Discovery of PD-L1 Peptide Inhibitors from Ascidian Enzymatic Hydrolysates by Affinity Ultrafiltration Coupled to NanoLC-MS/MS
Abstract
:1. Introduction
2. Results and Discussion
2.1. Anti-PD-1/PD-L1 Effects of Total Enzymatic Hydrolysates of Ascidians
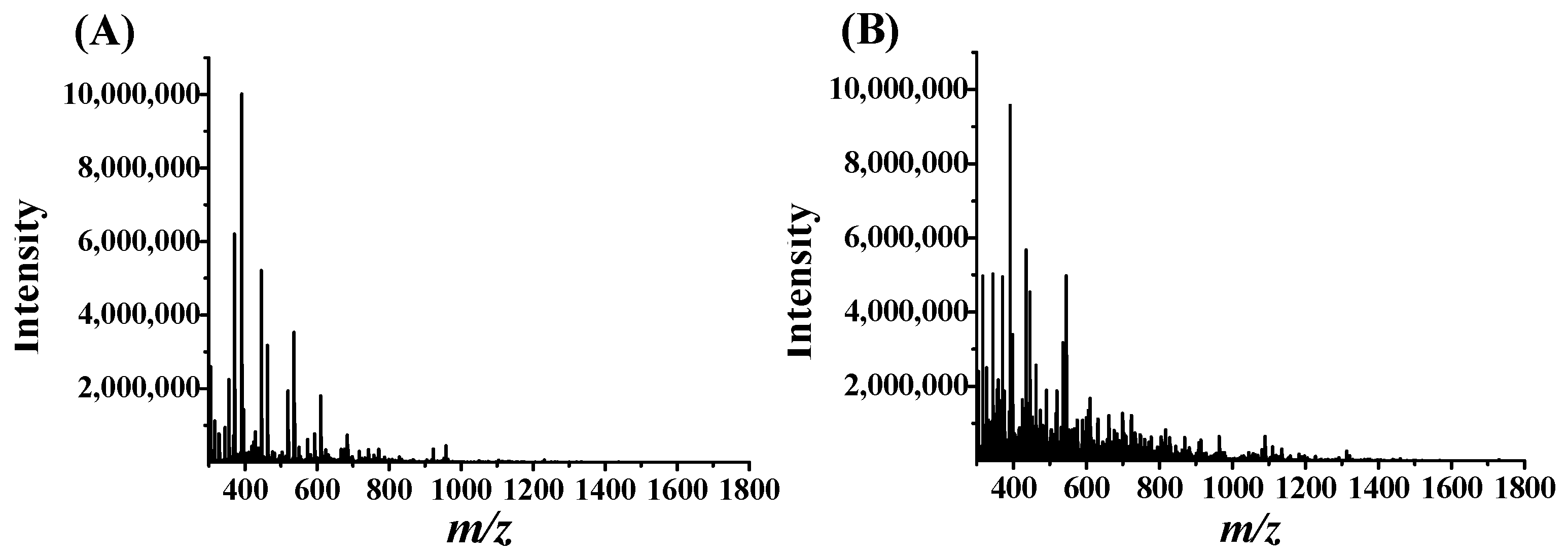
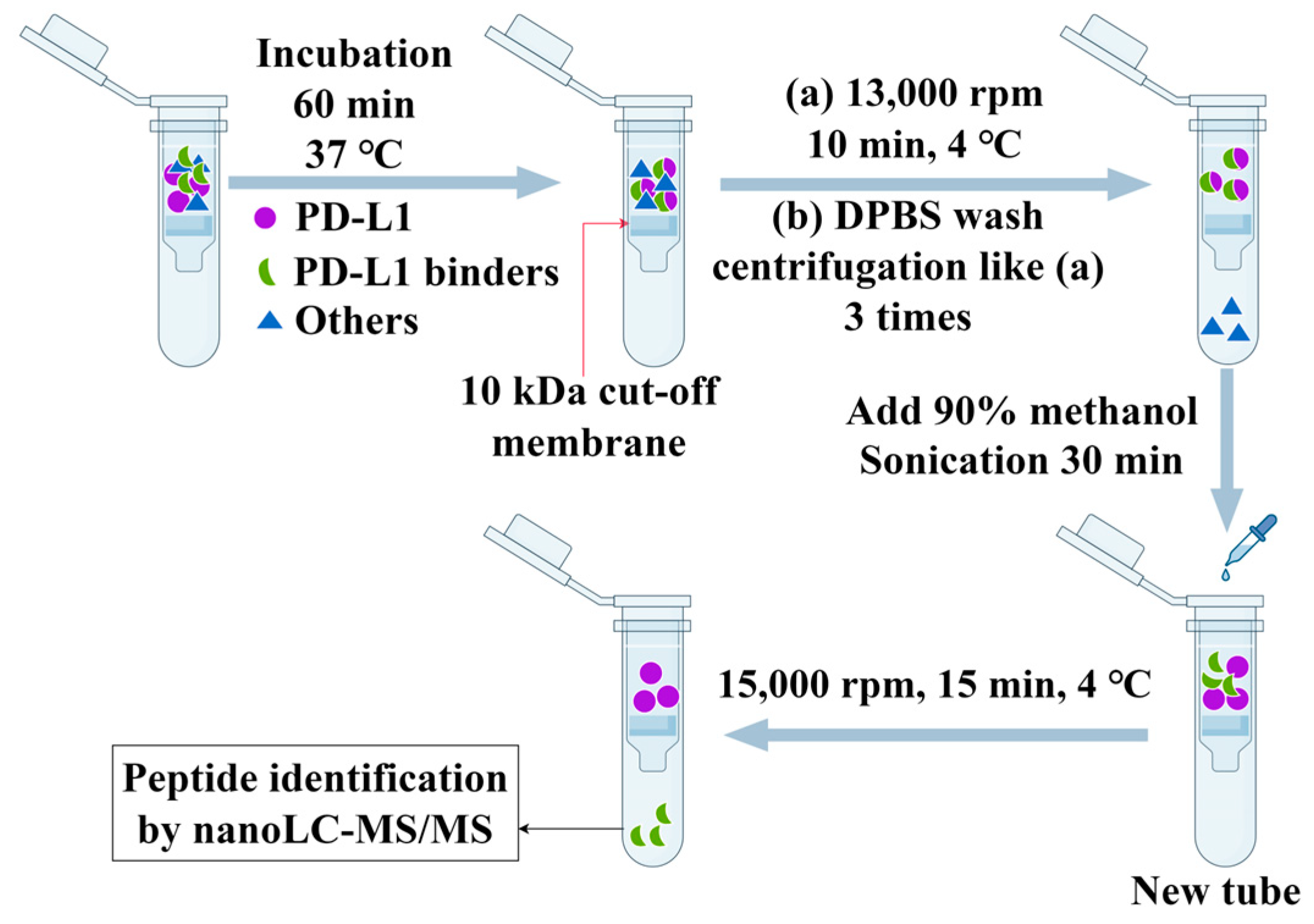
2.2. Filtration of PD-L1 Peptide Inhibitors Using AUF
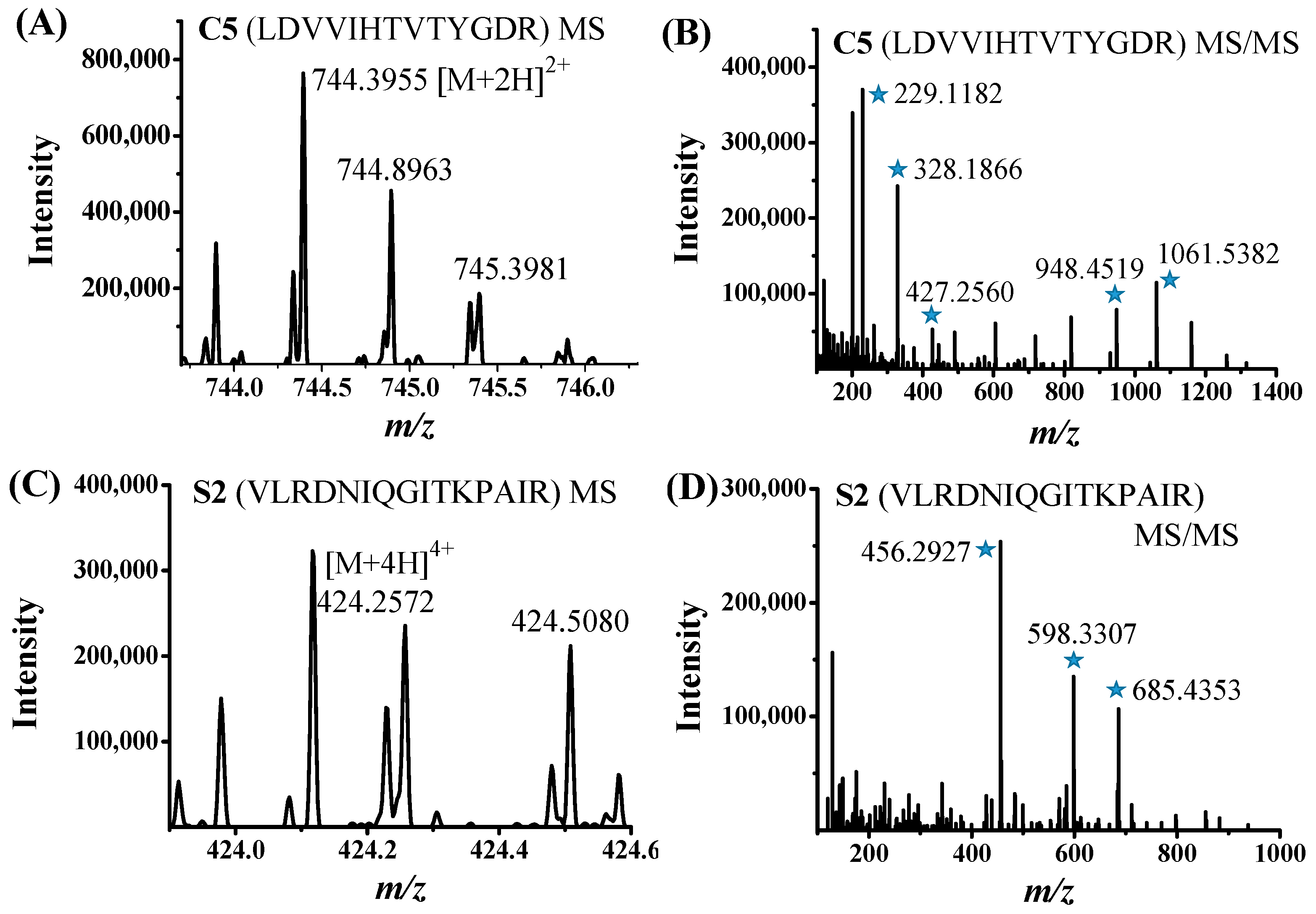
2.3. Identification of PD-L1 Peptide Binders by nanoLC-MS/MS
2.4. Anti-PD-1/PD-L1 Effects of PD-L1 Peptide Binders
2.5. Predicted Binding Sites of C5 and S2 Towards PD-L1
3. Materials and Methods
3.1. Preparation of Trypsin Hydrolysates of Ascidians
3.2. AUF Experiment
3.3. NanoLC-MS/MS Analysis
3.4. Molecular Docking Using HPEPDOCK and MOE
3.5. HTRF Binding Assay
3.6. SPR Experiment
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Rauf, A.; Khalil, A.A.; Khan, M.; Anwar, S.; Alamri, A.; Alqarni, A.M.; Alghamdi, A.; Alshammari, F.; Rengasamy, K.R.R.; Wan, C. Can be marine bioactive peptides (MBAs) lead the future of foodomics for human health? Crit. Rev. Food Sci. Nutr. 2021, 62, 7072–7116. [Google Scholar]
- Tamilselvi, M.; Sivakumar, V.; Ali, H.A.J.; Thilaga, R.D. Preparation of pickle from Herdmania pallida, simple ascidian. World J. Dairy Food Sci. 2010, 5, 88–92. [Google Scholar]
- Lambert, G. Ecology and natural history of the protochordates. Can. J. Zool. 2005, 83, 34–50. [Google Scholar] [CrossRef]
- Gulliksen, B.; Skjæveland, S.H. The sea-star, Asterias rubens L., as predator on the ascidian, Ciona intestinalis (L.), in Borgenfjorden, North-Tröndelag, Norway. Sarsia 1973, 52, 15–20. [Google Scholar]
- Watters, D.J. Ascidian Toxins with Potential for Drug Development. Mar. Drugs 2018, 16, 162. [Google Scholar] [CrossRef] [PubMed]
- Dou, X.; Dong, B. Origins and Bioactivities of Natural Compounds Derived from Marine Ascidians and Their Symbionts. Mar. Drugs 2019, 17, 670. [Google Scholar] [CrossRef]
- Conte, M.; Fontana, E.; Nebbioso, A.; Altucci, L. Marine-Derived Secondary Metabolites as Promising Epigenetic Bio-Compounds for Anticancer Therapy. Mar. Drugs 2020, 19, 15. [Google Scholar] [CrossRef]
- Mauro, M.; Lazzara, V.; Punginelli, D.; Arizza, V.; Vazzana, M. Antitumoral compounds from vertebrate sister group: A review of Mediterranean ascidians. Dev. Comp. Immunol. 2020, 108, 103669. [Google Scholar]
- Rinehart, K.L.; Holt, T.G.; Fregeau, N.L.; Stroh, J.G.; Keifer, P.A.; Sun, F.; Li, L.H.; Martin, D.G. Ecteinascidins 729, 743, 745, 759A, 759B, and 770: Potent antitumor agents from the Caribbean tunicate Ecteinascidia turbinata. J. Org. Chem. 1990, 55, 4512–4515. [Google Scholar]
- Holt, T.G. The Isolation and Structural Characterization of the Ecteinascidins. Ph.D. Dissertation, University of Illinois at Urbana-Champaign, Urbana, IL, USA, 1986. [Google Scholar]
- Cuevas, C.; Francesch, A. Development of Yondelis® (trabectedin, ET-743). A semisynthetic process solves the supply problem. Nat. Product. Rep. 2009, 26, 322–337. [Google Scholar] [CrossRef]
- Schwartsmann, G.; Brondani da Rocha, A.; Berlinck, R.G.; Jimeno, J. Marine organisms as a source of new anticancer agents. Lancet Oncol. 2001, 2, 221–225. [Google Scholar] [CrossRef] [PubMed]
- Arumugam, V.; Venkatesan, M.; Ramachandran, S.; Sundaresan, U. Bioactive peptides from marine ascidians and future drug development–A review. Int. J. Pept. Res. Ther. 2018, 24, 13–18. [Google Scholar] [CrossRef]
- Dyshlovoy, S.A.; Honecker, F. Marine Compounds and Cancer: Updates 2020. Mar. Drugs 2020, 18, 643. [Google Scholar] [CrossRef]
- Broggini, M.; Marchini, S.V.; Galliera, E.; Borsotti, P.; Taraboletti, G.; Erba, E.; Sironi, M.; Jimeno, J.; Faircloth, G.T.; Giavazzi, R.; et al. Aplidine, a new anticancer agent of marine origin, inhibits vascular endothelial growth factor (VEGF) secretion and blocks VEGF-VEGFR-1 (flt-1) autocrine loop in human leukemia cells MOLT-4. Leukemia 2003, 17, 52–59. [Google Scholar] [CrossRef]
- Martins, A.; Vieira, H.; Gaspar, H.; Santos, S. Marketed marine natural products in the pharmaceutical and cosmeceutical industries: Tips for success. Mar. Drugs 2014, 12, 1066–1101. [Google Scholar] [CrossRef]
- Palanisamy, S.K.; Giacobbe, S.; Sundaresan, U. Marine ascidians potential source for new class of anti-cancer drugs. World J. Pharm. Pharm. Sci. 2015, 4, 474–485. [Google Scholar]
- Schmitz, F.J.; Bowden, B.F.; Toth, S.I.; Attaway, D.H.; Zaborsky, O.R. Marine Biotechnology, Volume 1, Pharmaceutical and Bioactive Natural Products; Plenum Press: New York, NY, USA, 1993. [Google Scholar]
- Suarez-Jimenez, G.-M.; Burgos-Hernandez, A.; Ezquerra-Brauer, J.-M. Bioactive peptides and depsipeptides with anticancer potential: Sources from marine animals. Mar. Drugs 2012, 10, 963–986. [Google Scholar] [CrossRef]
- McKeever, B.; Pattenden, G. Total synthesis of trunkamide A, a novel thiazoline-based prenylated cyclopeptide metabolite from Lissoclinum sp. Tetrahedron 2003, 59, 2713–2727. [Google Scholar] [CrossRef]
- Vervoort, H.; Fenical, W.; Epifanio, R.A. Tamandarins A and B: New cytotoxic depsipeptides from a Brazilian ascidian of the family Didemnidae. J. Org. Chem. 2000, 65, 782–792. [Google Scholar] [CrossRef]
- Donia, M.S.; Wang, B.; Dunbar, D.C.; Desai, P.V.; Patny, A.; Avery, M.; Hamann, M.T. Mollamides B and C, Cyclic hexapeptides from the indonesian tunicate Didemnum molle. J. Nat. Prod. 2008, 71, 941–945. [Google Scholar] [CrossRef]
- Carroll, A.R.; Bowden, B.F.; Coll, J.C.; Hockless, D.C.; Skelton, B.W.; White, A.H. Studies of Australian ascidians. IV. Mollamide, a cytotoxic cyclic heptapeptide from the compound ascidian Didemnum molle. Aust. J. Chem. 1994, 47, 61–69. [Google Scholar]
- Zhang, Q.-T.; Liu, Z.-D.; Wang, Z.; Wang, T.; Wang, N.; Wang, N.; Zhang, B.; Zhao, Y.-F. Recent Advances in Small Peptides of Marine Origin in Cancer Therapy. Mar. Drugs 2021, 19, 115. [Google Scholar] [CrossRef]
- Edler, M.C.; Fernandez, A.M.; Lassota, P.; Ireland, C.M.; Barrows, L.R. Inhibition of tubulin polymerization by vitilevuamide, a bicyclic marine peptide, at a site distinct from colchicine, the vinca alkaloids, and dolastatin 10. Biochem. Pharmacol. 2002, 63, 707–715. [Google Scholar] [PubMed]
- Urda, C.; Fernández, R.; Rodríguez, J.; Pérez, M.; Jiménez, C.; Cuevas, C. Bistratamides M and N, Oxazole-Thiazole Containing Cyclic Hexapeptides Isolated from Lissoclinum bistratum Interaction of Zinc (II) with Bistratamide K. Mar. Drugs 2017, 15, 209. [Google Scholar] [CrossRef]
- Won, T.H.; Kim, C.-K.; Lee, S.-H.; Rho, B.J.; Lee, S.K.; Oh, D.-C.; Oh, K.-B.; Shin, J. Amino Acid-Derived Metabolites from the Ascidian Aplidium sp. Mar. Drugs 2015, 13, 3836–3848. [Google Scholar] [CrossRef]
- Hansen, I.K.O.; Isaksson, J.; Poth, A.G.; Hansen, K.O.; Andersen, A.J.C.; Richard, C.S.M.; Blencke, H.M.; Stensvag, K.; Craik, D.J.; Haug, T. Isolation and Characterization of Antimicrobial Peptides with Unusual Disulfide Connectivity from the Colonial Ascidian Synoicum turgens. Mar. Drugs 2020, 18, 51. [Google Scholar] [CrossRef]
- Brönstrup, M.; Sasse, F. Natural products targeting the elongation phase of eukaryotic protein biosynthesis. Nat. Product. Rep. 2020, 37, 752–762. [Google Scholar]
- Vadevoo, S.M.P.; Gurung, S.; Khan, F.; Haque, M.E.; Gunassekaran, G.R.; Chi, L.; Permpoon, U.; Lee, B. Peptide-based targeted therapeutics and apoptosis imaging probes for cancer therapy. Arch. Pharm. Res. 2019, 42, 150–158. [Google Scholar]
- Keir, M.E.; Butte, M.J.; Freeman, G.J.; Sharpe, A.H. PD-1 and its ligands in tolerance and immunity. Annu. Rev. Immunol. 2008, 26, 677–704. [Google Scholar]
- Wang, Z.; Liang, H.; Cao, H.; Zhang, B.; Li, J.; Wang, W.; Qin, S.; Wang, Y.; Xuan, L.; Lai, L.; et al. Efficient ligand discovery from natural herbs by integrating virtual screening, affinity mass spectrometry and targeted metabolomics. Analyst 2019, 144, 2881–2890. [Google Scholar]
- Pai, S.I.; Faquin, W.C. Programmed cell death ligand 1 as a biomarker in head and neck cancer. Cancer Cytopathol. 2017, 125, 529–533. [Google Scholar] [CrossRef] [PubMed]
- Sznol, M.; Chen, L. Antagonist antibodies to PD-1 and B7-H1 (PD-L1) in the treatment of advanced human cancer. Clin. Cancer Res. 2013, 19, 1021–1034. [Google Scholar] [CrossRef] [PubMed]
- Zahavi, D.; Weiner, L. Monoclonal Antibodies in Cancer Therapy. Antibodies 2020, 9, 34. [Google Scholar] [CrossRef] [PubMed]
- Bojko, M.; Węgrzyn, K.; Sikorska, E.; Kocikowski, M.; Parys, M.; Battin, C.; Steinberger, P.; Kogut, M.M.; Winnicki, M.; Sieradzan, A.K.; et al. Design, synthesis and biological evaluation of PD-1 derived peptides as inhibitors of PD-1/PD-L1 complex formation for cancer therapy. Bioorg. Chem. 2022, 128, 106047. [Google Scholar] [CrossRef]
- Li, Q.; Quan, L.; Lyu, J.; He, Z.; Wang, X.; Meng, J.; Zhao, Z.; Zhu, L.; Liu, X.; Li, H. Discovery of peptide inhibitors targeting human programmed death 1 (PD-1) receptor. Oncotarget 2016, 7, 64967–64976. [Google Scholar] [CrossRef]
- Cheng, X.; Veverka, V.; Radhakrishnan, A.; Waters, L.C.; Muskett, F.W.; Morgan, S.H.; Huo, J.; Yu, C.; Evans, E.J.; Leslie, A.J.; et al. Structure and interactions of the human programmed cell death 1 receptor. J. Biol. Chem. 2013, 288, 11771–11785. [Google Scholar] [CrossRef]
- Gong, J.; Chehrazi-Raffle, A.; Reddi, S.; Salgia, R. Development of PD-1 and PD-L1 inhibitors as a form of cancer immunotherapy: A comprehensive review of registration trials and future considerations. J. Immunother. Cancer 2018, 6, 8. [Google Scholar] [CrossRef]
- Zou, S.; Liu, J.; Sun, Z.; Feng, X.; Wang, Z.; Jin, Y.; Yang, Z. Discovery of hPRDX5-based peptide inhibitors blocking PD-1/PD-L1 interaction through in silico proteolysis and rational design. Cancer Chemother. Pharmacol. 2020, 85, 185–193. [Google Scholar] [CrossRef]
- Chang, H.N.; Liu, B.Y.; Qi, Y.K.; Zhou, Y.; Chen, Y.P.; Pan, K.M.; Li, W.W.; Zhou, X.M.; Ma, W.W.; Fu, C.Y.; et al. Blocking of the PD-1/PD-L1 interaction by a D-peptide antagonist for cancer immunotherapy. Angew. Chem. 2015, 54, 11760–11764. [Google Scholar] [CrossRef]
- Li, C.; Zhang, N.; Zhou, J.; Ding, C.; Jin, Y.; Cui, X.; Pu, K.; Zhu, Y. Peptide blocking of PD-1/PD-L1 interaction for cancer immunotherapy. Cancer Immunol. Res. 2018, 6, 178–188. [Google Scholar] [CrossRef]
- Liu, H.; Zhao, Z.; Zhang, L.; Li, Y.; Jain, A.; Barve, A.; Jin, W.; Liu, Y.; Fetse, J.; Cheng, K. Discovery of low-molecular weight anti-PD-L1 peptides for cancer immunotherapy. J. Immunother. Cancer 2019, 7, 270. [Google Scholar] [PubMed]
- Ganesan, A.; Ahmed, M.; Okoye, I.; Arutyunova, E.; Babu, D.; Turnbull, W.L.; Kundu, J.K.; Shields, J.; Agopsowicz, K.C.; Xu, L.; et al. Comprehensive in vitro characterization of PD-L1 small molecule inhibitors. Sci. Rep. 2019, 9, 12392. [Google Scholar]
- Fetse, J.; Zhao, Z.; Liu, H.; Mamani, U.F.; Mustafa, B. Discovery of Cyclic Peptide Inhibitors Targeting PD-L1 for Cancer Immunotherapy. J. Med. Chem. 2022, 65, 12002–12013. [Google Scholar] [PubMed]
- Zhai, W.; Zhou, X.; Zhai, M.; Li, W.; Ran, Y.; Sun, Y.; Du, J.; Zhao, W.; Xing, L.; Qi, Y.; et al. Blocking of the PD-1/PD-L1 interaction by a novel cyclic peptide inhibitor for cancer immunotherapy. Sci. China Life Sci. 2021, 64, 548–562. [Google Scholar] [PubMed]
- Sun, H.; Chen, D.; Zhan, S.; Wu, W.; Xu, H.; Luo, C.; Su, H.; Feng, Y.; Shao, W.; Wan, A.; et al. Design and Discovery of Natural Cyclopeptide Skeleton Based Programmed Death Ligand 1 Inhibitor as Immune Modulator for Cancer Therapy. J. Med. Chem. 2020, 63, 11286–11301. [Google Scholar]
- Park, S.; Shevlin, E.; Vedvyas, Y.; Zaman, M.; Park, S.; Hsu, Y.S.; Min, I.M.; Jin, M.M. Micromolar affinity CAR T cells to ICAM-1 achieves rapid tumor elimination while avoiding systemic toxicity. Sci. Rep. 2017, 7, 14366. [Google Scholar]
- Hou, X.; Sun, M.; Bao, T.; Xie, X.; Wei, F.; Wang, S. Recent advances in screening active components from natural products based on bioaffinity techniques. Acta Pharm. Sin. B 2020, 10, 1800–1813. [Google Scholar]
- Mooney, C.; Haslam, N.J.; Pollastri, G.; Shields, D.C. Towards the improved discovery and design of functional peptides: Common features of diverse classes permit generalized prediction of bioactivity. PLoS ONE 2012, 7, e45012. [Google Scholar]
- Zhou, P.; Jin, B.; Li, H.; Huang, S.Y. HPEPDOCK: A web server for blind peptide-protein docking based on a hierarchical algorithm. Nucleic Acids Res. 2018, 46, W443–W450. [Google Scholar]
- Chalamaiah, M.; Yu, W.; Wu, J. Immunomodulatory and anticancer protein hydrolysates (peptides) from food proteins: A review. Food Chem. 2017, 15, 205–222. [Google Scholar]
- Pavlicevic, M.; Maestri, E.; Marmiroli, M. Marine Bioactive Peptides-An Overview of Generation, Structure and Application with a Focus on Food Sources. Mar. Drugs 2020, 18, 424. [Google Scholar] [CrossRef] [PubMed]
- Huang, Q.; Zang, X.; Zhang, Z.; Yu, H.; Ding, B.; Li, Z.; Cheng, S.; Zhang, X.; Ali, M.R.K.; Qiu, X.; et al. Study on endogenous inhibitors against PD-L1: cAMP as a potential candidate. Int. J. Biol. Macromol. 2023, 230, 123266. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Hu, J.S.; Xu, J.L.; Shao, C.L.; Wang, G.Y. Biological and Chemical Diversity of Ascidian-Associated Microorganisms. Mar. Drugs 2018, 16, 362. [Google Scholar] [CrossRef] [PubMed]
- Socha, A.M.; Garcia, D.; Sheffer, R.; Rowley, D.C. Antibiotic bisanthraquinones produced by a streptomycete isolated from a cyanobacterium associated with Ecteinascidia turbinata. J. Nat. Prod. 2006, 69, 1070–1073. [Google Scholar] [CrossRef]
- Harunari, E.; Imada, C.; Igarashi, Y.; Fukuda, T.; Terahara, T.; Kobayashi, T. Hyaluromycin, a new hyaluronidase inhibitor of polyketide origin from marine Streptomyces sp. Mar. Drugs 2014, 12, 491–507. [Google Scholar] [CrossRef]
- Paciotti, R.; Agamennone, M.; Coletti, C.; Storchi, L. Characterization of PD-L1 binding sites by a combined FMO/GRID-DRY approach. J. Comput. Aided Mol. Des. 2020, 34, 897–914. [Google Scholar] [CrossRef]
- Zak, K.M.; Grudnik, P.; Guzik, K.; Zieba, B.J.; Musielak, B.; Domling, A.; Dubin, G.; Holak, T.A. Structural basis for small molecule targeting of the programmed death ligand 1 (PD-L1). Oncotarget 2016, 7, 30323–30335. [Google Scholar] [CrossRef]



| ID | Sequence | Candidate Protein Name (Organism) | Mass (Da) |
|---|---|---|---|
| C1 | TTGIVFDSGDGVSHTVPIYEGYALPHAILR | Actin, and CsCA1 (Transparent ascidian) | 3186.55 |
| C2 | KDLYANTVLSGGSTMFPGIADR | Actin (Transparent ascidian) | 2313.57 |
| C3 | TVTFEEFLPMLAQIK | EF-hand domain-containing protein (Transparent ascidian) | 1767.10 |
| C4 | AGFAGDDAPRAVFPSIVGRPR | Actin (Rotaria magnacalcarata, Rotifer, Caenorhabditis latens), Beta-actin (Nematostella vectensis (Starlet sea anemone)), and ATP-binding cassette sub-family B member 10 (Asian catfish) | 2156.39 |
| C5 | LDVVIHTVTYGDR | Uncharacterized protein (Streptomyces albireticuli, Streptomyces sp. ISL-11, Streptomyces tubercidicus) a | 1487.67 |
| C6 | QYEDMFGEDLVDR | Annexin (Transparent ascidian) | 1616.70 |
| C7 | QVNMANHLSKDSR | RING-type E3 ubiquitin transferase (Transparent ascidian) | 1499.65 |
| C8 | PLAIDLLHPSPEEEK | 40S ribosomal protein S27 (Transparent ascidian) | 1687.88 |
| C9 | ISVDVNIPADAIIGR | Transglutaminase-like domain-containing protein (Transparent ascidian) a | 1552.79 |
| C10 | PGAGGKSSTYGR | Transcription elongation factor SPT5 (Transparent ascidian) | 1137.20 |
| C11 | LLDPEDIVVERPDEK | Spectrin beta chain (Transparent ascidian) | 1766.94 |
| C12 | FVDHIMDDQVVEDLTIK | Dolichyl-diphosphooligosaccharide-protein glycosyltransferase subunit 1 (Transparent ascidian) | 2017.27 |
| C13 | HLDIPQMLDAQELVDMAKPDER | Calponin-homology (CH) domain-containing protein, and Actinin, alpha 1 (Transparent ascidian) | 2564.87 |
| C14 | DYEEAVMQNDATVGQLK | Myosin motor domain-containing protein (Transparent ascidian) | 1911.05 |
| C15 | SNAVGITWSPVEGAEK | Uncharacterized LOC100184933 (Transparent ascidian) | 1644.79 |
| C16 | VSYEESIEQLEIVK | Myosin motor domain-containing protein (Transparent ascidian) | 1665.85 |
| C17 | NLLAVAAETDITFPEAEK | 60S acidic ribosomal protein P0, and Large ribosomal subunit protein uL10 (Transparent ascidian) | 1932.12 |
| C18 | IGTSGGLGLDPGTVVITDK | Nucleoside phosphorylase domain-containing protein (Transparent ascidian) | 1800.03 |
| C19 | NDLTLQLQAEQDNLADAEER | Myosin motor domain-containing protein (Transparent ascidian) a | 2286.35 |
| C20 | TIAMDSTER | ATP synthase subunit beta (Transparent ascidian) | 1023.11 |
| C21 | TLHPEVDEDLIER | Obscurin (Transparent ascidian) | 1565.68 |
| C22 | HGIVEDWDLMEK | Actin-related protein 3 (Transparent ascidian) | 1471.64 |
| C23 | EAIANVQDQIADLDK | Large ribosomal subunit protein uL13 (Transparent ascidian) | 1642.77 |
| C24 | LEMQEIQLTEAK | Tropomyosin (Transparent ascidian) a | 1432.64 |
| C25 | LEHINHEK | Eukaryotic translation initiation factor 3 subunit A (Transparent ascidian) | 1019.13 |
| C26 | EILIEDEGELKDIYETFPIDLK | NTR domain-containing protein (Transparent ascidian) | 2622.92 |
| C27 | DVASALGDLINATK | FERM domain-containing protein (Transparent ascidian) | 1387.52 |
| C28 | GINLPGIEVDLPAVSEK | Pyruvate kinase (Transparent ascidian) | 1750.99 |
| C29 | LHEEEIEDLKEQIK | IF rod domain-containing protein (Transparent ascidian) a | 1752.93 |
| C30 | TTNVVVADLSESK | Alcohol dehydrogenase-like N-terminal domain-containing protein (Transparent ascidian) | 1362.48 |
| C31 | KKDEEEIEELR | Troponin T (Transparent ascidian) | 1417.52 |
| C32 | AGVLADLEDK | Myosin type-2 heavy chain 2 (Penicillium cataractarum) a, Myosin motor domain-containing protein, Myosin tail, and Myosin type II heavy chain (Penicillium) a | 1030.12 |
| C33 | EILQGESNVQEVK | Serine/threonine-protein phosphatase (Transparent ascidian) | 1472.62 |
| C34 | IEDLSGGELQR | ATP-binding cassette sub-family E member 1 (Transparent ascidian) | 1216.30 |
| C35 | EILIEDEGELK | NTR domain-containing protein (Transparent ascidian) | 1287.42 |
| C36 | SELDDELGR | IF rod domain-containing protein (Transparent ascidian) | 1033.03 |
| C37 | DGFIDKEDLK | EF-hand domain-containing protein, and Myosin regulatory light chain (Transparent ascidian) | 1179.28 |
| C38 | TSLEEQLEEEEESR | Myosin-10, Myosin tail domain-containing protein (Transparent ascidian) | 1707.69 |
| C39 | IVPPEDGDDEK | RRM domain-containing protein (Transparent ascidian) | 1213.25 |
| C40 | EGIFEEEIK | Tropomyosin (Transparent ascidian) | 1093.20 |
| C41 | DAEEIEKDEQVAAEK | Small ribosomal subunit protein uS2 (Transparent ascidian) | 1703.76 |
| C42 | AEQAEADKK | Beta-lactamase-related domain-containing protein (Patagioenas fasciata monilis), and DAK2 domain fusion protein YloV (Lachnospiraceae bacterium M18-1) | 989.03 |
| ID | Sequence | Energy Score | Ranker Score | Isoelectric Point |
|---|---|---|---|---|
| C1 | TTGIVFDSGDGVSHTVPIYEGYALPHAILR | −213.328 | 0.136 | 5.07 |
| C2 | KDLYANTVLSGGSTMFPGIADR | −205.126 | 0.049 | 6.56 |
| C3 | TVTFEEFLPMLAQIK | −199.778 | 0.218 | 4.15 |
| C4 | AGFAGDDAPRAVFPSIVGRPR | −195.369 | 0.532 | 10.65 |
| C5 | LDVVIHTVTYGDR | −187.135 | 0.118 | 5.04 |
| C6 | QYEDMFGEDLVDR | −185.745 | 0.092 | 3.25 |
| C7 | QVNMANHLSKDSR | −183.074 | 0.184 | 9.83 |
| C8 | PLAIDLLHPSPEEEK | −179.346 | 0.17 | 4.04 |
| C9 | ISVDVNIPADAIIGR | −175.185 | 0.386 | 3.71 |
| C10 | PGAGGKSSTYGR | −173.527 | 0.343 | 10.71 |
| C11 | LLDPEDIVVERPDEK | −172.11 | 0.401 | 3.61 |
| C12 | FVDHIMDDQVVEDLTIK | −170.133 | 0.097 | 3.61 |
| C13 | HLDIPQMLDAQELVDMAKPDER | −169.437 | 0.123 | 3.85 |
| C14 | DYEEAVMQNDATVGQLK | −168.407 | 0.412 | 3.43 |
| C15 | SNAVGITWSPVEGAEK | −167.176 | 0.08 | 4.15 |
| C16 | VSYEESIEQLEIVK | −166.506 | 0.202 | 3.67 |
| C17 | NLLAVAAETDITFPEAEK | −161.999 | 0.28 | 3.54 |
| C18 | IGTSGGLGLDPGTVVITDK | −161.344 | 0.072 | 3.71 |
| C19 | NDLTLQLQAEQDNLADAEER | −160.497 | 0.209 | 3.19 |
| C20 | TIAMDSTER | −158.804 | 0.083 | 3.93 |
| C21 | TLHPEVDEDLIER | −158.752 | 0.082 | 3.79 |
| C22 | HGIVEDWDLMEK | −158.583 | 0.11 | 3.93 |
| C23 | EAIANVQDQIADLDK | −155.919 | 0.126 | 3.32 |
| C24 | LEMQEIQLTEAK | −155.906 | 0.083 | 3.85 |
| C25 | LEHINHEK | −153.529 | 0.17 | 6.06 |
| C26 | EILIEDEGELKDIYETFPIDLK | −152.172 | 0.068 | 3.48 |
| C27 | DVASALGDLINATK | −149.681 | 0.088 | 3.71 |
| C28 | GINLPGIEVDLPAVSEK | −146.833 | 0.088 | 3.69 |
| C29 | LHEEEIEDLKEQIK | −144.492 | 0.298 | 4.08 |
| C30 | TTNVVVADLSESK | −142.701 | 0.079 | 3.93 |
| C31 | KKDEEEIEELR | −142.135 | 0.133 | 4.08 |
| C32 | AGVLADLEDK | −141.924 | 0.094 | 3.54 |
| C33 | EILQGESNVQEVK | −141.694 | 0.136 | 3.85 |
| C34 | IEDLSGGELQR | −140.846 | 0.056 | 3.69 |
| C35 | EILIEDEGELK | −135.713 | 0.119 | 3.44 |
| C36 | SELDDELGR | −134.834 | 0.095 | 3.43 |
| C37 | DGFIDKEDLK | −133.987 | 0.303 | 3.82 |
| C38 | TSLEEQLEEEEESR | −132.64 | 0.249 | 3.37 |
| C39 | IVPPEDGDDEK | −131.107 | 0.032 | 3.25 |
| C40 | EGIFEEEIK | −129.198 | 0.139 | 3.67 |
| C41 | DAEEIEKDEQVAAEK | −124.001 | 0.112 | 3.61 |
| C42 | AEQAEADKK | −119.713 | 0.067 | 4.32 |
| ID | Sequence | Candidate Protein Name (Organism) | Mass |
|---|---|---|---|
| S1 | FLEIFTQR | Annexin (Transparent ascidian) | 1053.22 |
| S2 | VLRDNIQGITKPAIR | Histone H4 (Transparent ascidian) | 1694.01 |
| S3 | AFILPEGVSAER | Body wall muscle protein HR-29 (Sea squirt) | 1288.44 |
| S4 | VWLDPNETSMISNANSR | Ribosomal protein L19 (Transparent ascidian) a | 1934.08 |
| S5 | VDTLMVRNNLR | Tetratricopeptide repeat protein (Streptomyces sp. Rer75) a, and Oxidoreductase (Actinomadura sp. WAC 06369) a | 1330.55 |
| S6 | VAPEEHPVLLTEAPLNPKANR | Beta-actin, and Alpha-actin (Rotaria magnacalcarata), Actin (Rotaria socialis) | 2295.58 |
| S7 | AVMSLQMEMQQIMK | Glutamine--fructose-6-phosphate transaminase (Transparent ascidian) a | 1668.08 |
| S8 | GYEEWLISEMR | Actinin, alpha 1, Calponin-homology (CH) domain-containing protein, and EF-hand domain-containing protein (Transparent ascidian) a | 1412.57 |
| S9 | TVQTLNLEIDSMR | IF rod domain-containing protein, and Glial fibrillary acidic protein (Transparent ascidian) a | 1519.72 |
| S10 | VLGSGTNLDSAR | L-lactate dehydrogenase (Transparent ascidian) | 1189.26 |
| S11 | FTGMLSMLDDPEPFAR | Body wall muscle protein HR-29 (Sea squirt) | 1827.06 |
| S12 | EELDMEHRRSK | Uncharacterized protein (Transparent ascidian) | 1429.55 |
| S13 | DGILQIDAPVAVAIDNK | Body wall muscle protein HR-29 (Sea squirt) | 1751.98 |
| S14 | MTEQWMK | Carboxylic ester hydrolase (Transparent ascidian) | 953.14 |
| S15 | LVMVEAELERGEER | Tropomyosin (Transparent ascidian) | 1659.85 |
| S16 | LLEAQIATGGLIDPR | Uncharacterized protein (Transparent ascidian) | 1566.80 |
| S17 | KLETLQEELELLK | IF rod domain-containing protein, and Glial fibrillary acidic protein (Transparent ascidian) a | 1585.83 |
| S18 | GVDLDQLLDMSR | Small ribosomal subunit protein uS19 (Transparent ascidian) | 1361.50 |
| S19 | ANAEVANWR | Myosin motor domain-containing protein (Transparent ascidian) | 1030.09 |
| S20 | MAGTSDCVKR | Uncharacterized LOC100177244 (Transparent ascidian) | 1067.24 |
| S21 | LSGGDIESYLLEK | Myosin motor domain-containing protein (Transparent ascidian) | 1423.56 |
| S22 | NIKEGDIVKR | ATP synthase subunit alpha (Transparent ascidian) | 1171.37 |
| S23 | VVSQTEDVR | Glial fibrillary acidic protein (Transparent ascidian) | 1032.11 |
| S24 | EDDVQQMNPPK | Myosin motor domain-containing protein (Transparent ascidian) a | 1300.40 |
| ID | Sequence | Energy Score | Ranker Score | Isoelectric Point |
|---|---|---|---|---|
| S1 | FLEIFTQR | −191.311 | 0.335 | 6.62 |
| S2 | VLRDNIQGITKPAIR | −182.923 | 0.15 | 11.23 |
| S3 | AFILPEGVSAER | −178.808 | 0.255 | 4.15 |
| S4 | VWLDPNETSMISNANSR | −175.773 | 0.248 | 3.93 |
| S5 | VDTLMVRNNLR | −175.323 | 0.154 | 10.65 |
| S6 | VAPEEHPVLLTEAPLNPKANR | −175.164 | 0.221 | 5.36 |
| S7 | AVMSLQMEMQQIMK | −174.227 | 0.215 | 6.91 |
| S8 | GYEEWLISEMR | −172.518 | 0.575 | 3.85 |
| S9 | TVQTLNLEIDSMR | −170.263 | 0.09 | 3.93 |
| S10 | VLGSGTNLDSAR | −169.041 | 0.185 | 6.61 |
| S11 | FTGMLSMLDDPEPFAR | −167.011 | 0.316 | 3.54 |
| S12 | EELDMEHRRSK | −165.086 | 0.128 | 5.39 |
| S13 | DGILQIDAPVAVAIDNK | −164.379 | 0.282 | 3.41 |
| S14 | MTEQWMK | −160.485 | 0.349 | 6.61 |
| S15 | LVMVEAELERGEER | −158.101 | 0.061 | 3.97 |
| S16 | LLEAQIATGGLIDPR | −157.057 | 0.246 | 3.93 |
| S17 | KLETLQEELELLK | −156.352 | 0.118 | 4.15 |
| S18 | GVDLDQLLDMSR | −155.986 | 0.374 | 3.41 |
| S19 | ANAEVANWR | −155.121 | 0.413 | 6.93 |
| S20 | MAGTSDCVKR | −153.381 | 0.307 | 8.6 |
| S21 | LSGGDIESYLLEK | −149.843 | 0.315 | 3.69 |
| S22 | NIKEGDIVKR | −149.659 | 0.127 | 9.53 |
| S23 | VVSQTEDVR | −144.2 | 0.063 | 3.93 |
| S24 | EDDVQQMNPPK | −134.958 | 0.22 | 3.54 |
| Ligand | PD-L1 b | Interaction | Distance (Å) | E (kcal/mol) | ||
|---|---|---|---|---|---|---|
| His6 | OE1 | AGlu60 | H-donor | 2.83 | −2.3 | |
| Thr7 | OE1 | AGlu60 | H-donor | 2.80 | −3.5 | |
| Tyr10 | O | AGlu60 | H-donor | 2.90 | −2.3 | |
| Arg13 | OE1 | AGlu60 | H-donor | 3.24 | −2.6 | |
| Arg13 | O | AGlu60 | H-donor | 2.72 | −3.1 | |
| Asp2 | NZ | BLys129 | H-acceptor | 2.62 | −12.7 | |
| C5 | Asp12 | OG1 | BThr127 | H-acceptor | 2.75 | −3.8 |
| Asp12 | NE | BArg125 | H-acceptor | 2.79 | −4.8 | |
| Asp12 | NH2 | BArg125 | H-acceptor | 3.12 | −2.2 | |
| Asp12 | NH2 | BArg125 | H-acceptor | 2.95 | −1.6 | |
| Asp2 | NZ | BLys129 | ionic | 2.62 | −7.6 | |
| Asp12 | NE | BArg125 | ionic | 2.79 | −6.0 | |
| Asp12 | NH2 | BArg125 | ionic | 3.12 | −3.7 | |
| Arg13 | OE1 | AGlu60 | ionic | 3.24 | −3.1 | |
| Val1 | OD2 | AAsp26 | H-donor | 2.78 | −14.1 | |
| Arg15 | O | AArg125 | H-donor | 2.82 | −4.8 | |
| Arg15 | OD2 | AAsp26 | H-donor | 2.75 | −4.6 | |
| Asp4 | OG1 | AThr127 | H-acceptor | 2.61 | −3.5 | |
| Gln7 | NZ | ALys129 | H-acceptor | 2.78 | −11.5 | |
| Thr10 | NE2 | BHis78 | H-acceptor | 2.88 | −3.0 | |
| Arg15 | N | AArg125 | H-acceptor | 3.43 | −1.5 | |
| S2 | Arg15 | NH2 | AArg125 | H-acceptor | 2.67 | −3.9 |
| Val1 | OD1 | AAsp26 | ionic | 2.69 | −7.0 | |
| Val1 | OD2 | AAsp26 | ionic | 2.78 | −6.1 | |
| Arg15 | OD2 | AAsp26 | ionic | 3.75 | −1.1 | |
| Arg15 | OD2 | AAsp26 | ionic | 2.75 | −6.4 | |
| Arg15 | NE | AArg125 | ionic | 3.93 | −0.6 | |
| Arg15 | NH2 | AArg125 | ionic | 3.89 | −0.7 | |
| Arg15 | NE | AArg125 | ionic | 3.43 | −2.2 | |
| Arg15 | NH2 | AArg125 | ionic | 2.67 | −7.1 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, Q.; Zang, X.; Jin, X.; Liu, Q.; Zhang, X.; Li, X.; Zhao, L.; Lv, Z. Discovery of PD-L1 Peptide Inhibitors from Ascidian Enzymatic Hydrolysates by Affinity Ultrafiltration Coupled to NanoLC-MS/MS. Mar. Drugs 2025, 23, 137. https://doi.org/10.3390/md23040137
Huang Q, Zang X, Jin X, Liu Q, Zhang X, Li X, Zhao L, Lv Z. Discovery of PD-L1 Peptide Inhibitors from Ascidian Enzymatic Hydrolysates by Affinity Ultrafiltration Coupled to NanoLC-MS/MS. Marine Drugs. 2025; 23(4):137. https://doi.org/10.3390/md23040137
Chicago/Turabian StyleHuang, Qiuyang, Xiaoling Zang, Xinyu Jin, Qian Liu, Xin Zhang, Xinyu Li, Lizhen Zhao, and Zhihua Lv. 2025. "Discovery of PD-L1 Peptide Inhibitors from Ascidian Enzymatic Hydrolysates by Affinity Ultrafiltration Coupled to NanoLC-MS/MS" Marine Drugs 23, no. 4: 137. https://doi.org/10.3390/md23040137
APA StyleHuang, Q., Zang, X., Jin, X., Liu, Q., Zhang, X., Li, X., Zhao, L., & Lv, Z. (2025). Discovery of PD-L1 Peptide Inhibitors from Ascidian Enzymatic Hydrolysates by Affinity Ultrafiltration Coupled to NanoLC-MS/MS. Marine Drugs, 23(4), 137. https://doi.org/10.3390/md23040137







