Biotechnological Potential of Oil-Tolerant Strains for Possible Use in Bioremediation
Abstract
:1. Introduction
2. Materials and Methods
2.1. Soil
2.2. Microbiological Analysis of Crude Oil-Contaminated Soil
2.3. Morphological and Biochemical Analyses
2.4. MALDI-TOF MS Analysis
3. Results and Discussion
3.1. Microbiological Analysis of Soil Contaminated with Crude Oil
3.2. Morphological and Biochemical Analysis of Microorganisms
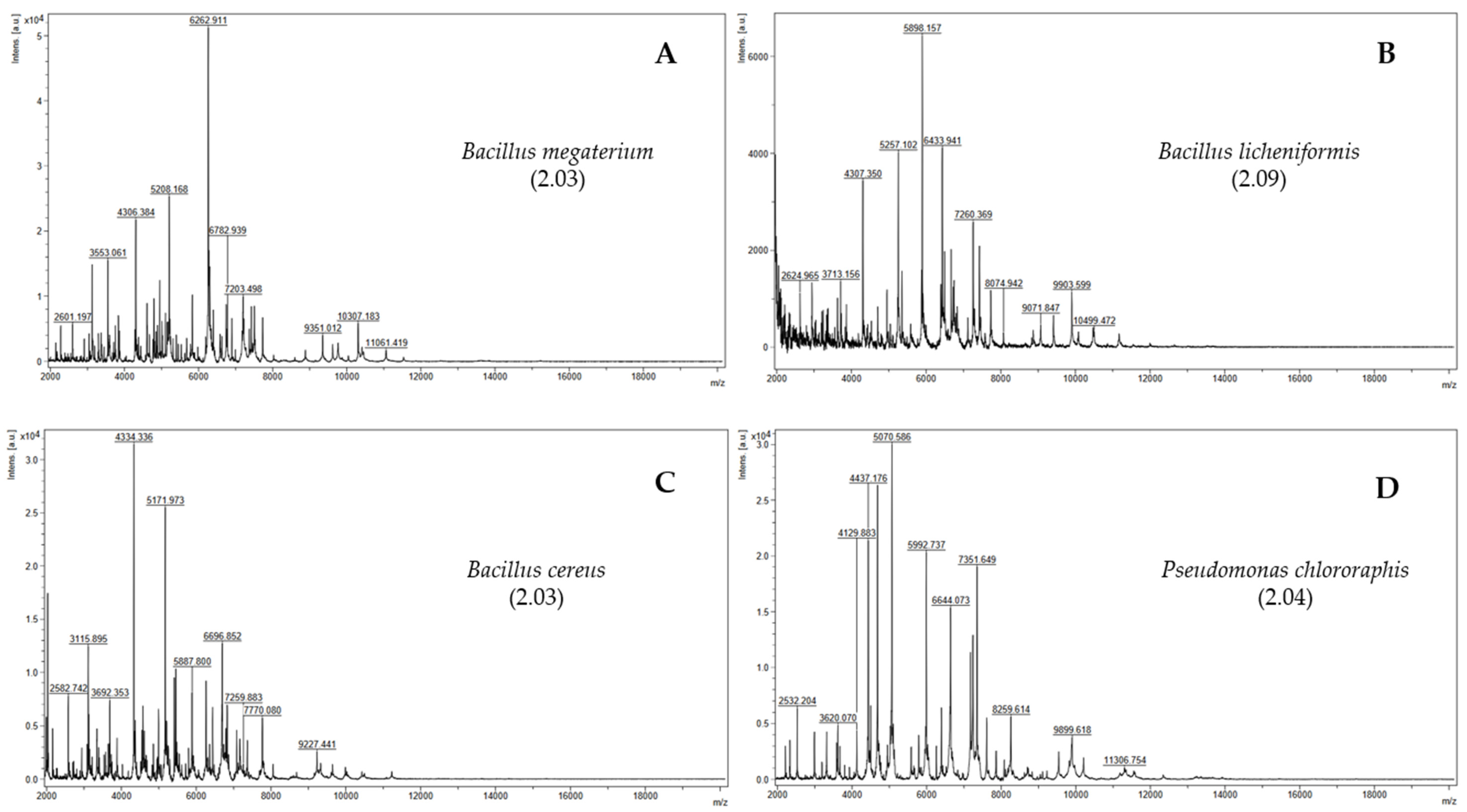
3.3. MALDI-TOF Analysis
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Lal, R.; Bouma, J.; Brevik, E.; Dawson, L.; Field, D.J.; Glaser, B.; Hatano, R.; Hartemink, A.E.; Kosaki, T.; Las-celles, B.; et al. Soils and Sustainable Development Goals of the United Nations: An In-ternational Union of Soil Sciences Perspective. Geoderma Reg. 2021, 25, e0039. [Google Scholar] [CrossRef]
- Lehmann, J.; Bossio, D.A.; Kögel-Knabner, I.; Rillig, M.C. The Concept and Future Prospects of Soil Health. Nat. Rev. Earth Environ. 2020, 1, 544–553. [Google Scholar] [CrossRef]
- Ossai, I.C.; Ahmed, A.; Hassan, A.; Hamid, F.S. Remediation of Soil and Water Contaminated with Petroleum Hydrocarbon: A Review. Environ. Technol. Innov. 2020, 17, 100526. [Google Scholar] [CrossRef]
- Mukjang, N.; Chitov, T.; Mhuantong, W.; Champreda, V.; Pathom-aree, W.; Sattayawat, P.; Bovonsombut, S. Bacterial Communities Associated with Crude Oil Bioremediation through Composting Approaches with Indigenous Bacterial Isolate. Life 2022, 12, 1712. [Google Scholar] [CrossRef]
- Koshlaf, E.; Ball, A.S. Soil Bioremediation Approaches for Petroleum Hydrocarbon Polluted Environments. AIMS Microbiol. 2017, 3, 25–49. [Google Scholar] [CrossRef]
- Pandolfo, E.; Barra Caracciolo, A.; Rolando, L. Recent Advances in Bacterial Degradation of Hydrocarbons. Water 2023, 15, 375. [Google Scholar] [CrossRef]
- Lim, M.W.; Von Lau, E.; Poh, P.E. A Comprehensive Guide of Remediation Technologies for Oil Contaminated Soil—Present Works and Future Directions. Mar. Pollut. Bull. 2016, 109, 14–45. [Google Scholar] [CrossRef]
- Mambwe, M.; Kalebaila, K.K.; Johnson, T. Remediation Technologies for Oil Contaminated Soil. Glob. J. Environ. Sci. Manag. 2021, 7, 419–438. [Google Scholar]
- Al-Dhabaan, F.A. Morphological, Biochemical and Molecular Identification of Petroleum Hydrocarbons Biodegradation Bacteria Isolated from Oil Polluted Soil in Dhahran, Saud Arabia. Saudi J. Biol. Sci. 2019, 26, 1247–1252. [Google Scholar] [CrossRef]
- Vidali, M. Bioremediation. An Overview. Pure Appl. Chem. 2001, 73, 1163–1172. [Google Scholar] [CrossRef]
- Łebkowska, M.; Zborowska, E.; Karwowska, E.; Miaśkiewicz-Peska, E.; Muszyński, A.; Tabernacka, A.; Naumczyk, J.; Jeczalik, M. Bioremediation of Soil Polluted with Fuels by Sequential Multiple Injection of Native Microorganisms: Field-Scale Processes in Poland. Ecol. Eng. 2011, 37, 1895–1900. [Google Scholar] [CrossRef]
- Szulc, A.; Ambrozewicz, D.; Sydow, M.; Ławniczak, Ł.; Piotrowska-Cyplik, A.; Marecik, R.; Chrzanowski, Ł. The Influence of Bioaugmentation and Biosurfactant Addition on Bioremediation Efficiency of Diesel-Oil Contaminated Soil: Feasibility during Field Studies. J. Environ. Manag. 2014, 132, 121–128. [Google Scholar] [CrossRef]
- Marigómez, I. Oil, Crude. In Encyclopedia of Toxicology, 3rd ed.; Wexler, P., Ed.; Academic Press: Cambridge, MA, USA, 2014; pp. 663–669. [Google Scholar]
- Sui, X.; Wang, X.; Li, Y.; Ji, H. Remediation of Petroleum-Contaminated Soils with Microbial and Microbial Combined Methods: Advances, Mechanisms and Challenges. Sustainability 2021, 13, 9267. [Google Scholar] [CrossRef]
- Varjani, S.J. Microbial Degradation of Petroleum Hydrocarbons. Bioresour. Technol. 2017, 223, 277–286. [Google Scholar] [CrossRef]
- Bidja Abena, M.T.; Li, T.; Shah, M.N.; Zhong, W. Biodegradation of Total Petroleum Hydrocarbons (TPH) in Highly Contaminated Soils by Natural Attenuation and Bioaugmentation. Chemosphere 2019, 234, 864–874. [Google Scholar] [CrossRef]
- Nwankwegu, A.S.; Onwosi, C.O. Bioremediation of Gasoline Contaminated Agricultural Soil by Bioaugmentation. Environ. Technol. Innov. 2017, 7, 1–11. [Google Scholar] [CrossRef]
- Koshlaf, E.; Shahsavari, E.; Aburto-Medina, A.; Taha, M.; Haleyur, N.; Makadia, T.H.; Morrison, P.D.; Ball, A.S. Bioremediation Potential of Diesel-Contaminated Libyan Soil. Ecotoxicol. Environ. Saf. 2016, 133, 297–305. [Google Scholar] [CrossRef]
- Atlas, R.M.; Bartha, R. Hydrocarbon Biodegradation and Oil Spill Bioremediation. In Advances in Microbial Ecology, 1st ed.; Marshall, K.C., Ed.; Springer: Boston, MA, USA, 1992; Volume 12, pp. 287–338. [Google Scholar]
- Rizzo, P.; Malerba, M.; Bucci, A.; Sanangelantoni, A.M.; Remelli, S.; Celico, F. Potential Enhancement of the In-Situ Bioremediation of Contaminated Sites through the Isolation and Screening of Bacterial Strains in Natural Hydrocarbon Springs. Water 2020, 12, 2090. [Google Scholar] [CrossRef]
- Wang, S.; Cheng, F.; Guo, S. Highly Efficient Screening and Optimal Combination of Functional Isolates for Bioremediation of Hydrocarbon-Polluted Soil. Environ. Res. 2023, 219, 115064. [Google Scholar] [CrossRef] [PubMed]
- APHA. Standard Methods for the Examination of Water and Wastewater; APHA: Washington, DC, USA, 2012. [Google Scholar]
- Leboffe, M.J.; Pierce, B.E. Microbiology: Laboratory Theory and Application, 2nd ed.; Morton Publishing Company: Englewood, CO, USA, 2012. [Google Scholar]
- Vuković Domanovac, M.; Šabić Runjavec, M.; Meštrović, E. Bioaugmentation Effect of Aeromonas Hydrophila and Pseudomonas Putida on Kinetics of Activated Sludge Process in Treating Pharmaceutical Industrial Wastewater. J. Chem. Technol. Biotechnol. 2019, 94, 2721–2728. [Google Scholar] [CrossRef]
- Kuppusamy, S.; Maddela, N.R.; Megharaj, M.; Venkateswarlu, K. Total Petroleum Hydrocarbons: Environmental Fate, Toxicity, and Remediation, 1st ed.; Springer International Publishing: Cham, Switzerland, 2019. [Google Scholar]
- Wang, L.; Cheng, Y.; Naidu, R.; Bowman, M. The Key Factors for the Fate and Transport of Petroleum Hydrocarbons in Soil with Related in/Ex Situ Measurement Methods: An Overview. Front. Environ. Sci. 2021, 9, 756404. [Google Scholar] [CrossRef]
- Abatenh, E.; Gizaw, B.; Tsegaye, Z.; Wassie, M. Application of microorganisms in bioremediation—Review. Environ. Microbiol. 2017, 1, 2–9. [Google Scholar]
- Maier, R.M. Bacterial Growth. In Environmental Microbiology, 2nd ed.; Maier, R.N., Pepper, I.L., Gerba, C.P., Eds.; Academic Press: Cambridge, MA, USA, 2009; pp. 37–54. [Google Scholar]
- Ujowundu, C.O.; Kalu, F.N.; Nwaoguikpe, R.N.; Kalu, O.I.; Ihejirika, C.E.; Nwosunjoku, E.C.; Okechukwu, R.I. Biochemical and Physical Characterization of Diesel Petroleum Contaminated Soil in Southeastern Nigeria. Res. J. Chem. Sci. 2011, 8, 57–62. [Google Scholar]
- Das, S.; Dash, H.R. Microbial Bioremediation: A Potential Tool for Restoration of Contaminated Areas. In Microbial Biodegradation and Bioremediation; Das, S., Ed.; Elsevier: Amsterdam, The Netherlands, 2014; pp. 2–21. [Google Scholar]
- Suja, F.; Rahim, F.; Taha, M.R.; Hambali, N.; Rizal Razali, M.; Khalid, A.; Hamzah, A. Effects of Local Microbial Bioaugmentation and Biostimulation on the Bioremediation of Total Petroleum Hydrocarbons (TPH) in Crude Oil Contaminated Soil Based on Laboratory and Field Observations. Int. Biodeterior. Biodegrad. 2014, 90, 115–122. [Google Scholar] [CrossRef]
- Barrow, G.I.; Feltham, R.K.A. Cowan and Steel’s Manual for the Identification of Medical Bacteria, 3rd ed.; Cambridge University Press: Cambridge, UK, 1993. [Google Scholar]
- Kaushal, J.; Mehandia, S.; Singh, G.; Raina, A.; Arya, S.K. Catalase Enzyme: Application in Bioremediation and Food Industry. Biocatal. Agric. Biotechnol. 2018, 16, 192–199. [Google Scholar] [CrossRef]
- Grgas, D.; Štefanac, T.; Galant, M.; Brozinčević, A.; Štrkalj, A.; Dragičević, T.L. Biološka denitrifikacija. Croat. J. Food Technol. Biotechnol. Nutr. 2021, 16, 28–34. [Google Scholar]
- Martinez-Urtaza, J.; Lozano-Leon, A.; Viña-Feas, A.; De Novoa, J.; Garcia-Martin, O. Differences in the API 20E Biochemical Patterns of Clinical and Environmental Vibrio Parahaemolyticus Isolates. FEMS Microbiol. Lett. 2006, 255, 75–81. [Google Scholar] [CrossRef] [PubMed]
- Lee, B.H. Structure, function and applications of microbial β-galactosidase (lactase). In Carbohydrate-Active Enzymes, 1st ed.; Park, K.-H., Ed.; Woodhead Publishing: Sawston, UK, 2008; pp. 77–110. [Google Scholar]
- Sharma, N.; Sharma, N.; Sharma, S.; Sharma, P.; Devi, B. Identification, morphological, biochemical, and genetic characterization of microorganisms. In Basic Biotechniques for Bioprocess and Bioentrepreneurship, 1st ed.; Bhatt, A.K., Bhatia, R.K., Bhalla, T.C., Eds.; Academic Press: Cambridge, MA, USA, 2023; pp. 47–84. [Google Scholar]
- Härtig, E.; Jahn, D. Regulation of the anaerobic metabolism in Bacillus subtilis. In Advances in Microbial Physiology, 1st ed.; Poole, R.K., Ed.; Academic Press: Cambridge, MA, USA, 2012; Volume 61, pp. 195–216. [Google Scholar]
- Rajoka, M.I.; Malik, K.A. Cellulomonas. In Encyclopedia of Food Microbiology, 1st ed.; Batt, C.A., Robinson, R.K., Eds.; Academic Press: Cambridge, MA, USA, 1999; pp. 365–371. [Google Scholar]
- Tittsler, R.P.; Sandholzer, L.A.; Dentistry, R.; York, N. The Fermentation of cellobiose by bacteria. J. Bacteriol. 1936, 31, 301–307. [Google Scholar] [CrossRef]
- Randell, P. It’s a MALDI but it’s a Goodie: MALDI-TOF mass spectrometry for microbial identification. Thorax 2014, 69, 776–778. [Google Scholar] [CrossRef]
- Schleifer, K.-H. Phylum XIII. Firmicutes Gibbons and Murray 1978, 5 (Firmacutes [Sic] Gibbons and Murray 1978, 5). In Bergey’s Manual of Systematic Bacteriology, 2nd ed.; De Vos, P., Garrity, G.M., Jones, D., Krieg, N.R., Ludwig, W., Rainey, F.A., Schleifer, K.-H., Whitman, W.B., Eds.; Springer: New York, NY, USA, USA, 2009; Volume 3, pp. 19–1317. [Google Scholar]
- Kuri, M.L.; Roy, S. Biodegradation of Petroleum Oil by a Novel Bacillus megaterium Strain Isolated from Contaminated Soil of Neemrana, Alwar, Rajasthan, India. Int. J. Biotechnol. Bioeng. 2019, 10, 17–27. [Google Scholar]
- Yudono, B.; Said, M.; Sabaruddin; Napoleon, A.; Utami, M.B. Kinetics of Petroleum-Contaminated Soil Biodegraded by an Indigenous Bacteria Bacillus megaterium. Hayati 2010, 17, 155–160. [Google Scholar] [CrossRef]
- Khanpour-Alikelayeh, E.; Partovinia, A.; Talebi, A.; Kermanian, H. Investigation of Bacillus licheniformis in the Biodegradation of Iranian Heavy Crude Oil: A Two-Stage Sequential Approach Containing Factor-Screening and Optimization. Ecotoxicol. Environ. Saf. 2020, 205, 111103. [Google Scholar] [CrossRef] [PubMed]
- El-Sheshtawy, H.S.; Ahmed, W. Bioremediation of Crude Oil by Bacillus licheniformis in the Presence of Different Concentration Nanoparticles and Produced Biosurfactant. Int. J. Environ. Sci. Technol. 2017, 14, 1603–1614. [Google Scholar] [CrossRef]
- Karlapudi, A.P.; Venkateswarulu, T.C.; Tammineedi, J.; Kanumuri, L.; Ravuru, B.K.; Ramu Dirisala, V.; Kodali, V.P. Role of Biosurfactants in Bioremediation of Oil Pollution—A Review. Petroleum 2018, 4, 241–249. [Google Scholar] [CrossRef]
- Schneider, K.R.; Parish, M.E.; Goodrich, R.M.; Cookingham, T. Preventing Foodborne Illness: Bacillus cereus and Bacillus anthracis; University of Florida—IFAS Exension: Gainesville, FL, USA, 2017; pp. 1–5. [Google Scholar]
- Deng, Z.; Jiang, Y.; Chen, K.; Gao, F.; Liu, X. Petroleum Depletion Property and Microbial Community Shift After Bioremediation Using Bacillus Halotolerans T-04 and Bacillus cereus 1-1. Front. Microbiol. 2020, 11, 353. [Google Scholar] [CrossRef] [PubMed]
- Vinothini, C.; Sudhakar, S.; Ravikumar, R. Biodegradation of petroleum and crude oil by Pseudomonas putida and Bacillus cereus. Int. J. Curr. Microbiol. App. Sci. 2015, 4, 318–329. [Google Scholar]
- Scales, B.S.; Dickson, R.P.; Lipuma, J.J.; Huffnagle, G.B. Microbiology, Genomics, and Clinical Significance of the Pseudomonas fluorescens Species Complex, an Unappreciated Colonizer of Humans. Clin. Microbiol. Rev. 2014, 27, 927–948. [Google Scholar] [CrossRef]
- Gutiérrez, E.J.; Abraham, M.D.R.; Baltazar, J.C.; Vázquez, G.; Delgadillo, E.; Tirado, D. Pseudomonas fluorescens: A Bioaugmentation Strategy for Oil-Contaminated and Nutrient-Poor Soil. Int. J. Environ. Res. Public Health 2020, 17, 6959. [Google Scholar] [CrossRef]
- Barathi, S.; Vasudevan, N. Bioremediation of Crude Oil Contaminated Soil by Bioaugmentation of Pseudomonas Fluorescens NS1. J. Environ. Sci. Health A Toxic Hazard. Subst Environ. Eng. 2003, 38, 1857–1866. [Google Scholar] [CrossRef]
- Abdelhaleem, H.A.R.; Zein, H.S.; Azeiz, A.; Sharaf, A.N.; Abdelhadi, A.A. Identification and Characterization of Novel Bacterial Polyaromatic Hydrocarbon-Degrading Enzymes as Potential Tools for Cleaning up Hydrocarbon Pollutants from Different Environmental Sources. Environ. Toxicol. Pharmacol. 2019, 67, 108–116. [Google Scholar] [CrossRef]
- Liu, H.; Yang, G.; Jia, H.; Sun, B. Crude Oil Degradation by a Novel Strain Pseudomonas aeruginosa AQNU-1 Isolated from an Oil-Contaminated Lake Wetland. Processes 2022, 10, 307. [Google Scholar] [CrossRef]
- Annie, G.; Sheela, A.M.; Ilamathi, R. Fate of Crude Oil in Soil Treated with Pseudomonas putida Immobilized on Coconut Coirpith a Lowcost Biocarrier. Soil Sediment Contam. 2020, 29, 770–787. [Google Scholar] [CrossRef]

| Isolate # | Colony Characteristics | Cell Characteristics | |||||
|---|---|---|---|---|---|---|---|
| Growth on Nutrient Medium | Morphology | Color | Microscopic View, M = 1000× | Gram Reaction | Schaeffer–Fulton Reaction | Shape | |
| A | 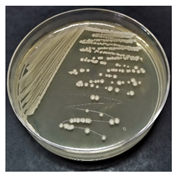 | Convex, round, shiny, entire edge, medium-sized | Cream, smooth, opaque | 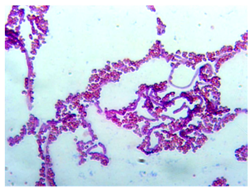 | Gram-positive | Endospores | Large bacilli |
| B | 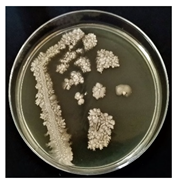 | Large, irregular, rough texture | White, creamy |  | Gram-positive | Endospores | Bacilli |
| C | 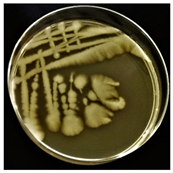 | Large, wavy edge | White, opaque | 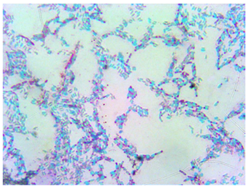 | Gram-positive | Endospores | Large rods |
| D |  | Small, round, raised | Transparent, creamy |  | Gram-negative | Not forming endospores | Small rods |
| Test # | Biochemical Test | Isolate | |||
|---|---|---|---|---|---|
| A | B | C | D | ||
| 1 | Catalase | + | + | + | + |
| 2 | Oxidase | − | − | + | + |
| 3 | Nitrate reductase | − | − | + | − |
| 4 | ß-galactosidase | + | + | − | − |
| 5 | Arginine dihydrolase | + | + | + | + |
| 6 | Utilization of trisodium citrate | + | + | + | + |
| 7 | Tryptophan deaminase | + | + | + | + |
| 8 | Formation of indole | − | + | − | − |
| 9 | Formation of acetoin | + | + | + | + |
| 10 | Gelatinase | + | + | + | − |
| 11 | Fermentation/oxidation of D-glucose | − | + | + | − |
| 12 | Fermentation/oxidation of D-mannitol | + | + | − | − |
| 13 | Fermentation/oxidation of L-arabinose | − | + | − | − |
| 14 | Fermentation/oxidation of D-sucrose | − | + | − | − |
| 15 | Fermentation/oxidation of D-melibiose | − | − | − | − |
| 16 | Fermentation/oxidation of amygdalin | − | + | − | − |
| 17 | Glycerol | + | − | − | / |
| 18 | L-arabinose | + | − | − | / |
| 19 | Ribose | + | − | + | / |
| 20 | D-xylose | + | + | − | / |
| 21 | Galactose | + | + | − | / |
| 22 | D-fructose | + | − | + | / |
| 23 | D-mannose | − | − | + | / |
| 24 | Rhamnose | − | + | − | / |
| 25 | Methyl-D-glucoside | − | + | − | / |
| 26 | N-acetyl-glucosamine | + | − | + | / |
| 27 | Arbutin | + | − | + | / |
| 28 | Aesculin | + | + | + | / |
| 29 | Salicin | + | − | + | / |
| 30 | Cellobiose | + | + | + | / |
| 31 | Maltose | + | + | + | / |
| 32 | Lactose | + | + | − | / |
| 33 | Trehalose | + | − | + | / |
| 34 | Inulin | − | + | − | / |
| 35 | D-raffinose | + | − | − | / |
| 36 | Starch | + | − | + | / |
| 37 | Glycogen | + | − | + | / |
| 38 | Xylitol | + | − | − | / |
| 39 | Gentibiosis | + | − | − | / |
| 40 | Turanose | + | + | − | / |
| Microorganism | Isolation Source | Function | Growth Conditions | Experimental Conditions | Efficiency | Reference |
|---|---|---|---|---|---|---|
| Bacillus megaterium Bacillus licheniformis Bacillus cereus Pseudomonas chlororaphis | Crude oil-contaminated soil | Crude oil degradation | 15 g of crude oil-contaminated soil, 135 mL of MSM, 25 °C, 150 rpm, aeration, 28 days | 15 g of crude oil-contaminated soil, 135 mL of MSM, 25 °C, 150 rpm, aeration, 28 days | Bacterial population—0 day 6.45 × 107 CFU/g and 1.15 × 109 CFU/g; max value −1.34 × 108 CFU/g and 1.75 × 109 CFU/g | This research |
| Bacillus subtilis | Oil-contaminated soil | Crude oil degradation | 5 g of oil-contaminated soil, 500 mL of MSM, 2000 µL of crude oil, room temperature, shaking, 7 days | 5 g of sterile soil, contaminated oil solution, 1% inoculum, 30 °C, 30 days | Hydrocarbons—70% | [9] |
| Oil-contaminated sludge, soil and sea water | Anthracene, catechol and alpha-naphthalene degradation | 1 mL suspension (sludge/soil/water, distilled water, MSM), 100 mL of MSM, 100 mg/L anthracene, catechol and alpha-naphthalene, 30 °C, 150 rpm | MSM medium, anthracene, catechol and alpha-naphthalene, 10% enriched cultures, 30 °C, 150 rpm, 5 days | Anthracene—99% Catechol—1% Alpha-naphthalene—41% | [54] | |
| Bacillus cereus | Oil-contaminated soil | Crude oil degradation | 5 g of oil-contaminated soil, 500 mL of MSM, 2000 of µL crude oil, room temperature, shaking, 7 days | 5 g of sterile soil, contaminated oil solution, 1% inoculum, 30 °C, 30 days | Hydrocarbons—50% | [9] |
| Oil-contaminated sludge, soil and sea water | Anthracene, catechol and alpha-naphthalene degradation | 1 mL suspension (sludge/soil/water, distilled water, MSM), 100 mL of MSM, 100 mg/L anthracene, catechol and alpha-naphthalene, 30 °C, 150 rpm | MSM medium, anthracene, catechol and alpha-naphthalene, 10% enriched cultures, 30 °C, shaking at 150 rpm, 5 days | Anthracene—98% Catechol—17% Alpha-naphthalene—38% | [54] | |
| Pseudomonas aeruginosa | Oil-contaminated soil | Crude oil degradation | 5 g of oil-contaminated soil, 500 mL of MSM, 2000 µL of crude oil, room temperature, shaking, 7 days | 5 g of sterile soil, contaminated oil solution, 1% inoculum, 30 °C, 30 days | Hydrocarbons—68% | [9] |
| Water from oil-contaminated lake wetland | Crude oil degradation | 5% water, 100 mL of MSM, 1 g/L crude oil, 37 °C, 180 rpm, sealed flask, 1 month; 5 mL of inoculum to 100 mL of fresh MSM | 5 mL of inoculum, 100 mL of MSM, 1% crude oil, 37 °C, 180 rpm/0 rpm, sealed flask | C13–C35—75–100% Various PAHs—10–100% | [55] | |
| Pseudomonas putida | National Collection of Industrial Microorganisms, Pune, India | Crude oil degradation | 30 mL suspension (P. putida, 60 mL of MSM), 300 mL of MSM, paraffin, 180 rpm, 3 days | 500 g of soil, 29 mL of crude oil, Pseudomonas putida immobilized in coconut coir pith, room temperature, C:N:P = 100:5:1, aeration, 60 days | Bacterial population—0 day 35.0 × 106 CFU/g Various hydrocarbons—31.6–45.8% | [56] |
| Microbial consortia: Bacillus sp. Pseudomonas putida Pseudomonas mendocin | Soil polluted with diesel oil and engine oil | Biosurfactant production, diesel and engine oil degradation | Soil, mineral Grundmann medium solidified with agar, 1% diesel oil, 26 °C, 72 h | Soil biopiles 4 m (width) × 33 m (length) × 0.5 m (height), microbial consortia, aeration, 15 °C–30 °C | Bacterial population—2 day 1.6 × 107 CFU/g; max value—9.9 × 108 CFU/g TPH—95% C10–C22—82% | [11] |
| Microbial consortia: Bacillus lentus Bacillus megaterium Bacillus pumilus Bacillus cereus Bacillus subtilis Bacillus sphaericus | Biofilter for phenol and petroleum hydrocarbon- contaminated wastewater treatment | Diesel and crude oil degradation | Individual strains, nutrient agar, 30 °C, 24 h. Individual colonies, nutrient broth, 30 °C, 48 h. | 250 g of soil, microbial consortium, pea straw (3% w/w), water holding capacity of 50%, 12 weeks | Bacterial population—0 day 104 CFU/g; max value—106 CFU/g TPH degraders—0 day 102 CFU/g; max value 104 CFU/g TPH degradation—96.1% | [18] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bertović, B.; Šabić Runjavec, M.; Todorović, N.; Zgrebec, I.; Vuković Domanovac, M. Biotechnological Potential of Oil-Tolerant Strains for Possible Use in Bioremediation. Sustainability 2024, 16, 563. https://doi.org/10.3390/su16020563
Bertović B, Šabić Runjavec M, Todorović N, Zgrebec I, Vuković Domanovac M. Biotechnological Potential of Oil-Tolerant Strains for Possible Use in Bioremediation. Sustainability. 2024; 16(2):563. https://doi.org/10.3390/su16020563
Chicago/Turabian StyleBertović, Barbara, Monika Šabić Runjavec, Nolla Todorović, Ivan Zgrebec, and Marija Vuković Domanovac. 2024. "Biotechnological Potential of Oil-Tolerant Strains for Possible Use in Bioremediation" Sustainability 16, no. 2: 563. https://doi.org/10.3390/su16020563







