Infection of Epstein–Barr Virus in Type III Latency Modulates Biogenesis of Exosomes and the Expression Profile of Exosomal miRNAs in the Burkitt Lymphoma Mutu Cell Lines
Abstract
:1. Introduction
2. Results
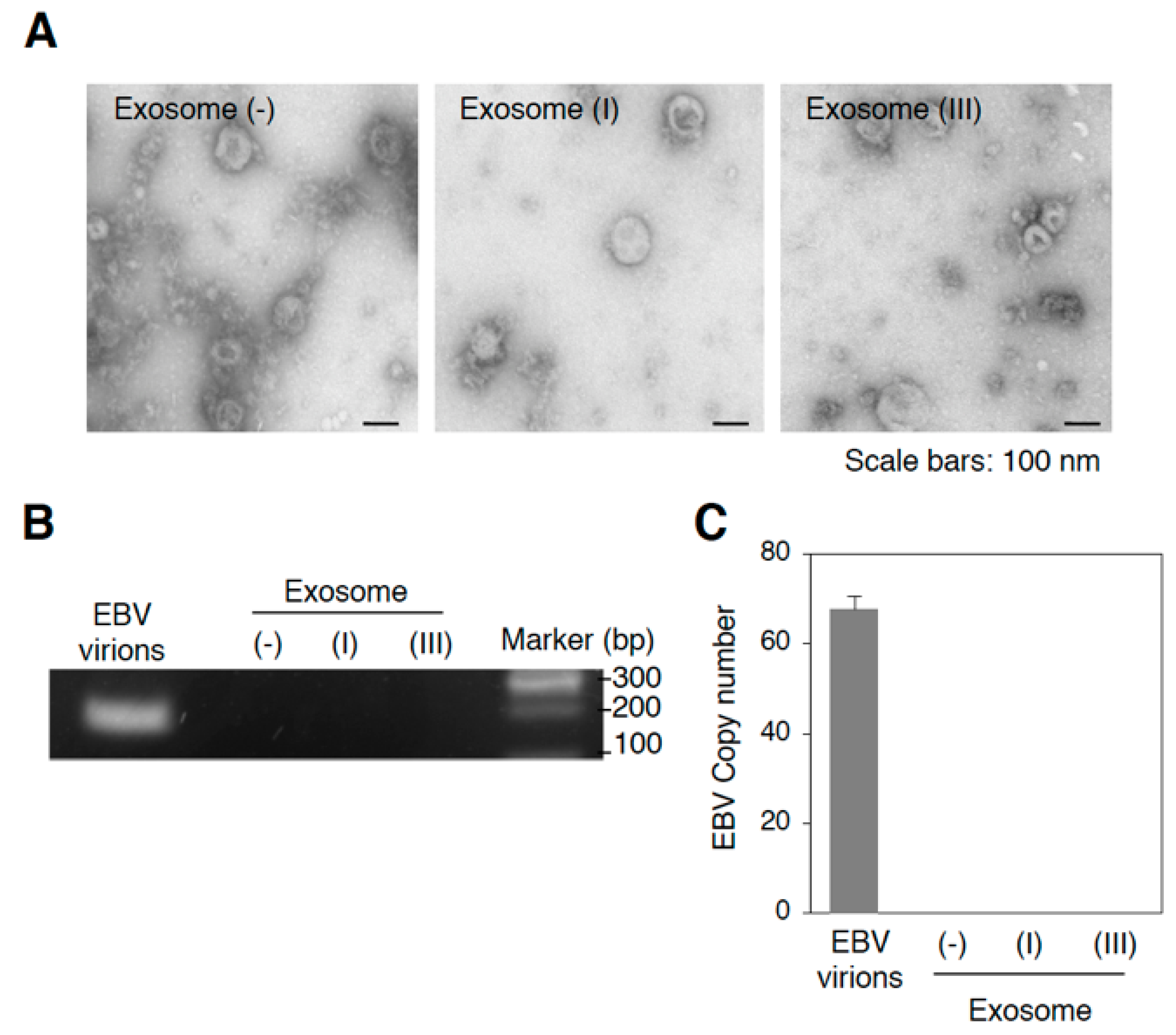
2.1. Infection with EBV Is Sufficient to Accelerate the Biogenesis of Exosomes in Mutu III Cells
2.2. More EBV miRNAs Are in Mutu III Cells and Exosomes (III) Than in EBV-Negative and Type I Latently Infected Cells and Their Exosomes
2.3. Characterization of Expression Profile of Cellular and Exosomal miRNAs of Mutu Cells
2.4. The Role of EXOmotifs in Sorting of Specific miRNA into Exosomes
2.5. Exosome-Mediated Transfer of EBV miRNAs to Target Epithelial Cells
3. Discussion
4. Materials and Methods
4.1. Cell Culture
4.2. Purification of Exosomes
4.3. Electron Microscopy
4.4. Quantitative PCR
4.5. Fluorescent In Situ Hybridization (FISH)
4.6. Immunofluorescence Staining
4.7. RNA Extraction
4.8. Next-Generation Sequencing (NGS)
4.9. Exosome Transfer Assay
4.10. Stem-Loop Real-Time PCR
4.11. Accession Number
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| EBV | Epstein–Barr virus |
| BL | Burkitt lymphoma |
| NGS | Next-generation sequence |
| miRNA | microRNA |
| MVB | multivesicular body |
References
- Longnecker, R.M.; Kieff, E.; Cohen, J.I. Epstein–Barr virus. In Fields Virology, 6th ed.; Knipe, M., Howley, P.M., Eds.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2013; pp. 1898–1959. [Google Scholar]
- Raab-Traub, N.; Dittmer, D.P. Viral effects on the content and function of extracellular vesicles. Nat. Rev. Microbiol. 2017, 15, 559–572. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meckes, D.G., Jr. Exosomal communication goes viral. J. Virol. 2015, 89, 5200–5203. [Google Scholar] [CrossRef] [PubMed]
- Wollert, T.; Hurley, J.H. Molecular mechanism of multivesicular body biogenesis by escrt complexes. Nature 2010, 464, 864–869. [Google Scholar] [CrossRef] [PubMed]
- Johnstone, R.M.; Adam, M.; Hammond, J.R.; Orr, L.; Turbide, C. Vesicle formation during reticulocyte maturation. Association of plasma membrane activities with released vesicles (exosomes). J. Biol. Chem. 1987, 262, 9412–9420. [Google Scholar] [PubMed]
- Van Niel, G.; D’Angelo, G.; Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Biol. 2018, 19, 213–228. [Google Scholar] [CrossRef] [PubMed]
- Raposo, G.; Stoorvogel, W. Extracellular vesicles: Exosomes, microvesicles, and friends. J. Cell Biol. 2013, 200, 373–383. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lasser, C.; Alikhani, V.S.; Ekstrom, K.; Eldh, M.; Paredes, P.T.; Bossios, A.; Sjostrand, M.; Gabrielsson, S.; Lotvall, J.; Valadi, H. Human saliva, plasma and breast milk exosomes contain RNA: Uptake by macrophages. J. Transl. Med. 2011, 9, 9. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tkach, M.; Thery, C. Communication by extracellular vesicles: Where we are and where we need to go. Cell 2016, 164, 1226–1232. [Google Scholar] [CrossRef] [PubMed]
- Vojtech, L.; Woo, S.; Hughes, S.; Levy, C.; Ballweber, L.; Sauteraud, R.P.; Strobl, J.; Westerberg, K.; Gottardo, R.; Tewari, M.; et al. Exosomes in human semen carry a distinctive repertoire of small non-coding RNAs with potential regulatory functions. Nucleic Acids Res. 2014, 42, 7290–7304. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Valadi, H.; Ekstrom, K.; Bossios, A.; Sjostrand, M.; Lee, J.J.; Lotvall, J.O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659. [Google Scholar] [CrossRef] [PubMed]
- Manterola, L.; Guruceaga, E.; Gallego Perez-Larraya, J.; Gonzalez-Huarriz, M.; Jauregui, P.; Tejada, S.; Diez-Valle, R.; Segura, V.; Sampron, N.; Barrena, C.; et al. A small noncoding RNA signature found in exosomes of gbm patient serum as a diagnostic tool. Neuro Oncol. 2014, 16, 520–527. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Zeringer, E.; Barta, T.; Schageman, J.; Cheng, A.; Vlassov, A.V. Analysis of the RNA content of the exosomes derived from blood serum and urine and its potential as biomarkers. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2014, 369, 20130502. [Google Scholar] [CrossRef] [PubMed]
- Batagov, A.O.; Kurochkin, I.V. Exosomes secreted by human cells transport largely mRNA fragments that are enriched in the 3’-untranslated regions. Biol. Direct. 2013, 8, 12. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. Metazoan microRNAs. Cell 2018, 173, 20–51. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Kuzembayeva, M.; Hayes, M.; Sugden, B. Multiple functions are mediated by the miRNAs of Epstein–Barr virus. Curr. Opin. Virol. 2014, 7, 61–65. [Google Scholar] [CrossRef] [PubMed]
- Vereide, D.T.; Seto, E.; Chiu, Y.F.; Hayes, M.; Tagawa, T.; Grundhoff, A.; Hammerschmidt, W.; Sugden, B. Epstein–Barr virus maintains lymphomas via its miRNAs. Oncogene 2014, 33, 1258–1264. [Google Scholar] [CrossRef] [PubMed]
- Tagawa, T.; Albanese, M.; Bouvet, M.; Moosmann, A.; Mautner, J.; Heissmeyer, V.; Zielinski, C.; Lutter, D.; Hoser, J.; Hastreiter, M.; et al. Epstein–Barr viral miRNAs inhibit antiviral cd4+ t cell responses targeting il-12 and peptide processing. J. Exp. Med. 2016, 213, 2065–2080. [Google Scholar] [CrossRef] [PubMed]
- Seto, E.; Moosmann, A.; Gromminger, S.; Walz, N.; Grundhoff, A.; Hammerschmidt, W. Micro RNAs of Epstein–Barr virus promote cell cycle progression and prevent apoptosis of primary human b cells. PLoS Pathog. 2010, 6, e1001063. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Feederle, R.; Linnstaedt, S.D.; Bannert, H.; Lips, H.; Bencun, M.; Cullen, B.R.; Delecluse, H.J. A viral microRNA cluster strongly potentiates the transforming properties of a human herpesvirus. PLoS Pathog. 2011, 7, e1001294. [Google Scholar] [CrossRef] [PubMed]
- Bernhardt, K.; Haar, J.; Tsai, M.H.; Poirey, R.; Feederle, R.; Delecluse, H.J. A viral microRNA cluster regulates the expression of pten, p27 and of a bcl-2 homolog. PLoS Pathog. 2016, 12, e1005405. [Google Scholar] [CrossRef] [PubMed]
- Albanese, M.; Tagawa, T.; Bouvet, M.; Maliqi, L.; Lutter, D.; Hoser, J.; Hastreiter, M.; Hayes, M.; Sugden, B.; Martin, L.; et al. Epstein–Barr virus microRNAs reduce immune surveillance by virus-specific cd8+ t cells. Proc. Natl. Acad. Sci. USA 2016, 113, E6467–E6475. [Google Scholar] [CrossRef] [PubMed]
- Shinozaki-Ushiku, A.; Kunita, A.; Isogai, M.; Hibiya, T.; Ushiku, T.; Takada, K.; Fukayama, M. Profiling of virus-encoded microRNAs in Epstein–Barr virus-associated gastric carcinoma and their roles in gastric carcinogenesis. J. Virol. 2015, 89, 5581–5591. [Google Scholar] [CrossRef] [PubMed]
- Marquitz, A.R.; Mathur, A.; Chugh, P.E.; Dittmer, D.P.; Raab-Traub, N. Expression profile of microRNAs in Epstein–Barr virus-infected ags gastric carcinoma cells. J. Virol. 2014, 88, 1389–1393. [Google Scholar] [CrossRef] [PubMed]
- Kang, D.; Skalsky, R.L.; Cullen, B.R. Ebv bart microRNAs target multiple pro-apoptotic cellular genes to promote epithelial cell survival. PLoS Pathog. 2015, 11, e1004979. [Google Scholar] [CrossRef] [PubMed]
- Kanda, T.; Miyata, M.; Kano, M.; Kondo, S.; Yoshizaki, T.; Iizasa, H. Clustered microRNAs of the Epstein–Barr virus cooperatively downregulate an epithelial cell-specific metastasis suppressor. J. Virol. 2015, 89, 2684–2697. [Google Scholar] [CrossRef] [PubMed]
- Cai, L.M.; Lyu, X.M.; Luo, W.R.; Cui, X.F.; Ye, Y.F.; Yuan, C.C.; Peng, Q.X.; Wu, D.H.; Liu, T.F.; Wang, E.; et al. Ebv-mir-bart7-3p promotes the emt and metastasis of nasopharyngeal carcinoma cells by suppressing the tumor suppressor pten. Oncogene 2015, 34, 2156–2166. [Google Scholar] [CrossRef] [PubMed]
- Pratt, Z.L.; Kuzembayeva, M.; Sengupta, S.; Sugden, B. The microRNAs of Epstein–Barr virus are expressed at dramatically differing levels among cell lines. Virology 2009, 386, 387–397. [Google Scholar] [CrossRef] [PubMed]
- Gallo, A.; Vella, S.; Miele, M.; Timoneri, F.; Di Bella, M.; Bosi, S.; Sciveres, M.; Conaldi, P.G. Global profiling of viral and cellular non-coding RNAs in Epstein–Barr virus-induced lymphoblastoid cell lines and released exosome cargos. Cancer Lett. 2017, 388, 334–343. [Google Scholar] [CrossRef] [PubMed]
- Cameron, J.E.; Fewell, C.; Yin, Q.; McBride, J.; Wang, X.; Lin, Z.; Flemington, E.K. Epstein–Barr virus growth/latency iii program alters cellular microRNA expression. Virology 2008, 382, 257–266. [Google Scholar] [CrossRef] [PubMed]
- Frixa, T.; Donzelli, S.; Blandino, G. Oncogenic microRNAs: Key players in malignant transformation. Cancers 2015, 7, 2466–2485. [Google Scholar] [CrossRef] [PubMed]
- Pegtel, D.M.; van de Garde, M.D.; Middeldorp, J.M. Viral miRNAs exploiting the endosomal-exosomal pathway for intercellular cross-talk and immune evasion. Biochim. Biophys. Acta 2011, 1809, 715–721. [Google Scholar] [CrossRef] [PubMed]
- Pegtel, D.M.; Cosmopoulos, K.; Thorley-Lawson, D.A.; van Eijndhoven, M.A.; Hopmans, E.S.; Lindenberg, J.L.; de Gruijl, T.D.; Wurdinger, T.; Middeldorp, J.M. Functional delivery of viral miRNAs via exosomes. Proc. Natl. Acad. Sci. USA 2010, 107, 6328–6333. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, Z.; Swan, K.; Zhang, X.; Cao, S.; Brett, Z.; Drury, S.; Strong, M.J.; Fewell, C.; Puetter, A.; Wang, X.; et al. Secreted oral epithelial cell membrane vesicles induce Epstein–Barr virus reactivation in latently infected b cells. J. Virol. 2016, 90, 3469–3479. [Google Scholar] [CrossRef] [PubMed]
- Janas, T.; Janas, M.M.; Sapon, K.; Janas, T. Mechanisms of RNA loading into exosomes. FEBS Lett. 2015, 589, 1391–1398. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, J.; Li, S.; Li, L.; Li, M.; Guo, C.; Yao, J.; Mi, S. Exosome and exosomal microRNA: Trafficking, sorting, and function. Genom. Proteom. Bioinform. 2015, 13, 17–24. [Google Scholar] [CrossRef] [PubMed]
- Wong, A.M.; Kong, K.L.; Tsang, J.W.; Kwong, D.L.; Guan, X.Y. Profiling of Epstein–Barr virus-encoded microRNAs in nasopharyngeal carcinoma reveals potential biomarkers and oncomirs. Cancer 2012, 118, 698–710. [Google Scholar] [CrossRef] [PubMed]
- Qiu, J.; Cosmopoulos, K.; Pegtel, M.; Hopmans, E.; Murray, P.; Middeldorp, J.; Shapiro, M.; Thorley-Lawson, D.A. A novel persistence associated ebv miRNA expression profile is disrupted in neoplasia. PLoS Pathog. 2011, 7, e1002193. [Google Scholar] [CrossRef] [PubMed]
- Piccaluga, P.P.; Navari, M.; De Falco, G.; Ambrosio, M.R.; Lazzi, S.; Fuligni, F.; Bellan, C.; Rossi, M.; Sapienza, M.R.; Laginestra, M.A.; et al. Virus-encoded microRNA contributes to the molecular profile of ebv-positive burkitt lymphomas. Oncotarget 2016, 7, 224–240. [Google Scholar] [CrossRef] [PubMed]
- Navari, M.; Etebari, M.; De Falco, G.; Ambrosio, M.R.; Gibellini, D.; Leoncini, L.; Piccaluga, P.P. The presence of Epstein–Barr virus significantly impacts the transcriptional profile in immunodeficiency-associated burkitt lymphoma. Front. Microbiol. 2015, 6, 556. [Google Scholar] [CrossRef] [PubMed]
- Cosmopoulos, K.; Pegtel, M.; Hawkins, J.; Moffett, H.; Novina, C.; Middeldorp, J.; Thorley-Lawson, D.A. Comprehensive profiling of Epstein–Barr virus microRNAs in nasopharyngeal carcinoma. J. Virol. 2009, 83, 2357–2367. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Andrade, T.A.; Evangelista, A.F.; Campos, A.H.; Poles, W.A.; Borges, N.M.; Camillo, C.M.; Soares, F.A.; Vassallo, J.; Paes, R.P.; Zerbini, M.C.; et al. A microRNA signature profile in ebv+ diffuse large b-cell lymphoma of the elderly. Oncotarget 2014, 5, 11813–11826. [Google Scholar] [CrossRef] [PubMed]
- Amoroso, R.; Fitzsimmons, L.; Thomas, W.A.; Kelly, G.L.; Rowe, M.; Bell, A.I. Quantitative studies of Epstein–Barr virus-encoded microRNAs provide novel insights into their regulation. J. Virol. 2011, 85, 996–1010. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, K.; Sekizuka, T.; Uehara, T.; Hishima, T.; Mine, S.; Fukumoto, H.; Sato, Y.; Hasegawa, H.; Kuroda, M.; Katano, H. Next-generation sequencing of miRNAs in clinical samples of Epstein–Barr virus-associated b-cell lymphomas. Cancer Med. 2017, 6, 605–618. [Google Scholar] [CrossRef] [PubMed]
- Motsch, N.; Alles, J.; Imig, J.; Zhu, J.; Barth, S.; Reineke, T.; Tinguely, M.; Cogliatti, S.; Dueck, A.; Meister, G.; et al. MicroRNA profiling of Epstein–Barr virus-associated NK/T-cell lymphomas by deep sequencing. PLoS ONE 2012, 7, e42193. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Imig, J.; Motsch, N.; Zhu, J.Y.; Barth, S.; Okoniewski, M.; Reineke, T.; Tinguely, M.; Faggioni, A.; Trivedi, P.; Meister, G.; et al. MicroRNA profiling in Epstein–Barr virus-associated b-cell lymphoma. Nucleic Acids Res. 2011, 39, 1880–1893. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.J.; Chen, G.H.; Chen, Y.H.; Liu, C.Y.; Chang, K.P.; Chang, Y.S.; Chen, H.C. Characterization of Epstein–Barr virus miRNAome in nasopharyngeal carcinoma by deep sequencing. PLoS ONE 2010, 5, e12745. [Google Scholar] [CrossRef] [PubMed]
- Alles, J.; Menegatti, J.; Motsch, N.; Hart, M.; Eichner, N.; Reinhardt, R.; Meister, G.; Grasser, F.A. MiRNA expression profiling of Epstein–Barr virus-associated nktl cell lines by illumina deep sequencing. FEBS Open Biol. 2016, 6, 251–263. [Google Scholar] [CrossRef] [PubMed]
- Hoshina, S.; Sekizuka, T.; Kataoka, M.; Hasegawa, H.; Hamada, H.; Kuroda, M.; Katano, H. Profile of exosomal and intracellular microRNA in gamma-herpesvirus-infected lymphoma cell lines. PLoS ONE 2016, 11, e0162574. [Google Scholar] [CrossRef] [PubMed]
- Takada, K.; Horinouchi, K.; Ono, Y.; Aya, T.; Osato, T.; Takahashi, M.; Hayasaka, S. An Epstein–Barr virus-producer line akata: Establishment of the cell line and analysis of viral DNA. Virus Genes 1991, 5, 147–156. [Google Scholar] [CrossRef] [PubMed]
- Escola, J.M.; Kleijmeer, M.J.; Stoorvogel, W.; Griffith, J.M.; Yoshie, O.; Geuze, H.J. Selective enrichment of tetraspan proteins on the internal vesicles of multivesicular endosomes and on exosomes secreted by human b-lymphocytes. J. Biol. Chem. 1998, 273, 20121–20127. [Google Scholar] [CrossRef] [PubMed]
- Baietti, M.F.; Zhang, Z.; Mortier, E.; Melchior, A.; Degeest, G.; Geeraerts, A.; Ivarsson, Y.; Depoortere, F.; Coomans, C.; Vermeiren, E.; et al. Syndecan-syntenin-alix regulates the biogenesis of exosomes. Nat. Cell Biol. 2012, 14, 677–685. [Google Scholar] [CrossRef] [PubMed]
- Lotvall, J.; Hill, A.F.; Hochberg, F.; Buzas, E.I.; Di Vizio, D.; Gardiner, C.; Gho, Y.S.; Kurochkin, I.V.; Mathivanan, S.; Quesenberry, P.; et al. Minimal experimental requirements for definition of extracellular vesicles and their functions: A position statement from the international society for extracellular vesicles. J. Extracell. Vesicles 2014, 3, 26913. [Google Scholar] [CrossRef] [PubMed]
- Verweij, F.J.; van Eijndhoven, M.A.; Hopmans, E.S.; Vendrig, T.; Wurdinger, T.; Cahir-McFarland, E.; Kieff, E.; Geerts, D.; van der Kant, R.; Neefjes, J.; et al. Lmp1 association with cd63 in endosomes and secretion via exosomes limits constitutive nf-kappab activation. EMBO J. 2011, 30, 2115–2129. [Google Scholar] [CrossRef] [PubMed]
- Nanbo, A.; Kawanishi, E.; Yoshida, R.; Yoshiyama, H. Exosomes derived from Epstein–Barr virus-infected cells are internalized via caveola-dependent endocytosis and promote phenotypic modulation in target cells. J. Virol. 2013, 87, 10334–10347. [Google Scholar] [CrossRef] [PubMed]
- Meckes, D.G., Jr.; Shair, K.H.; Marquitz, A.R.; Kung, C.P.; Edwards, R.H.; Raab-Traub, N. Human tumor virus utilizes exosomes for intercellular communication. Proc. Natl. Acad. Sci. USA 2010, 107, 20370–20375. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hurwitz, S.N.; Nkosi, D.; Conlon, M.M.; York, S.B.; Liu, X.; Tremblay, D.C.; Meckes, D.G., Jr. Cd63 regulates Epstein–Barr virus lmp1 exosomal packaging, enhancement of vesicle production, and noncanonical nf-kappab signaling. J. Virol. 2017, 91. [Google Scholar] [CrossRef] [PubMed]
- Villarroya-Beltri, C.; Gutierrez-Vazquez, C.; Sanchez-Madrid, F.; Mittelbrunn, M. Analysis of microRNA and protein transfer by exosomes during an immune synapse. Methods Mol. Biol. 2013, 1024, 41–51. [Google Scholar] [PubMed]
- Trajkovic, K.; Hsu, C.; Chiantia, S.; Rajendran, L.; Wenzel, D.; Wieland, F.; Schwille, P.; Brugger, B.; Simons, M. Ceramide triggers budding of exosome vesicles into multivesicular endosomes. Science 2008, 319, 1244–1247. [Google Scholar] [CrossRef] [PubMed]
- Hurwitz, S.N.; Cheerathodi, M.R.; Nkosi, D.; York, S.B.; Meckes, D.G., Jr. Tetraspanin cd63 bridges autophagic and endosomal processes to regulate exosomal secretion and intracellular signaling of Epstein–Barr virus lmp1. J. Virol. 2018, 92. [Google Scholar] [CrossRef] [PubMed]
- Nanbo, A.; Sugden, A.; Sugden, B. The coupling of synthesis and partitioning of ebv’s plasmid replicon is revealed in live cells. EMBO J. 2007, 26, 4252–4262. [Google Scholar] [CrossRef] [PubMed]
- Kosaka, N.; Iguchi, H.; Hagiwara, K.; Yoshioka, Y.; Takeshita, F.; Ochiya, T. Neutral sphingomyelinase 2 (nsmase2)-dependent exosomal transfer of angiogenic microRNAs regulate cancer cell metastasis. J. Biol. Chem. 2013, 288, 10849–10859. [Google Scholar] [CrossRef] [PubMed]
- Pekow, J.; Meckel, K.; Dougherty, U.; Butun, F.; Mustafi, R.; Lim, J.; Crofton, C.; Chen, X.; Joseph, L.; Bissonnette, M. Tumor suppressors mir-143 and mir-145 and predicted target proteins api5, erk5, k-ras, and irs-1 are differentially expressed in proximal and distal colon. Am. J. Physiol. Gastrointest. Liver Physiol. 2015, 308, G179–G187. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Gu, S.; Cao, H.; Li, Z.; Xiang, Z.; Hu, K.; Han, X. Mir-877-3p targets smad7 and is associated with myofibroblast differentiation and bleomycin-induced lung fibrosis. Sci. Rep. 2016, 6, 30122. [Google Scholar] [CrossRef] [PubMed]
- Shi, Q.; Xu, X.; Liu, Q.; Luo, F.; Shi, J.; He, X. MicroRNA-877 acts as a tumor suppressor by directly targeting eef2k in renal cell carcinoma. Oncol. Lett. 2016, 11, 1474–1480. [Google Scholar] [CrossRef] [PubMed]
- Chowdhari, S.; Saini, N. Hsa-mir-4516 mediated downregulation of stat3/cdk6/ube2n plays a role in puva induced apoptosis in keratinocytes. J. Cell. Physiol. 2014, 229, 1630–1638. [Google Scholar] [CrossRef] [PubMed]
- Reza, A.M.; Choi, Y.J.; Yasuda, H.; Kim, J.H. Human adipose mesenchymal stem cell-derived exosomal-miRNAs are critical factors for inducing anti-proliferation signalling to a2780 and skov-3 ovarian cancer cells. Sci. Rep. 2016, 6, 38498. [Google Scholar] [CrossRef] [PubMed]
- Swaminathan, S.; Hu, X.; Zheng, X.; Kriga, Y.; Shetty, J.; Zhao, Y.; Stephens, R.; Tran, B.; Baseler, M.W.; Yang, J.; et al. Interleukin-27 treated human macrophages induce the expression of novel microRNAs which may mediate anti-viral properties. Biochem. Biophys. Res. Commun. 2013, 434, 228–234. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gregory, C.D.; Rowe, M.; Rickinson, A.B. Different Epstein–Barr virus-b cell interactions in phenotypically distinct clones of a burkitt’s lymphoma cell line. J. Gen. Virol. 1990, 71 Pt 7, 1481–1495. [Google Scholar] [CrossRef] [PubMed]
- Kitagawa, N.; Goto, M.; Kurozumi, K.; Maruo, S.; Fukayama, M.; Naoe, T.; Yasukawa, M.; Hino, K.; Suzuki, T.; Todo, S.; et al. Epstein–Barr virus-encoded poly(a)(-) RNA supports burkitt’s lymphoma growth through interleukin-10 induction. EMBO J. 2000, 19, 6742–6750. [Google Scholar] [CrossRef] [PubMed]
- Menezes, J.; Leibold, W.; Klein, G.; Clements, G. Establishment and characterization of an Epstein–Barr virus (EBC)-negative lymphoblastoid b cell line (BJA-b) from an exceptional, ebv-genome-negative african burkitt’s lymphoma. Biomedicine 1975, 22, 276–284. [Google Scholar] [PubMed]
- Matsuo, T.; Heller, M.; Petti, L.; O’Shiro, E.; Kieff, E. Persistence of the entire Epstein–Barr virus genome integrated into human lymphocyte DNA. Science 1984, 226, 1322–1325. [Google Scholar] [CrossRef] [PubMed]
- Lawrence, J.B.; Villnave, C.A.; Singer, R.H. Sensitive, high-resolution chromatin and chromosome mapping in situ: Presence and orientation of two closely integrated copies of ebv in a lymphoma line. Cell 1988, 52, 51–61. [Google Scholar] [PubMed]
- Yoshiyama, H.; Imai, S.; Shimizu, N.; Takada, K. Epstein–Barr virus infection of human gastric carcinoma cells: Implication of the existence of a new virus receptor different from cd21. J. Virol. 1997, 71, 5688–5691. [Google Scholar] [PubMed]
- Barranco, S.C.; Townsend, C.M., Jr.; Casartelli, C.; Macik, B.G.; Burger, N.L.; Boerwinkle, W.R.; Gourley, W.K. Establishment and characterization of an in vitro model system for human adenocarcinoma of the stomach. Cancer Res. 1983, 43, 1703–1709. [Google Scholar] [PubMed]
- Takada, K.; Ono, Y. Synchronous and sequential activation of latently infected Epstein–Barr virus genomes. J. Virol. 1989, 63, 445–449. [Google Scholar] [PubMed]
- Takada, K. Cross-linking of cell surface immunoglobulins induces Epstein–Barr virus in burkitt lymphoma lines. Int. J. Cancer 1984, 33, 27–32. [Google Scholar] [CrossRef] [PubMed]
- Vereide, D.T.; Sugden, B. Lymphomas differ in their dependence on Epstein–Barr virus. Blood 2011, 117, 1977–1985. [Google Scholar] [CrossRef] [PubMed]
- Nanbo, A.; Terada, H.; Kachi, K.; Takada, K.; Matsuda, T. Roles of cell signaling pathways in cell-to-cell contact-mediated Epstein–Barr virus transmission. J. Virol. 2012, 86, 9285–9296. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.C.; Liem, A.; Lambert, P.F.; Sugden, B. Dissecting the regulation of ebv’s bart miRNAs in carcinomas. Virology 2017, 505, 148–154. [Google Scholar] [CrossRef] [PubMed]




| Samples | Raw Reads | Annotated with Database (% Total Annotated Reads) | |||||
|---|---|---|---|---|---|---|---|
| Annotated Reads (% Raw Reads) | miRBase | Homo_Sapiens. GRCh38.ncrna | Human-Trna | ||||
| Homo sapiens | EBV | ||||||
| Mutu− | cell | 119,769 | 42,692 (35.65) | 11,958 (28.01) | 1 (0.00) | 28,749 (67.34) | 1984 (4.65) |
| exosome | 66,018 | 25,690 (38.91) | 12,040 (46.87) | 1 (0.00) | 13,051 (50.80) | 598 (2.33) | |
| Mutu I | cell | 26,388 | 10,934 (41.44) | 6459 (59.07) | 461 (7.14) | 3612 (33.03) | 402 (3.68) |
| exosome | 535,527 | 64,085 (11.97) | 32,041 (50.00) | 1233 (3.85) | 28,267 (44.11) | 2544 (3.97) | |
| Mutu III | cell | 1,559,462 | 1,087,053 (69.71) | 769,785 (70.81) | 157,016 (14.44) | 137,381 (12.64) | 22,871 (2.10) |
| exosome | 2,445,816 | 522,266 (21.35) | 80,434 (15.40) | 6489 (8.07) | 422,846 (80.96) | 12,497 (2.39) | |
| Mutu− | Mutu I | Mutu III | ||||
|---|---|---|---|---|---|---|
| miRNA | % *,1 | miRNA | % | miRNA | % | |
| 1 | miR-92a-1//miR-92a-2-3p | 18.20 | miR-92a-1//miR-92a-2-3p | 34.31 | miR-148a-3p | 22.06 |
| 2 | miR-378a-3p | 13.65 | miR-182-5p | 7.90 | miR-182-5p | 18.70 |
| 3 | miR-28-3p | 6.33 | miR-BART10-3p | 3.74 | miR-92a-1//miR-92a-2-3p | 15.03 |
| 4 | miR-181a-2//miR-181a-1-5p | 6.15 | miR-378a-3p | 3.62 | miR-BART10-3p | 8.48 |
| 5 | miR-182-5p | 5.98 | miR-148a-3p | 3.06 | miR-181a-2//miR-181a-1-5p | 4.27 |
| 6 | miR-486-1//miR-486-2-5p | 4.66 | miR-25-3p | 3.04 | miR-181b-1//miR-181b-2-5p | 2.89 |
| 7 | miR-181b-1//miR-181b-2-5p | 4.33 | let-7f-1//let-7f-2-5p | 2.68 | miR-25-3p | 2.71 |
| 8 | miR-148a-3p | 3.73 | miR-191-5p | 2.13 | miR-BART6-3p | 2.35 |
| 9 | miR-25-3p | 3.00 | miR-28-3p | 2.04 | miR-378a-3p | 2.02 |
| 10 | miR-101-1//miR-101-2-3p | 2.74 | miR-103a-2//miR-103a-1-3p | 1.95 | miR-BART17-5p | 1.62 |
| 11 | miR-1275-5p | 2.64 | miR-21-3p | 1.95 | miR-BART7-5p | 1.40 |
| 12 | miR-21-3p | 2.29 | let-7a-1//let-7a-2//let-7a-3-5p | 1.91 | miR-191-5p | 1.34 |
| 13 | miR-103a-2//miR-103a-1-3p | 2.18 | miR-30e-3p | 1.79 | miR-21-5p | 1.13 |
| 14 | miR-30e-3p | 2.11 | miR-181a-2//miR-181a-1-5p | 1.60 | miR-30e-3p | 1.10 |
| 15 | miR-191-5p | 1.85 | miR-128-1-5p | 1.51 | miR-21-3p | 1.04 |
| 16 | miR-140-3p | 1.27 | miR-101-1//miR-101-2-3p | 1.48 | let-7f-1//let-7f-2-5p | 0.97 |
| 17 | miR-423-5p | 0.98 | miR-486-1//miR-486-2-5p | 1.44 | miR-BART22-3p | 0.92 |
| 18 | miR-128-1-5p | 0.85 | miR-181b-1//miR-181b-2-5p | 1.41 | miR-BART11-3p | 0.91 |
| 19 | miR-423-3p | 0.65 | miR-106b-3p | 1.18 | miR-181a-2-3p | 0.78 |
| 20 | miR-210-3p | 0.65 | miR-BART7-5p | 1.07 | miR-103a-2//miR-103a-1-3p | 0.78 |
| 21 | miR-769-5p | 0.62 | miR-320a-3p | 1.03 | miR-155-5p | 0.75 |
| 22 | let-7i-5p | 0.61 | let-7i-5p | 0.92 | miR-101-1//miR-101-2-3p | 0.73 |
| 23 | miR-30d-5p | 0.60 | miR-1307-3p | 0.88 | miR-BART13-5p | 0.61 |
| 24 | miR-106b-3p | 0.59 | miR-140-3p | 0.76 | miR-148a-5p | 0.44 |
| 25 | miR-186-5p | 0.58 | miR-423-5p | 0.70 | miR-30d-5p | 0.39 |
| 26 | miR-142-3p | 0.57 | miR-30d-5p | 0.68 | let-7a-1//let-7a-2//let-7a-3-5p | 0.37 |
| 27 | miR-184-3p | 0.51 | miR-1275-5p | 0.67 | miR-106b-3p | 0.33 |
| 28 | miR-320a-3p | 0.50 | miR-877-5p | 0.64 | let-7c-5p | 0.27 |
| 29 | miR-21-5p | 0.46 | miR-16-1//miR-16-2-5p | 0.58 | miR-125b-2-3p | 0.27 |
| 30 | let-7a-1//let-7a-2//let-7a-3-5p | 0.44 | miR-186-5p | 0.56 | miR-423-5p | 0.26 |
| 31 | miR-1307-3p | 0.42 | miR-577-5p | 0.56 | miR-26a-1//miR-26a-2-5p | 0.21 |
| 32 | miR-143-3p | 0.41 | miR-142-3p | 0.53 | miR-7974-3p | 0.18 |
| 33 | miR-181a-1-3p | 0.38 | miR-BART11-3p | 0.51 | miR-146b-5p | 0.18 |
| 34 | miR-296-3p | 0.37 | miR-BART6-3p | 0.45 | miR-486-1//miR-486-2-5p | 0.17 |
| 35 | miR-1285-1//miR-1285-2-3p | 0.32 | let-7g-5p | 0.42 | let-7i-5p | 0.17 |
| 36 | miR-27b-3p | 0.31 | miR-93-5p | 0.39 | miR-27b-3p | 0.17 |
| 37 | miR-151a-5p | 0.27 | miR-423-3p | 0.38 | miR-16-1//miR-16-2-5p | 0.17 |
| 38 | miR-1260a-5p | 0.25 | miR-143-3p | 0.33 | miR-181a-1-3p | 0.14 |
| 39 | miR-877-5p | 0.25 | miR-151a-5p | 0.30 | miR-532-5p | 0.14 |
| 40 | miR-130b-3p | 0.23 | miR-98-5p | 0.29 | miR-92a-1-5p | 0.14 |
| Exosome (−) | Exosome (I) | Exosome (III) | |||||||
|---|---|---|---|---|---|---|---|---|---|
| miRNA | % *,1 | -Fold *,2 | miRNA | % | -Fold | miRNA | % | -Fold | |
| 1 | miR-92a-1//miR-92a-2-3p | 43.77 | 2.40 | miR-92a-1//miR-92a-2-3p | 11.8 | 0.35 | miR-92a-1//miR-92a-2-3p | 35.16 | 2.34 |
| 2 | miR-10b-5p | 8.16 | - | miR-486-1//miR-486-2-5p | 9.39 | 6.53 | miR-486-1//miR-486-2-5p | 19.55 | 112.19 |
| 3 | miR-378a-3p | 7.51 | 0.55 | miR-181a-2//miR-181a-1-5p | 7.29 | 4.54 | miR-182-5p | 5.00 | 0.27 |
| 4 | miR-486-1//miR-486-2-5p | 6.60 | 1.42 | miR-378a-3p | 6.65 | 1.84 | miR-10b-5p | 4.40 | 699.66 |
| 5 | miR-25-3p | 5.67 | 1.89 | miR-186-5p | 5.03 | 8.98 | miR-BART10-3p | 4.20 | 0.50 |
| 6 | miR-181b-1//miR-181b-2-5p | 3.77 | 0.87 | miR-16-1//miR-16-2-5p | 4.82 | 8.37 | miR-181a-2//miR-181a-1-5p | 3.37 | 0.79 |
| 7 | miR-28-3p | 2.90 | 0.46 | miR-191-5p | 4.35 | 2.04 | miR-148a-3p | 2.76 | 0.13 |
| 8 | miR-182-5p | 2.68 | 0.45 | miR-27b-3p | 2.81 | 11.59 | miR-7704-5p | 2.34 | 758.35 |
| 9 | miR-181a-2//miR-181a-1-5p | 1.80 | 0.29 | miR-28-3p | 2.70 | 1.32 | miR-378a-3p | 2.30 | 1.14 |
| 10 | miR-143-3p | 1.33 | 3.27 | miR-30e-5p | 2.61 | 11.51 | miR-25-3p | 2.28 | 0.84 |
| 11 | miR-191-5p | 1.07 | 0.58 | miR-142-5p | 2.50 | 20.64 | miR-181b-1//miR-181b-2-5p | 2.18 | 0.75 |
| 12 | miR-148a-3p | 0.91 | 0.24 | miR-143-3p | 2.32 | 6.98 | miR-191-5p | 1.90 | 1.42 |
| 13 | miR-320a-3p | 0.79 | 1.59 | miR-26a-1//miR-26a-2-5p | 2.05 | 22.54 | miR-423-5p | 1.59 | 6.02 |
| 14 | miR-423-5p | 0.70 | 0.71 | miR-21-5p | 2.04 | 9.62 | miR-BART17-5p | 1.45 | 0.90 |
| 15 | miR-21-3p | 0.63 | 0.28 | miR-30d-5p | 1.99 | 2.92 | miR-BART11-3p | 1.39 | 1.53 |
| 16 | miR-30e-3p | 0.63 | 0.30 | miR-10b-5p | 1.93 | 25.44 | miR-BART13-5p | 0.79 | 1.29 |
| 17 | let-7a-1//let-7a-2//let-7a-3-5p | 0.61 | 1.39 | miR-25-3p | 1.84 | 0.61 | miR-BART6-3p | 0.73 | 0.31 |
| 18 | miR-106b-3p | 0.57 | 0.97 | miR-22-3p | 1.63 | 7.18 | miR-30d-5p | 0.62 | 1.58 |
| 19 | miR-877-5p | 0.57 | 2.24 | miR-93-5p | 1.36 | 3.45 | miR-181a-2-3p | 0.40 | 0.50 |
| 20 | miR-103a-2//miR-103a-1-3p | 0.51 | 0.24 | let-7a-1//let-7a-2//let-7a-3-5p | 1.33 | 0.70 | let-7f-1//let-7f-2-5p | 0.38 | 0.39 |
| 21 | miR-577-5p | 0.49 | 3.84 | miR-148a-3p | 1.15 | 0.38 | miR-103a-2//miR-103a-1-3p | 0.36 | 0.46 |
| 22 | miR-101-1//miR-101-2-3p | 0.48 | 0.17 | miR-15a-5p | 1.07 | 17.69 | miR-BART7-5p | 0.36 | 0.26 |
| 23 | let-7f-1//let-7f-2-5p | 0.44 | 3.48 | miR-BART11-3p | 0.98 | 1.90 | miR-1307-3p | 0.36 | 4.03 |
| 24 | miR-423-3p | 0.41 | 0.63 | miR-BART7-5p | 0.93 | 0.87 | miR-155-5p | 0.32 | 0.43 |
| 25 | miR-140-3p | 0.39 | 0.31 | let-7f-1//let-7f-2-5p | 0.89 | 0.33 | miR-4516-5p | 0.31 | 406.30 |
| 26 | miR-432-5p | 0.35 | 0.57 | miR-127-3p | 0.84 | 55.84 | miR-106b-3p | 0.26 | 0.79 |
| 27 | let-7i-5p | 0.33 | 1.46 | miR-151a-3p | 0.76 | 8.32 | miR-BHRF1-1-5p | 0.26 | 8.01 |
| 28 | miR-130b-3p | 0.30 | 0.73 | miR-BART10-3p | 0.68 | 0.18 | miR-877-5p | 0.24 | 22.33 |
| 29 | miR-1307-3p | 0.27 | 0.46 | miR-182-5p | 0.68 | 0.09 | miR-320a-3p | 0.23 | 8.79 |
| 30 | miR-186-5p | 0.27 | 1.73 | miR-126-5p | 0.63 | 10.49 | let-7a-1//let-7a-2//let-7a-3-5p | 0.20 | 0.55 |
| 31 | miR-93-5p | 0.25 | 7.05 | miR-1246-5p | 0.63 | 41.65 | miR-21-3p | 0.19 | 0.18 |
| 32 | miR-4516-5p | 0.21 | 0.35 | miR-101-1//miR-101-2-3p | 0.58 | 0.39 | miR-6087-5p | 0.19 | - |
| 33 | miR-30d-5p | 0.19 | 0.97 | miR-151a-5p | 0.57 | 1.89 | miR-BART22-3p | 0.18 | 0.19 |
| 34 | miR-1304-3p | 0.17 | 0.27 | miR-103a-2//miR-103a-1-3p | 0.57 | 0.29 | miR-423-3p | 0.17 | 2.11 |
| 35 | miR-1246-5p | 0.17 | 19.26 | miR-192-5p | 0.54 | 8.97 | miR-30e-3p | 0.17 | 0.15 |
| 36 | miR-210-3p | 0.17 | 0.26 | miR-128-1//miR-128-2-3p | 0.47 | 30.86 | miR-125b-2-3p | 0.16 | 0.58 |
| 37 | miR-16-1//miR-16-2-5p | 0.16 | 1.07 | miR-425-5p | 0.44 | 29.35 | let-7c-5p | 0.15 | 0.55 |
| 38 | miR-320b-1//miR-320b-2-3p | 0.15 | 8.54 | let-7g-5p | 0.44 | 1.04 | miR-21-5p | 0.14 | 0.12 |
| 39 | miR-660-5p | 0.15 | 4.27 | miR-1307-5p | 0.43 | 14.05 | miR-BART19-5p | 0.14 | 1.99 |
| 40 | miR-1285-1//miR-1285-2-3p | 0.15 | 0.49 | miR-181b-1//miR-181b-2-5p | 0.42 | 0.30 | miR-143-3p | 0.13 | 25.43 |
| miRNA | % Mature Annotated | Concentrate (-Fold) | Sequence | EXOmotifs | ||
|---|---|---|---|---|---|---|
| Cell | Exosome | |||||
| Cell | miR-6087-5p | N/D | 13.92 | - | TGAGGCGGGGGGGCGAGC | 8 |
| miR-7704-5p | 0.003 | 2.34 | 758.35 | CGGGGTCGGCGGCGACGTG | 3 | |
| miR-10b-5p | 0.006 | 4.40 | 699.66 | TACCCTGTAGAACCGAATTTGTG | 1 | |
| miR-4516-5p | 0.001 | 0.31 | 406.30 | GGGAGAAGGGTCGGGGC | 3 | |
| miR-486-1//miR-486-2-5p | 0.174 | 19.55 | 112.19 | TCCTGTACTGAGCTGCCCCGAG | 4 | |
| EBV | miR-BART7-3p | 0.015 | 0.13 | 8.53 | CATCATAGTCCAGTGTCCAGGG | 2 |
| miR-BHRF1-1-5p | 0.033 | 0.26 | 8.01 | TAACCTGATCAGCCCCGGAGTT | 2 | |
| miR-BART2-5p | 0.015 | 0.09 | 6.27 | TATTTTCTGCATTCGCCCTTGC | 1 | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nanbo, A.; Katano, H.; Kataoka, M.; Hoshina, S.; Sekizuka, T.; Kuroda, M.; Ohba, Y. Infection of Epstein–Barr Virus in Type III Latency Modulates Biogenesis of Exosomes and the Expression Profile of Exosomal miRNAs in the Burkitt Lymphoma Mutu Cell Lines. Cancers 2018, 10, 237. https://doi.org/10.3390/cancers10070237
Nanbo A, Katano H, Kataoka M, Hoshina S, Sekizuka T, Kuroda M, Ohba Y. Infection of Epstein–Barr Virus in Type III Latency Modulates Biogenesis of Exosomes and the Expression Profile of Exosomal miRNAs in the Burkitt Lymphoma Mutu Cell Lines. Cancers. 2018; 10(7):237. https://doi.org/10.3390/cancers10070237
Chicago/Turabian StyleNanbo, Asuka, Harutaka Katano, Michiyo Kataoka, Shiho Hoshina, Tsuyoshi Sekizuka, Makoto Kuroda, and Yusuke Ohba. 2018. "Infection of Epstein–Barr Virus in Type III Latency Modulates Biogenesis of Exosomes and the Expression Profile of Exosomal miRNAs in the Burkitt Lymphoma Mutu Cell Lines" Cancers 10, no. 7: 237. https://doi.org/10.3390/cancers10070237





