1. Introduction
Cinnamomum kanehirae Hayata, a native tree in Taiwan, plays a crucial role in the life cycle of the medicinal fungus
Taiwanofungus camphoratus. It thrives in mountainous regions at altitudes ranging from 200 to 2000 m within Taiwan’s broad-leaved forests [
1,
2]. Due to the economic importance of
T. camphoratus,
C. kanehirae logs have been extensively harvested for cultivation, leading to a drastic reduction in the total timber volume in Taiwan [
3]. As a result of excessive logging,
C. kanehirae is now classified as endangered according to the IUCN Red List [
3].
The natural regeneration of
C. kanehirae faces various challenges, including seed predation and notably low seed germination rates [
4]. Most regeneration relies on stump resprouting [
2,
3], which is not a sustainable long-term solution. Given these challenges, there is an urgent need for effective management strategies to restore the
C. kanehirae population. Kao and Huang [
5] collected the clones of
C. kanehirae and successfully regenerated the population using cutting. Cutting may well be a good silvicultural practice for the conservation of
C. kanehirae. However, the lack of axial roots results in growth arrest and a low survival rate [
6].
The development of roots is essential for the successful establishment of seedlings during forest restoration. Mycorrhization, a technique gaining global significance in nursery seedling cultivation, has demonstrated its ability to enhance plant growth and enhance the overall plant performance when it is relocated to field settings [
7]. It has also been demonstrated that inoculating seedlings or cuttings with different mycorrhizal fungi during the early stages of cultivation can affect their performance after being transplanted for reforestation [
7]. Previous studies have shown that inoculating beneficial microorganisms in the nursery can enhance seedling growth and nutrient efficiency [
8,
9,
10].
While mycorrhization has garnered global recognition for its effectiveness in enhancing plant growth and overall performance [
11], the utilization of mycorrhizal fungi in nursery settings presents challenges [
12]. In particular, in vitro propagation of these fungi remains a formidable hurdle.
Dark septate endophytes (DSEs) represent a fascinating group of asexual filamentous ascomycetous fungi, renowned for their propensity to colonize the root systems of plants [
13]. These fungi are characterized by their melanized hyphae and microsclerotia [
14]. What sets DSEs apart is their ability to significantly enhance nutrient uptake, including vital elements such as carbon (C), nitrogen (N), and phosphorus (P), in host plants. Furthermore, their presence was linked to improved plant health and productivity, as well as an enhanced ability to withstand various biotic and abiotic stresses (reviewed in Akhtar et al. [
15]). DSE colonization has been shown to particularly benefit the underground parts of host plants [
16]. DSEs emerge as a promising alternative due to their inherent adaptability for cultivation and scalability facilitated by advanced fermentation technology.
The root of
C. kanehirae had been demonstrated belong dark septate endophyte [
17]. CkDB5 endophyte, isolated from the root of
C. kanehirae, had promoted the root growth of
C. kanehirae, and be a dark septate endophytic fungus in laboratory [
18]. In this study, we identified and characterized four fungal strains from the roots of
C. kanehirae. Subsequently, we conducted on-site inoculation experiments, meticulously exploring the effects of DSE colonization on the growth of
C. kanehirae cuttings in nursery. Additionally, we pioneered the development of a cutting-edge molecular detection method to ascertain the presence of the most effective DSE strain. The integration of dark septate endophytes (DSEs) into forest restoration strategies has the potential to revolutionize the field. By circumventing the hurdles associated with the in vitro propagation of mycorrhizal fungi, we unlock the considerable benefits that DSEs offer in terms of plant growth, health, and resilience. Our findings not only offer a glimpse into the future of sustainable forest restoration but also hold the promise of fostering a more harmonious and thriving ecosystem.
2. Materials and Methods
2.1. Endophyte and Cutting Materials
Four endophytes (DB5, YC14, YL14, and YC2) isolated from the roots of Cinnamomum kanehirae were used in this study. All the endophytes were maintained on 2% malt extract at 25 °C and deposited into the Tree Mycorrhiza Laboratory of National Chiayi University. Two-year-old cuttings of C. kanehirae were selected and incubated in Jhongsing Nursery, Chiayi, Taiwan (120°41′56.22″ E, 23°29′35.70″ N). The cuttings were incubated in containers for high survival rates of planting.
2.2. Characterization of the Endophytes
To identify the endophytes, the genomic DNA was extracted from the fresh mycelia using a NucleoSpin® PlantIIDNA extraction kit. The internal transcribed spacer (ITS) region of the ribosomal DNA (rDNA) was amplified using the primers V9G/LR1 [
19,
20,
21] (de Hoog and Gerrits van den Ende 1998; Vilgalys and Hester 1990) or ITS5/ITS4 (White et al., 1990). DNA sequencing was performed at Tri-I Biotech, Taiwan, using the same primers. The sequences were assembled, and related sequences were identified using BLAST searches.
The fungal isolates were screened for the production of IAA, as described by Gravel et al. [
22]. The fungal isolates were inoculated into 250 mL flasks containing 100 mL of 2% malt extract broth and 0.1% L-tryptophan and incubated at ambient temperature for 15 days with agitation. After incubation, the mediums were centrifuged at 4000 rpm for 20 min to remove the fungal biomass. Each cultural filtrate (2 mL) was mixed with 8 mL Salkowski reagent (250 mL distilled water, 150 mL H
2SO
4, 7.5 mL 5 M FeCl
3·6H
2O) and incubated in the dark for 30 min. The IAA concentration was detected using a spectrophotometer (A
535) and compared with the standard IAA concentration.
Four fungal isolates were preliminarily screened for their potential to produce extracellular enzymes. The enzymatic activities of laccase, cellulases, and peroxidase were systematically analyzed in this study. Laccase activity was evaluated via the decolorization of Congo red on potato dextrose agar [
23]. To assess the cellulolytic activity, the fungal isolate was cultured on a solid medium supplemented with 0.5% carboxymethyl cellulose and then incubated at 23 °C in darkness for 7 days. Subsequently, a 0.1% Congo red solution was applied, and the plates were subjected to a 10-min rinse with 5 M of NaCl. The presence of yellow zones around the colonies of fungal endophytes served as an indicative marker of the cellulolytic activity [
24]. For the detection of the peroxidase activity, the fungal isolate was incubated in MEA media at 23 °C in the dark for 7 days. Following this incubation period, a solution containing pyrogallol and 4% hydrogen peroxide was introduced into the media to assess the peroxidase activity. The development of a brownish-orange color in the medium indicated the secretion of peroxidase [
25,
26,
27].
2.3. DSE Inoculation, Plant Growth Conditions, and DSE Root Colonization
The cuttings of C. kanehirae were inoculated with four dark septate endophytes, respectively. The four DSE isolates were cultured on a malt extract agar (MEA) plate for 7 days and then transferred into a 500 mL 2% malt extract broth for 30 days of incubation at 23 °C in the dark. After 30 days of incubation, the DSE mycelium was collected and blended with distilled water as an inoculum. The inoculation basically included three steps: drill holes, fill them with inoculum, and cover the holes with nursery substrate. A Phillips head screwdriver was used to drill the holes and avoid hurting the roots of the cuttings. There were three holes for each cutting and 10 mL of the inoculum was added to each hole. There were four DSE treatments and one control. Each had 10 cuttings. The cuttings were grown and watered at Jhongsing Nursery for 7 months without fertilization.
After 7 months, root colonization by the DSE isolates was observed according to McGonigle et al. [
28]. The sampled roots were washed thoroughly with tap water, cut into 1 cm long segments, and cleaned with 10% KOH for 48 h, then 3% H
2O
2 for 48 h. Finally, the root segments were stained with trypan blue lactophenol and observed under a light microscope.
2.4. Plant Growth and Physiological Parameters
To measure the influences of the DSEs on the growth performance of the cuttings, the net growth in height according to all treatments was recorded after transplantation and before harvest. The plant biomass (root, stem, and leaf) was measured before and after drying the samples in an oven at 70 °C to a constant weight.
To measure the chlorophyll concentration, fully expanded leaves (counted from the apex) were collected. Then, 0.05 g of each sample (fresh material) was ground with 10 mL of 85% acetone. The suspension was filtered and measured using a spectrophotometer at wavelengths of 663 nm (chlorophyll a) and 645 nm (chlorophyll b). The concentrations of chlorophyll a and chlorophyll b were calculated as follows: chlorophyll a (mg/g) = (12.7 × OD663 − 2.69 × OD645) × (V/1000W); chlorophyll b (22.9 × OD645 − 4.68 × OD663) × (V/1000W); chlorophyll a + b = (20.2 × OD645 + 8.02 × OD663) × (V/1000W). V represents the volume of ground leaf–acetone liquid (mL); W represents the fresh weight (g).
2.5. Detection of DSE from Roots of C. kanehirae
To gain a better understanding of the relative host responses to the DSE fungi, we use “DSE responsiveness (R)” from Mandyam et al. [
13] to describe the DSE colonization as a means to assist in evaluating variable plant growth responses.
To determine the presence of DSEs, specific primers for the best-performing DSE were designed for PCR detection. The primers were designed from the variable regions of the ITS. To design the DB5-specific primers, a multiple-sequence alignment was made. According to the alignment, the DB5-specific primers were designed from unique regions which were then assessed using the NCBI Primer-BLAST tool. The oligonucleotide primer pairs DB5-1F (5’-TGTCATCTGTGCTGAACCCC-3’) and DB5-1R (5’-TTGGAGTGTGTAATGGCGCT-3’) were designed, and these primers shared little or no homology with other fungal rDNA sequences. Based on the sequencing information, the primer pair was DB5-specific, and the predicted amplification size of the PCR product was 329 bp. The primers were synthesized by Mission Biotech (Taipei, Taiwan).
To ensure optimal PCR conditions, preliminary experiments were conducted to establish the reproducibility of PCR amplification, with the aim of maximizing yield. The PCR cycling conditions consisted of an initial denaturation step at 95 °C for 5 min, followed by 29 cycles of denaturation at 95 °C for 30 s, annealing at 55°C for 30 s, extension at 72 °C for 30 s, and a final extension step at 72 °C for 5 min. Standard PCR assays were employed to evaluate the sensitivity of the DB5-1F and DB5-1R primers for DB5 detection.
For sensitivity testing, the total DNA from DB5 was serially diluted in 10-fold increments, ranging from 10 ng to 1 fg, and used as template DNA for PCR amplification. A 25 μL PCR mixture was prepared, containing 25 ng of DNA, 1.0 nM of each primer, 1.0 unit of Taq DNA polymerase, and 100 μM dNTP in a PCR reaction buffer.
To investigate the presence of DB5, root samples from
C. kanehirae in both the control and DB5-treated conditions were collected. The DNA was extracted from 1 cm root tissue specimens using the CTAB method [
29], and PCR amplification with the DB5-1F/DB5-1R primer pairs was employed to detect the presence of DB5.
2.6. Statistical Analysis
Prior to statistical analysis, the Kolmogorov–Smirnov test was used to test the normality of data. The means of the separate experiments ± standard error were derived from all collected data. Measurements of IAA concentration, plant growth, and physiological parameters were subjected to a variance analysis (ANOVA) in order to determine the effects of DSE inoculation. Differences between means were tested using Duncan’s new multiple-range test (p ≤ 0.05). The analysis of variance and multiple mean comparisons were carried out on the data using the software Statistical Package for the Social Sciences (SPSS 12.0) (Chicago, IL, USA).
3. Results
3.1. Identification and Characterization of the Endophytes
Taxonomic affinities, including the most closely matched sequences, were assigned to DB5, YC14, YL14, and YC2 based on BLAST sequence similarity analysis. Based on a megablast search of NCBI’s GenBank nucleotide database, the closest hit of the DB5 ITS sequence was Melnikomyces longisporum (GenBank MT731290; Identities = 464/549 (85%), Gaps = 26/549 (4%)). The closest hits of YC14 were close to Acrocalymma vagum (GenBank MK968525; Identities = 624/624 (100%), Gaps = 0/624 (0%)). The closest hits of YL14 were close to Wiesneriomyces laurinus (GenBank KR822217; Identities = 509/542 (94%), Gaps = 15/542 (2%)). YC2 was closely matched to the Tricholomataceae sp. (GenBank MW695325; Identities = 591/611 (97%), Gaps = 2/611 (0%)).
Four endophytic fungal isolates were screened for the presence of extracellular enzymes such as laccase, cellulase, and peroxidase, which was grown on a specific medium in Materials and methods. The four selected endophytic fungi showed laccase activity, while only DB5 exhibited peroxidase activity. The four endophytic fungal isolates were not involved in cellulolytic activity. No IAA was detected in the liquid culture of YC2, YC14, or YL14. Only DB5 produced IAA in medium and the concentration of IAA was 2.51 ± 0.69 ppm.
3.2. Morphology and Colonization of C. kanehirae Cuttings
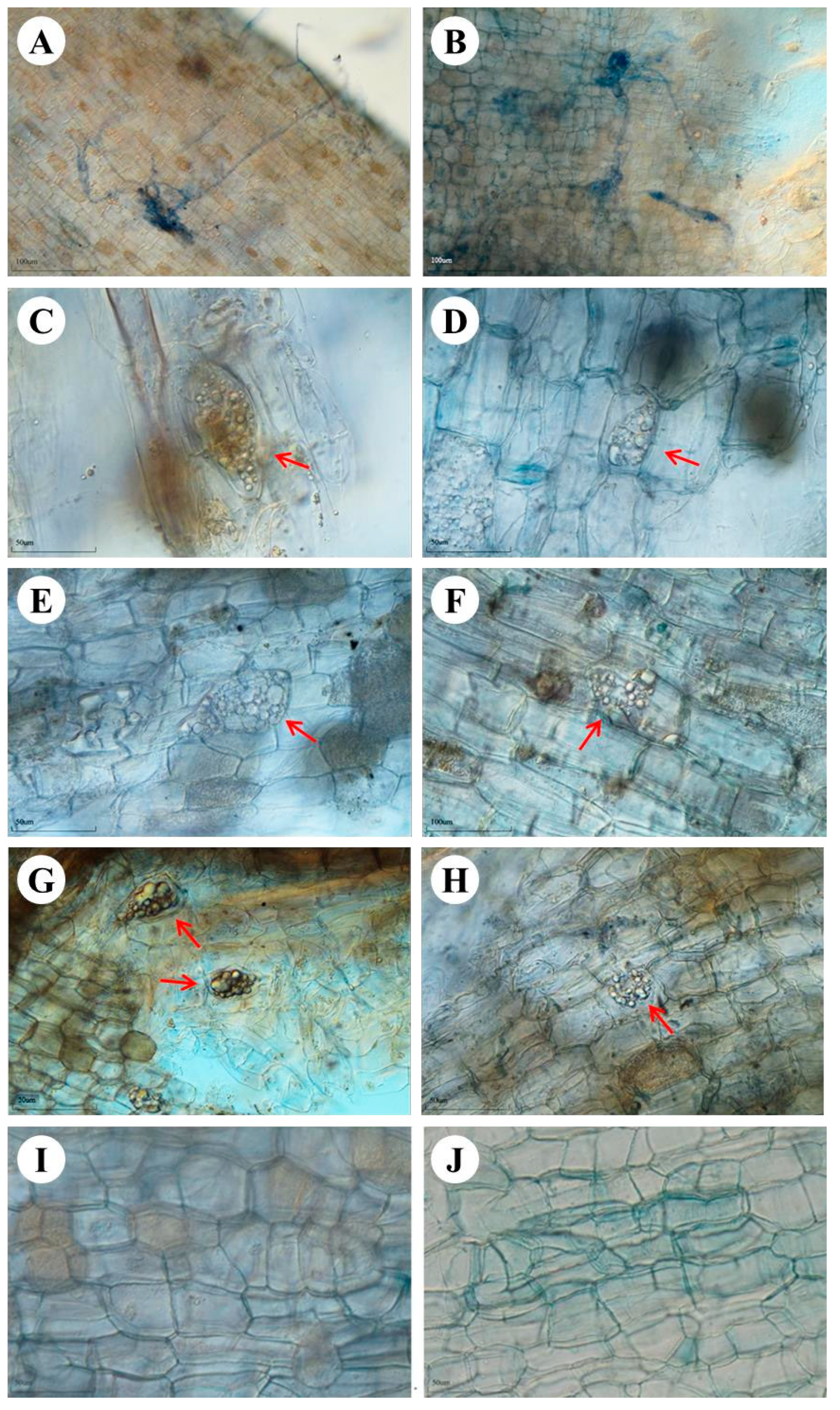
After 7 months, root colonization by the fungal isolates was observed. The light microscopic analysis also showed the active association and habitation of these fungal isolates inside the plant’s roots (
Figure 1). The hyphae from the epidermal region to the cortex cells form a dense network at the end of the cortex cells in YC2 inoculation (
Figure 1A,B). Microsclerotia were not observed with YC2 inoculation. Microsclerotia, a feature of DSEs, were found in the roots with YC14 (
Figure 1C,D), YL14 (
Figure 1E,F), and DB5 (
Figure 1G,H) inoculation, respectively. In the control treatment, microsclerotia and fungal hyphae were not observed.
The nucleotide sequences of the rDNA ITS region of DB5 were aligned with sequences from related Melnikomyces species. Subsequently, the primer pair DB5-1F (5′-TGT CATCTGTGCTGAACCCC-3′) and DB5-1R (5′-TTGGAGTGTGTAATGGCGCT-3′) was custom-designed based on this sequence information. The primer pair was tailored to be DB5-specific, and the expected amplification product size was 329 bp. Employing optimized conditions with an annealing temperature set at 55 °C, the DB5-1F/DB5-1R primer pair consistently produced an amplification product of the expected size from the Melnikomyces sp. DB5.
When pure cultures of the Melnikomyces sp. DB5 were employed for DNA extraction and amplification, the minimum initial DNA amount required to detect the specific PCR product, as visualized by ethidium bromide staining after 30 amplification cycles, was approximately 250–300 bp. This observation underscores the high sensitivity of this technique and its potential for the detection of DSE.
Upon applying the DB5-1F/DB5-1R primer pair to assess the presence of DB5 in the C. kanehirae cuttings, positive detection signals were observed exclusively in the cuttings subjected to DB5 treatment, while no such signals were observed in the control treatments. This provides clear evidence of successful DB5 inoculation in the nursery settings.
3.3. Effects of DSE on C. kanehirae Cuttings
To assess the role of DSEs on the cutting growth, the endophytes were inoculated onto the host plants,
C. kanehirae. After 7 months of incubation, all the cuttings survived. The net height growths of the cuttings inoculated with YL14, DB5, YC14, YC2, and the control were 7.2 ± 3.0, 5.6 ± 2.2, 5.4 ± 1.6, 3.9 ± 1.9, and 4.6 ± 2.5 cm, respectively. Besides YC2, the cuttings inoculated with the DSEs showed greater net height growths than the control. However, the differences were not significant: only the cuttings inoculated with strain YL14 showed significant improvements in net height growth (
Figure 2).
The fresh weight of shoots (WSF) in the cuttings inoculated with YL14, DB5, YC14, YC2, and the control was 17.69 ± 3.21, 17.40 ± 2.86, 16.79 ± 3.41, 14.08 ± 2.32, and 13.34 ± 2.23 g, respectively (
Table 1;
Figure 3). The fresh weight of the roots (WRF) in the cuttings inoculated with YL14, DB5, YC14, YC2, and the control was 21.37 ± 7.09, 19.33 ± 9.63, 24.53 ± 10.44, 16.41 ± 6.03, and 8.47 ± 3.80 g, respectively (
Table 1;
Figure 3). We found that the cuttings inoculated with the DSEs have a better performance in the fresh weight of the shoots, roots, and whole plant (
Table 1;
Figure 3). Similar trends were found in the dry weight of the plants (
Table 1;
Figure 4).
After harvesting, the chlorophyll concentration was also analyzed. The Chl a concentrations in the cuttings inoculated with YL14, DB5, YC14, YC2, and the control were 1.06 ± 0.01, 1.22 ± 0.03, 1.12 ± 0.01, 1.01 ± 0.47, and 0.82 ± 0.24 mg g
−1, respectively (
Table 2). The cuttings inoculated with the DSEs showed greater Chl a concentrations than the control. However, the differences were not significant; only the cuttings inoculated with strain DB5 showed a significant higher Chl a concentration (
Table 1). For Chl b concentration, all DSE treatments had a significant higher concentration than the control. The Chl b concentrations in the cuttings inoculated with YL14, DB5, YC14, YC2, and the control were 0.35 ± 0.01, 0.35 ± 0.01, 0.34 ± 0.05, 0.30 ± 0.03, and 0.24 ± 0.20 mg g
−1, respectively (
Table 2). Similar results were also observed for Chl b and Chl a + b concentration (
Table 2).
The DSE responsiveness (R) of YC2, YC14, YL14, and DB5 was 0.02, 0.25, 0.35, and 0.36, respectively. Overall, the seedlings mostly responded positively to DSE colonization. However, DB5 was the best strain among the four isolates. To confirm the presence of the DSE, we designed specific primers for DB5 (DB5-1F/DB5-1R) for PCR detection. Our findings revealed the successful detection of DB5 in the roots of infected cuttings (
Figure 5). This observation strongly supports the existence of DB5 in the roots of
C. kanehirae cuttings. The specific primer pair (DB5-1R/DB5-1F) designed for the DNA sequence of the DB5 strain exhibited excellent specificity and sensitivity in the PCR reaction. These results underscore the reliability of the specific primer set in accurately identifying the presence of DB5 in the examined samples.
4. Discussion
In this study, the effects of four endophytes (YC2, YC14, YL14, and DB5) on the growth of C. kanehirae cuttings were evaluated. The results indicate that the inoculation of these endophytes had a positive influence on various growth parameters of C. kanehirae, including plant height, biomass, and photosynthesis rates. Notably, microsclerotia formation observed in the root associations of the YC14, YL14, and DB5 inoculations indicated successful colonization by dark septate endophytes (DSEs).
Among them, DB5 demonstrated remarkable efficacy in the growth of
C. kanehirae cuttings and was found to secrete peroxidase and indole acetic acid (IAA). Several endophytic fungi, such as
Alternaria alternate,
Phialophora mustea, and
Phialocephala fortini, have been reported to synthesize IAA, exerting a stimulatory effect on plant growth [
30,
31]. The colonization of
C. kanehirae by DSEs is shown to have a significant impact on plant physiology via the modulation of phytohormone production. The presence of DB5 significantly enhanced the growth of
C. kanehirae cuttings in the nursery setting, potentially due to its higher indole acetic acid synthesis.
Positive effects were also demonstrated in plot experiments conducted in a greenhouse, as reported by Lin [
18]. This consistency across different experimental environments underscores the robustness and reliability of the observed benefits associated with the endophytes. The nursery experiments further support the notion that DB5 is a promising candidate for enhancing the growth and development of
C. kanehirae, making it a valuable component in silviculture practices.
The identification of DB5 revealed it to be a
Melnikomyces sp., closely related to
M. longisporum. The genus
Melnikomyces, introduced by Crous et al. [
32], is a relatively new taxonomic entity, comprising only three known species,
M. vietnamensis,
M. thailandicus, and
M. longisporum.
Melnikomyces vietnamensis was initially collected from dry leaves in Vietnam [
32],
M. thailandicus was isolated from soil near a waterfall [
33], and
M. longisporum was identified from forest litter in China [
34]. In this study,
Melnikomyces sp. DB5 was isolated from the roots of
C. kanehirae, thereby expanding the known ecological range of
Melnikomyces species. This represents the first documented instance of dark septate endophytes belonging to the
Melnikomyces genus.
The diverse habitats and ecological functions observed among Melnikomyces species highlight the potential significance of these fungi in various ecological niches. This study offers a novel perspective by demonstrating the presence of Melnikomyces sp. as a dark septate endophyte in the root system of C. kanehirae, adding to the growing body of knowledge concerning the diversity and roles of Melnikomyces within ecosystems.
In our study, the endophyte YC14 was identified as
Acrocalymma vagum, which was found to be associated with the roots of
C. kanehirae and demonstrated a positive impact on plant growth. Previous research has also recognized
A. vagum as a dark septate endophyte in the roots of Chinese licorice [
35] and tobacco [
36]. Notably,
A. vagum has been reported to exhibit the ability to reduce the heavy metal content in tobacco [
36] and promote plant growth in Chinese licorice [
35].
A. vagum has been documented in association with various plants from diverse ecosystems, colonizing not only non-grass herbaceous plants but also woody species. Given its potential to positively affect plant nutrient uptake, performance, and survival and induce resistance in host plants,
A. vagum provides valuable insights into the mechanisms underlying this less understood plant–fungal symbiosis.
In contrast, YL14 was identified as a
Wiesneriomyces sp.
Wiesneriomyces is known to have a cosmopolitan distribution and has been documented in fallen leaves of several plant species, including
Beilschmiedia,
Knightia,
Nothofagus,
Pittosporum, and Arecaceae (Palmae) [
37,
38]. This study presents the first report of its potential positive effect on plant growth. The discovery of
A. vagum and
Wiesneriomyces in association with
C. kanehirae and their impact on plant growth underscores the importance of exploring the multifaceted interactions between plants and their fungal symbionts. These findings contribute to our understanding of the diverse roles played by endophytes in shaping plant health and ecosystem dynamics.
The inoculation of DB5 has shown significant promise in augmenting the growth of C. kanehirae cuttings, marking it as a potentially valuable approach to the conservation of this endangered tree species in forestry nurseries. Furthermore, we have successfully devised a molecular detection method to confirm the presence of DB5, and our findings unequivocally establish its colonization within the roots of the inoculated cuttings. This molecular detection method holds considerable potential as a certification tool for validating successful DB5 inoculation in nurseries, thereby streamlining its application in conservation efforts.
5. Conclusions
In this study, we successfully isolated and identified four fungal strains from the roots of C. kanehirae, while also assessing their influence on host growth and nutrient acquisition. Our results revealed a range of plant-growth-promoting activities, including an increased plant height, biomass, and photosynthesis rates. Notably, the fungal isolate DB5, identified as a Melnikomyces sp., exhibited enhanced performance and was found to secrete peroxidase and indole acetic acid. Furthermore, the development of a molecular detection method for DB5 represents a significant advancement in our ability to monitor and confirm successful endophyte inoculation. We propose that the application of the endophyte inoculation approach, coupled with the use of this molecular detection method, should be strongly encouraged in forestry nurseries. This integrated approach has the potential to greatly enhance silvicultural practices and contribute to the conservation efforts aimed at safeguarding C. kanehirae, an endangered tree species.