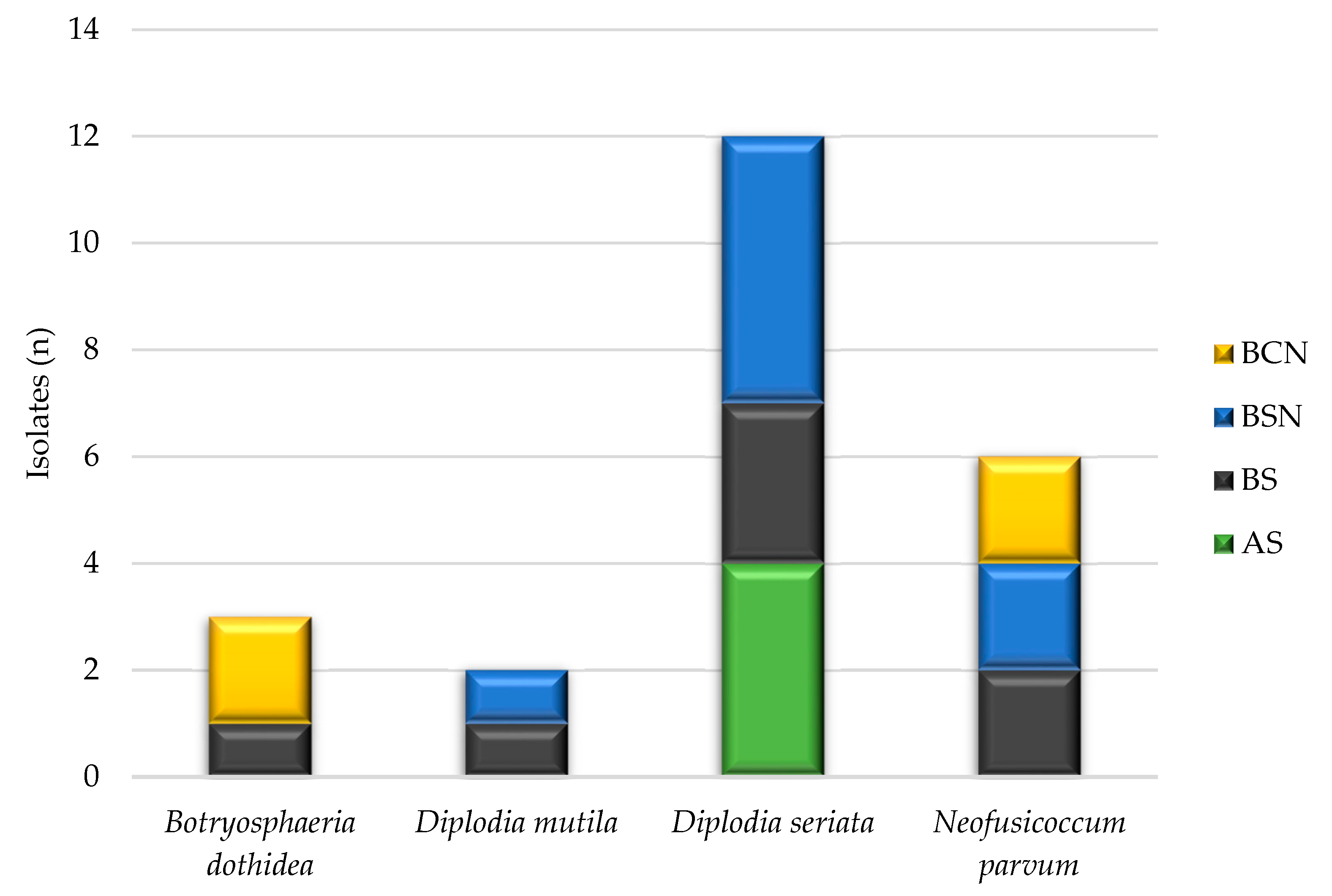
Wood necroses associated with specific pathogens are displayed in Figure 4. Three different shapes of inner necrosis were observed in transverse sections of trunk and arm from symptomatic grapevines: black spots (BS); black sectorial necrosis (BSN); black central necrosis (BCN). Botryosphaeriaceae isolates were inhabiting mostly the BSN (35%), followed by BS and BCS with 31% and 17%, respectively. The remaining 17% of the obtained Botryosphaeriaceae isolates originated from asymptomatic wood tissues from the young Welschriesling vineyard.
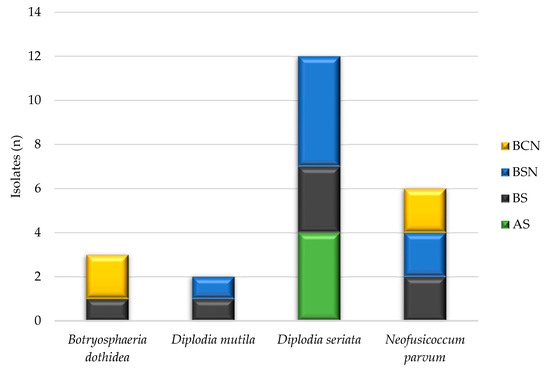
Figure 4.
The association between the Botryosphaeriaceae isolates and the type of wood necrosis. AS = asymptomatic; BS = black spots; BSN = black sectorial necrosis; BCN = black central necrosis.
4. Discussion
This study provides the initial comprehensive evaluation of the occurrence of Botryosphaeriaceae species in grapevines within Czech vineyards. Among 22 Botryosphaeriaceae strains obtained, four species belonging to the three genera were detected, among which Diplodia seriata De Not. comprised 50%, Neofusicoccum parvum (Pennycook and Samuels) Crous, Slippers, and A.J.L. Phillips 27%, Botryosphaeria dothidea (Moug.) Ces. and De Not. 14%, and Diplodia mutila (Fr.) Mont. 9%. These species have already been isolated from grapevines worldwide and their pathogenicity has been confirmed [29,30,31,32,33,34,35]. The most isolated species in the Czech Republic was D. seriata. This finding is in accordance with previous studies that have identified D. seriata as the predominant fungus associated with the decline of mature vines in Iran [36], Mexico [37], Hungary [38], and Tunisia [39].
In our study, the pathogen D. seriata was also isolated from the asymptomatic material from the young (3-year-old) vineyard, suggesting latent infection from propagation process in grapevine nursery. This result is consistent with previous studies that reported infection by Botryosphaeriaceae fungi in grapevine nurseries. Fourie et al. reported the presence of latent infection caused by Botryosphaeriaceae fungi in rootstock mother plants in South Africa [40]. Aroca et al. reported presence of three Botryosphaeriaceae fungi in grapevine propagation material in Spain, namely, Botryosphaeria dothidea, Diplodia seriata, and Neofusicoccum parvum [41]. Eichmeier et al. also reported the presence of the same three Botryosphaeriaceae fungi in young grapevine seedlings in Spain [42].
To the best of our knowledge, only two studies have been performed to date on detection of GTDs in the Czech Republic. The initial investigation was conducted by a study of Baranek et al. [43]. The authors examined two grapevine cultivars, namely, ‘Chardonnay’ and ‘Cabernet Sauvignon’, and identified a total of 21 fungal taxa. Among these taxa, only one species, Botryosphaeria dothidea, was classified under the Botryosphaeriaceae family. Subsequently, an incidence of Dactylonectria torresensis, a causal agent of black-foot disease, was reported from Czech vineyards [44].
Multiple Botryosphaeriaceae species do not have specificity in host range and have the ability to transition from their original indigenous hosts to agricultural crops cultivated in proximity [45]. Excluding grapevine, two Botryosphaeriaceae spp. were recently reported causing dieback of highbush blueberry from the Czech Republic, namely, Lasiodiplodia theobromae and Neofusicoccum parvum [46,47].
5. Conclusions
This study provided an investigation of the Botryosphaeriaceae fungi associated with GTDs in four Czech vineyards. Four pathogenic Botryosphaeriaceae spp. have been identified based on phylogenetic analyses, and a correlation between fungal isolates, grapevine cultivar, and type of wood necroses was described in this study. The detection of the pathogen Diplodia seriata in young asymptomatic grapevine plants represents an urgent matter for Czech viticulture. Producing healthy propagation material is an essential requirement. We propose incorporating molecular detection techniques into nurseries to reveal hidden fungal infection. We also highly recommend implementing preventative treatment during the grapevine propagation process using hot water treatment [48], novel nanomaterials [49], or phenolic compounds [50].
Author Contributions
Conceptualization, A.E. and A.B.-T.; Methodology, A.B.-T.; Isolation, A.E.M., D.A.T., J.P., K.S. and M.S.; DNA extraction, D.A.T., J.P. and M.S.; Molecular work, M.S.; Phylogeny analyses, M.S.; Visualization, M.S.; Writing—Original Draft Preparation, M.S. and D.A.T.; Review and Editing, A.B.-T., A.E. and D.A.T.; Supervision, A.E.; Project Administration, M.S. and A.E.; Funding Acquisition, A.E. and M.S. All authors have read and agreed to the published version of the manuscript.
Funding
The project was supported by the Internal Grant of Mendel University in Brno, No. IGA-ZF/2021-SI1003 and by Ministerstvo kultury, Program NAKI III: DH23P03OVV053.
Data Availability Statement
Newly generated sequences were deposited in the NCBI GenBank database under the accession numbers shown in Table 3. The alignments and corresponding trees are available on Figshare (10.6084/m9.figshare.22837472).
Conflicts of Interest
The authors declare no conflict of interest.
References
- FAOSTAT. Food and Agriculture Organization of the United Nations; FAOSTAT Database: Rome, Italy, 2023. [Google Scholar]
- Fontaine, F.; Gramaje, D.; Armengol, J.; Smart, R.; Nagy, Z.A.; Borgo, M.; Rego, C.; Corio-Costet, M.-F. Grapevine Trunk Diseases. A Review; OIV Publications: Paris, France, 2016; p. 24. [Google Scholar]
- Bertsch, C.; Ramírez-Suero, M.; Magninrobert, M.; Larignon, P.; Chong, J.; Abou-Mansour, E.; Spagnolo, A.; Clément, C.; Fontaine, F. Grapevine trunk diseases: Complex and still poorly understood. Plant Pathol. 2013, 62, 243–265. [Google Scholar] [CrossRef]
- Hofstetter, V.; Buyck, B.; Croll, D.; Viret, O.; Couloux, A.; Gindro, K. What if esca disease of grapevine were not a fungal disease? Fungal Divers. 2012, 54, 51–67. [Google Scholar] [CrossRef]
- Phillips, A.J.L.; Alves, A.; Abdollahzadeh, J.; Slippers, B.; Wingfield, M.J.; Groenewald, J.Z.; Crous, P.W. The Botryosphaeriaceae: Genera and species known from culture. Stud. Mycol. 2013, 76, 51–167. [Google Scholar] [CrossRef]
- Gramaje, D.; Úrbez-Torres, J.R.; Sosnowski, M.R. Managing Grapevine Trunk Diseases With Respect to Etiology and Epidemiology: Current Strategies and Future Prospects. Plant Dis. 2017, 102, 12–39. [Google Scholar] [CrossRef] [PubMed]
- Urbez-Torres, J. The status of Botryosphaeriaceae species infecting grapevines. Phytopathol. Mediterr. 2011, 50, 5–45. [Google Scholar]
- Aroca, Á.; Gramaje, D.; Armengol, J.; García-Jiménez, J.; Raposo, R. Evaluation of the grapevine nursery propagation process as a source of Phaeoacremonium spp. and Phaeomoniella chlamydospora and occurrence of trunk disease pathogens in rootstock mother vines in Spain. Eur. J. Plant Pathol. 2010, 126, 165–174. [Google Scholar] [CrossRef]
- Pintos, C.; Redondo, V.; Costas, D.; Aguín, O.; Mansilla, P. Fungi associated with grapevine trunk diseases in nursery-produced Vitis vinifera plants. Phytopathol. Mediterr. 2018, 57, 407–424. [Google Scholar] [CrossRef]
- Carbone, M.J.; Gelabert, M.; Moreira, V.; Mondino, P.; Alaniz, S. Grapevine nursery propagation material as source of fungal trunk disease pathogens in Uruguay. Front. Fungal Biol. 2022, 3. [Google Scholar] [CrossRef]
- Billones-Baaijens, R.; Ridgway, H.J.; Jones, E.E.; Jaspers, M.V. Inoculum sources of Botryosphaeriaceae species in New Zealand grapevine nurseries. Eur. J. Plant Pathol. 2013, 135, 159–174. [Google Scholar] [CrossRef]
- Slippers, B.; Wingfield, M.J. Botryosphaeriaceae as endophytes and latent pathogens of woody plants: Diversity, ecology and impact. Fungal Biol. Rev. 2007, 21, 90–106. [Google Scholar] [CrossRef]
- Marsberg, A.; Kemler, M.; Jami, F.; Nagel, J.H.; Postma-Smidt, A.; Naidoo, S.; Wingfield, M.J.; Crous, P.W.; Spatafora, J.W.; Hesse, C.N.; et al. Botryosphaeria dothidea: A latent pathogen of global importance to woody plant health. Mol. Plant Pathol. 2017, 18, 477–488. [Google Scholar] [CrossRef] [PubMed]
- Berraf-Tebbal, A.; Mahamedi, A.E.; Aigoun-Mouhous, W.; Spetik, M.; Čechová, J.; Pokluda, R.; Baránek, M.; Eichmeier, A.; Alves, A. Lasiodiplodia mitidjana sp. nov. and other Botryosphaeriaceae species causing branch canker and dieback of Citrus sinensis in Algeria. PLoS ONE 2020, 15, e0232448. [Google Scholar] [CrossRef] [PubMed]
- Jensen, A.B.; Aronstein, K.; Flores, J.M.; Vojvodic, S.; Palacio, M.A.; Spivak, M. Standard methods for fungal brood disease research. J. Apic. Res. 2019, 52, 1–20. [Google Scholar] [CrossRef]
- Eichmeier, A.; Pecenka, J.; Spetik, M.; Necas, T.; Ondrasek, I.; Armengol, J.; León, M.; Berlanas, C.; Gramaje, D. Fungal Trunk Pathogens Associated with Juglans regia in the Czech Republic. Plant Dis. 2019, 104, 761–771. [Google Scholar] [CrossRef]
- Spetik, M.; Berraf-Tebbal, A.; Gramaje, D.; Mahamedi, A.E.; Stusková, K.; Burgova, J.; Eichmeier, A. Paecilomyces clematidis (Eurotiales, Thermoascaceae): A new species from Clematis root. Phytotaxa 2022, 559, 238–246. [Google Scholar] [CrossRef]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: San Diego, CA, USA, 1990; pp. 315–322. [Google Scholar]
- Carbone, I.; Kohn, L.M. A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 1999, 91, 553–556. [Google Scholar] [CrossRef]
- O’Donnell, K.; Cigelnik, E. Two Divergent Intragenomic rDNA ITS2 Types within a Monophyletic Lineage of the Fungus Fusarium Are Nonorthologous. Mol. Phylogenetics Evol. 1997, 7, 103–116. [Google Scholar] [CrossRef]
- Glass, N.L.; Donaldson, G.C. Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl. Environ. Microbiol. 1995, 61, 1323–1330. [Google Scholar] [CrossRef]
- Li, W.; Cowley, A.; Uludag, M.; Gur, T.; McWilliam, H.; Squizzato, S.; Park, Y.M.; Buso, N.; Lopez, R. The EMBL-EBI bioinformatics web and programmatic tools framework. Nucleic Acids Res. 2015, 43, 580–584. [Google Scholar] [CrossRef]
- Vaidya, G.; Lohman, D.J.; Meier, R. SequenceMatrix: Concatenation software for the fast assembly of multi-gene datasets with character set and codon information. Cladistics 2011, 27, 171–180. [Google Scholar] [CrossRef]
- Minh, B.Q.; Schmidt, H.A.; Chernomor, O.; Schrempf, D.; Woodhams, M.D.; von Haeseler, A.; Lanfear, R. IQ-TREE 2: New Models and Efficient Methods for Phylogenetic Inference in the Genomic Era. Mol. Biol. Evol. 2020, 37, 1530–1534. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef]
- Ronquist, F.; Teslenko, M.; van der Mark, P.; Ayres, D.L.; Darling, A.; Höhna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient Bayesian Phylogenetic Inference and Model Choice across a Large Model Space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef]
- Posada, D. jModelTest: Phylogenetic Model Averaging. Mol. Biol. Evol. 2008, 25, 1253–1256. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef] [PubMed]
- Úrbez-Torres, J.R.; Gubler, W.D. Pathogenicity of Botryosphaeriaceae Species Isolated from Grapevine Cankers in California. Plant Dis. 2009, 93, 584–592. [Google Scholar] [CrossRef] [PubMed]
- Akgul, D.S.; Savas, N.G.; Eskalen, A. First Report of Wood Canker Caused by Botryosphaeria dothidea, Diplodia seriata, Neofusicoccum parvum, and Lasiodiplodia theobromae on Grapevine in Turkey. Plant Dis. 2013, 98, 568. [Google Scholar] [CrossRef]
- Arkam, M.; Alves, A.; Lopes, A.; Čechová, J.; Pokluda, R.; Eichmeier, A.; Zitouni, A.; Mahamedi, A.E.; Berraf-Tebbal, A. Diversity of Botryosphaeriaceae causing grapevine trunk diseases and their spatial distribution under different climatic conditions in Algeria. Eur. J. Plant Pathol. 2021, 161, 933–952. [Google Scholar] [CrossRef]
- Billones-Baaijens, R.; Savocchia, S. A review of Botryosphaeriaceae species associated with grapevine trunk diseases in Australia and New Zealand. Australas. Plant Pathol. 2019, 48, 3–18. [Google Scholar] [CrossRef]
- Yan, J.-Y.; Xie, Y.; Zhang, W.; Wang, Y.; Liu, J.-K.; Hyde, K.D.; Seem, R.C.; Zhang, G.-Z.; Wang, Z.-Y.; Yao, S.-W.; et al. Species of Botryosphaeriaceae involved in grapevine dieback in China. Fungal Divers. 2013, 61, 221–236. [Google Scholar] [CrossRef]
- Pitt, W.M.; Huang, R.; Steel, C.C.; Savocchia, S. Identification, distribution and current taxonomy of Botryosphaeriaceae species associated with grapevine decline in New South Wales and South Australia. Aust. J. Grape Wine Res. 2010, 16, 258–271. [Google Scholar] [CrossRef]
- Phillips, A.J.L. Botryosphaeria species associated with diseases of grapevines in Portugal. Phytopathol. Mediterr. 2002, 41, 3–18. [Google Scholar] [CrossRef]
- Mohammadi, H.; Gramaje, D.; Banihashemi, Z.; Armengol, J. Characterization of Diplodia seriata and Neofusicoccum parvum associated with grapevine decline in Iran. J. Agric. Sci. Technol. 2013, 15, 603–616. [Google Scholar]
- Úrbez-Torres, J.R.; Leavitt, G.M.; Guerrero, J.C.; Guevara, J.; Gubler, W.D. Identification and Pathogenicity of Lasiodiplodia theobromae and Diplodia seriata, the Causal Agents of Bot Canker Disease of Grapevines in Mexico. Plant Dis. 2008, 92, 519–529. [Google Scholar] [CrossRef]
- Kovács, C.; Balling, P.; Bihari, Z.; Nagy, A.; Sándor, E. Incidence of grapevine trunk diseases is influenced by soil, topology and vineyard age, but not by Diplodia seriata infection rate in the Tokaj Wine Region, Hungary. Phytoparasitica 2017, 45, 21–32. [Google Scholar] [CrossRef]
- Chebil, S.; Fersi, R.; Bouzid, M.; Quaglino, F.; Chenenaoui, S.; Melki, I.; Durante, G.; Zacchi, E.; Bahri, B.A.; Bianco, P.A.; et al. Fungi from the Diaporthaceae and Botryosphaeriaceae families associated with grapevine decline in Tunisia. Cienc. E Investig. Agrar. 2017, 44, 127–138. [Google Scholar] [CrossRef]
- Fourie, P.H.; Halleen, F. Occurrence of grapevine trunk disease pathogens in rootstock mother plants in South Africa. Australas. Plant Pathol. 2004, 33, 313–315. [Google Scholar] [CrossRef]
- Aroca, A.; García-Figueres, F.; Bracamonte, L.; Luque, J.; Raposo, R. A Survey of Trunk Disease Pathogens within Rootstocks of Grapevines in Spain. Eur. J. Plant Pathol. 2006, 115, 195–202. [Google Scholar] [CrossRef]
- Eichmeier, A.; Pečenka, J.; Peňázová, E.; Baránek, M.; Català-García, S.; León, M.; Armengol, J.; Gramaje, D. High-throughput amplicon sequencing-based analysis of active fungal communities inhabiting grapevine after hot-water treatments reveals unexpectedly high fungal diversity. Fungal Ecol. 2018, 36, 26–38. [Google Scholar] [CrossRef]
- Baránek, M.; Armengol, J.; Holleinová, V.; Pečenka, J.; Calzarano, F.; Peňázová, E.; Vachůn, M.; Eichmeier, A. Incidence of symptoms and fungal pathogens associated with grapevine trunk diseases in Czech vineyards: First example from a north-eastern European grape-growing region. Phytopathol. Mediterr. 2018, 57, 449–458. [Google Scholar] [CrossRef]
- Pečenka, J.; Eichmeier, A.; Peňázová, E.; Baránek, M.; León, M.; Armengol, J. First Report of Dactylonectria torresensis Causing Black-Foot Disease on Grapevines in the Czech Republic. Plant Dis. 2018, 102, 2038. [Google Scholar] [CrossRef]
- Mondragón, F.A.; Rodríguez-Alvarado, G.; Gmez-Dorantes, N.; Guerra-Santos, J.J.; Fernández-Pavía, S.P. Botryosphaeriaceae: A complex, diverse and cosmopolitan family of fungi. Rev. Mex. Cienc. Agrícolas 2021, 12, 643–654. [Google Scholar] [CrossRef]
- Spetik, M.; Cechova, J.; Eichmeier, A. First report of Neofusicoccum parvum causing stem blight and dieback of highbush blueberry in the Czech Republic. Plant Dis. 2023. [Google Scholar] [CrossRef] [PubMed]
- Pečenka, J.; Tekielska, D.A.; Kocanová, M.; Peňázová, E.; Berraf-Tebbal, A.; Eichmeier, A. First Report of Lasiodiplodia theobromae Causing Decline of Blueberry (Vaccinium corymbosum) in the Czech Republic. Plant Dis. 2020, 105, 215. [Google Scholar] [CrossRef] [PubMed]
- Elena, G.; DI Bella, V.; Armengol, J.; Luque, J. Viability of Botryosphaeriaceae species pathogenic to grapevine after hot water treatment. Phytopathol. Mediterr. 2015, 54, 325–334. [Google Scholar] [CrossRef]
- Štůsková, K.; Pečenka, J.; Tekielska, D.A.; Špetík, M.; Bytešníková, Z.; Švec, P.; Ondreáš, F.; Ridošková, A.; Richtera, L.; Adam, V.; et al. The in vitro effects of selected substances and nanomaterials against Diaporthe eres, Diplodia seriata and Eutypa lata. Ann. Appl. Biol. 2023, 182, 226–237. [Google Scholar] [CrossRef]
- Špetík, M.; Balík, J.; Híc, P.; Hakalová, E.; Štůsková, K.; Frejlichová, L.; Tříska, J.; Eichmeier, A. Lignans Extract from Knotwood of Norway Spruce—A Possible New Weapon against GTDs. J. Fungi 2022, 8, 357. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).