Exploring the Role of the Processing Body in Plant Abiotic Stress Response
Abstract
:1. Introduction
2. Components of P-Body
2.1. The Conserved Protein Components of P-Bodies in Plants
2.2. The Dynamic Protein Components of P-Body in Plants
3. Assembly of P-Body
3.1. Liquid–Liquid Phase Separation (LLPS) Drives P-Body Assembly
3.2. P-Body Specific Sequence and Mechanism of Assembly
3.3. RNA Decay Is the Main Reason for the Formation of P-Body
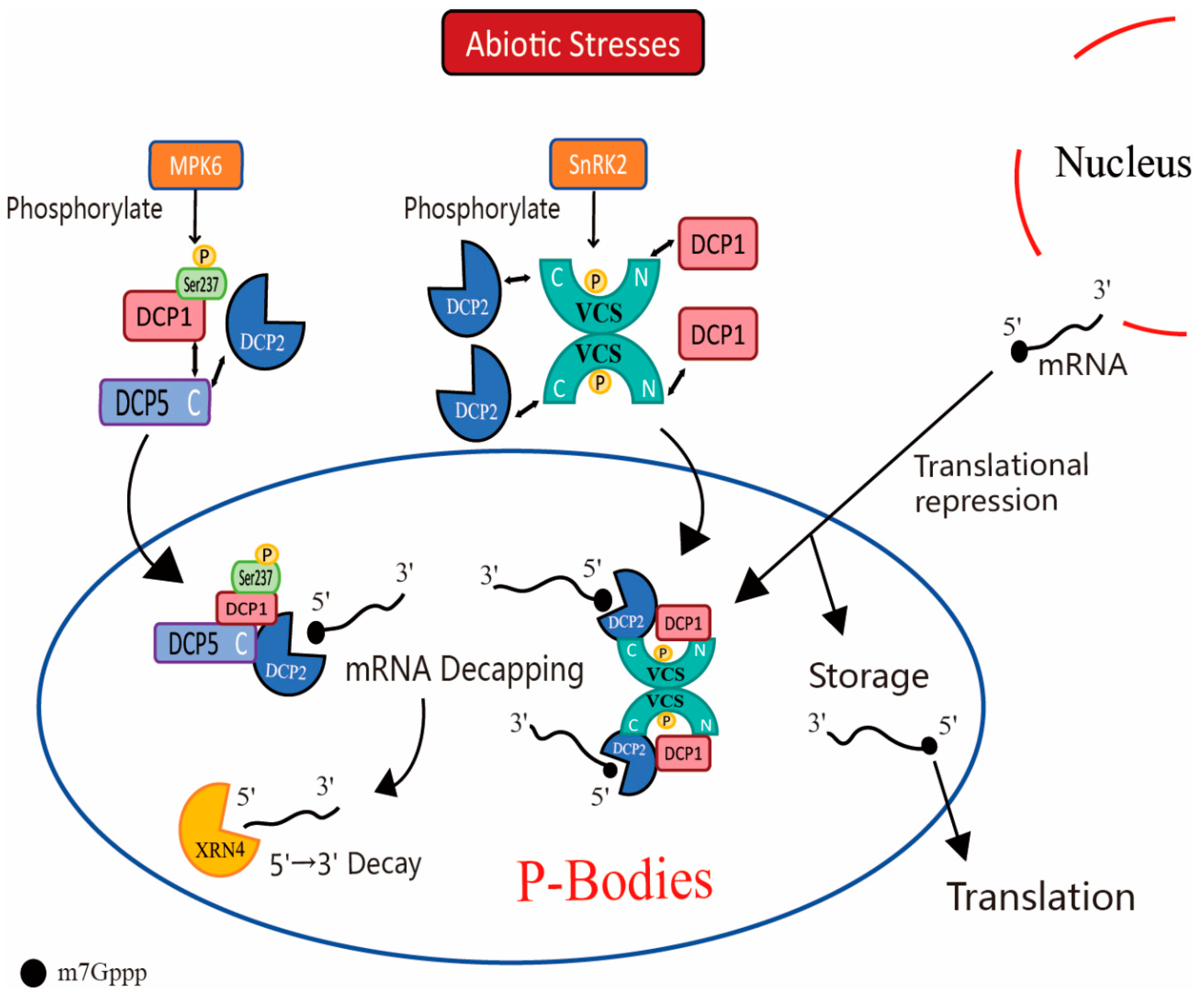
4. P-Body and Plant Stress Responses
4.1. Response of P-Body to Drought and Salt Stress
4.2. Response of P-Body to Cold or Heat Stress
4.3. P-Body Involving in Ethylene Signaling Pathway
5. Conclusions and Perspectives
Author Contributions
Funding
Conflicts of Interest
References
- Alshareedah, I.; Moosa, M.M.; Raju, M.; Potoyan, D.A.; Banerjee, P.R. Phase transition of RNA−protein complexes into ordered hollow condensates. Proc. Natl. Acad. Sci. USA 2020, 117, 15650–15658. [Google Scholar] [CrossRef] [PubMed]
- Li, X.-H.; Chavali, P.L.; Pancsa, R.; Chavali, S.; Babu, M.M. Function and Regulation of Phase-Separated Biological Condensates. Biochemistry 2018, 57, 2452–2461. [Google Scholar] [CrossRef] [PubMed]
- Sheth, U.; Parker, R. Decapping and Decay of Messenger RNA Occur in Cytoplasmic Processing Bodies. Science 2003, 300, 805–808. [Google Scholar] [CrossRef] [PubMed]
- Teixeira, D.; Sheth, U.; Valencia-Sanchez, M.A.; Brengues, M.; Parker, R. Processing bodies require RNA for assembly and contain nontranslating mRNAs. RNA 2005, 11, 371–382. [Google Scholar] [CrossRef]
- Parker, R.; Sheth, U. P Bodies and the Control of mRNA Translation and Degradation. Mol. Cell 2007, 25, 635–646. [Google Scholar] [CrossRef]
- Nott, T.J.; Petsalaki, E.; Farber, P.; Jervis, D.; Fussner, E.; Plochowietz, A.; Craggs, T.D.; Bazett-Jones, D.P.; Pawson, T.; Forman-Kay, J.D.; et al. Phase Transition of a Disordered Nuage Protein Generates Environmentally Responsive Membraneless Organelles. Mol. Cell 2015, 57, 936–947. [Google Scholar] [CrossRef]
- Cummings, C.S.; Obermeyer, A.C. Phase Separation Behavior of Supercharged Proteins and Polyelectrolytes. Biochemistry 2017, 57, 314–323. [Google Scholar] [CrossRef]
- Zhang, Y.-L.; Guo, Y.-H.; Wang, Y.-G.; Liu, L.; Qian, C. P-body and Its Involvement in Post-transcriptional Gene Regulation. Chin. J. Biochem. Mol. Biol. 2008, 24, 685–691. [Google Scholar] [CrossRef]
- Stoecklin, G.; Mayo, T.; Anderson, P. ARE-mRNA degradation requires the 5′–3′ decay pathway. Embo Rep. 2006, 7, 72–77. [Google Scholar] [CrossRef]
- Kawa, D.; Testerink, C. Regulation of mRNA decay in plant responses to salt and osmotic stress. Cell. Mol. Life Sci. 2016, 74, 1165–1176. [Google Scholar] [CrossRef]
- Brengues, M.; Teixeira, D.; Parker, R. Movement of Eukaryotic mRNAs Between Polysomes and Cytoplasmic Processing Bodies. Science 2005, 310, 486–489. [Google Scholar] [CrossRef]
- Nsengimana, B.; Khan, F.A.; Ngowi, E.E.; Zhou, X.; Jin, Y.; Jia, Y.; Wei, W.; Ji, S. Processing body (P-body) and its mediators in cancer. Mol. Cell. Biochem. 2022, 477, 1217–1238. [Google Scholar] [CrossRef] [PubMed]
- Yu, B.; LEI, A.-M. The Function of Maternal P-bodies During the Maturation of Mouse(Mus musculus) Oocyte. J. Agric. Biotechnol. 2016, 24, 1781–1789. [Google Scholar]
- Zhang, L.; Wang, Y.; Lang, X.; Chen, S.; Gu, S. Processing bodies and virus infection. Int. J. Genet. 2017, 40, 4. [Google Scholar]
- Xu, J.; Yang, J.-Y.; Niu, Q.-W.; Chua, N.-H. Arabidopsis DCP2, DCP1, and VARICOSE Form a Decapping Complex Required for Postembryonic Development. Plant Cell 2006, 18, 3386–3398. [Google Scholar] [CrossRef]
- Stephan, L.; Jakoby, M.; Hülskamp, M. Evolutionary Comparison of the Developmental/Physiological Phenotype and the Molecular Behavior of SPIRRIG Between Arabidopsis thaliana and Arabis alpina. Front. Plant Sci. 2021, 11, 596065. [Google Scholar] [CrossRef] [PubMed]
- Mo, B.-X.; Ye, H.; Ou, Z.-H.; Liu, L.; Huang, B.; Zhu, S.-L. Stress-induced processing body formation in tobacco suspension cells. J. Shenzhen Univ. Sci. Eng. 2012, 29, 80–84. [Google Scholar] [CrossRef]
- Ye, J.; Yang, J.; Sun, Y.; Zhao, P.; Gao, S.; Jung, C.; Qu, J.; Fang, R.; Chua, N.-H. Geminivirus Activates ASYMMETRIC LEAVES 2 to Accelerate Cytoplasmic DCP2-Mediated mRNA Turnover and Weakens RNA Silencing in Arabidopsis. PLOS Pathog. 2015, 11, e1005196. [Google Scholar] [CrossRef]
- Luo, Y.; Na, Z.; Slavoff, S.A. P-Bodies: Composition, Properties, and Functions. Biochemistry 2018, 57, 2424–2431. [Google Scholar] [CrossRef]
- Coller, J.; Parker, R. Eukaryotic mRNA Decapping. Annu. Rev. Biochem. 2004, 73, 861–890. [Google Scholar] [CrossRef]
- Fenger-Grøn, M.; Fillman, C.; Norrild, B.; Lykke-Andersen, J. Multiple Processing Body Factors and the ARE Binding Protein TTP Activate mRNA Decapping. Mol. Cell 2005, 20, 905–915. [Google Scholar] [CrossRef] [PubMed]
- Perea-Resa, C.; Carrasco-López, C.; Catalá, R.; Turečková, V.; Novak, O.; Zhang, W.; Sieburth, L.E.; Jiménez-Gómez, J.M.; Salinas, J. The LSM1-7 Complex Differentially Regulates Arabidopsis Tolerance to Abiotic Stress Conditions by Promoting Selective mRNA Decapping. Plant Cell 2016, 28, 505–520. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Chua, N.-H. Arabidopsis Decapping 5 Is Required for mRNA Decapping, P-Body Formation, and Translational Repression during Postembryonic Development. Plant Cell 2009, 21, 3270–3279. [Google Scholar] [CrossRef]
- Gong, L. Subcellular Localization of P-Body in Arabidopsis thaliana and Identification of New Protein Components. Master’s Thesis, Shanghai Normal University, Shanghai, China, 2023. [Google Scholar] [CrossRef]
- Guo, C.; Chen, L.; Cui, Y.; Tang, M.; Guo, Y.; Yi, Y.; Li, Y.; Liu, L.; Chen, L. RNA Binding Protein OsTZF7 Traffics Between the Nucleus and Processing Bodies/Stress Granules and Positively Regulates Drought Stress in Rice. Front. Plant Sci. 2022, 13, 802337. [Google Scholar] [CrossRef]
- Seong, S.Y.; Shim, J.S.; Bang, S.W.; Kim, J.-K. Overexpression of OsC3H10, a CCCH-Zinc Finger, Improves Drought Tolerance in Rice by Regulating Stress-Related Genes. Plants 2020, 9, 1298. [Google Scholar] [CrossRef]
- Jan, A.; Maruyama, K.; Todaka, D.; Kidokoro, S.; Abo, M.; Yoshimura, E.; Shinozaki, K.; Nakashima, K.; Yamaguchi-Shinozaki, K. OsTZF1, a CCCH-Tandem Zinc Finger Protein, Confers Delayed Senescence and Stress Tolerance in Rice by Regulating Stress-Related Genes. Plant Physiol. 2013, 161, 1202–1216. [Google Scholar] [CrossRef]
- Tong, J.; Ren, Z.; Sun, L.; Zhou, S.; Yuan, W.; Hui, Y.; Ci, D.; Wang, W.; Fan, L.-M.; Wu, Z.; et al. ALBA proteins confer thermotolerance through stabilizing HSF messenger RNAs in cytoplasmic granules. Nat. Plants 2022, 8, 778–791. [Google Scholar] [CrossRef] [PubMed]
- Mokin, Y.I.; Ilyinsky, N.S.; Nesterov, S.V.; Smirnov, E.Y.; Sergeeva, O.S.; Romanovich, A.E.; Kuznetsova, I.M.; Turoverov, K.K.; Uversky, V.N.; Fonin, A.V. Stress-granules, P-bodies, and cell aging: A bioinformatics study. Biochem. Biophys. Res. Commun. 2024, 694, 149404. [Google Scholar] [CrossRef]
- Martin, E.W.; Holehouse, A.S. Intrinsically disordered protein regions and phase separation: Sequence determinants of assembly or lack thereof. Emerg. Top. Life Sci. 2020, 4, 307–329. [Google Scholar] [CrossRef]
- Statt, A.; Casademunt, H.; Brangwynne, C.P.; Panagiotopoulos, A.Z. Model for disordered proteins with strongly sequence-dependent liquid phase behavior. J. Chem. Phys. 2020, 152, 075101. [Google Scholar] [CrossRef]
- Cheng, C.L.; Wong, M.K.; Hochstrasser, M. Yeast Nst1 is a novel component of P-bodies and is a specific suppressor of proteasome base assembly defects. Mol. Biol. Cell 2021, 32, ar6. [Google Scholar] [CrossRef]
- Cairo, A.; Vargova, A.; Shukla, N.; Capitao, C.; Mikulkova, P.; Valuchova, S.; Pecinkova, J.; Bulankova, P.; Riha, K. Meiotic exit in Arabidopsis is driven by P-body–mediated inhibition of translation. Science 2022, 377, 629–634. [Google Scholar] [CrossRef]
- Jonas, S.; Izaurralde, E. The role of disordered protein regions in the assembly of decapping complexes and RNP granules. Genes Dev. 2013, 27, 2628–2641. [Google Scholar] [CrossRef] [PubMed]
- A Fromm, S.; Truffault, V.; Kamenz, J.; E Braun, J.; A Hoffmann, N.; Izaurralde, E.; Sprangers, R. The structural basis of Edc3- and Scd6-mediated activation of the Dcp1:Dcp2 mRNA decapping complex. EMBO J. 2011, 31, 279–290. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Wang, C.; Wang, Y.; Ma, J.; Wang, T.; Tao, Z.; Liu, P.; Li, S.; Hu, Y.; Gu, A.; et al. The P-body component DECAPPING5 and the floral repressor SISTER OF FCA regulate FLOWERING LOCUS C transcription in Arabidopsis. Plant Cell 2023, 35, 3303–3324. [Google Scholar] [CrossRef] [PubMed]
- Sankaranarayanan, M.; Emenecker, R.J.; Wilby, E.L.; Jahnel, M.; Trussina, I.R.; Wayland, M.; Alberti, S.; Holehouse, A.S.; Weil, T.T. Adaptable P body physical states differentially regulate bicoid mRNA storage during early Drosophila development. Dev. Cell 2021, 56, 2886–2901.e6. [Google Scholar] [CrossRef]
- Na, Z.; Luo, Y.; Cui, D.S.; Khitun, A.; Smelyansky, S.; Loria, J.P.; Slavoff, S.A. Phosphorylation of a Human Microprotein Promotes Dissociation of Biomolecular Condensates. J. Am. Chem. Soc. 2021, 143, 12675–12687. [Google Scholar] [CrossRef]
- Magliozzi, J.O.; Moseley, J.B. Pak1 kinase controls cell shape through ribonucleoprotein granules. eLife 2021, 10, e67648. [Google Scholar] [CrossRef]
- Hsiao, W.-Y.; Wang, Y.-T.; Wang, S.-W. Fission Yeast Puf2, a Pumilio and FBF Family RNA-Binding Protein, Links Stress Granules to Processing Bodies. Mol. Cell. Biol. 2020, 40, e00589-19. [Google Scholar] [CrossRef]
- Protter, D.S.W.; Rao, B.S.; Van Treeck, B.; Lin, Y.; Mizoue, L.; Rosen, M.K.; Parker, R. Intrinsically Disordered Regions Can Contribute Promiscuous Interactions to RNP Granule Assembly. Cell Rep. 2018, 22, 1401–1412. [Google Scholar] [CrossRef]
- Sanders, D.W.; Kedersha, N.; Lee, D.S.W.; Strom, A.R.; Drake, V.; Riback, J.A.; Bracha, D.; Eeftens, J.M.; Iwanicki, A.; Wang, A.; et al. Competing Protein-RNA Interaction Networks Control Multiphase Intracellular Organization. Cell 2020, 181, 306–324.e28. [Google Scholar] [CrossRef] [PubMed]
- Cassani, M.; Seydoux, G. Specialized germline P-bodies are required to specify germ cell fate in Caenorhabditis elegans embryos. Development 2022, 149, dev200920. [Google Scholar] [CrossRef]
- Jain, S.; Parker, R. The discovery and analysis of P Bodies. Adv. Exp. Med. Biol. 2013, 768, 23–43. [Google Scholar] [CrossRef] [PubMed]
- Decker, C.J.; Teixeira, D.; Parker, R. Edc3p and a glutamine/asparagine-rich domain of Lsm4p function in processing body assembly in Saccharomyces cerevisiae. J. Cell Biol. 2007, 179, 437–449. [Google Scholar] [CrossRef] [PubMed]
- Youn, J.-Y.; Dyakov, B.J.; Zhang, J.; Knight, J.D.; Vernon, R.M.; Forman-Kay, J.D.; Gingras, A.-C. Properties of Stress Granule and P-Body Proteomes. Mol. Cell 2019, 76, 286–294. [Google Scholar] [CrossRef]
- Eulalio, A.; Behm-Ansmant, I.; Schweizer, D.; Izaurralde, E. P-Body Formation Is a Consequence, Not the Cause, of RNA-Mediated Gene Silencing. Mol. Cell. Biol. 2007, 27, 3970–3981. [Google Scholar] [CrossRef]
- Aizer, A.; Kafri, P.; Kalo, A.; Shav-Tal, Y. The P Body Protein Dcp1a Is Hyper-phosphorylated during Mitosis. PLoS ONE 2013, 8, e49783. [Google Scholar] [CrossRef]
- Bhullar, D.S.; Sheahan, M.B.; Rose, R.J. RNA processing body (P-body) dynamics in mesophyll protoplasts re-initiating cell division. Protoplasma 2016, 254, 1627–1637. [Google Scholar] [CrossRef]
- Sweet, T.J.; Boyer, B.; Hu, W.; Baker, K.E.; Coller, J. Microtubule disruption stimulates P-body formation. RNA 2007, 13, 493–502. [Google Scholar] [CrossRef]
- Kleer, M.; Mulloy, R.P.; Robinson, C.-A.; Evseev, D.; Bui-Marinos, M.P.; Castle, E.L.; Banerjee, A.; Mubareka, S.; Mossman, K.; Corcoran, J.A. Human coronaviruses disassemble processing bodies. PLOS Pathog. 2022, 18, e1010724. [Google Scholar] [CrossRef]
- Hurst, Z.; Liu, W.; Shi, Q.; Herman, P.K. A distinct P-body-like granule is induced in response to the disruption of microtubule integrity in Saccharomyces cerevisiae. Genetics 2022, 222, iyac105. [Google Scholar] [CrossRef] [PubMed]
- Fang, J.-C.; Tsai, Y.-C.; Chou, W.-L.; Liu, H.-Y.; Chang, C.-C.; Wu, S.-J.; Lu, C.-A. A CCR4-associated factor 1, OsCAF1B, confers tolerance of low-temperature stress to rice seedlings. Plant Mol. Biol. 2020, 105, 177–192. [Google Scholar] [CrossRef] [PubMed]
- Chou, W.-L.; Huang, L.-F.; Fang, J.-C.; Yeh, C.-H.; Hong, C.-Y.; Wu, S.-J.; Lu, C.-A. Divergence of the expression and subcellular localization of CCR4-associated factor 1 (CAF1) deadenylase proteins in Oryza sativa. Plant Mol. Biol. 2014, 85, 443–458. [Google Scholar] [CrossRef]
- Xu, J.; Chua, N.-H. Dehydration stress activates Arabidopsis MPK6 to signal DCP1 phosphorylation. EMBO J. 2012, 31, 1975–1984. [Google Scholar] [CrossRef] [PubMed]
- Soma, F.; Mogami, J.; Yoshida, T.; Abekura, M.; Takahashi, F.; Kidokoro, S.; Mizoi, J.; Shinozaki, K.; Yamaguchi-Shinozaki, K. ABA-unresponsive SnRK2 protein kinases regulate mRNA decay under osmotic stress in plants. Nat. Plants 2017, 3, 16204. [Google Scholar] [CrossRef] [PubMed]
- Pomeranz, M.C.; Hah, C.; Lin, P.-C.; Kang, S.G.; Finer, J.J.; Blackshear, P.J.; Jang, J.-C. The Arabidopsis Tandem Zinc Finger Protein AtTZF1 Traffics between the Nucleus and Cytoplasmic Foci and Binds Both DNA and RNA. Plant Physiol. 2009, 152, 151–165. [Google Scholar] [CrossRef]
- Lin, P.; Pomeranz, M.C.; Jikumaru, Y.; Kang, S.G.; Hah, C.; Fujioka, S.; Kamiya, Y.; Jang, J. The Arabidopsis tandem zinc finger protein AtTZF1 affects ABA- and GA-mediated growth, stress and gene expression responses. Plant J. 2010, 65, 253–268. [Google Scholar] [CrossRef]
- Motomura, K.; Le, Q.T.; Hamada, T.; Kutsuna, N.; Mano, S.; Nishimura, M.; Watanabe, Y. Diffuse Decapping Enzyme DCP2 Accumulates in DCP1 Foci Under Heat Stress in Arabidopsis thaliana. Plant Cell Physiol. 2014, 56, 107–115. [Google Scholar] [CrossRef]
- Merret, R.; Descombin, J.; Juan, Y.-T.; Favory, J.-J.; Carpentier, M.-C.; Chaparro, C.; Charng, Y.-Y.; Deragon, J.-M.; Bousquet-Antonelli, C. XRN4 and LARP1 Are Required for a Heat-Triggered mRNA Decay Pathway Involved in Plant Acclimation and Survival during Thermal Stress. Cell Rep. 2013, 5, 1279–1293. [Google Scholar] [CrossRef]
- Nakaminami, K.; Seki, M. RNA Regulation in Plant Cold Stress Response. Adv. Exp. Med. Biol. 2018, 1081, 23–44. [Google Scholar] [CrossRef]
- Weber, C.; Nover, L.; Fauth, M. Plant stress granules and mRNA processing bodies are distinct from heat stress granules. Plant J. 2008, 56, 517–530. [Google Scholar] [CrossRef]
- Hamada, T.; Yako, M.; Minegishi, M.; Sato, M.; Kamei, Y.; Yanagawa, Y.; Toyooka, K.; Watanabe, Y.; Hara-Nishimura, I. Stress granule formation is induced by a threshold temperature rather than a temperature difference in Arabidopsis. J. Cell Sci. 2018, 131, jcs216051. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez-Beltran, E.; Moschou, P.N.; Smertenko, A.P.; Bozhkov, P.V. Tudor Staphylococcal Nuclease Links Formation of Stress Granules and Processing Bodies with mRNA Catabolism in Arabidopsis. Plant Cell 2015, 27, 926–943. [Google Scholar] [CrossRef]
- Xu, W.; Jian, S.; Li, J.; Wang, Y.; Zhang, M.; Xia, K. Genomic Identification of CCCH-Type Zinc Finger Protein Genes Reveals the Role of HuTZF3 in Tolerance of Heat and Salt Stress of Pitaya (Hylocereus polyrhizus). Int. J. Mol. Sci. 2023, 24, 6359. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Ma, M.; Feng, Y.; Li, H.; Wang, Y.; Ma, Y.; Li, M.; An, F.; Guo, H. EIN2-Directed Translational Regulation of Ethylene Signaling in Arabidopsis. Cell 2015, 163, 670–683. [Google Scholar] [CrossRef] [PubMed]
- Ji, Y.; Guo, H. From Endoplasmic Reticulum (ER) to Nucleus: EIN2 Bridges the Gap in Ethylene Signaling. Mol. Plant 2013, 6, 11–14. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.-H.; Han, J.-Q.; Ma, B.; Cao, W.-Q.; Li, X.-K.; Xiong, Q.; Zhao, H.; Zhao, R.; Zhang, X.; Zhou, Y.; et al. A translational regulator MHZ9 modulates ethylene signaling in rice. Nat. Commun. 2023, 14, 4674. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Song, J.; Wang, L.; Yang, Z.M.; Sun, D. Identification of a DEAD-box RNA Helicase BnRH6 Reveals Its Involvement in Salt Stress Response in Rapeseed (Brassica napus). Int. J. Mol. Sci. 2022, 24, 2. [Google Scholar] [CrossRef]
- Bogamuwa, S.; Jang, J.-C. Plant Tandem CCCH Zinc Finger Proteins Interact with ABA, Drought, and Stress Response Regulators in Processing-Bodies and Stress Granules. PLoS ONE 2016, 11, e0151574. [Google Scholar] [CrossRef]
- Yu, X.; Li, B.; Jang, G.-J.; Jiang, S.; Jiang, D.; Jang, J.-C.; Wu, S.-H.; Shan, L.; He, P. Orchestration of Processing Body Dynamics and mRNA Decay in Arabidopsis Immunity. Cell Rep. 2019, 28, 2194–2205.e6. [Google Scholar] [CrossRef]
- Xu, L.; Liu, T.; Xiong, X.; Liu, W.; Yu, Y.; Cao, J. AtC3H18L is a stop-codon read-through gene and encodes a novel non-tandem CCCH zinc-finger protein that can form cytoplasmic foci similar to mRNP granules. Biochem. Biophys. Res. Commun. 2020, 528, 140–145. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.; Liang, J.; Li, T.; Zhang, D.; Teng, N. A LlMYB305-LlC3H18-LlWRKY33 module regulates thermotolerance in lily. Mol. Hortic. 2023, 3, 15. [Google Scholar] [CrossRef]
| Protein | Uniprot | Experimental Method | Species | References |
|---|---|---|---|---|
| DCP1 | ACZ94554.1 | Prediction | D. melanogaster | [34] |
| DCP2 | AFH04447.1 | Prediction | D. melanogaster | [34,35] |
| DCP5 | OAP16255.1 | predicted by D2P2 | A. thaliana | [36] |
| Me31B | EDW88764.1 | Prediction | Drosophila | [37] |
| NBDY | NP_001335058.1 | 1H-15N HSQC | H. sapiens | [38] |
| Sts5 | BAA23619.1 | Prediction | S. pombe | [39] |
| Puf2 | NP_595389.2 | Prediction | S. pombe | [40] |
| Puf3 | NP_593141.2 | Prediction | S. pombe | [40] |
| Puf4 | CAB03616.1 | Prediction | S. pombe | [40] |
| SMG7 | OAO95115.1 | Prediction | A. thaliana | [33] |
| Lsm4 | QHB08189.1 | Prediction | S. cerevisiae | [41] |
| Dhh1 | CAA98734.1 | Prediction | S. cerevisiae | [41] |
| Pop2 | CAA96333.1 | Prediction | S. cerevisiae | [41] |
| Ccr4 | AAB24455.1 | Prediction | S. cerevisiae | [41] |
| Nst1 | QHB11286.1 | Prediction | S. cerevisiae | [32] |
| G3BP | CAG38772.1 | Prediction | H. sapiens | [42] |
| MEG-1 | NP_510319.1 | Prediction | C. elegans | [43] |
| MEG-2 | NP_510318.2 | Prediction | C. elegans | [43] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, Z.; Xu, Z.; Liu, X.; Chen, G.; Hu, C.; Chen, M.; Liu, Y. Exploring the Role of the Processing Body in Plant Abiotic Stress Response. Curr. Issues Mol. Biol. 2024, 46, 9844-9855. https://doi.org/10.3390/cimb46090585
Huang Z, Xu Z, Liu X, Chen G, Hu C, Chen M, Liu Y. Exploring the Role of the Processing Body in Plant Abiotic Stress Response. Current Issues in Molecular Biology. 2024; 46(9):9844-9855. https://doi.org/10.3390/cimb46090585
Chicago/Turabian StyleHuang, Zhehao, Zhi Xu, Xiuqing Liu, Gangmin Chen, Chensi Hu, Menglu Chen, and Yun Liu. 2024. "Exploring the Role of the Processing Body in Plant Abiotic Stress Response" Current Issues in Molecular Biology 46, no. 9: 9844-9855. https://doi.org/10.3390/cimb46090585





