Identification of Breast Cancer Subtype-Specific Biomarkers by Integrating Copy Number Alterations and Gene Expression Profiles
Abstract
1. Introduction
2. Materials and Methods
2.1. Data
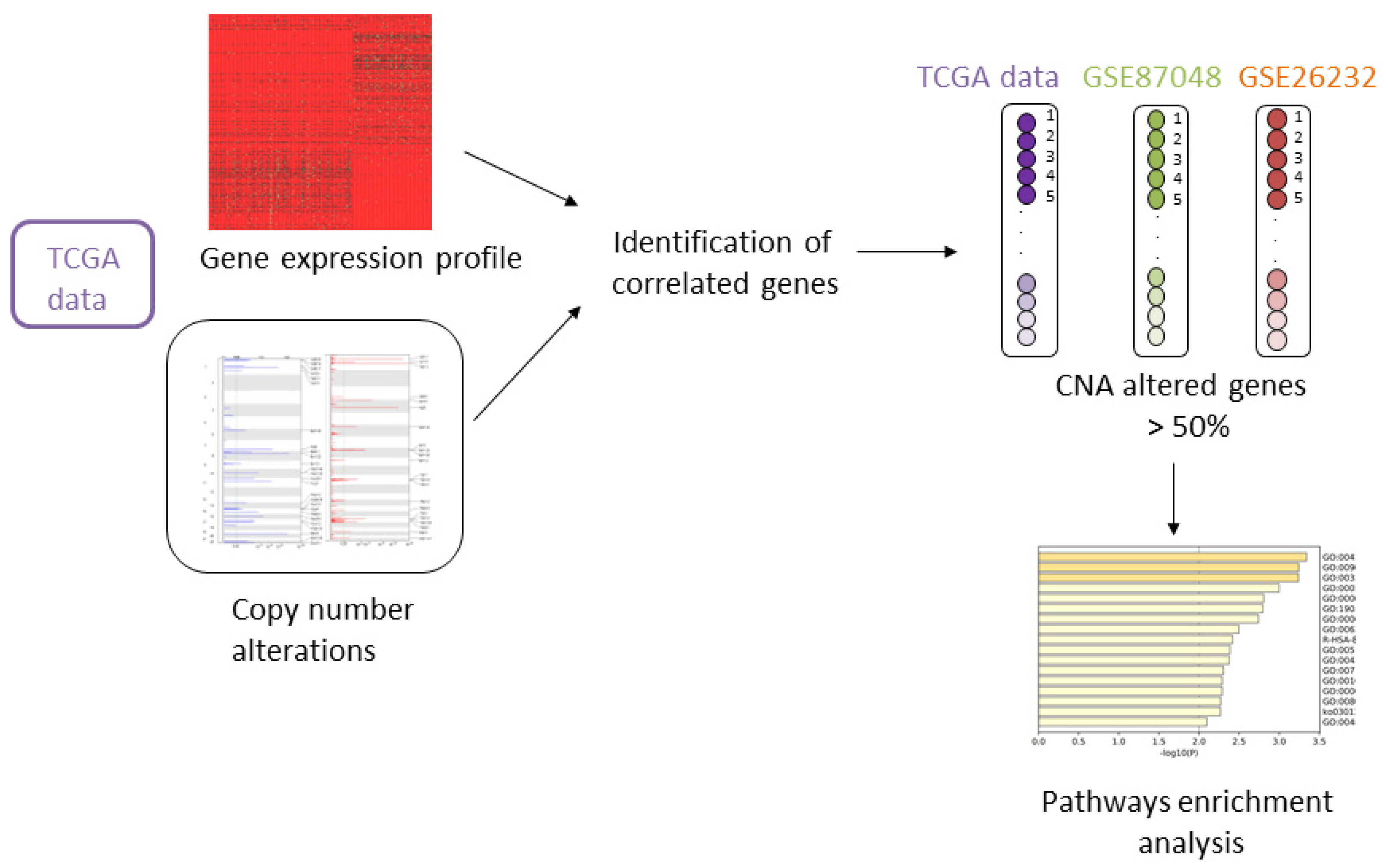
2.2. Integrating Copy Number and Gene Expression
2.3. Identification of Subtype-Specific CNAs
2.4. Survival Analysis
2.5. Pathway Analysis
3. Results
3.1. Correlation Analysis in TCGA Data
3.1.1. Correlation Analysis: Common Genes among Breast Cancer Subtypes
3.1.2. Correlation Analysis: Subtype-Specific Genes
3.2. Survival Analysis
3.3. Analysis of GEO Datasets
3.4. Pathway Enrichment Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Timms, K.M.; Abkevich, V.; Hughes, E.; Neff, C.; Reid, J.; Morris, B.; Kalva, S.; Potter, J.; Tran, T.V.; Chen, J.; et al. Association of BRCA1/2defects with genomic scores predictive of DNA damage repair deficiency among breast cancer subtypes. Breast Cancer Res. 2014, 16, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Sorlie, T.; Perou, C.M.; Tibshirani, R.; Aas, T.; Geisler, S.; Johnsen, H.; Hastie, T.; Eisen, M.B.; van de Rijn, M.; Jeffrey, S.S.; et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc. Natl. Acad. Sci. USA 2001, 98, 10869–10874. [Google Scholar] [CrossRef] [PubMed]
- Pusztai, L.; Mazouni, C.; Anderson, K.; Wu, Y.; Symmans, W.F. Molecular Classification of Breast Cancer: Limitations and Potential. Oncology 2006, 11, 868–877. [Google Scholar] [CrossRef]
- Cava, C.; Zoppis, I.F.; Mauri, G.; Ripamonti, M.; Gallivanone, F.; Salvatore, C.; Gilardi, M.C.; Castiglioni, I. Combination of gene expression and genome copy number alteration has a prognostic value for breast cancer. In Proceedings of the 2013 35th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Osaka, Japan, 3–7 July 2013; pp. 608–611. [Google Scholar] [CrossRef]
- Cava, C.; Bertoli, G.; Castiglioni, I. In silico identification of drug target pathways in breast cancer subtypes using pathway cross-talk inhibition. J. Transl. Med. 2018, 16, 154. [Google Scholar] [CrossRef]
- Pinkel, D.; Albertson, D.G. Array comparative genomic hybridization and its applications in cancer. Nat. Genet. 2005, 37, S11–S17. [Google Scholar] [CrossRef]
- Li, D.; Xia, H.; Li, Z.-Y.; Hua, L.; Li, L. Identification of Novel Breast Cancer Subtype-Specific Biomarkers by Integrating Genomics Analysis of DNA Copy Number Aberrations and miRNA-mRNA Dual Expression Profiling. BioMed Res. Int. 2015, 2015, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Cava, C.; Bertoli, G.; Colaprico, A.; Bontempi, G.; Mauri, G.; Castiglioni, I. In-Silico Integration Approach to Identify a Key miRNA Regulating a Gene Network in Aggressive Prostate Cancer. Int. J. Mol. Sci. 2018, 19, 910. [Google Scholar] [CrossRef]
- Cava, C.; Sabetian, S.; Castiglioni, I. Patient-Specific Network for Personalized Breast Cancer Therapy with Multi-Omics Data. Entropy 2021, 23, 225. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Cheng, F.; Zhao, Z. Tissue-Specific Signaling Networks Rewired by Major Somatic Mutations in Human Cancer Revealed by Proteome-Wide Discovery. Cancer Res. 2017, 77, 2810–2821. [Google Scholar] [CrossRef] [PubMed]
- Porta-Pardo, E.; Garcia-Alonso, L.; Hrabe, T.; Dopazo, J.; Godzik, A. A Pan-Cancer Catalogue of Cancer Driver Protein Interaction Interfaces. PLoS Comput. Biol. 2015, 11, e1004518. [Google Scholar] [CrossRef] [PubMed]
- Korthauer, K.D.; Kendziorski, C. MADGiC: A model-based approach for identifying driver genes in cancer. Bioinformatics 2015, 31, 1526–1535. [Google Scholar] [CrossRef]
- Shrestha, R.; Hodzic, E.; Sauerwald, T.; Dao, P.; Wang, K.; Yeung, J.; Anderson, S.; Vandin, F.; Haffari, G.; Collins, C.C.; et al. HIT’nDRIVE: Patient-specific multidriver gene prioritization for precision oncology. Genome Res. 2017, 27, 1573–1588. [Google Scholar] [CrossRef]
- Ding, J.; McConechy, M.K.; Horlings, H.M.; Ha, G.; Chan, F.C.; Funnell, T.; Mullaly, S.C.; Reimand, J.; Bashashati, A.; Bader, G.D.; et al. Systematic analysis of somatic mutations impacting gene expression in 12 tumour types. Nat. Commun. 2015, 6, 8554. [Google Scholar] [CrossRef]
- Suo, C.; Hrydziuszko, O.; Lee, D.; Pramana, S.; Saputra, D.; Joshi, H.; Calza, S.; Pawitan, Y. Integration of somatic mutation, expression and functional data reveals potential driver genes predictive of breast cancer survival. Bioinformatics 2015, 31, 2607–2613. [Google Scholar] [CrossRef]
- Bhattacharya, A.; Bense, R.D.; Urzúa-Traslaviña, C.G.; De Vries, E.G.E.; Van Vugt, M.A.T.M.; Fehrmann, R.S.N. Transcriptional effects of copy number alterations in a large set of human cancers. Nat. Commun. 2020, 11, 1–12. [Google Scholar] [CrossRef]
- Fehrmann, R.S.N.; Karjalainen, J.M.; Krajewska, M.; Westra, H.-J.; Maloney, D.J.; Simeonov, A.; Pers, T.H.; Hirschhorn, J.N.; Jansen, R.C.; Schultes, E.A.; et al. Gene expression analysis identifies global gene dosage sensitivity in cancer. Nat. Genet. 2015, 47, 115–125. [Google Scholar] [CrossRef] [PubMed]
- Fleck, J.L.; Pavel, A.B.; Cassandras, C.G. Integrating mutation and gene expression cross-sectional data to infer cancer progression. BMC Syst. Biol. 2016, 10, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Cava, C.; Zoppis, I.; Gariboldi, M.; Castiglioni, I.; Mauri, G.; Antoniotti, M. Combined analysis of chromosomal instabilities and gene expression for colon cancer progression inference. J Clin Bioinform. 2014, 4, 2. [Google Scholar] [CrossRef] [PubMed]
- Colaprico, A.; Silva, T.C.; Olsen, C.; Garofano, L.; Cava, C.; Garolini, D.; Sabedot, T.S.; Malta, T.M.; Pagnotta, S.M.; Castiglioni, I.; et al. TCGAbiolinks: An R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Res. 2016, 44, e71. [Google Scholar] [CrossRef]
- Ng, S.; Collisson, E.A.; Sokolov, A.; Goldstein, T.; Gonzalez-Perez, A.; Lopez-Bigas, N.; Benz, C.; Haussler, D.; Stuart, J.M. PARADIGM-SHIFT predicts the function of mutations in multiple cancers using pathway impact analysis. Bioinformatics 2012, 28, i640–i646. [Google Scholar] [CrossRef] [PubMed]
- Bussey, K.J.; Chin, K.; Lababidi, S.; Reimers, M.; Reinhold, W.C.; Kuo, W.-L.; Gwadry, F.; Ajay; Kouros-Mehr, H.; Fridlyand, J.; et al. Integrating data on DNA copy number with gene expression levels and drug sensitivities in the NCI-60 cell line panel. Mol. Cancer Ther. 2006, 5, 853–867. [Google Scholar] [CrossRef]
- Sales, G.; Coppe, A.; Bisognin, A.; Biasiolo, M.; Bortoluzzi, S.; Romualdi, C. MAGIA, a web-based tool for miRNA and Genes Integrated Analysis. Nucleic Acids Res. 2010, 38, W352–W359. [Google Scholar] [CrossRef]
- Lygirou, V.; Latosinska, A.; Makridakis, M.; Mullen, W.; Delles, C.; Schanstra, J.P.; Zoidakis, J.; Pieske, B.; Mischak, H.; Vlahou, A. Plasma proteomic analysis reveals altered protein abundances in cardiovascular disease. J. Transl. Med. 2018, 16, 1–12. [Google Scholar] [CrossRef]
- Zhao, Z.; Jinde, S.; Koike, S.; Tada, M.; Satomura, Y.; Yoshikawa, A.; Nishimura, Y.; Takizawa, R.; Kinoshita, A.; Sakakibara, E.; et al. Altered expression of microRNA-223 in the plasma of patients with first-episode schizophrenia and its possible relation to neuronal migration-related genes. Transl. Psychiatry 2019, 9, 1–11. [Google Scholar] [CrossRef]
- Nagy, Á.; Lánczky, A.; Menyhárt, O.; Győrffy, B. Validation of miRNA prognostic power in hepatocellular carcinoma using expression data of independent datasets. Sci. Rep. 2018, 8, 1–9. [Google Scholar] [CrossRef]
- Fabregat, A.; Jupe, S.; Matthews, L.; Sidiropoulos, K.; Gillespie, M.; Garapati, P.; Haw, R.; Jassal, B.; Korninger, F.; May, B.; et al. The Reactome Pathway Knowledgebase. Nucleic Acids Res. 2018, 46, D649–D655. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.; Kruse, J.-P.; Tang, Y.; Jung, S.Y.; Qin, J.; Gu, W. Negative regulation of the deacetylase SIRT1 by DBC1. Nat. Cell Biol. 2008, 451, 587–590. [Google Scholar] [CrossRef]
- Best, S.A.; Nwaobasi, A.N.; Schmults, C.D.; Ramsey, M.R. CCAR2 Is Required for Proliferation and Tumor Maintenance in Human Squamous Cell Carcinoma. J. Investig. Dermatol. 2017, 137, 506–512. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Liao, J.; Wu, S.; Fan, J.; Peng, Z.; Wang, Z. Overexpression of DBC1, correlated with poor prognosis, is a potential therapeutic target for hepatocellular carcinoma. Biochem. Biophys. Res. Commun. 2017, 494, 511–517. [Google Scholar] [CrossRef]
- Richardson, J.; Shaaban, A.M.; Kamal, M.; Alisary, R.; Walker, C.; Ellis, I.O.; Speirs, V.; Green, A.R.; Bell, S.M. Microcephalin is a new novel prognostic indicator in breast cancer associated with BRCA1 inactivation. Breast Cancer Res. Treat. 2010, 127, 639–648. [Google Scholar] [CrossRef][Green Version]
- Tervasmäki, A.; Mantere, T.; Eshraghi, L.; Laurila, N.; Tuppurainen, H.; Ronkainen, V.; Koivuluoma, S.; Devarajan, R.; Peltoketo, H.; Pylkäs, K. Tumor suppressor MCPH1 regulates gene expression profiles related to malignant conversion and chromosomal assembly. Int. J. Cancer 2019, 145, 2070–2081. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharya, N.; Mukherjee, N.; Singh, R.K.; Sinha, S.; Alam, N.; Roy, A.; Roychoudhury, S.; Panda, C.K. Frequent Alterations of MCPH1 and ATM are Associated with Primary Breast Carcinoma: Clinical and Prognostic Implications. Ann. Surg. Oncol. 2012, 20, 424–432. [Google Scholar] [CrossRef] [PubMed]
- Mimori, K.; Inoue, H.; Shiraishi, T.; Ueo, H.; Mafune, K.-I.; Tanaka, Y.; Mori, M. A Single-Nucleotide Polymorphism of SMARCB1 in Human Breast Cancers. Genomics 2002, 80, 254–258. [Google Scholar] [CrossRef]
- Sévenet, N.; Lellouch-Tubiana, A.; Schofield, D.; Hoang-Xuan, K.; Gessler, M.; Birnbaum, D.; Jeanpierre, C.; Jouvet, A.; Delattre, O. Spectrum of hSNF5IINI1 Somatic Mutations in Human Cancer and Genotype-Phenotype Correlations. Hum. Mol. Genet. 1999, 8, 2359–2368. [Google Scholar] [CrossRef] [PubMed]
- Silva, T.M.; Cirenajwis, H.; Wallace, H.M.; Oredsson, S.; Persson, L. A role for antizyme inhibitor in cell proliferation. Amino Acids 2015, 47, 1341–1352. [Google Scholar] [CrossRef] [PubMed]
- Chu, P.-Y.; Wang, S.-M.; Chen, P.-M.; Tang, F.-Y.; Chiang, E.-P.I. Expression of MTDH and IL-10 Is an Independent Predictor of Worse Prognosis in ER-Negative or PR-Negative Breast Cancer Patients. J. Clin. Med. 2020, 9, 3153. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, Y.T.-K.; Moon, J.Y.; Ediriweera, M.K.; Cho, S.K. Phenethyl Isothiocyanate Suppresses Stemness in the Chemo- and Radio-Resistant Triple-Negative Breast Cancer Cell Line MDA-MB-231/IR Via Downregulation of Metadherin. Cancers 2020, 12, 268. [Google Scholar] [CrossRef] [PubMed]
- Dupont, W.D.; Bs, J.P.B.; Bs, K.M.B.; Rn, P.A.S.; Bs, W.D.P.; Sanders, M.E.; Page, D.L.; Smith, J.R. Protein phosphatase 2A subunit gene haplotypes and proliferative breast disease modify breast cancer risk. Cancer 2009, 116, 8–19. [Google Scholar] [CrossRef] [PubMed]
- Ali, K.; Mahjabeen, I.; Sabir, M.; Baig, R.M.; Zafeer, M.; Faheem, M.; Kayani, M.A. Germline variations of apurinic/apyrimidinic endonuclease 1 (APEX1) detected in female breast cancer patients. Asian Pac. J. Cancer Prev. 2014, 15, 7589–7595. [Google Scholar] [CrossRef][Green Version]
- Coughlin, S.S. Epidemiology of Breast Cancer in Women. Adv. Exp. Med. Biol. 2019, 1152, 9–29. [Google Scholar] [CrossRef]
- Oh, J.H.; Lee, J.-Y.; Kim, K.H.; Kim, C.Y.; Jeong, D.S.; Cho, Y.; Nam, K.T.; Kim, M.H. Elevated GCN5 expression confers tamoxifen resistance by upregulating AIB1 expression in ER-positive breast cancer. Cancer Lett. 2020, 495, 145–155. [Google Scholar] [CrossRef]
- Taghavi, A.; Akbari, M.E.; Hashemi-Bahremani, M.; Nafissi, N.; Khalilnezhad, A.; Poorhosseini, S.M.; Hashemi-Gorji, F.; Yassaee, V.R. Gene expression profiling of the 8q22-24 position in human breast cancer: TSPYL5, MTDH, ATAD2 and CCNE2 genes are implicated in oncogenesis, while WISP1 and EXT1 genes may predict a risk of metastasis. Oncol. Lett. 2016, 12, 3845–3855. [Google Scholar] [CrossRef] [PubMed]
- Osawa, T.; Shimamura, T.; Saito, K.; Hasegawa, Y.; Ishii, N.; Nishida, M.; Ando, R.; Kondo, A.; Anwar, M.; Tsuchida, R.; et al. Phosphoethanolamine Accumulation Protects Cancer Cells under Glutamine Starvation through Downregulation of PCYT2. Cell Rep. 2019, 29, 89–103.e7. [Google Scholar] [CrossRef]
- Zhu, L.; Bakovic, M. Breast cancer cells adapt to metabolic stress by increasing ethanolamine phospholipid synthesis and CTP:ethanolaminephosphate cytidylyltransferase-Pcyt2 activity. Biochem. Cell Biol. 2012, 90, 188–199. [Google Scholar] [CrossRef]
- Pluta, P.; Cebula-Obrzut, B.; Ehemann, V.; Pluta, A.; Wierzbowska, A.; Piekarski, J.; Bilski, A.; Nejc, D.; Kordek, R.; Robak, T.; et al. Correlation of Smac/DIABLO protein expression with the clinico-pathological features of breast cancer patients. Neoplasma 2011, 58, 430–435. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zhang, J.; Cao, M.; Dong, J.; Li, C.; Xu, W.; Zhan, Y.; Wang, X.; Yu, M.; Ge, C.; Ge, Z.; et al. ABRO1 suppresses tumourigenesis and regulates the DNA damage response by stabilizing p53. Nat. Commun. 2014, 5, 5059. [Google Scholar] [CrossRef] [PubMed]
- Cashman, R.; Cohen, H.; Ben-Hamo, R.; Zilberberg, A.; Efroni, S. SENP5 mediates breast cancer invasion via a TGFβRI SUMOylation cascade. Oncotarget 2014, 5, 1071–1082. [Google Scholar] [CrossRef]
- Adamopoulos, P.G.; Raptis, G.D.; Kontos, C.K.; Scorilas, A. Discovery and expression analysis of novel transcripts of the human SR-related CTD-associated factor 1 (SCAF1) gene in human cancer cells using Next-Generation Sequencing. Gene 2018, 670, 155–165. [Google Scholar] [CrossRef]
- Alshareeda, A.T.; Negm, O.H.; Green, A.R.; Nolan, C.; Tighe, P.; AlBarakati, N.; Sultana, R.; Madhusudan, S.; Ellis, I.O.; Rakha, E.A. SUMOylation proteins in breast cancer. Breast Cancer Res. Treat. 2014, 144, 519–530. [Google Scholar] [CrossRef]
- Wazir, U.; Jiang, W.G.; Yasaei, H.; Linne, H.; Newbold, R.F.; Mokbel, K. P14ARF is down-regulated during tumour progression and predicts the clinical outcome in human breast cancer. Anticancer. Res. 2013, 33, 2185–2189. [Google Scholar]
- Bao, J.; Zhu, L.; Zhu, Q.; Su, J.; Liu, M.; Huang, W. SREBP-1 is an independent prognostic marker and promotes invasion and migration in breast cancer. Oncol. Lett. 2016, 12, 2409–2416. [Google Scholar] [CrossRef] [PubMed]



| Molecular Subtype | TCGA | GSE87048 | GSE26232 |
|---|---|---|---|
| LumA | 548 | 37 | 12 |
| LumB | 206 | 20 | 6 |
| Her2 | 82 | 12 | 2 |
| Basal | 188 | 8 | 17 |
| Total | 1024 | 77 | 37 |
| Pathway | p-Value | |
|---|---|---|
| 29 Luminal A-Specific Genes | ||
| SUMOylation of SUMOylation proteins | 0.003 | |
| SUMOylation of ubiquitinylation proteins | 0.004 | |
| Defective intrinsic pathway for apoptosis due to p14ARF loss of function | 0.004 | |
| 90 Luminal B-Specific Genes | ||
| Cellular senescence | 0.005 | |
| Oxidative stress induced senescence | 0.001 | |
| Disassembly of the destruction complex and recruitment of AXIN to the membrane | 0.001 | |
| 40 HER2-Specific Genes | ||
| Aryl hydrocarbon receptor signaling | 0.0003 | |
| VxPx cargo-targeting to cilium | 0.003 | |
| Amplification of signal from the kinetochores | 0.004 | |
| 23 Basal-Specific Genes | ||
| Regulation of cholesterol biosynthesis by SREBP (SREBF) | 0.0002 | |
| Activation of gene expression by SREBF (SREBP) | 0.004 | |
| Metabolism of steroids | 0.004 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cava, C.; Pisati, M.; Frasca, M.; Castiglioni, I. Identification of Breast Cancer Subtype-Specific Biomarkers by Integrating Copy Number Alterations and Gene Expression Profiles. Medicina 2021, 57, 261. https://doi.org/10.3390/medicina57030261
Cava C, Pisati M, Frasca M, Castiglioni I. Identification of Breast Cancer Subtype-Specific Biomarkers by Integrating Copy Number Alterations and Gene Expression Profiles. Medicina. 2021; 57(3):261. https://doi.org/10.3390/medicina57030261
Chicago/Turabian StyleCava, Claudia, Mirko Pisati, Marco Frasca, and Isabella Castiglioni. 2021. "Identification of Breast Cancer Subtype-Specific Biomarkers by Integrating Copy Number Alterations and Gene Expression Profiles" Medicina 57, no. 3: 261. https://doi.org/10.3390/medicina57030261
APA StyleCava, C., Pisati, M., Frasca, M., & Castiglioni, I. (2021). Identification of Breast Cancer Subtype-Specific Biomarkers by Integrating Copy Number Alterations and Gene Expression Profiles. Medicina, 57(3), 261. https://doi.org/10.3390/medicina57030261







