2S-BUSGAN: A Novel Generative Adversarial Network for Realistic Breast Ultrasound Image with Corresponding Tumor Contour Based on Small Datasets
Abstract
:1. Introduction
- Image quality: BUS images exhibit diverse and random morphological features of tumors and texture features of surrounding tissues, leading to unrealistic details and a lack of structural legitimacy in the generated images.
- Data constraints: The limited size of the BUS dataset affects both the performance of deep learning models and GANs. Since GANs rely on the underlying data distribution, the small dataset needs to accurately represent the true data distribution, resulting in insufficient diversity and limited information in the generated results.
- Application limitations: The current GAN augmentation methods have a significant constraint when applied to different tasks. They are typically tailored for specific tasks, such as classification or segmentation. However, if these augmented data need to be utilized for other tasks, the data must be relabeled, or a new GAN model must be trained, leading to increased costs. Consequently, a universal generation-based augmentation method is meaningful.
2. Materials and Methods
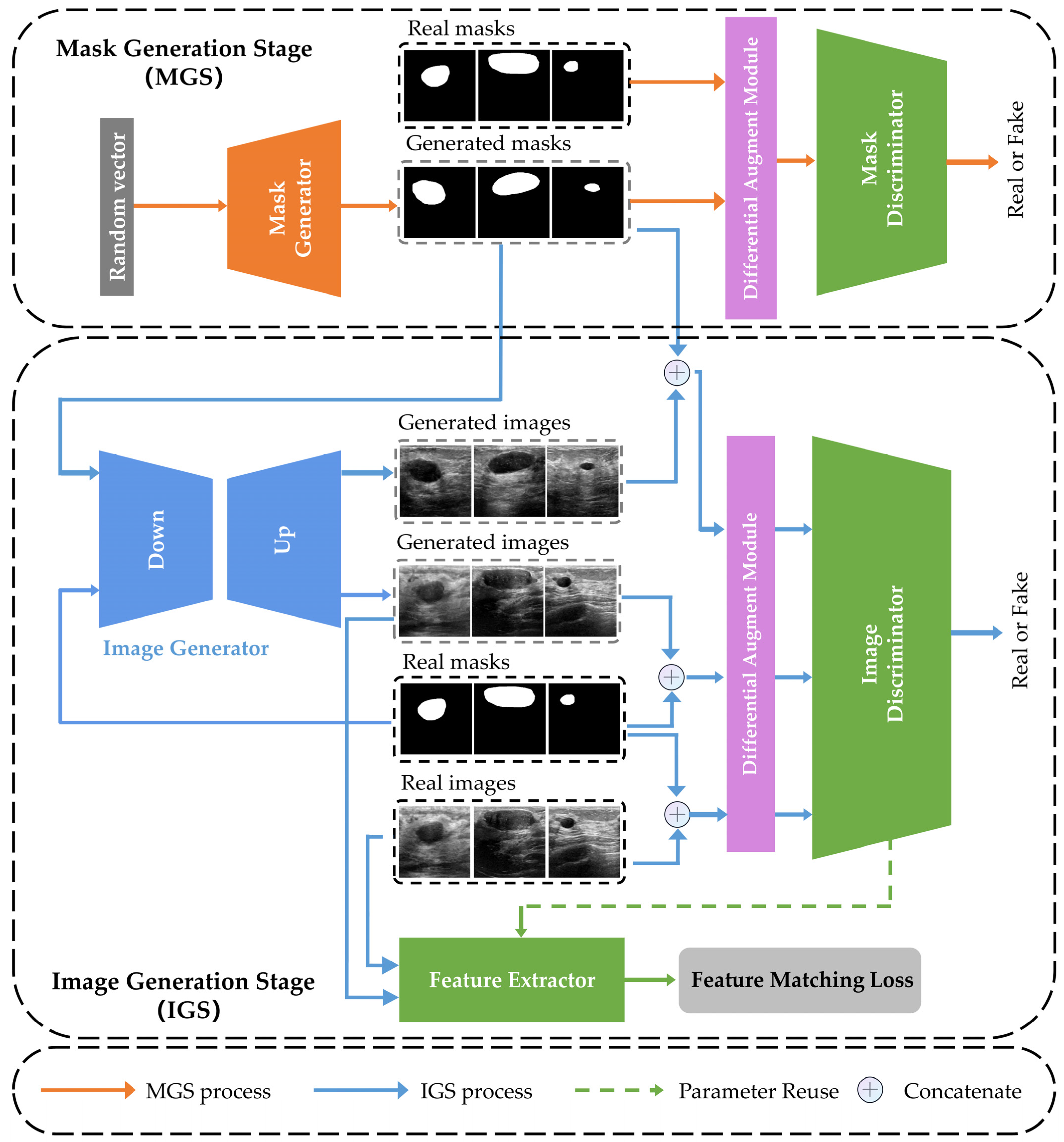
2.1. Overview of 2s-BUSGAN
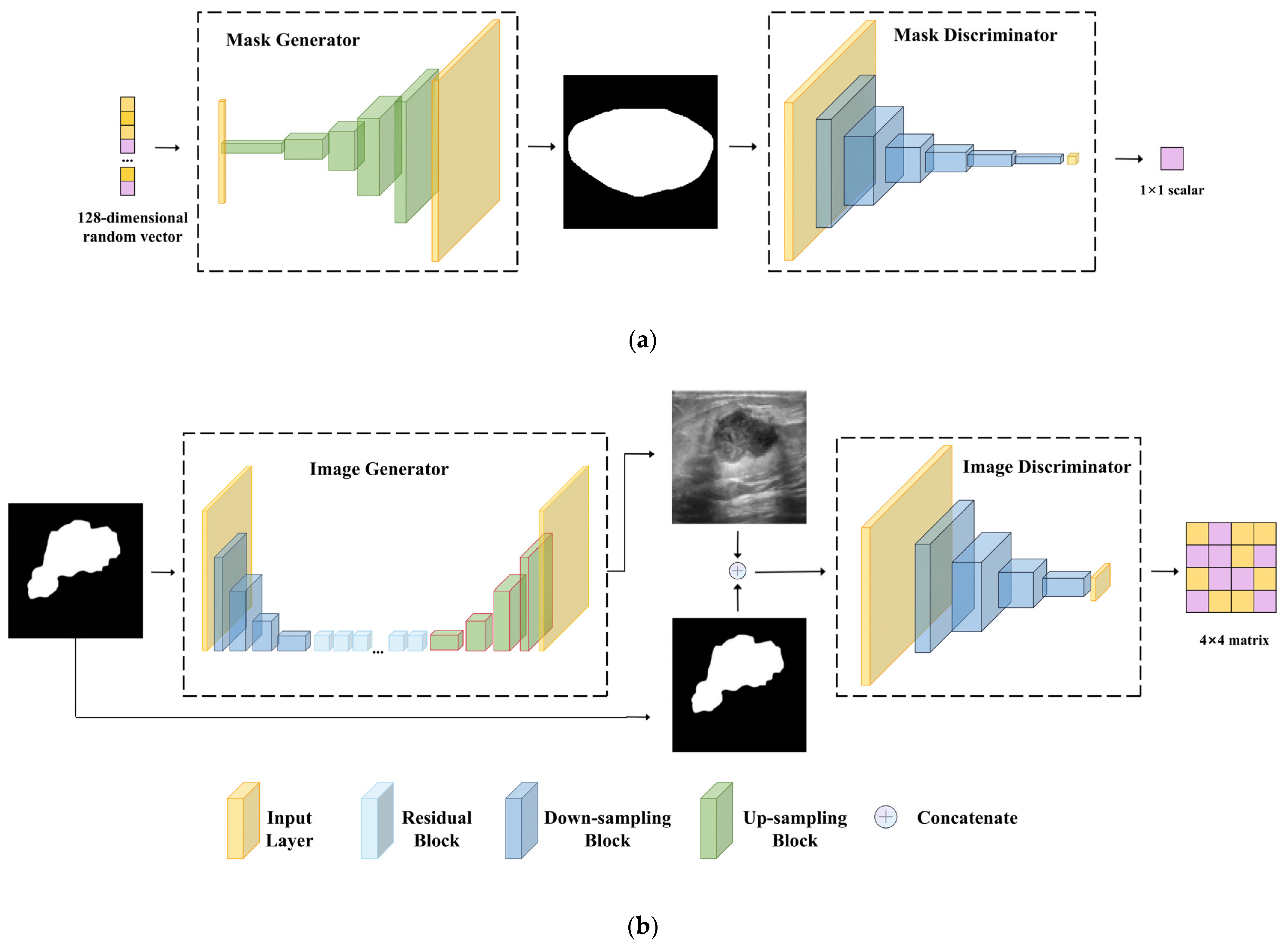
2.2. Mask Generation Stage (MGS)
2.3. Image Generation Stage (IGS)
2.4. Feature Matching Loss (FML)
2.5. Differentiable Augmentation Module (DAM)
3. Experiments
3.1. Datasets
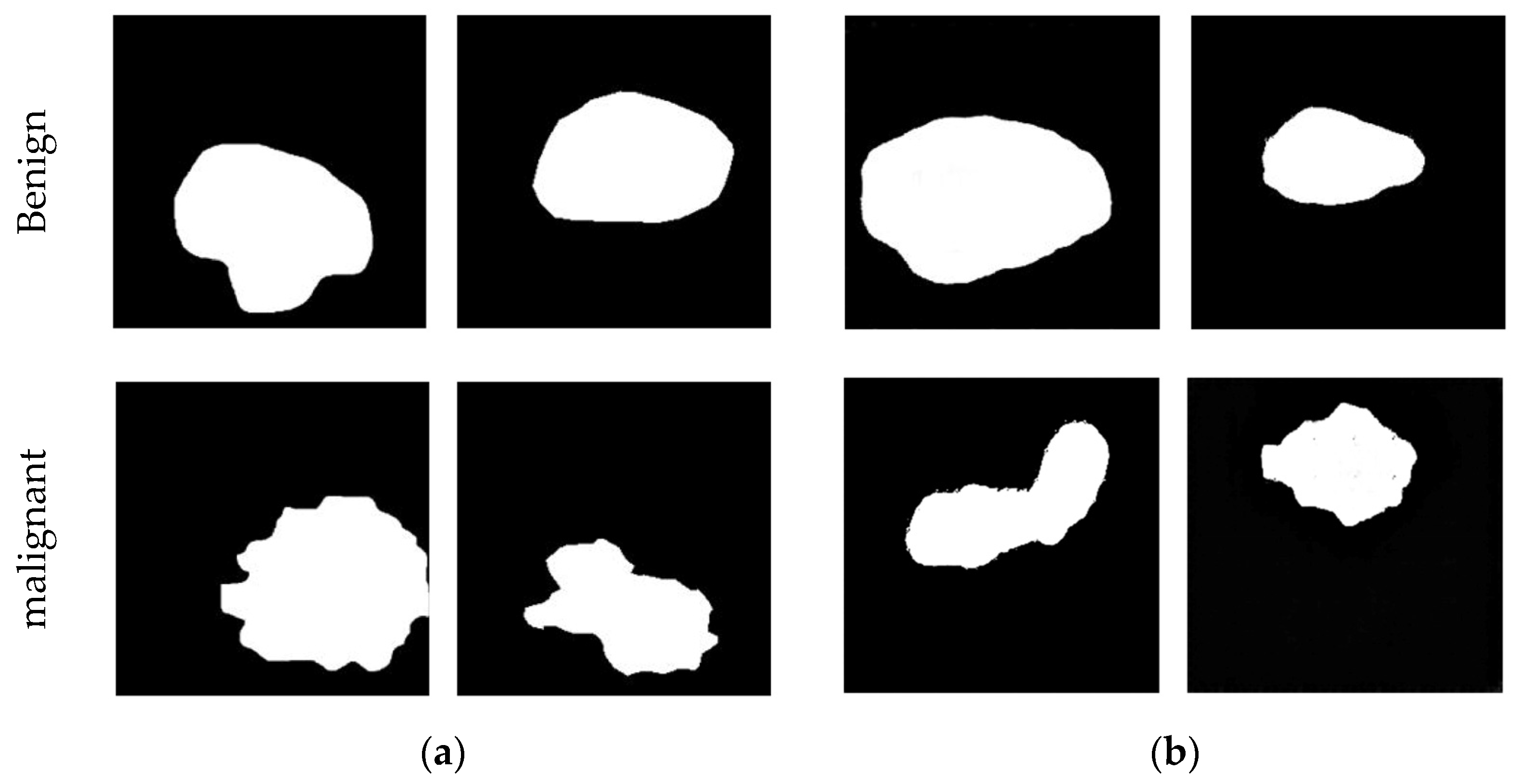
- BUSI [19,35]. This dataset was curated by the National Cancer Center of Cairo University in Egypt in 2018. The dataset consists of 780 PNG images obtained using the LOGIQ E9 and LOGIQ E9 agile ultrasound systems. These images were acquired from 600 female patients aged between 25 and 75. Among the images, 487 represent benign tumors, 210 represent malignant tumors, and 133 contain normal breast tissue. The labels for benign and malignant tumors were determined based on pathological examination results from biopsy samples. The average image size is 500 × 500 pixels, and each case includes the original ultrasound image data and the corresponding breast tumor boundary image annotated by an expert imaging diagnostic physician. The annotated tumor contour images serve as the gold standard for segmentation in the experiments, providing a reference for the training process and evaluation of results.
- Collected. The China-Japan Friendship Hospital in Beijing collected and organized the dataset in 2018. Before data collection, informed written consent was obtained from each patient after a comprehensive explanation of the study’s procedures, purpose, and nature. Multiple acquisition systems were used to capture the images in this dataset. It comprises a total of 920 images, including 470 benign cases and 450 malignant cases. The benign or malignant classification labels are derived from pathological examinations based on puncture biopsy results. A professional imaging diagnostic physician has annotated each data case. Furthermore, the dataset includes notable misdiagnoses and missed diagnosis cases, providing valuable examples for analysis and evaluation.
3.2. Implementation Details
3.3. Generation Experiments
3.3.1. MGS Mask Synthesis
3.3.2. IGS Image Synthesis
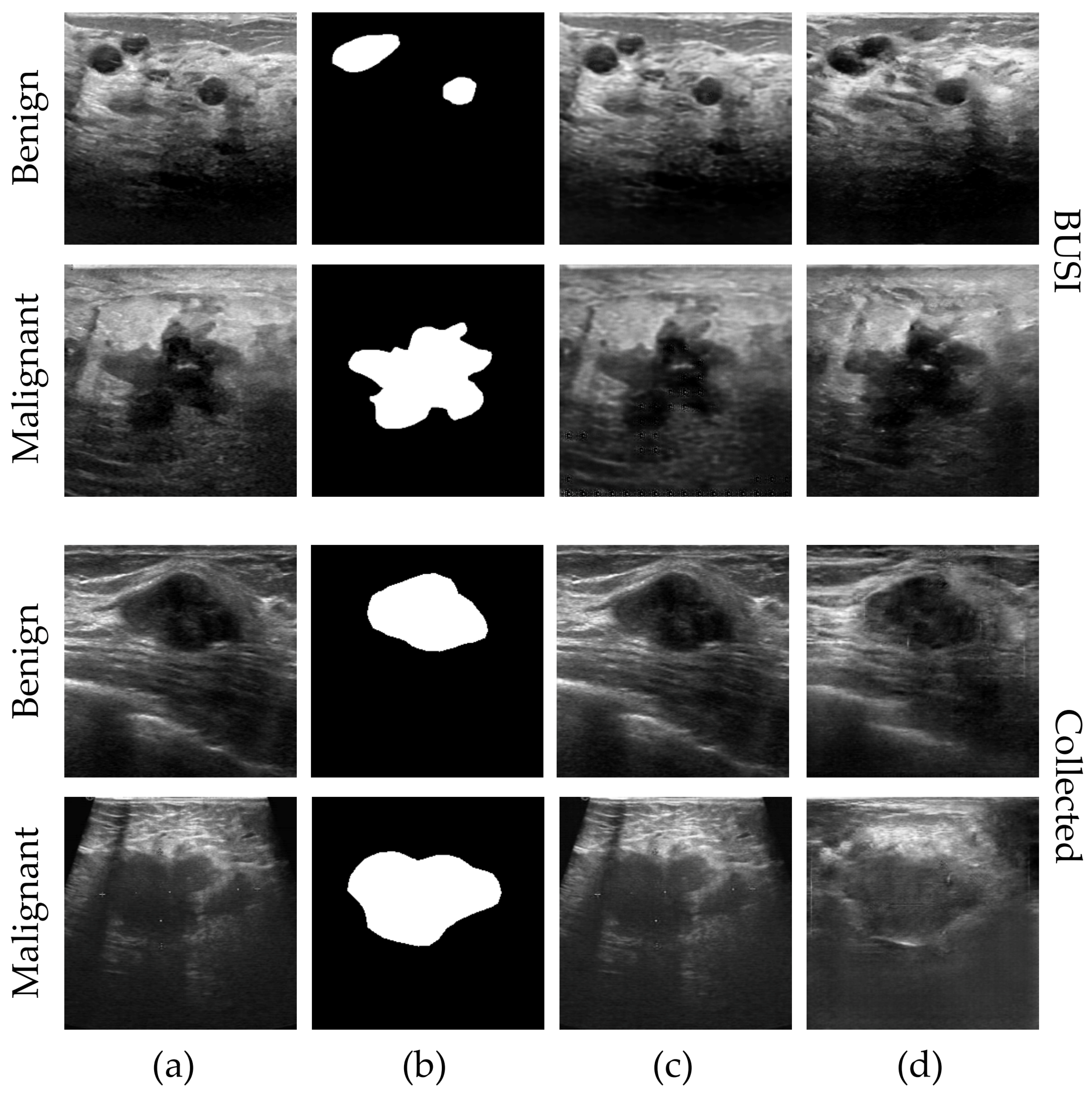
3.3.3. 2s-BUSGAN Image Synthesis
3.4. Evaluation by Doctors
3.5. Augmentation Experiments for BUS Segmentation and Classification
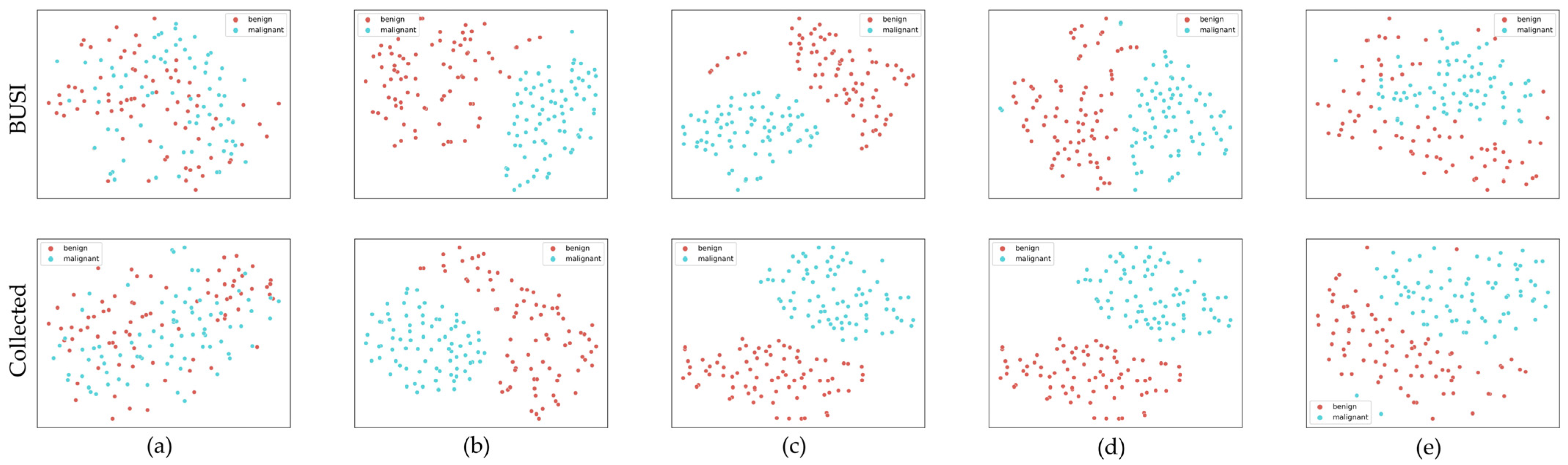
3.5.1. Classification Experiments
3.5.2. Segmentation Experiments
4. Results
4.1. Generation Experiments Results
4.1.1. MGS Mask Synthesis Results
4.1.2. IGS Image Synthesis Results
4.1.3. 2s-BUSGAN Image Synthesis Results
4.2. Results of Evaluation by Doctors
4.3. Results of Augmentation Experiments for BUS Segmentation and Classification
4.3.1. Classification Experiments Results
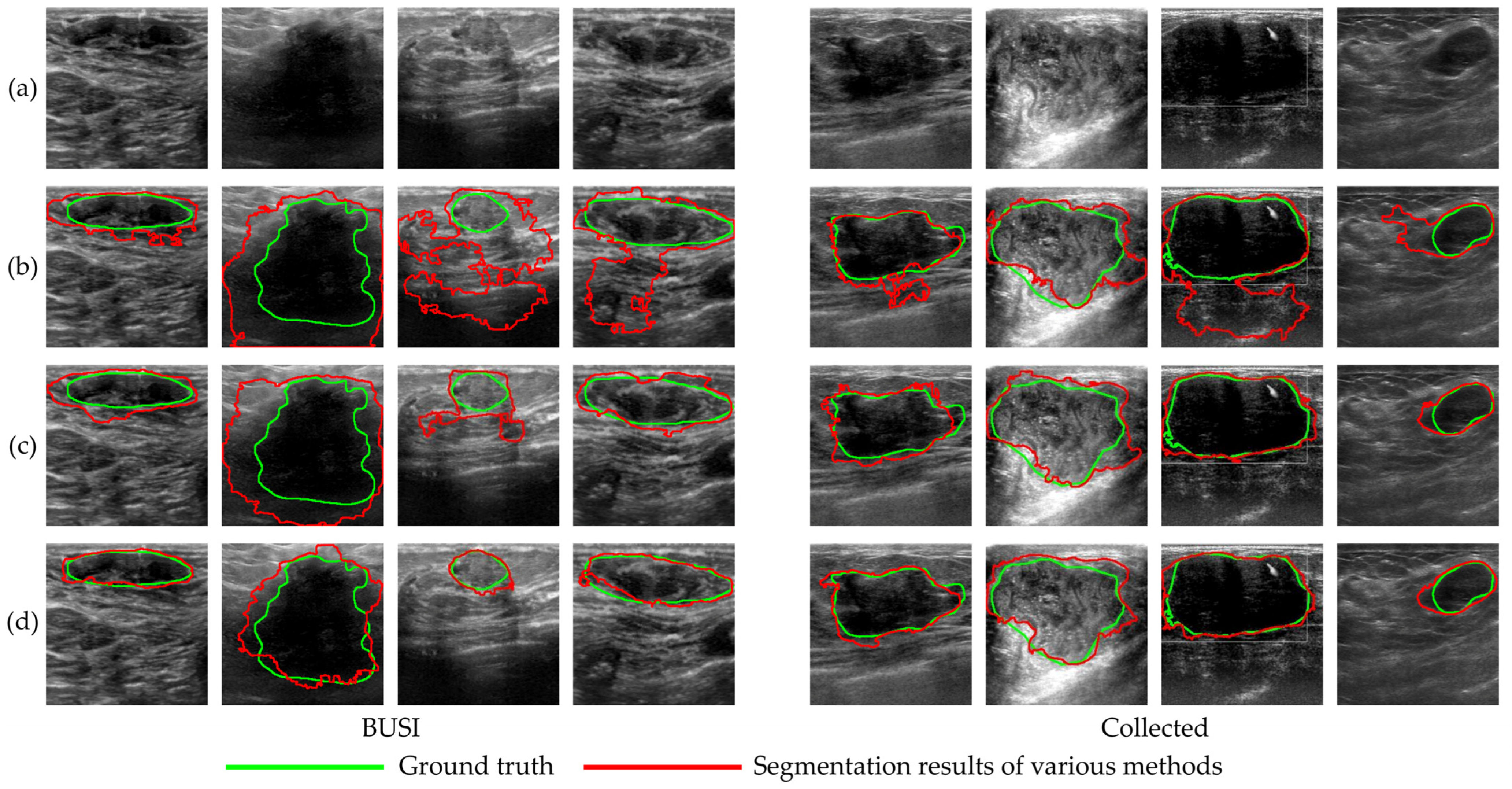
4.3.2. Segmentation Experiments Results
5. Discussion
5.1. Generation Experiments Discussion
5.1.1. MGS Mask Synthesis Discussion
5.1.2. IGS Image Synthesis Discussion
5.1.3. 2s-BUSGAN Image Synthesis Discussion
5.2. Discussion of Evaluation by Doctors
5.3. Discussion of Augmentation Experiments for BUS Segmentation and Classification
5.3.1. Classification Experiments Discussion
5.3.2. Segmentation Experiments Discussion
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Xia, C.; Dong, X.; Li, H.; Cao, M.; Sun, D.; He, S.; Yang, F.; Yan, X.; Zhang, S.; Li, N.; et al. Cancer statistics in China and United States, 2022: Profiles, trends, and determinants. Chin. Med. J. 2022, 135, 584–590. [Google Scholar] [CrossRef] [PubMed]
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Zhai, D.; Hu, B.; Gong, X.; Zou, H.; Luo, J. ASS-GAN: Asymmetric semi-supervised GAN for breast ultrasound image segmentation. Neurocomputing 2022, 493, 204–216. [Google Scholar] [CrossRef]
- Chugh, G.; Kumar, S.; Singh, N. Survey on Machine Learning and Deep Learning Applications in Breast Cancer Diagnosis. Cogn. Comput. 2021, 13, 1451–1470. [Google Scholar] [CrossRef]
- Ilesanmi, A.E.; Chaumrattanakul, U.; Makhanov, S.S. A method for segmentation of tumors in breast ultrasound images using the variant enhanced deep learning. Biocybern. Biomed. Eng. 2021, 41, 802–818. [Google Scholar] [CrossRef]
- Yap, M.H.; Goyal, M.; Osman, F.; Marti, R.; Denton, E.; Juette, A.; Zwiggelaar, R. Breast ultrasound region of interest detection and lesion localisation. Artif. Intell. Med. 2020, 107, 101880. [Google Scholar] [CrossRef]
- Haibo, H.; Garcia, E.A. Learning from Imbalanced Data. IEEE Trans. Knowl. Data Eng. 2009, 21, 1263–1284. [Google Scholar] [CrossRef]
- Krawczyk, B.; Galar, M.; Jeleń, Ł.; Herrera, F. Evolutionary undersampling boosting for imbalanced classification of breast cancer malignancy. Appl. Soft Comput. 2016, 38, 714–726. [Google Scholar] [CrossRef]
- Pang, T.; Wong, J.H.D.; Ng, W.L.; Chan, C.S. Semi-supervised GAN-based Radiomics Model for Data Augmentation in Breast Ultrasound Mass Classification. Comput. Methods Programs Biomed. 2021, 203, 106018. [Google Scholar] [CrossRef]
- Frid-Adar, M.; Diamant, I.; Klang, E.; Amitai, M.; Goldberger, J.; Greenspan, H. GAN-based synthetic medical image augmentation for increased CNN performance in liver lesion classification. Neurocomputing 2018, 321, 321–331. [Google Scholar] [CrossRef]
- Chawla, N.V.; Bowyer, K.W.; Hall, L.O.; Kegelmeyer, W.P. SMOTE: Synthetic minority over-sampling technique. J. Artif. Intell. Res. 2002, 16, 321–357. [Google Scholar] [CrossRef]
- Inoue, H.J. Data augmentation by pairing samples for images classification. arXiv 2018, arXiv:1801.02929. [Google Scholar]
- Goodfellow, I.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y.J.C. Generative adversarial networks. Commun. ACM 2020, 63, 139–144. [Google Scholar] [CrossRef]
- Radford, A.; Metz, L.; Chintala, S.J. Unsupervised representation learning with deep convolutional generative adversarial networks. arXiv 2015, arXiv:1511.06434. [Google Scholar]
- Mao, X.; Li, Q.; Xie, H.; Lau, R.Y.; Wang, Z.; Paul Smolley, S. Least squares generative adversarial networks. In Proceedings of the IEEE International Conference on Computer Vision, Venice, Italy, 22–29 October 2017; pp. 2794–2802. [Google Scholar]
- Arjovsky, M.; Chintala, S.; Bottou, L. Wasserstein Generative Adversarial Networks. In Proceedings of the 34th International Conference on Machine Learning, Sydney, Australia, 11 August 2017; Doina, P., Yee Whye, T., Eds.; Proceedings of Machine Learning Research (PMLR): Ann Arbor, MI, USA, 2017; Volume 70, pp. 214–223. [Google Scholar]
- Han, L.; Huang, Y.; Dou, H.; Wang, S.; Ahamad, S.; Luo, H.; Liu, Q.; Fan, J.; Zhang, J. Semi-supervised segmentation of lesion from breast ultrasound images with attentional generative adversarial network. Comput. Methods Programs Biomed. 2020, 189, 105275. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Lu, W.; Wei, X.; Zhu, J.; Jiang, H.; Liu, Z.; Gao, J.; Li, X.; Yu, J.; Yu, M.; et al. A Progressive Generative Adversarial Method for Structurally Inadequate Medical Image Data Augmentation. IEEE J. Biomed. Health Inform. 2022, 26, 7–16. [Google Scholar] [CrossRef]
- Al-Dhabyani, W.; Gomaa, M.; Khaled, H.; Fahmy, A. Deep Learning Approaches for Data Augmentation and Classification of Breast Masses using Ultrasound Images. Int. J. Adv. Comput. Sci. Appl. 2019, 10, 618–627. [Google Scholar] [CrossRef]
- Saha, S.; Sheikh, N. Ultrasound image classification using ACGAN with small training dataset. In Proceedings of the Recent Trends in Signal and Image Processing: ISSIP 2020, Kolkata, India, 18–19 March 2020; Springer: Berlin/Heidelberg, Germany, 2021; pp. 85–93. [Google Scholar]
- Odena, A.; Olah, C.; Shlens, J. Conditional image synthesis with auxiliary classifier gans. In Proceedings of the International Conference on Machine Learning, Sydney, Australia, 6–11 August 2017; PMLR: Ann Arbor, MI, USA, 2017; pp. 2642–2651. [Google Scholar]
- Zhao, S.; Liu, Z.; Lin, J.; Zhu, J.-Y.; Han, S.J.A. Differentiable augmentation for data-efficient gan training. Adv. Neural Inf. Process. Syst. 2020, 33, 7559–7570. [Google Scholar]
- Ioffe, S.; Szegedy, C. Batch normalization: Accelerating deep network training by reducing internal covariate shift. In Proceedings of the International Conference on Machine Learning, Lille, France, 6–11 July 2015; PMLR: Ann Arbor, MI, USA, 2015; pp. 448–456. [Google Scholar]
- Glorot, X.; Bordes, A.; Bengio, Y. Deep sparse rectifier neural networks. In Proceedings of the Fourteenth International Conference on Artificial Intelligence and Statistics, Fort Lauderdale, FL, USA, 11–13 April 2011; JMLR Workshop and Conference Proceedings. PMLR: Ann Arbor, MI, USA, 2011; pp. 315–323. [Google Scholar]
- Miyato, T.; Kataoka, T.; Koyama, M.; Yoshida, Y.J. Spectral normalization for generative adversarial networks. arXiv 2018, arXiv:1802.05957. [Google Scholar]
- Xu, B.; Wang, N.; Chen, T.; Li, M.J. Empirical evaluation of rectified activations in convolutional network. arXiv 2015, arXiv:1505.00853. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Ulyanov, D.; Vedaldi, A.; Lempitsky, V.J. Instance normalization: The missing ingredient for fast stylization. arXiv 2016, arXiv:1607.08022. [Google Scholar]
- Isola, P.; Zhu, J.-Y.; Zhou, T.; Efros, A.A. Image-to-Image Translation with Conditional Adversarial Networks. In Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017; pp. 5967–5976. [Google Scholar]
- Zhang, R.; Isola, P.; Efros, A.A.; Shechtman, E.; Wang, O. The unreasonable effectiveness of deep features as a perceptual metric. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–22 June 2018; pp. 586–595. [Google Scholar]
- Dosovitskiy, A.; Brox, T.J.A. Generating images with perceptual similarity metrics based on deep networks. Adv. Neural Inf. Process. Syst. 2016, 29. [Google Scholar] [CrossRef]
- Gatys, L.A.; Ecker, A.S.; Bethge, M. Image style transfer using convolutional neural networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 2414–2423. [Google Scholar]
- Johnson, J.; Alahi, A.; Li, F.-F. Perceptual losses for real-time style transfer and super-resolution. In Proceedings of the Computer Vision–ECCV 2016: 14th European Conference, Amsterdam, The Netherlands, 11–14 October 2016; Proceedings, Part II 14. Springer: Berlin/Heidelberg, Germany, 2016; pp. 694–711. [Google Scholar]
- Ledig, C.; Theis, L.; Huszár, F.; Caballero, J.; Cunningham, A.; Acosta, A.; Aitken, A.; Tejani, A.; Totz, J.; Wang, Z. Photo-realistic single image super-resolution using a generative adversarial network. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 4681–4690. [Google Scholar]
- Al-Dhabyani, W.; Gomaa, M.; Khaled, H.; Fahmy, A. Dataset of breast ultrasound images. Data Brief 2020, 28, 104863. [Google Scholar] [CrossRef] [PubMed]
- Kingma, D.P.; Ba, J.J. Adam: A method for stochastic optimization. arXiv 2014, arXiv:1412.6980. [Google Scholar]
- Salimans, T.; Goodfellow, I.; Zaremba, W.; Cheung, V.; Radford, A.; Chen, X.J.A. Improved techniques for training gans. Adv. Neural Inf. Process. Syst. 2016, 29. [Google Scholar] [CrossRef]
- Borji, A.J.C.V.; Understanding, I. Pros and cons of gan evaluation measures. Comput. Vis. Image Underst. 2019, 179, 41–65. [Google Scholar] [CrossRef]
- Wang, Z.; Simoncelli, E.P.; Bovik, A.C. Multiscale structural similarity for image quality assessment. In Proceedings of the Thrity-Seventh Asilomar Conference on Signals, Systems & Computers, Pacific Grove, CA, USA, 9–12 November 2003; pp. 1398–1402. [Google Scholar]
- Han, T.; Nebelung, S.; Haarburger, C.; Horst, N.; Reinartz, S.; Merhof, D.; Kiessling, F.; Schulz, V.; Truhn, D.J.S. Breaking medical data sharing boundaries by using synthesized radiographs. Sci. Adv. 2020, 6, eabb7973. [Google Scholar] [CrossRef]
- Van der Maaten, L.; Hinton, G.J.J. Visualizing data using t-SNE. J. Mach. Learn. Res. 2008, 9, 2579–2605. [Google Scholar]
- Tang, Z.; Gao, Y.; Karlinsky, L.; Sattigeri, P.; Feris, R.; Metaxas, D. OnlineAugment: Online data augmentation with less domain knowledge. In Proceedings of the Computer Vision–ECCV 2020: 16th European Conference, Glasgow, UK, 23–28 August 2020; Proceedings, Part VII 16. Springer: Berlin/Heidelberg, Germany, 2020; pp. 313–329. [Google Scholar]
- Simonyan, K.; Zisserman, A.J. Very deep convolutional networks for large-scale image recognition. arXiv 2014, arXiv:1409.1556. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Proceedings of the Medical Image Computing and Computer-Assisted Intervention–MICCAI 2015: 18th International Conference, Munich, Germany, 5–9 October 2015; Proceedings, Part III 18. Springer: Berlin/Heidelberg, Germany, 2015; pp. 234–241. [Google Scholar]






| Dataset | Setting | Benign | Malignant | ||||
|---|---|---|---|---|---|---|---|
| FID ↓ | KID × 100 ↓ | MS-SSIM ↓ | FID ↓ | KID × 100 ↓ | MS-SSIM ↓ | ||
| Collected | IGS w/o FML | 167.1353 | 158.602 | 0.222 | 419.3311 | 580.4763 | 0.223 |
| IGS w/o DAM | 142.3543 | 116.565 | 0.1422 | 188.7287 | 187.7055 | 0.1307 | |
| IGS | 89.1111 | 43.5631 | 0.1286 | 93.6413 | 48.6419 | 0.1285 | |
| BUSI | IGS w/o FML | 208.5485 | 203.7132 | 0.2066 | 189.5309 | 184.7761 | 0.2618 |
| IGS w/o DAM | 153.3376 | 130.2609 | 0.177 | 178.1268 | 179.8315 | 0.2843 | |
| IGS | 72.0245 | 27.7052 | 0.1509 | 101.0019 | 62.3815 | 0.2089 | |
| Dataset | Setting | Benign | Malignant | ||||
|---|---|---|---|---|---|---|---|
| FID ↓ | KID × 100 ↓ | MS-SSIM ↓ | FID ↓ | KID × 100 ↓ | MS-SSIM ↓ | ||
| Collected | DCGAN | 101.3506 | 60.6894 | 0.209 | 96.2566 | 63.4413 | 0.1736 |
| WGAN | 186.4802 | 176.8394 | 0.3502 | 160.9212 | 144.1009 | 0.2556 | |
| LSGAN | 76.8730 | 41.055 | 0.2201 | 70.7381 | 33.7014 | 0.1908 | |
| 2s-BUSGAN | 89.1111 | 43.5631 | 0.1422 | 93.6413 | 48.6419 | 0.1307 | |
| BUSI | DCGAN | 62.2625 | 32.8401 | 0.1957 | 128.2199 | 108.9818 | 0.3507 |
| WGAN | 120.3191 | 109.0968 | 0.1727 | 135.7969 | 119.5491 | 0.3150 | |
| LSGAN | 72.7607 | 38.9978 | 0.2074 | 92.5337 | 57.3108 | 0.3028 | |
| 2s-BUSGAN | 72.0245 | 27.7052 | 0.1509 | 101.0019 | 62.3815 | 0.2089 | |
| Collected | BUSI | |||||
|---|---|---|---|---|---|---|
| GT | LSGAN | 2s-BUSGAN | GT | LSGAN | 2s-BUSGAN | |
| Doctor 1 | 0.75 | 0.60 | 0.57 | 0.85 | 0.47 | 0.47 |
| Doctor 2 | 0.80 | 0.67 | 0.63 | 0.88 | 0.47 | 0.37 |
| Doctor 3 | 0.70 | 0.67 | 0.60 | 0.83 | 0.53 | 043 |
| All doctors | 0.75 | 0.65 | 0.60 | 0.85 | 0.49 | 0.42 |
| Collected | BUSI | |||||
|---|---|---|---|---|---|---|
| GT | LSGAN | 2s-BUSGAN | GT | LSGAN | 2s-BUSGAN | |
| Doctor 1 | 0.70 | 0.53 | 0.57 | 0.83 | 1.00 | 0.83 |
| Doctor 2 | 0.63 | 0.57 | 0.63 | 0.82 | 0.93 | 0.73 |
| Doctor 3 | 0.65 | 0.50 | 0.63 | 0.85 | 0.90 | 0.8 |
| All doctors | 0.66 | 0.53 | 0.61 | 0.83 | 0.92 | 0.79 |
| Dataset | Setting | Precision | Recall | f1-Score | Accuracy |
|---|---|---|---|---|---|
| Collected | Baseline | 0.63 ± 0.01 | 0.627 ± 0.006 | 0.623 ± 0.006 | 0.624 ± 0.007 |
| TA | 0.64 ± 0.01 | 0.637 ± 0.012 | 0.637 ± 0.012 | 0.636 ± 0.010 | |
| TA + LSGAN | 0.66 ± 0.00 | 0.657 ± 0.006 | 0.667 ± 0.006 | 0.661 ± 0.005 | |
| TA + 2s-BUSGAN | 0.69 ± 0.026 | 0.69 ± 0.026 | 0.687 ± 0.021 | 0.688 ± 0.025 | |
| BUSI | Baseline | 0.82 ± 0.017 | 0.817 ± 0.021 | 0.813 ± 0.023 | 0.816 ± 0.019 |
| TA | 0.833 ± 0.012 | 0.833 ± 0.012 | 0.833 ± 0.012 | 0.836 ± 0.013 | |
| TA + LSGAN | 0.84 ± 0.00 | 0.823 ± 0.001 | 0.823 ± 0.006 | 0.821 ± 0.009 | |
| TA + 2s-BUSGAN | 0.857 ± 0.025 | 0.847 ± 0.015 | 0.847 ± 0.015 | 0.846 ± 0.015 |
| Setting | BUSI | Collected | ||
|---|---|---|---|---|
| Benign | Malignant | Benign | Malignant | |
| Baseline | 0.7332 | 0.7354 | 0.6468 | 0.7021 |
| TA | 0.7422 | 0.7395 | 0.6588 | 0.7215 |
| 2s-BUSGAN + TA | 0.7591 | 0.7565 | 0.6786 | 0.7316 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Luo, J.; Zhang, H.; Zhuang, Y.; Han, L.; Chen, K.; Hua, Z.; Li, C.; Lin, J. 2S-BUSGAN: A Novel Generative Adversarial Network for Realistic Breast Ultrasound Image with Corresponding Tumor Contour Based on Small Datasets. Sensors 2023, 23, 8614. https://doi.org/10.3390/s23208614
Luo J, Zhang H, Zhuang Y, Han L, Chen K, Hua Z, Li C, Lin J. 2S-BUSGAN: A Novel Generative Adversarial Network for Realistic Breast Ultrasound Image with Corresponding Tumor Contour Based on Small Datasets. Sensors. 2023; 23(20):8614. https://doi.org/10.3390/s23208614
Chicago/Turabian StyleLuo, Jie, Heqing Zhang, Yan Zhuang, Lin Han, Ke Chen, Zhan Hua, Cheng Li, and Jiangli Lin. 2023. "2S-BUSGAN: A Novel Generative Adversarial Network for Realistic Breast Ultrasound Image with Corresponding Tumor Contour Based on Small Datasets" Sensors 23, no. 20: 8614. https://doi.org/10.3390/s23208614
APA StyleLuo, J., Zhang, H., Zhuang, Y., Han, L., Chen, K., Hua, Z., Li, C., & Lin, J. (2023). 2S-BUSGAN: A Novel Generative Adversarial Network for Realistic Breast Ultrasound Image with Corresponding Tumor Contour Based on Small Datasets. Sensors, 23(20), 8614. https://doi.org/10.3390/s23208614





