Ubiquilin Networking in Cancers
Abstract
:1. Introduction
2. UBQLN’s Genetic Variants in Cancers
3. UBQLNs and p53
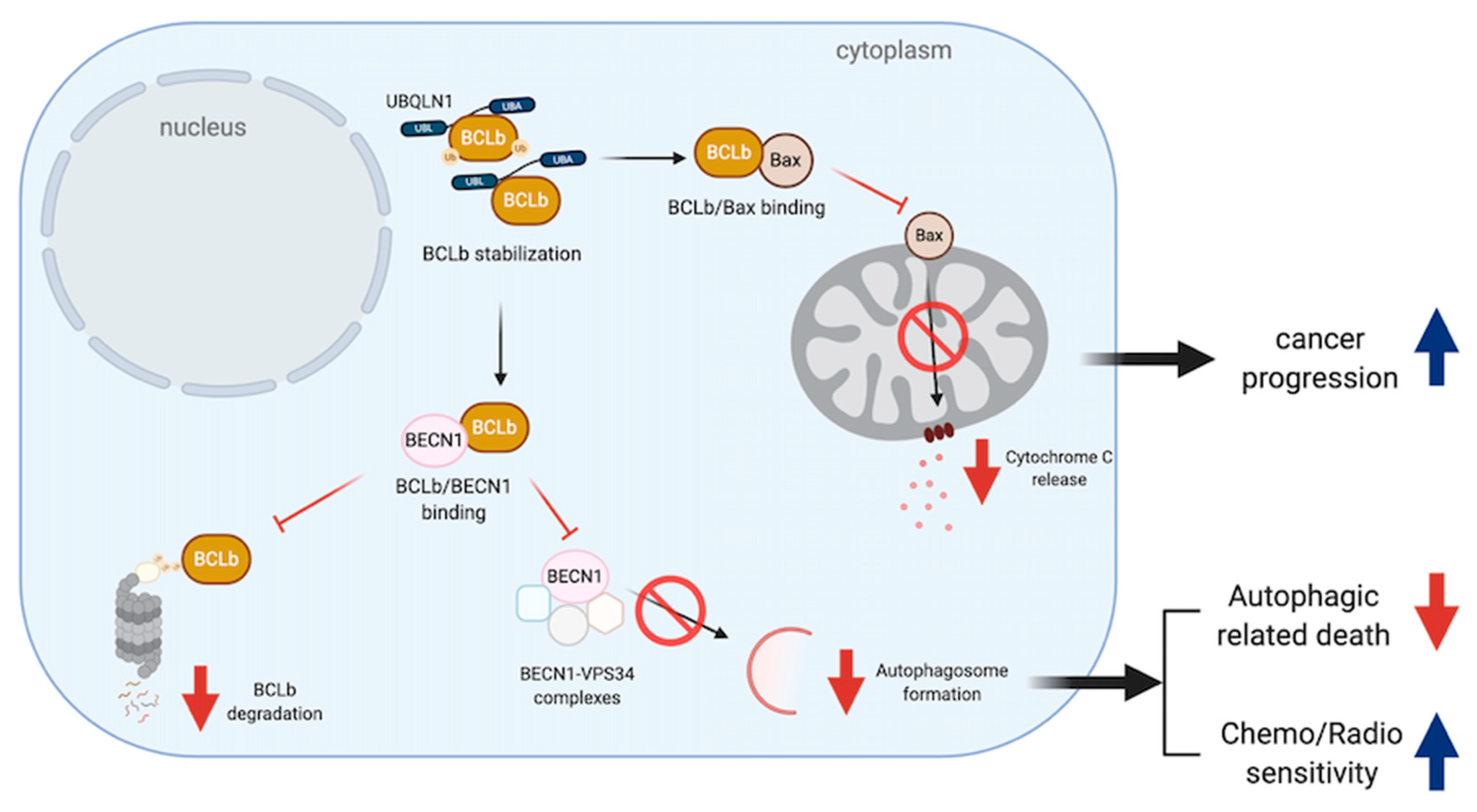
4. UBQLNs and BCL2 Family Proteins
5. UBQLNs and Membrane Receptor Proteins
6. UBQLNs and EMT Markers
7. UBQLNs and DNA Repair Proteins
8. Emerging Evidence of Ubiquilins/ncRNAs Axis in Cancer
9. Conclusions and Future Perspectives
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Seok Ko, H.; Uehara, T.; Tsuruma, K.; Nomura, Y. Ubiquilin interacts with ubiquitylated proteins and proteasome through its ubiquitin-associated and ubiquitin-like domains. FEBS Lett. 2004, 566, 110–114. [Google Scholar] [CrossRef]
- Lee, D.Y.; Brown, E.J. Ubiquilins in the crosstalk among proteolytic pathways. Biol. Chem. 2012, 393, 441–447. [Google Scholar] [CrossRef] [PubMed]
- N’Diaye, E.N.; Kajihara, K.K.; Hsieh, I.; Morisaki, H.; Debnath, J.; Brown, E.J. PLIC proteins or ubiquilins regulate autophagy-dependent cell survival during nutrient starvation. EMBO Rep. 2009, 10, 173–179. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rothenberg, C.; Srinivasan, D.; Mah, L.; Kaushik, S.; Peterhoff, C.M.; Ugolino, J.; Fang, S.; Cuervo, A.M.; Nixon, R.A.; Monteiro, M.J. Ubiquilin functions in autophagy and is degraded by chaperone-mediated autophagy. Hum. Mol. Genet. 2010, 19, 3219–3232. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Şentürk, M.; Lin, G.; Zuo, Z.; Mao, D.; Watson, E.; Mikos, A.G.; Bellen, H.J. Ubiquilins Regulate Autophagic Flux through mTOR Signaling and Lysosomal Acidification. Nat. Cell Biol. 2019, 21, 384. [Google Scholar] [CrossRef] [PubMed]
- Lim, P.J.; Danner, R.; Liang, J.; Doong, H.; Harman, C.; Srinivasan, D.; Rothenberg, C.; Wang, H.; Ye, Y.; Fang, S.; et al. Ubiquilin and p97/VCP bind erasin, forming a complex involved in ERAD. J. Cell Biol. 2009, 187, 201–217. [Google Scholar] [CrossRef] [Green Version]
- Marín, I. The ubiquilin gene family: Evolutionary patterns and functional insights. BMC Evol. Biol. 2014, 14, 63. [Google Scholar] [CrossRef] [Green Version]
- Haapasalo, A.; Viswanathan, J.; Bertram, L.; Soininen, H.; Tanzi, R.E.; Hiltunen, M. Emerging role of Alzheimer’s disease-associated ubiquilin-1 in protein aggregation: Figure 1. Biochem. Soc. Trans. 2010, 38, 150–155. [Google Scholar] [CrossRef] [Green Version]
- El Ayadi, A.; Stieren, E.S.; Barral, J.M.; Boehning, D. Ubiquilin-1 and protein quality control in Alzheimer disease. Prion 2013, 7, 164–169. [Google Scholar] [CrossRef] [Green Version]
- Ling, S.-C.C.; Polymenidou, M.; Cleveland, D.W.W. Converging mechanisms in als and FTD: Disrupted RNA and protein homeostasis. Neuron 2013, 79, 416–438. [Google Scholar] [CrossRef] [Green Version]
- Zhang, K.Y.; Yang, S.; Warraich, S.T.; Blair, I.P. Ubiquilin 2: A component of the ubiquitin-proteasome system with an emerging role in neurodegeneration. Int. J. Biochem. Cell Biol. 2014, 50, 123–126. [Google Scholar] [CrossRef] [PubMed]
- Jantrapirom, S.; Lo Piccolo, L.; Yamaguchi, M. Non-Proteasomal UbL-UbA Family of Proteins in Neurodegeneration. Int. J. Mol. Sci. 2019, 20, 1893. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jachimowicz, R.D.; Beleggia, F.; Isensee, J.; Velpula, B.B.; Goergens, J.; Bustos, M.A.; Doll, M.A.; Shenoy, A.; Checa-Rodriguez, C.; Wiederstein, J.L.; et al. UBQLN4 Represses Homologous Recombination and Is Overexpressed in Aggressive Tumors. Cell 2019, 176, 505–519. [Google Scholar] [CrossRef] [Green Version]
- Gao, J.; Aksoy, B.A.; Dogrusoz, U.; Dresdner, G.; Gross, B.; Sumer, S.O.; Sun, Y.; Jacobsen, A.; Sinha, R.; Larsson, E.; et al. Integrative Analysis of Complex Cancer Genomics and Clinical Profiles Using the cBioPortal. Sci. Signal 2013, 6, pl1. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cerami1, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio Cancer Genomics Portal: An Open Platform for Exploring Multidimensional Cancer Genomics Data. Cancer Discov. 2012, 2, 401. [Google Scholar] [CrossRef] [Green Version]
- Shimada, K.; Fujii, T.; Tatsumi, Y.; Anai, S.; Fujimoto, K.; Konishi, N. Ubiquilin2 as a novel marker for detection of urothelial carcinoma cells in urine. Diagn. Cytopathol. 2016, 44, 3–9. [Google Scholar] [CrossRef]
- Chen, G.; Wang, X.; Yu, J.; Varambally, S.; Yu, J.; Thomas, D.G.; Lin, M.-Y.; Vishnu, P.; Wang, Z.; Wang, R.; et al. Autoantibody Profiles Reveal Ubiquilin 1 as a Humoral Immune Response Target in Lung Adenocarcinoma. Cancer Res. 2007, 67, 3461–3467. [Google Scholar] [CrossRef] [Green Version]
- Patel, K.; Farlow, E.C.; Kim, A.W.; Lee, B.S.; Basu, S.; Coon, J.S.; DeCresce, D.; Thimothy, L.; Walters, K.A.; Fhied, C.; et al. Enhancement of a multianalyte serum biomarker panel to identify lymph node metastases in non-small cell lung cancer with circulating autoantibody biomarkers. Int. J. Cancer 2011, 129, 133–142. [Google Scholar] [CrossRef]
- Bao, J.; Jiang, X.; Zhu, X.; Dai, G.; Dou, R.; Liu, X.; Sheng, H.; Liang, Z.; Yu, H. Clinical significance of ubiquilin 1 in gastric cancer. Medicine 2018, 97, e9701. [Google Scholar] [CrossRef]
- Wang, Y.; Lu, J.; Zhao, X.; Feng, Y.; Lv, S.; Mu, Y.; Wang, D.; Fu, H.; Chen, Y.; Li, Y. Prognostic significance of Ubiquilin1 expression in invasive breast cancer. Cancer Biomark. 2015, 15, 635–643. [Google Scholar] [CrossRef]
- Hackelford, R.E.; Kaufmann, W.K.; Paules, R.S. Cell cycle control, checkpoint mechanisms, and genotoxic stress. Environ. Health Perspect. 1999, 107, 5–24. [Google Scholar]
- Bartek, J.; Lukas, J. DNA damage checkpoints: From initiation to recovery or adaptation. Curr. Opin. Cell Biol. 2007, 19, 238–245. [Google Scholar] [CrossRef]
- Eastman, A. Cell cycle checkpoints and their impact on anticancer therapeutic strategies. J. Cell. Biochem. 2004, 91, 223–231. [Google Scholar] [CrossRef] [PubMed]
- Nakanishi, M.; Shimada, M.; Niida, H. Genetic instability in cancer cells by impaired cell cycle checkpoints. Cancer Sci. 2006, 97, 984–989. [Google Scholar] [CrossRef] [PubMed]
- Mijit, M.; Caracciolo, V.; Melillo, A.; Amicarelli, F.; Giordano, A. Role of p53 in the Regulation of Cellular Senescence. Biomolecules 2020, 10, 420. [Google Scholar] [CrossRef] [Green Version]
- Vanzo, R.; Bartkova, J.; Merchut-Maya, J.M.; Hall, A.; Bouchal, J.; Dyrskjøt, L.; Frankel, L.B.; Gorgoulis, V.; Maya-Mendoza, A.; Jäättelä, M.; et al. Autophagy role(s) in response to oncogenes and DNA replication stress. Cell Death Differ. 2020, 27, 1134–1153. [Google Scholar] [CrossRef]
- Childs, B.G.; Durik, M.; Baker, D.J.; van Deursen, J.M. Cellular senescence in aging and age-related disease: From mechanisms to therapy. Nat. Med. 2015, 21, 1424–1435. [Google Scholar] [CrossRef] [Green Version]
- Feng, P.; Scott, C.W.; Cho, N.-H.; Nakamura, H.; Chung, Y.-H.; Monteiro, M.J.; Jung, J.U. Kaposi’s Sarcoma-Associated Herpesvirus K7 Protein Targets a Ubiquitin-Like/Ubiquitin-Associated Domain-Containing Protein To Promote Protein Degradation. Mol. Cell. Biol. 2004, 24, 3938–3948. [Google Scholar] [CrossRef] [Green Version]
- Huang, S.; Li, Y.; Yuan, X.; Zhao, M.; Wang, J.; Li, Y.; Li, Y.; Lin, H.; Zhang, Q.; Wang, W.; et al. The UbL-UBA Ubiquilin4 protein functions as a tumor suppressor in gastric cancer by p53-dependent and p53-independent regulation of p21. Cell Death Differ. 2019, 26, 516–530. [Google Scholar] [CrossRef] [Green Version]
- Knights, C.D.; Catania, J.; Di Giovanni, S.; Muratoglu, S.; Perez, R.; Swartzbeck, A.; Quong, A.A.; Zhang, X.; Beerman, T.; Pestell, R.G.; et al. Distinct p53 acetylation cassettes differentially influence gene-expression patterns and cell fate. J. Cell Biol. 2006, 173, 533–544. [Google Scholar] [CrossRef]
- Nakayama, K.I.; Nakayama, K. Regulation of the cell cycle by SCF-type ubiquitin ligases. Semin. Cell Dev. Biol. 2005, 16, 323–333. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.W.; Lee, M.S.; Camus, S.; Ghim, J.; Yang, M.R.; Oh, W.; Ha, N.C.; Lane, D.P.; Song, J. Differential regulation of p53 and p21 by MKRN1 E3 ligase controls cell cycle arrest and apoptosis. EMBO J. 2009, 28, 2100–2113. [Google Scholar] [CrossRef] [Green Version]
- Soria, G.; Gottifredi, V. PCNA-coupled p21 degradation after DNA damage: The exception that confirms the rule? DNA Repair (Amst). 2010, 9, 354–364. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Amador, V.; Ge, S.; Santamaría, P.G.; Guardavaccaro, D.; Pagano, M. APC/CCdc20 Controls the Ubiquitin-Mediated Degradation of p21 in Prometaphase. Mol. Cell 2007, 27, 462–473. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Han, J.; Kim, Y.L.; Lee, K.W.; Her, N.G.; Ha, T.K.; Yoon, S.; Jeong, S.I.; Lee, J.H.; Kang, M.J.; Lee, M.G.; et al. ZNF313 is a novel cell cycle activator with an E3 ligase activity inhibiting cellular senescence by destabilizing p21WAF1. Cell Death Differ. 2013, 20, 1055–1067. [Google Scholar] [CrossRef] [Green Version]
- Gozuacik, D.; Kimchi, A. Autophagy as a cell death and tumor suppressor mechanism. Oncogene 2004. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Apel, A.; Herr, I.; Schwarz, H.; Rodemann, H.P.; Mayer, A. Blocked Autophagy Sensitizes Resistant Carcinoma Cells to Radiation Therapy. Cancer Res. 2008, 68, 1485–1494. [Google Scholar] [CrossRef] [Green Version]
- Hjerpe, R.; Bett, J.S.; Keuss, M.J.; Solovyova, A.; McWilliams, T.G.; Johnson, C.; Sahu, I.; Varghese, J.; Wood, N.; Wightman, M.; et al. UBQLN2 Mediates Autophagy-Independent Protein Aggregate Clearance by the Proteasome. Cell 2016, 166, 935–949. [Google Scholar] [CrossRef] [Green Version]
- Jantrapirom, S.; Lo Piccolo, L.; Yoshida, H.; Yamaguchi, M. Depletion of Ubiquilin induces an augmentation in soluble ubiquitinated Drosophila TDP-43 to drive neurotoxicity in the fly. Biochim. Biophys. Acta—Mol. Basis Dis. 2018, 1864, 3038–3049. [Google Scholar] [CrossRef]
- Jantrapirom, S.; Lo Piccolo, L.; Yoshida, H.; Yamaguchi, M. A new Drosophila model of Ubiquilin knockdown shows the effect of impaired proteostasis on locomotive and learning abilities. Exp. Cell Res. 2018, 362, 461–471. [Google Scholar] [CrossRef]
- Sun, Q.; Liu, T.; Yuan, Y.; Guo, Z.; Xie, G.; Du, S.; Lin, X.; Xu, Z.; Liu, M.; Wang, W.; et al. MiR-200c inhibits autophagy and enhances radiosensitivity in breast cancer cells by targeting UBQLN1. Int. J. Cancer 2015, 136, 1003–1012. [Google Scholar] [CrossRef] [PubMed]
- Ryan, K.M. P53 and autophagy in cancer: Guardian of the genome meets guardian of the proteome. Eur. J. Cancer 2011, 47, 44–50. [Google Scholar] [CrossRef] [PubMed]
- Budanov, A.V.; Karin, M. p53 Target Genes Sestrin1 and Sestrin2 Connect Genotoxic Stress and mTOR Signaling. Cell 2008, 134, 451–460. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kurlawala, Z.; Shah, P.P.; Shah, C.; Beverly, L.J. The STI and UBA Domains of UBQLN1 Are Critical Determinants of Substrate Interaction and Proteostasis. J. Cell. Biochem. 2017, 118, 2261–2270. [Google Scholar] [CrossRef] [PubMed]
- Nougarede, A.; Popgeorgiev, N.; Kassem, L.; Omarjee, S.; Borel, S.; Mikaelian, I.; Lopez, J.; Gadet, R.; Marcillat, O.; Treilleux, I.; et al. Breast Cancer Targeting through Inhibition of the Endoplasmic Reticulum-Based Apoptosis Regulator Nrh/BCL2L10. Cancer Res. 2018, 78, 1404–1417. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beverly, L.J.; Lockwood, W.W.; Shah, P.P.; Erdjument-Bromage, H.; Varmus, H. Ubiquitination, localization, and stability of an anti-apoptotic BCL2-like protein, BCL2L10/BCLb, are regulated by Ubiquilin1. Proc. Natl. Acad. Sci. USA 2012, 109, e119–e126. [Google Scholar] [CrossRef] [Green Version]
- Van de Kooij, B.; Rooswinkel, R.W.; Kok, F.; Herrebout, M.; de Vries, E.; Paauwe, M.; Janssen, G.M.C.; van Veelen, P.A.; Borst, J. Polyubiquitination and proteasomal turnover controls the anti-apoptotic activity of Bcl-B. Oncogene 2013, 32, 5439–5448. [Google Scholar] [CrossRef] [Green Version]
- Beverly, L.J.; Varmus, H.E. MYC-induced myeloid leukemogenesis is accelerated by all six members of the antiapoptotic BCL family. Oncogene 2009, 28, 1274–1279. [Google Scholar] [CrossRef] [Green Version]
- Krajewska, M.; Kitada, S.; Winter, J.N.; Variakojis, D.; Lichtenstein, A.; Zhai, D.; Cuddy, M.; Huang, X.; Luciano, F.; Baker, C.H.; et al. Bcl-B Expression in Human Epithelial and Nonepithelial Malignancies. Clin. Cancer Res. 2008, 14, 3011–3021. [Google Scholar] [CrossRef] [Green Version]
- Ke, N.; Godzik, A.; Reed, J.C. Bcl-B, a Novel Bcl-2 Family Member That Differentially Binds and Regulates Bax and Bak. J. Biol. Chem. 2001, 276, 12481–12484. [Google Scholar] [CrossRef] [Green Version]
- Robert, G.; Gastaldi, C.; Puissant, A.; Hamouda, A.; Jacquel, A.; Dufies, M.; Belhacene, N.; Colosetti, P.; Reed, J.C.; Auberger, P.; et al. The anti-apoptotic Bcl-B protein inhibits BECN1-dependent autophagic cell death. Autophagy 2012, 8, 637–649. [Google Scholar] [CrossRef] [Green Version]
- He, J.; Deng, L.; Liu, H.; Chen, T.; Chen, S.; Xia, S.; Liu, Y. BCL2L10/BECN1 modulates hepatoma cells autophagy by regulating PI3K/AKT signaling pathway. Aging 2019, 11, 350. [Google Scholar] [CrossRef] [PubMed]
- Tsukamoto, S.; Shimada, K.; Honoki, K.; Kido, A.; Akahane, M.; Tanaka, Y.; Konishi, N. Ubiquilin 2 enhances osteosarcoma progression through resistance to hypoxic stress. Oncol. Rep. 2015, 33, 1799–1806. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dhanasekaran, D.N.; Reddy, E.P. JNK signaling in apoptosis. Oncogene 2008, 27, 6245–6251. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cevenini, A.; Orrù, S.; Mancini, A.; Alfieri, A.; Buono, P.; Imperlini, E. Molecular signatures of the insulin-like growth factor 1-mediated epithelial-mesenchymal transition in breast, lung and gastric cancers. Int. J. Mol. Sci. 2018, 19, 2411. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Alencar, S.A.; Lopes, J.C.D. A comprehensive in silico analysis of the functional and structural impact of SNPs in the IGF1R gene. J. Biomed. Biotechnol. 2010, 2010, 715139. [Google Scholar] [CrossRef] [Green Version]
- Brahmkhatri, V.P.; Prasanna, C.; Atreya, H.S. Insulin-Like Growth Factor System in Cancer: Novel Targeted Therapies. Biomed Res. Int. 2015, 2015, 538019. [Google Scholar] [CrossRef] [Green Version]
- Vishwamitra, D.; George, S.K.; Shi, P.; Kaseb, A.O.; Amin, H.M. Type I insulin-like growth factor receptor signaling in hematological malignancies. Oncotarget 2017, 8, 1814. [Google Scholar] [CrossRef] [Green Version]
- Boone, D.N.; Lee, A. V Targeting the insulin-like growth factor receptor: Developing biomarkers from gene expression profiling. Crit. Rev. Oncog. 2012, 17, 161–173. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tognon, C.E.; Sorensen, P.H.B. Targeting the insulin-like growth factor 1 receptor (IGF1R) signaling pathway for cancer therapy. Expert Opin. Ther. Targets 2012, 16, 33–48. [Google Scholar] [CrossRef]
- Yoneyama, Y.; Lanzerstorfer, P.; Niwa, H.; Umehara, T.; Shibano, T.; Yokoyama, S.; Chida, K.; Weghuber, J.; Hakuno, F.; Takahashi, S.-I. IRS-1 acts as an endocytic regulator of IGF-I receptor to facilitate sustained IGF signaling. Elife 2018, 7, e32893. [Google Scholar] [CrossRef] [PubMed]
- Dufresne, A.M.; Smith, R.J. The Adapter Protein GRB10 Is an Endogenous Negative Regulator of Insulin-Like Growth Factor Signaling. Endocrinology 2005, 146, 4399–4409. [Google Scholar] [CrossRef] [Green Version]
- Buckley, D.A.; Cheng, A.; Kiely, P.A.; Tremblay, M.L.; O’Connor, R. Regulation of insulin-like growth factor type I (IGF-I) receptor kinase activity by protein tyrosine phosphatase 1B (PTP-1B) and enhanced IGF-I-mediated suppression of apoptosis and motility in PTP-1B-deficient fibroblasts. Mol. Cell. Biol. 2002, 22, 1998–2010. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kurlawala, Z.; Dunaway, R.; Shah, P.P.; Gosney, J.A.; Siskind, L.J.; Ceresa, B.P.; Beverly, L.J. Regulation of insulin-like growth factor receptors by Ubiquilin1. Biochem. J. 2017, 474, 4105–4118. [Google Scholar] [CrossRef]
- Saliba, R.S.; Pangalos, M.; Moss, S.J. The ubiquitin-like protein Plic-1 enhances the membrane insertion of GABAA receptors by increasing their stability within the endoplasmic reticulum. J. Biol. Chem. 2008, 283, 18538–18544. [Google Scholar] [CrossRef] [Green Version]
- Bedford, F.K.; Kittler, J.T.; Muller, E.; Thomas, P.; Uren, J.M.; Merlo, D.; Wisden, W.; Triller, A.; Smart, T.G.; Moss, S.J. GABAA receptor cell surface number and subunit stability are regulated by the ubiquitin-like protein Plic-1. Nat. Neurosci. 2001, 4, 908–916. [Google Scholar] [CrossRef] [PubMed]
- Ficklin, M.B.; Zhao, S.; Feng, G. Ubiquilin-1 Regulates Nicotine-induced Up-regulation of Neuronal Nicotinic Acetylcholine Receptors. J. Biol. Chem. 2005, 280, 34088–34095. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- N’Diaye, E.-N.; Hanyaloglu, A.C.; Kajihara, K.K.; Puthenveedu, M.A.; Wu, P.; von Zastrow, M.; Brown, E.J. The Ubiquitin-like Protein PLIC-2 Is a Negative Regulator of G Protein-coupled Receptor Endocytosis. Mol. Biol. Cell 2008, 19, 1252–1260. [Google Scholar] [CrossRef] [Green Version]
- Whiteley, A.M.; Prado, M.A.; Peng, I.; Abbas, A.R.; Haley, B.; Paulo, J.A.; Reichelt, M.; Katakam, A.; Sagolla, M.; Modrusan, Z.; et al. Ubiquilin1 promotes antigen-receptor mediated proliferation by eliminating mislocalized mitochondrial proteins. Elife 2017, 6, e26435. [Google Scholar] [CrossRef] [PubMed]
- Wrobel, L.; Topf, U.; Bragoszewski, P.; Wiese, S.; Sztolsztener, M.E.; Oeljeklaus, S.; Varabyova, A.; Lirski, M.; Chroscicki, P.; Mroczek, S.; et al. Mistargeted mitochondrial proteins activate a proteostatic response in the cytosol. Nature 2015, 524, 485–488. [Google Scholar] [CrossRef]
- Wang, X.; Chen, X.J. A cytosolic network suppressing mitochondria-mediated proteostatic stress and cell death. Nature 2015, 524, 481–484. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Topf, U.; Wrobel, L.; Chacinska, A. Chatty Mitochondria: Keeping Balance in Cellular Protein Homeostasis. Trends Cell Biol. 2016, 26, 577–586. [Google Scholar] [CrossRef] [PubMed]
- Thiery, J.P.; Acloque, H.; Huang, R.Y.J.; Nieto, M.A. Epithelial-Mesenchymal Transitions in Development and Disease. Cell 2009, 139, 871–890. [Google Scholar] [CrossRef] [PubMed]
- Kalluri, R.; Weinberg, R.A. The basics of epithelial-mesenchymal transition. J. Clin. Invest. 2009, 119, 1420. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Strouhalova, K.; Přechová, M.; Gandalovičová, A.; Brábek, J.; Gregor, M.; Rosel, D. Vimentin intermediate filaments as potential target for cancer treatment. Cancers 2020, 12, 184. [Google Scholar] [CrossRef] [Green Version]
- Loh, C.-Y.; Chai, J.Y.; Tang, T.F.; Wong, W.F.; Sethi, G.; Shanmugam, M.K.; Chong, P.P.; Looi, C.Y. The E-Cadherin and N-Cadherin Switch in Epithelial-to-Mesenchymal Transition: Signaling, Therapeutic Implications, and Challenges. Cells 2019, 8, 1118. [Google Scholar] [CrossRef] [Green Version]
- Cano, A.; Pérez-Moreno, M.A.; Rodrigo, I.; Locascio, A.; Blanco, M.J.; del Barrio, M.G.; Portillo, F.; Nieto, M.A. The transcription factor Snail controls epithelial–mesenchymal transitions by repressing E-cadherin expression. Nat. Cell Biol. 2000, 2, 76–83. [Google Scholar] [CrossRef]
- Wong, T.-S.; Gao, W.; Chan, J.Y.-W. Transcription Regulation of E-Cadherin by Zinc Finger E-Box Binding Homeobox Proteins in Solid Tumors. Biomed Res. Int. 2014, 2014. [Google Scholar] [CrossRef] [Green Version]
- Vergara, D.; Simeone, P.; Franck, J.; Trerotola, M.; Giudetti, A.; Capobianco, L.; Tinelli, A.; Bellomo, C.; Fournier, I.; Gaballo, A.; et al. Translating epithelial mesenchymal transition markers into the clinic: Novel insights from proteomics. EuPA Open Proteom. 2016, 10, 31–41. [Google Scholar] [CrossRef] [Green Version]
- Yamazaki, T.; Liu, L.; Lazarev, D.; Al-Zain, A.; Fomin, V.; Yeung, P.L.; Chambers, S.M.; Lu, C.-W.; Studer, L.; Manley, J.L. TCF3 alternative splicing controlled by hnRNP H/F regulates E-cadherin expression and hESC pluripotency. Genes Dev. 2018, 32, 1161–1174. [Google Scholar] [CrossRef] [Green Version]
- Vesuna, F.; van Diest, P.; Chen, J.H.; Raman, V. Twist is a transcriptional repressor of E-cadherin gene expression in breast cancer. Biochem. Biophys. Res. Commun. 2008, 367, 235–241. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lahiri, S.K.; Zhao, J. Krüppel-like factor 8 emerges as an important regulator of cancer. Am. J. Transl. Res. 2012, 4, 357. [Google Scholar]
- Zhang, Y.; Xu, L.; Li, A.; Han, X. The roles of ZEB1 in tumorigenic progression and epigenetic modifications. Biomed. Pharmacother. 2019, 110, 400–408. [Google Scholar] [CrossRef]
- Byles, V.; Zhu, L.; Lovaas, J.D.; Chmilewski, L.K.; Wang, J.; Faller, D.V.; Dai, Y. SIRT1 induces EMT by cooperating with EMT transcription factors and enhances prostate cancer cell migration and metastasis. Oncogene 2012, 31, 4619–4629. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- HOU, L.; LI, Q.; YU, Y.; LI, M.; ZHANG, D. SET8 induces epithelial-mesenchymal transition and enhances prostate cancer cell metastasis by cooperating with ZEB1. Mol. Med. Rep. 2016, 13, 1681–1688. [Google Scholar] [CrossRef] [PubMed]
- Shah, P.P.; Lockwood, W.W.; Saurabh, K.; Kurlawala, Z.; Shannon, S.P.; Waigel, S.; Zacharias, W.; Beverly, L.J. Ubiquilin1 Represses Migration and Epithelial to Mesenchymal Transition of Human Non-small Cell Lung Cancer Cells. Oncogene 2015, 34, 1709. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Torgovnick, A.; Schumacher, B. DNA repair mechanisms in cancer development and therapy. Front. Genet. 2015, 6, 157. [Google Scholar] [CrossRef] [Green Version]
- Ciccia, A.; Elledge, S.J. The DNA Damage Response: Making It Safe to Play with Knives. Mol. Cell 2010, 40, 179–204. [Google Scholar] [CrossRef] [Green Version]
- Davis, A.J.; Chen, D.J. DNA double strand break repair via non-homologous end-joining. Transl. Cancer Res. 2013, 2, 130–143. [Google Scholar]
- Wright, W.D.; Shah, S.S.; Heyer, W.-D. Homologous recombination and the repair of DNA double-strand breaks. J. Biol. Chem. 2018, 293, 10524–10535. [Google Scholar] [CrossRef] [Green Version]
- Jachimowicz, R.D.; Reinhardt, H.C. UBQLN4 promotes non-homologous end joining by repressing DNA end-resection. Mol. Cell. Oncol. 2019, 6, 1575692. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kozlov, S.V.; Graham, M.E.; Jakob, B.; Tobias, F.; Kijas, A.W.; Tanuji, M.; Chen, P.; Robinson, P.J.; Taucher-Scholz, G.; Suzuki, K.; et al. Autophosphorylation and ATM Activation. J. Biol. Chem. 2011, 286, 9107–9119. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shiloh, Y.; Ziv, Y. The ATM protein kinase: Regulating the cellular response to genotoxic stress, and more. Nat. Rev. Mol. Cell Biol. 2013, 14, 197–210. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Gong, Y.; Peng, B.; Shi, R.; Fan, D.; Zhao, H.; Zhu, M.; Zhang, H.; Lou, Z.; Zhou, J.; et al. MRE11 UFMylation promotes ATM activation. Nucleic Acids Res. 2019, 47, 4124–4135. [Google Scholar] [CrossRef] [Green Version]
- Wei, L.; Sun, J.; Zhang, N.; Zheng, Y.; Wang, X.; Lv, L.; Liu, J.; Xu, Y.; Shen, Y.; Yang, M. Noncoding RNAs in gastric cancer: Implications for drug resistance. Mol. Cancer 2020, 19, 62. [Google Scholar] [CrossRef] [Green Version]
- Crudele, F.; Bianchi, N.; Reali, E.; Galasso, M.; Agnoletto, C.; Volinia, S. The network of non-coding RNAs and their molecular targets in breast cancer. Mol. Cancer 2020, 19, 61. [Google Scholar] [CrossRef] [Green Version]
- Bhat, A.A.; Younes, S.N.; Raza, S.S.; Zarif, L.; Nisar, S.; Ahmed, I.; Mir, R.; Kumar, S.; Sharawat, S.K.; Hashem, S.; et al. Role of non-coding RNA networks in leukemia progression, metastasis and drug resistance. Mol. Cancer 2020, 19, 57. [Google Scholar] [CrossRef]
- Almstrup, K.; Lobo, J.; Mørup, N.; Belge, G.; Rajpert-De Meyts, E.; Looijenga, L.H.J.; Dieckmann, K.-P. Application of miRNAs in the diagnosis and monitoring of testicular germ cell tumours. Nat. Rev. Urol. 2020, 17, 201–213. [Google Scholar] [CrossRef]
- Shimono, Y.; Zabala, M.; Cho, R.W.; Lobo, N.; Dalerba, P.; Qian, D.; Diehn, M.; Liu, H.; Panula, S.P.; Chiao, E.; et al. Downregulation of miRNA-200c Links Breast Cancer Stem Cells with Normal Stem Cells. Cell 2009, 138, 592–603. [Google Scholar] [CrossRef] [Green Version]
- Jurmeister, S.; Baumann, M.; Balwierz, A.; Keklikoglou, I.; Ward, A.; Uhlmann, S.; Zhang, J.D.; Wiemann, S.; Sahin, O. MicroRNA-200c Represses Migration and Invasion of Breast Cancer Cells by Targeting Actin-Regulatory Proteins FHOD1 and PPM1F. Mol. Cell. Biol. 2012, 32, 633–651. [Google Scholar] [CrossRef] [Green Version]
- Gregory, P.A.; Bert, A.G.; Paterson, E.L.; Barry, S.C.; Tsykin, A.; Farshid, G.; Vadas, M.A.; Khew-Goodall, Y.; Goodall, G.J. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat. Cell Biol. 2008, 10, 593–601. [Google Scholar] [CrossRef] [PubMed]
- Cochrane, D.R.; Howe, E.N.; Spoelstra, N.S.; Richer, J.K. Loss of miR-200c: A Marker of Aggressiveness and Chemoresistance in Female Reproductive Cancers. J. Oncol. 2010, 2010, 821717. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, F.; Liu, Q.; Hu, C.M. Epstein-Barr virus-encoded LMP1 increases MIR-155 expression, which promotes radioresistance of nasopharyngeal carcinoma via suppressing UBQLN1. Eur. Rev. Med. Pharmacol. Sci. 2015, 19, 4507–4515. [Google Scholar] [PubMed]
- Yadav, S.; Singh, N.; Shah, P.P.; Rowbotham, D.A.; Malik, D.; Srivastav, A.; Shankar, J.; Lam, W.L.; Lockwood, W.W.; Beverly, L.J. MIR155 Regulation of Ubiquilin1 and Ubiquilin2: Implications in Cellular Protection and Tumorigenesis. Neoplasia 2017, 19, 321. [Google Scholar] [CrossRef]
- Cai, X.; Cullen, B.R. The imprinted H19 noncoding RNA is a primary microRNA precursor. RNA 2007, 13, 313. [Google Scholar] [CrossRef] [Green Version]
- Keniry, A.; Oxley, D.; Monnier, P.; Kyba, M.; Dandolo, L.; Smits, G.; Reik, W. The H19 lincRNA is a developmental reservoir of miR-675 that suppresses growth and Igf1r. Nat. Cell Biol. 2012, 14, 659–665. [Google Scholar] [CrossRef]
- He, D.; Wang, J.; Zhang, C.; Shan, B.; Deng, X.; Li, B.; Zhou, Y.; Chen, W.; Hong, J.; Gao, Y.; et al. Down-regulation of miR-675-5p contributes to tumor progression and development by targeting pro-tumorigenic GPR55 in non-small cell lung cancer. Mol. Cancer 2015, 14, 73. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.; Zhang, Y.; Wei, H.; Zhang, X.; Wu, Y.; Gong, A.; Xia, Y.; Wang, W.; Xu, M. The mir-675-5p regulates the progression and development of pancreatic cancer via the UBQLN1-ZEB1-mir200 axis. Oncotarget 2017, 8, 24978. [Google Scholar] [CrossRef] [Green Version]
- Yu, Y.; Xu, P.; Cui, G.; Xu, X.; Li, K.; Chen, X.; Bao, J. UBQLN4 promotes progression of HCC via activating wnt-β-catenin pathway and is regulated by miR-370. Cancer Cell Int. 2020, 20, 3. [Google Scholar] [CrossRef] [Green Version]
- Huang, S.; Dong, X.; Wang, J.; Ding, J.; Li, Y.; Li, D.; Lin, H.; Wang, W.; Zhao, M.; Chang, Q.; et al. Overexpression of the Ubiquilin-4 (UBQLN4) is Associated with Cell Cycle Arrest and Apoptosis in Human Normal Gastric Epithelial Cell Lines GES-1 Cells by Activation of the ERK Signaling Pathway. Med. Sci. Monit. 2018, 24, 3564. [Google Scholar] [CrossRef]






© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jantrapirom, S.; Lo Piccolo, L.; Pruksakorn, D.; Potikanond, S.; Nimlamool, W. Ubiquilin Networking in Cancers. Cancers 2020, 12, 1586. https://doi.org/10.3390/cancers12061586
Jantrapirom S, Lo Piccolo L, Pruksakorn D, Potikanond S, Nimlamool W. Ubiquilin Networking in Cancers. Cancers. 2020; 12(6):1586. https://doi.org/10.3390/cancers12061586
Chicago/Turabian StyleJantrapirom, Salinee, Luca Lo Piccolo, Dumnoensun Pruksakorn, Saranyapin Potikanond, and Wutigri Nimlamool. 2020. "Ubiquilin Networking in Cancers" Cancers 12, no. 6: 1586. https://doi.org/10.3390/cancers12061586





