Sponge Whirl-Pak Sampling Method and Droplet Digital RT-PCR Assay for Monitoring of SARS-CoV-2 on Surfaces in Public and Working Environments
Abstract
:1. Introduction
2. Materials and Methods
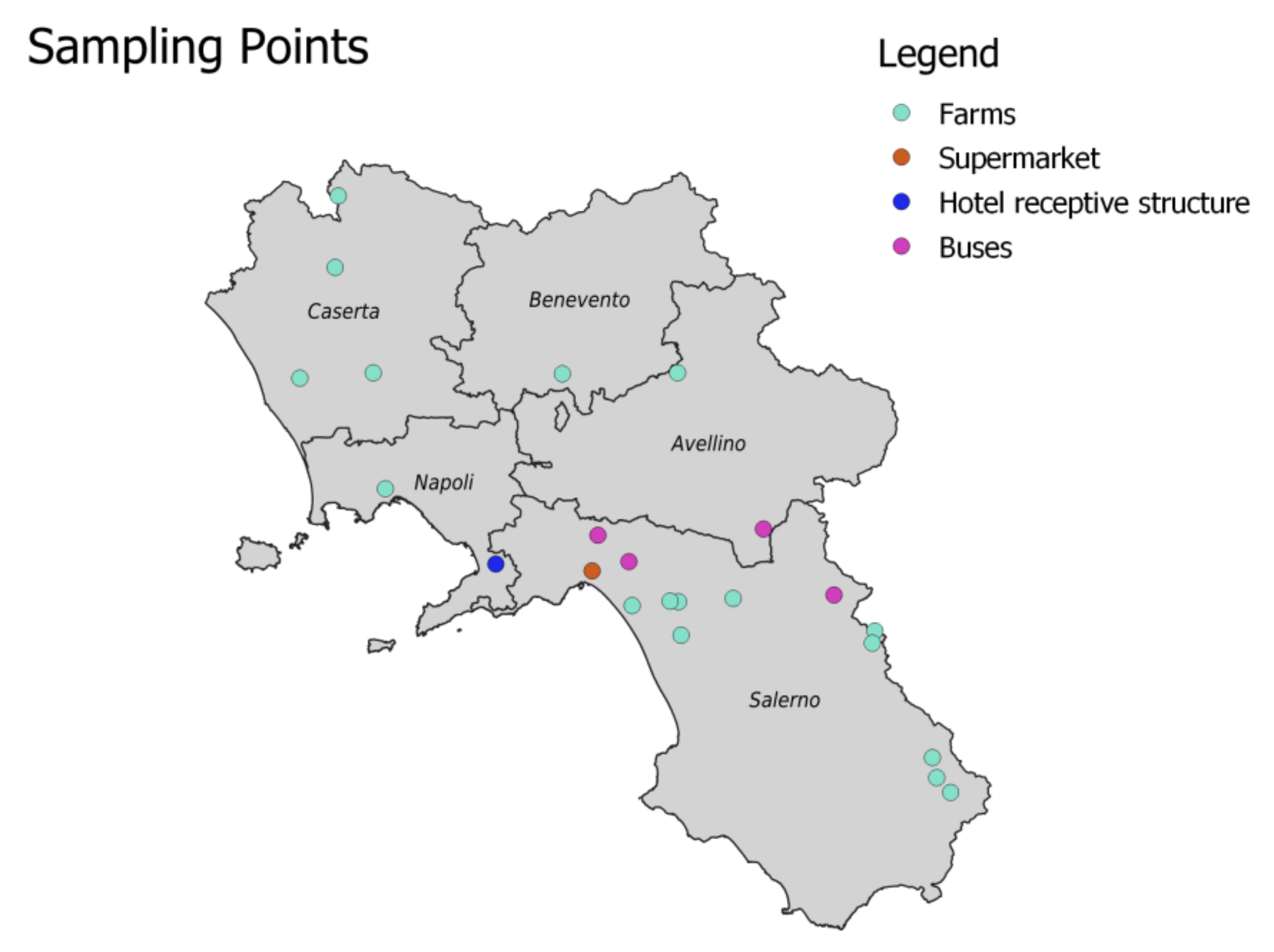
2.1. Sampling
2.2. Nucleic Acids Extraction
2.3. SARS-CoV-2 Detection by Real-Time RT-PCR
2.4. SARS-CoV-2 Detection in Environmental Samples by Droplet Digital RT-PCR (dd RT-PCR)
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- She, J.; Jiang, J.; Ye, L.; Hu, L.; Bai, C.; Song, Y. 2019 novel coronavirus of pneumonia in Wuhan, China: Emerging attack and management strategies. Clin. Transl. Med. 2020, 9, 19. [Google Scholar] [CrossRef] [PubMed]
- Andersen, K.G.; Rambaut, A.; Lipkin, W.I.; Holmes, E.C.; Garry, R.F. The proximal origin of SARS-CoV-2. Nat. Med. 2020, 26, 450–452. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Al-Qahtani, A.A. Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2): Emergence, history, basic and clinical aspects. Saudi J. Biol. Sci. 2020, 27, 2531–2538. [Google Scholar] [CrossRef] [PubMed]
- Cucinotta, D.; Vanelli, M. WHO Declares COVID-19 a Pandemic. Acta Bio Med. Atenei Parm. 2020, 91, 157–160. [Google Scholar] [CrossRef]
- Morawska, L.; Johnson, G.R.; Ristovski, Z.D.; Hargreaves, M.; Mengersen, K.; Corbett, S.; Chao, C.Y.H.; Li, Y.; Katoshevski, D. Size distribution and sites of origin of droplets expelled from the human respiratory tract during expiratory activities. J. Aerosol Sci. 2009, 40, 256–269. [Google Scholar] [CrossRef] [Green Version]
- Stadnytskyi, V.; Bax, C.E.; Bax, A.; Anfinrud, P. The airborne lifetime of small speech droplets and their potential importance in SARS-CoV-2 transmission. Proc. Natl. Acad. Sci. USA 2020, 117, 11875–11877. [Google Scholar] [CrossRef]
- Santarpia, J.L.; Rivera, D.N.; Herrera, V.L.; Morwitzer, M.J.; Creager, H.M.; Santarpia, G.W.; Crown, K.K.; Brett-Major, D.M.; Schnaubelt, E.R.; Broadhurst, M.J.; et al. Aerosol and surface contamination of SARS-CoV-2 observed in quarantine and isolation care. Sci. Rep. 2020, 10, 12732. [Google Scholar] [CrossRef]
- Kenneth McIntosh, M.; Martin, S.; Hirsch, M.; Allyson Bloom, M. Coronavirus disease 2019 (COVID-19). UpToDate Hirsch MS Bloom 2020, 5, 873. [Google Scholar]
- Kraay, A.N.M.; Hayashi, M.A.L.; Hernandez-Ceron, N.; Spicknall, I.H.; Eisenberg, M.C.; Meza, R.; Eisenberg, J.N.S. Fomite-mediated transmission as a sufficient pathway: A comparative analysis across three viral pathogens 11 Medical and Health Sciences 1117 Public Health and Health Services. BMC Infect. Dis. 2018, 18, 540. [Google Scholar]
- Rusin, P.; Maxwell, S.; Gerba, C. Comparative surface-to-hand and fingertip-to-mouth transfer efficiency of gram-positive bacteria, gram-negative bacteria, and phage. J. Appl. Microbiol. 2002, 93, 585–592. [Google Scholar] [CrossRef]
- Mohammad, N.S.; Nazli, R.; Zafar, H.; Fatima, S. Effects of lipid based multiple micronutrients supplement on the birth outcome of underweight pre-eclamptic women: A randomized clinical trial. Pakistan, J. Med. Sci. 2022, 38, 219–226. [Google Scholar] [CrossRef]
- Biryukov, J.; Boydston, J.A.; Dunning, R.A.; Yeager, J.J.; Wood, S.; Ferris, A.; Miller, D.; Weaver, W.; Zeitouni, N.E.; Freeburger, D.; et al. SARS-CoV-2 is rapidly inactivated at high temperature. Environ. Chem. Lett. 2021, 19, 1773–1777. [Google Scholar] [CrossRef]
- Biryukov, J.; Boydston, J.A.; Dunning, R.A.; Yeager, J.J.; Wood, S.; Reese, A.L.; Ferris, A.; Miller, D.; Weaver, W.; Zeitouni, N.E.; et al. Increasing Temperature and Relative Humidity Accelerates Inactivation of SARS-CoV-2 on Surfaces. mSphere 2020, 5, e00441-20. [Google Scholar] [CrossRef]
- Casanova, L.M.; Jeon, S.; Rutala, W.A.; Weber, D.J.; Sobsey, M.D. Effects of Air Temperature and Relative Humidity on Coronavirus Survival on Surfaces. Appl. Environ. Microbiol. 2010, 76, 2712–2717. [Google Scholar] [CrossRef] [Green Version]
- Wang, D.; Freed, D.C.; He, X.; Li, F.; Tang, A.; Cox, K.S.; Dubey, S.A.; Cole, S.; Medi, M.B.; Liu, Y.; et al. A replication-defective human cytomegalovirus vaccine for prevention of congenital infection. Sci. Transl. Med. 2016, 8, 747–766. [Google Scholar] [CrossRef]
- Raiteux, J.; Eschlimann, M.; Marangon, A.; Rogée, S.; Dadvisard, M.; Taysse, L.; Larigauderie, G. Inactivation of SARS-CoV-2 by Simulated Sunlight on Contaminated Surfaces. Microbiol. Spectr. 2021, 9, e00333-21. [Google Scholar] [CrossRef]
- Chin, A.W.H.; Chu, J.T.S.; Perera, M.R.A.; Hui, K.P.Y.; Yen, H.L.; Chan, M.C.W.; Peiris, M.; Poon, L.L.M. Stability of SARS-CoV-2 in different environmental conditions. Lancet Microbe 2020, 1, e10. [Google Scholar] [CrossRef]
- Van Doremalen, N.; Bushmaker, T.; Morris, D.H.; Holbrook, M.G.; Gamble, A.; Williamson, B.N.; Tamin, A.; Harcourt, J.L.; Thornburg, N.J.; Gerber, S.I.; et al. Aerosol and surface stability of HCoV-19 (SARS-CoV-2) compared to SARS-CoV-1. medRxiv 2020. [Google Scholar] [CrossRef] [Green Version]
- Bonil, L.; Lingas, G.; Coupeau, D.; Lucet, J.C.; Guedj, J.; Visseaux, B.; Muylkens, B. Survival of SARS-CoV-2 on non-porous materials in an experimental setting representative of fomites. Coatings 2021, 11, 371. [Google Scholar] [CrossRef]
- Hirose, R.; Miyazaki, H.; Bandou, R.; Watanabe, N.; Yoshida, T.; Daidoji, T.; Itoh, Y.; Nakaya, T. Stability of SARS-CoV-2 and influenza virus varies across different paper types. J. Infect. Chemother. 2022, 28, 252–256. [Google Scholar] [CrossRef]
- Di Novo, N.G.; Carotenuto, A.R.; Mensitieri, G.; Fraldi, M.; Pugno, N.M. Modeling of Virus Survival Time in Respiratory Droplets on Surfaces: A New Rational Approach for Antivirus Strategies. Front. Mater. 2021, 8, 56. [Google Scholar] [CrossRef]
- Wu, S.; Wang, Y.; Jin, X.; Tian, J.; Liu, J.; Mao, Y. Environmental contamination by SARS-CoV-2 in a designated hospital for coronavirus disease 2019. Am. J. Infect. Control 2020, 48, 910–914. [Google Scholar] [CrossRef]
- Chia, P.Y.; Coleman, K.K.; Tan, Y.K.; Ong, S.W.X.; Gum, M.; Lau, S.K.; Lim, X.F.; Lim, A.S.; Sutjipto, S.; Lee, P.H.; et al. Detection of air and surface contamination by SARS-CoV-2 in hospital rooms of infected patients. Nat. Commun. 2020, 11, 2800. [Google Scholar] [CrossRef]
- Li, Y.H.; Fan, Y.Z.; Jiang, L.; Wang, H.B. Aerosol and environmental surface monitoring for SARS-CoV-2 RNA in a designated hospital for severe COVID-19 patients. Epidemiol. Infect. 2020, 148, e154. [Google Scholar] [CrossRef]
- Razzini, K.; Castrica, M.; Menchetti, L.; Maggi, L.; Negroni, L.; Orfeo, N.V.; Pizzoccheri, A.; Stocco, M.; Muttini, S.; Balzaretti, C.M. SARS-CoV-2 RNA detection in the air and on surfaces in the COVID-19 ward of a hospital in Milan, Italy. Sci. Total Environ. 2020, 742, 140540. [Google Scholar] [CrossRef]
- Coil, D.A.; Albertson, T.; Banerjee, S.; Brennan, G.; Campbell, A.J.; Cohen, S.H.; Dandekar, S.; Díaz-Muñoz, S.L.; Eisen, J.A.; Goldstein, T.; et al. SARS-CoV-2 detection and genomic sequencing from hospital surface samples collected at UC Davis. PLoS ONE 2021, 16, e0253578. [Google Scholar] [CrossRef]
- Alam, S.; Darul Ehsan, S. Environmental surface sampling of SARS-CoV-2 in selected hospitals in Malaysia. Trop. Biomed. 2021, 38, 462–468. [Google Scholar] [CrossRef]
- Ben-Shmuel, A.; Brosh-Nissimov, T.; Glinert, I.; Bar-David, E.; Sittner, A.; Poni, R.; Cohen, R.; Achdout, H.; Tamir, H.; Yahalom-Ronen, Y.; et al. Detection and infectivity potential of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) environmental contamination in isolation units and quarantine facilities. Clin. Microbiol. Infect. 2020, 26, 1658–1662. [Google Scholar] [CrossRef]
- Moore, G.; Rickard, H.; Stevenson, D.; Aranega-Bou, P.; Pitman, J.; Crook, A.; Davies, K.; Spencer, A.; Burton, C.; Easterbrook, L.; et al. Detection of SARS-CoV-2 within the healthcare environment: A multi-centre study conducted during the first wave of the COVID-19 outbreak in England. J. Hosp. Infect. 2021, 108, 189–196. [Google Scholar] [CrossRef]
- Marcenac, P.; Park, G.W.; Duca, L.M.; Lewis, N.M.; Dietrich, E.A.; Barclay, L.; Tamin, A.; Harcourt, J.L.; Thornburg, N.J.; Rispens, J.; et al. Detection of SARS-CoV-2 on surfaces in households of persons with COVID-19. Int. J. Environ. Res. Public Health 2021, 18, 8154. [Google Scholar] [CrossRef]
- Hu, X.; Xing, Y.; Ni, W.; Zhang, F.; Lu, S.; Wang, Z.; Gao, R.; Jiang, F. Environmental contamination by SARS-CoV-2 of an imported case during incubation period. Sci. Total Environ. 2020, 742, 140620. [Google Scholar] [CrossRef] [PubMed]
- Ong, S.W.X.; Tan, Y.K.; Chia, P.Y.; Lee, T.H.; Ng, O.T.; Wong, M.S.Y.; Marimuthu, K. Air, Surface Environmental, and Personal Protective Equipment Contamination by Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) from a Symptomatic Patient. JAMA-J. Am. Med. Assoc. 2020, 323, 1610–1612. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yamagishi, T.; Ohnishi, M.; Matsunaga, N.; Kakimoto, K.; Kamiya, H.; Okamoto, K.; Suzuki, M.; Gu, Y.; Sakaguchi, M.; Tajima, T.; et al. Environmental sampling for severe acute respiratory syndrome coronavirus 2 during a COVID-19 outbreak on the diamond princess cruise ship. J. Infect. Dis. 2020, 222, 1098–1102. [Google Scholar] [CrossRef] [PubMed]
- Silva, S.J.R.; Nascimento, J.C.F.; Santos Reis, W.P.M.; Silva, C.T.A.; Silva, P.G.; Mendes, R.P.G.; Mendonça, A.A.; Santos, B.N.R.; Magalhães, J.J.F.; Kohl, A.; et al. Widespread contamination of SARS-CoV-2 on highly touched surfaces in Brazil during the second wave of the COVID-19 pandemic. Environ. Microbiol. 2021, 23, 7382–7395. [Google Scholar] [CrossRef] [PubMed]
- Harvey, A.P.; Fuhrmeister, E.R.; Cantrell, M.E.; Pitol, A.K.; Swarthout, J.M.; Powers, J.E.; Nadimpalli, M.L.; Julian, T.R.; Pickering, A.J. Longitudinal Monitoring of SARS-CoV-2 RNA on High-Touch Surfaces in a Community Setting. Environ. Sci. Technol. Lett. 2021, 8, 168–175. [Google Scholar] [CrossRef]
- Kozer, E.; Rinott, E.; Kozer, G.; Bar-Haim, A.; Benveniste-Levkovitz, P.; Klainer, H.; Perl, S.; Youngster, I. Presence of SARS-CoV-2 RNA on playground surfaces and water fountains. Epidemiol. Infect. 2021, 149, e67. [Google Scholar] [CrossRef]
- Guadalupe, J.J.; Rojas, M.I.; Pozo, G.; Erazo-Garcia, M.P.; Vega-Polo, P.; Terán-Velástegui, M.; Rohwer, F.; Torres, M.D.L. Presence of SARS-CoV-2 RNA on surfaces of public places and a transportation system located in a densely populated urban area in South America. Viruses 2022, 14, 19. [Google Scholar] [CrossRef]
- Moreno, T.; Pintó, R.M.; Bosch, A.; Moreno, N.; Alastuey, A.; Minguillón, M.C.; Anfruns-Estrada, E.; Guix, S.; Fuentes, C.; Buonanno, G.; et al. Tracing surface and airborne SARS-CoV-2 RNA inside public buses and subway trains. Environ. Int. 2021, 147, 106326. [Google Scholar] [CrossRef]
- Di Carlo, P.; Chiacchiaretta, P.; Sinjari, B.; Aruffo, E.; Stuppia, L.; De Laurenzi, V.; Di Tomo, P.; Pelusi, L.; Potenza, F.; Veronese, A.; et al. Air and surface measurements of SARS-CoV-2 inside a bus during normal operation. PLoS ONE 2020, 15, e0235943. [Google Scholar] [CrossRef]
- Caggiano, G.; Triggiano, F.; Apollonio, F.; Diella, G.; Lopuzzo, M.; D’ambrosio, M.; Fasano, F.; Stefanizzi, P.; Sorrenti, G.T.; Magarelli, P.; et al. SARS-CoV-2 rna and supermarket surfaces: A real or presumed threat? Int. J. Environ. Res. Public Health 2021, 18, 404. [Google Scholar] [CrossRef]
- Montagna, M.T.; De Giglio, O.; Calia, C.; Pousis, C.; Apollonio, F.; Campanale, C.; Diella, G.; Lopuzzo, M.; Marzella, A.; Triggiano, F.; et al. First Detection of Severe Acute Respiratory Syndrome Coronavirus 2 on the Surfaces of Tourist-Recreational Facilities in Italy. Int. J. Environ. Res. Public Health 2021, 18, 3252. [Google Scholar] [CrossRef]
- Caggiano, G.; Apollonio, F.; Triggiano, F.; Diella, G.; Stefanizzi, P.; Lopuzzo, M.; D’ambrosio, M.; Bartolomeo, N.; Barbuti, G.; Sorrenti, G.T.; et al. SARS-CoV-2 and public transport in Italy. Int. J. Environ. Res. Public Health 2021, 18, 1415. [Google Scholar] [CrossRef]
- ISO. UNI EN ISO 15216-2:2019; Microbiology of the Food Chain—Horizontal Method for Determination of Hepatitis A Virus and Norovirus Using Real-Time RT-PCR—Part 2: Method for Detection; International Organization for Standardization: Geneva, Switzerland, 2019. [Google Scholar]
- Corman, V.M.; Landt, O.; Kaiser, M.; Molenkamp, R.; Meijer, A.; Chu, D.K.; Bleicker, T.; Brünink, S.; Schneider, J.; Schmidt, M.L.; et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eurosurveillance 2020, 25, 23. [Google Scholar] [CrossRef] [Green Version]
- Dargahi, A.; Jeddi, F.; Vosoughi, M.; Karami, C.; Hadisi, A.; Ahamad Mokhtari, S.; Ghobadi, H.; Alighadri, M.; Haghighi, S.B.; Sadeghi, H. Investigation of SARS-CoV-2 virus in environmental surface. Environ. Res. 2021, 195, 110765. [Google Scholar] [CrossRef]
- Goldman, E. Exaggerated risk of transmission of COVID-19 by fomites. Lancet Infect. Dis. 2020, 20, 892–893. [Google Scholar] [CrossRef]
- Samudrala, P.K.; Kumar, P.; Choudhary, K.; Thakur, N.; Wadekar, G.S.; Dayaramani, R.; Agrawal, M.; Alexander, A. Virology, pathogenesis, diagnosis and in-line treatment of COVID-19. Eur. J. Pharmacol. 2020, 883, 173375. [Google Scholar] [CrossRef]
- Abdel-Hady, A.; Worth Calfee, M.; Aslett, D.; Lee, S.D.; Wyrzykowska-Ceradini, B.; Robbins Delafield, F.; May, K.; Touati, A. Alternative fast analysis method for cellulose sponge surface sampling wipes with low concentrations of Bacillus Spores. J. Microbiol. Methods 2019, 156, 5–8. [Google Scholar] [CrossRef]
- Mancusi, A.; Capuano, F.; Girardi, S.; Di Maro, O.; Suffredini, E.; Di Concilio, D.; Vassallo, L.; Cuomo, M.C.; Tafuro, M.; Signorelli, D.; et al. Detection of SARS-CoV-2 RNA in Bivalve Mollusks by Droplet Digital RT-PCR (dd RT-PCR). Int. J. Environ. Res. Public Health 2022, 19, 943. [Google Scholar] [CrossRef]
- Kraay, A.N.M.; Hayashi, M.A.L.; Berendes, D.M.; Sobolik, J.S.; Leon, J.S.; Lopman, B.A. Risk of fomite-mediated transmission of SARS-CoV-2 in child daycares, schools, and offices: A modeling study. medRxiv 2020. [Google Scholar] [CrossRef]
- Park, G.W.; Chhabra, P.; Vinjé, J. Swab Sampling Method for the Detection of Human Norovirus on Surfaces. J. Vis. Exp. 2017, 120, 55205. [Google Scholar] [CrossRef]
- Di Maria, F.; La Rosa, G.; Bonato, T.; Pivato, A.; Piazza, R.; Mancini, P.; Bonanno Ferraro, G.; Veneri, C.; Iaconelli, M.; Beccaloni, E.; et al. An innovative approach for the non-invasive surveillance of communities and early detection of SARS-CoV-2 via solid waste analysis. Sci. Total Environ. 2021, 801, 149743. [Google Scholar] [CrossRef]
- Hindson, C.M.; Chevillet, J.R.; Briggs, H.A.; Gallichotte, E.N.; Ruf, I.K.; Hindson, B.J.; Vessella, R.L.; Tewari, M. Absolute quantification by droplet digital PCR versus analog real-time PCR. Nat. Methods 2013, 10, 1003–1005. [Google Scholar] [CrossRef]
- Park, C.; Lee, J.; ul Hassan, Z.; Ku, K.B.; Kim, S.J.; Kim, H.G.; Park, E.C.; Park, G.S.; Park, D.; Baek, S.H.; et al. Comparison of digital pcr and quantitative pcr with various SARS-CoV-2 primer-probe sets. J. Microbiol. Biotechnol. 2021, 31, 358–367. [Google Scholar] [CrossRef]
- Tenhagen, B.A.; Arth, O.; Bandick, N.; Fetsch, A. Comparison of three sampling methods for the quantification of methicillin-resistant Staphylococcus aureus on the surface of pig carcases. Food Control 2011, 22, 643–645. [Google Scholar] [CrossRef]
- Nguyen, H.; Morgan, D.A.F.; Cull, S.; Benkovich, M.; Forwood, M.R. Sponge swabs increase sensitivity of sterility testing of processed bone and tendon allografts. J. Ind. Microbiol. Biotechnol. 2011, 38, 1127–1132. [Google Scholar] [CrossRef]
- Engelhardt, N.E.P.; Foster, N.F.; Hong, S.; Riley, T.V.; McGechie, D.B. Comparison of two environmental sampling tools for the detection of Clostridium difficile spores on hard bathroom surfaces in the hospital setting. J. Hosp. Infect. 2017, 96, 295–296. [Google Scholar] [CrossRef]
- Gallina, S.; Bianchi, D.M.; Ru, G.; Maurella, C.; Barzanti, P.; Baioni, E.; Virgilio, S.; Mioni, R.; Lanni, L.; Migliazzo, A.; et al. Microbiological recovery from bovine, swine, equine, and ovine carcasses: Comparison of excision, sponge and swab sampling methods. Food Control 2015, 50, 919–924. [Google Scholar] [CrossRef]
- Yu, F.; Yan, L.; Wang, N.; Yang, S.; Wang, L.; Tang, Y.; Gao, G.; Wang, S.; Ma, C.; Xie, R.; et al. Quantitative detection and viral load analysis of SARS-CoV-2 in infected patients. Clin. Infect. Dis. 2020, 71, 793–798. [Google Scholar] [CrossRef] [Green Version]
- Cassinari, K.; Alessandri-Gradt, E.; Chambon, P.; Charbonnier, F.; Gracias, S.; Beaussire, L.; Alexandre, K.; Sarafan-Vasseur, N.; Houdayer, C.; Etienne, M.; et al. Assessment of Multiplex Digital Droplet RT-PCR as a Diagnostic Tool for SARS-CoV-2 Detection in Nasopharyngeal Swabs and Saliva Samples. Clin. Chem. 2021, 67, 736–741. [Google Scholar] [CrossRef]
- Long, S.; Berkemeier, B. Ultrasensitive detection and quantification of viral nucleic acids with Raindance droplet digital PCR (ddPCR). Methods 2021, 201, 49–64. [Google Scholar] [CrossRef] [PubMed]
- Sedlak, R.H.; Cook, L.; Cheng, A.; Magaret, A.; Jerome, K.R. Clinical utility of droplet digital PCR for human cytomegalovirus. J. Clin. Microbiol. 2014, 52, 2844–2848. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dube, S.; Qin, J.; Ramakrishnan, R. Mathematical Analysis of Copy Number Variation in a DNA Sample Using Digital PCR on a Nanofluidic Device. PLoS ONE 2008, 3, e2876. [Google Scholar] [CrossRef] [PubMed]
- Teo, C.R.L.; Rajan-Babu, I.S.; Law, H.Y.; Lee, C.G.; Chong, S.S. Tolerance of Droplet-Digital PCR vs. Real-Time Quantitative PCR to Inhibitory Substances. Clin. Chem. 2013, 59, 1670–1672. [Google Scholar]
- Singh, M.; Sadat, A.; Abdi, R.; Colaruotolo, L.A.; Francavilla, A.; Petker, K.; Nasr, P.; Moraveji, M.; Cruz, G.; Huang, Y.; et al. Detection of SARS-CoV-2 on surfaces in food retailers in Ontario. Curr. Res. Food Sci. 2021, 4, 598–602. [Google Scholar] [CrossRef] [PubMed]
- Chaintoutis, S.C.; Thomou, Z.; Mouchtaropoulou, E.; Tsiolas, G.; Chassalevris, T.; Stylianaki, I.; Lagou, M.; Michailidou, S.; Moutou, E.; Koenen, J.J.H.; et al. Outbreaks of SARS-CoV-2 in naturally infected mink farms: Impact, transmission dynamics, genetic patterns, and environmental contamination. PLoS Pathog. 2021, 17, e1009883. [Google Scholar] [CrossRef]
- De Rooij, M.M.T.; Hakze-Van Der Honing, R.W.; Hulst, M.M.; Harders, F.; Engelsma, M.; Van De Hoef, W.; Meliefste, K.; Nieuwenweg, S.; Oude Munnink, B.B.; Van Schothorst, I.; et al. Occupational and environmental exposure to SARS-CoV-2 in and around infected mink farms. Occup. Environ. Med. 2021, 78, 893–899. [Google Scholar] [CrossRef]
- Montse, M.; José, L.D. Contamination of inert surfaces by SARS-CoV-2: Persistence, stability and infectivity. A review. Environ. Res. 2021, 193, 110559. [Google Scholar]
- Kampf, G.; Todt, D.; Pfaender, S.; Steinmann, E. Persistence of coronaviruses on inanimate surfaces and their inactivation with biocidal agents. J. Hosp. Infect. 2020, 104, 246–251. [Google Scholar] [CrossRef] [Green Version]
| Sampling Location | District | Environmental Samples | Animal Swabs | Human Swabs |
|---|---|---|---|---|
| Buffaloes farm_1 | Caserta | 1 | 20 | 7 |
| Buffaloes farm_2 | Caserta | 1 | 22 | 11 |
| Buffaloes farm_3 | Caserta | 1 | 23 | 12 |
| Buffaloes farm_4 | Caserta | 10 | 32 | 9 |
| Cattle farm | Salerno | - | 10 | 1 |
| Goat and cattle farm | Salerno | 2 | 17 | 2 |
| Goat farm_1 | Salerno | 1 | 6 | - |
| Goat farm_2 | Salerno | - | 10 | 1 |
| Hen farm_1 | Salerno | 1 | 10 | 5 |
| Hen farm_2 | Salerno | 1 | 11 | 10 |
| Horse stall_1 | Salerno | 4 | 6 | 3 |
| Horse stall_2 | Salerno | 6 | 14 | 4 |
| Pig farm | Salerno | 1 | 9 | 7 |
| Pig farm | Benevento | 1 | 16 | 6 |
| Pig farm | Avellino | 2 | 21 | 2 |
| Pigeon farm | Napoli | 1 | 12 | 2 |
| Rabbits farm | Salerno | 1 | 7 | 11 |
| Bus_1 | Salerno | 5 | - | - |
| Bus_2 | Salerno | 5 | - | - |
| Bus_3 | Salerno | 5 | - | - |
| Bus_4 | Salerno | 5 | - | - |
| Supermarket | Salerno | 14 | - | - |
| Hotel receptive structure | Napoli | 9 | - | - |
| Total | 77 | 246 | 93 |
| Primers and Probes | Sequence | Concentrations |
|---|---|---|
| RdRp_SARSr-F | GTGARATGGTCATGTGTGGCGG | 600 nM |
| RdRp_SARSr-R | CARATGTTAAASACACTATTAGCATA | 800 nM |
| RdRP_SARSr-P1 | FAM-CCAGGTGGWACRTCATCMGGTGATGC-BBQ | 100 nM |
| RdRp_SARSr-P2 | FAM-CAGGTGGAACCTCATCAGGAGATGC-BBQ | 100 nM |
| E_Sarbeco_F | ACAGGTACGTTAATAGTTAATAGCGT | 400 nM |
| E_Sarbeco_R | ATATTGCAGCAGTACGCACACA | 400 nM |
| E_Sarbeco_P1 | FAM-ACACTAGCCATCCTTACTGCGCTTCG-BBQ | 200 nM |
| N_Sarbeco_F | CACATTGGCACCCGCAATC | 600 nM |
| N_Sarbeco_R | GAGGAACGAGAAGAGGCTTG | 800 nM |
| N_Sarbeco_P | FAM-ACTTCCTCAAGGAACAACATTGCCA-BBQ | 200 nM |
| Sampling Location | Municipality | N° of Samples | dd RT-PCR | ||
|---|---|---|---|---|---|
| RdRp | E Gene | N Gene | |||
| Buffaloes farm_1 | Caserta | 1 | + | - | - |
| Buffaloes farm_2 | Caserta | 1 | - | - | - |
| Buffaloes farm_3 | Caserta | 1 | - | - | - |
| Buffaloes farm_4 | Caserta | 1 | + | - | - |
| Buffaloes farm_4 | Caserta | 2 | + | + | - |
| Buffaloes farm_4 | Caserta | 7 | - | - | - |
| Cattle farm | Salerno | 1 | - | - | - |
| Goat and cattle farm | Salerno | 1 | - | - | - |
| Goat farm_1 | Salerno | 1 | - | - | - |
| Goat farm_2 | Salerno | 1 | + | - | - |
| Hen farm_1 | Salerno | 1 | + | - | - |
| Hen farm_2 | Salerno | 1 | - | - | - |
| Horse stall_1 | Salerno | 2 | - | - | - |
| Horse stall_1 | Salerno | 2 | + | + | - |
| Horse stall_1 | Salerno | 2 | + | - | - |
| Horse stall_2 | Salerno | 4 | - | - | - |
| Pig farm | Salerno | 1 | - | - | - |
| Pig farm | Benevento | 1 | - | - | - |
| Pig farm | Avellino | 1 | - | - | - |
| Pigeon farm | Napoli | 1 | - | - | - |
| Rabbits farm | Salerno | 1 | - | + | - |
| subtotal | 34 | 10 (29%) | 5 (15%) | 0 (0%) | |
| Bus_1 | Salerno | 3 | - | - | - |
| Bus_1 | Salerno | 1 | + | - | - |
| Bus_1 | Salerno | 1 | + | + | - |
| Bus_2 | Salerno | 1 | + | - | - |
| Bus_2 | Salerno | 4 | - | - | - |
| Bus_3 | Salerno | 1 | + | + | - |
| Bus_3 | Salerno | 3 | - | - | - |
| Bus_3 | Salerno | 1 | + | - | - |
| Bus_4 | Salerno | 4 | - | - | - |
| Bus_4 | Salerno | 1 | + | - | - |
| subtotal | 20 | 6 (30%) | 2 (10%) | 0 (0%) | |
| Supermarket | Salerno | 11 | - | - | - |
| Supermarket | Salerno | 1 | - | + | - |
| Supermarket | Salerno | 1 | - | - | + |
| Supermarket | Salerno | 1 | + | - | - |
| subtotal | 14 | 1 (7%) | 1 (7%) | 1 (7%) | |
| Hotel receptive structure | Napoli | 9 | - | - | - |
| subtotal | 9 | 0 (0%) | 0 (0%) | 0 (0%) | |
| Total | 77 | 17 (22%) | 8 (10%) | 1 (1%) | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cardinale, D.; Tafuro, M.; Mancusi, A.; Girardi, S.; Capuano, F.; Proroga, Y.T.R.; Corrado, F.; D’Auria, J.L.; Coppola, A.; Rofrano, G.; et al. Sponge Whirl-Pak Sampling Method and Droplet Digital RT-PCR Assay for Monitoring of SARS-CoV-2 on Surfaces in Public and Working Environments. Int. J. Environ. Res. Public Health 2022, 19, 5861. https://doi.org/10.3390/ijerph19105861
Cardinale D, Tafuro M, Mancusi A, Girardi S, Capuano F, Proroga YTR, Corrado F, D’Auria JL, Coppola A, Rofrano G, et al. Sponge Whirl-Pak Sampling Method and Droplet Digital RT-PCR Assay for Monitoring of SARS-CoV-2 on Surfaces in Public and Working Environments. International Journal of Environmental Research and Public Health. 2022; 19(10):5861. https://doi.org/10.3390/ijerph19105861
Chicago/Turabian StyleCardinale, Davide, Maria Tafuro, Andrea Mancusi, Santa Girardi, Federico Capuano, Yolande Thérèse Rose Proroga, Federica Corrado, Jacopo Luigi D’Auria, Annachiara Coppola, Giuseppe Rofrano, and et al. 2022. "Sponge Whirl-Pak Sampling Method and Droplet Digital RT-PCR Assay for Monitoring of SARS-CoV-2 on Surfaces in Public and Working Environments" International Journal of Environmental Research and Public Health 19, no. 10: 5861. https://doi.org/10.3390/ijerph19105861







