MRI-Based Radiomics Analysis for the Pretreatment Prediction of Pathologic Complete Tumor Response to Neoadjuvant Systemic Therapy in Breast Cancer Patients: A Multicenter Study
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
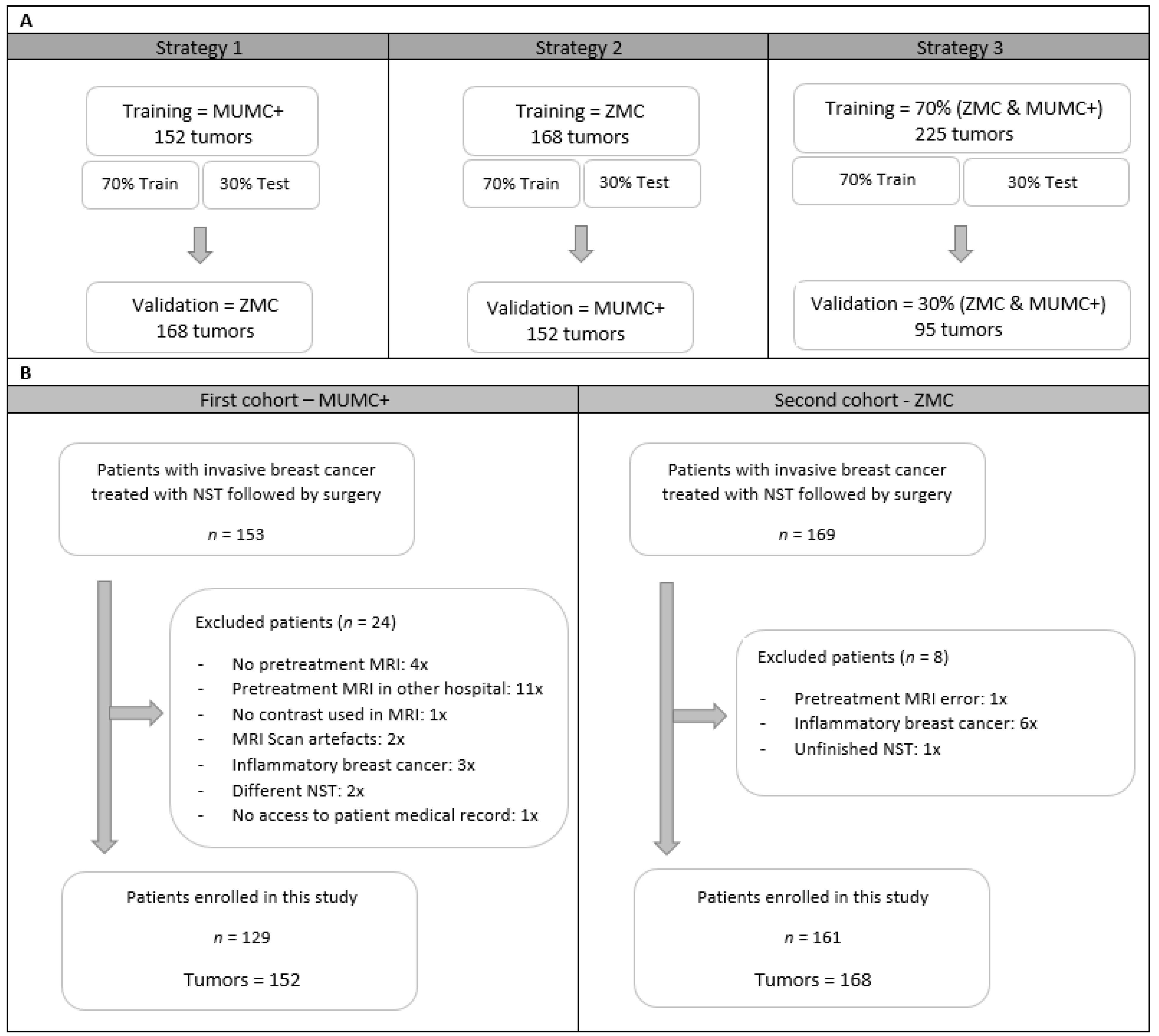
2.1. Study Population
2.2. Study Strategy
2.3. Clinical and Pathological Data
2.4. Imaging Data
2.5. Tumor Segmentation
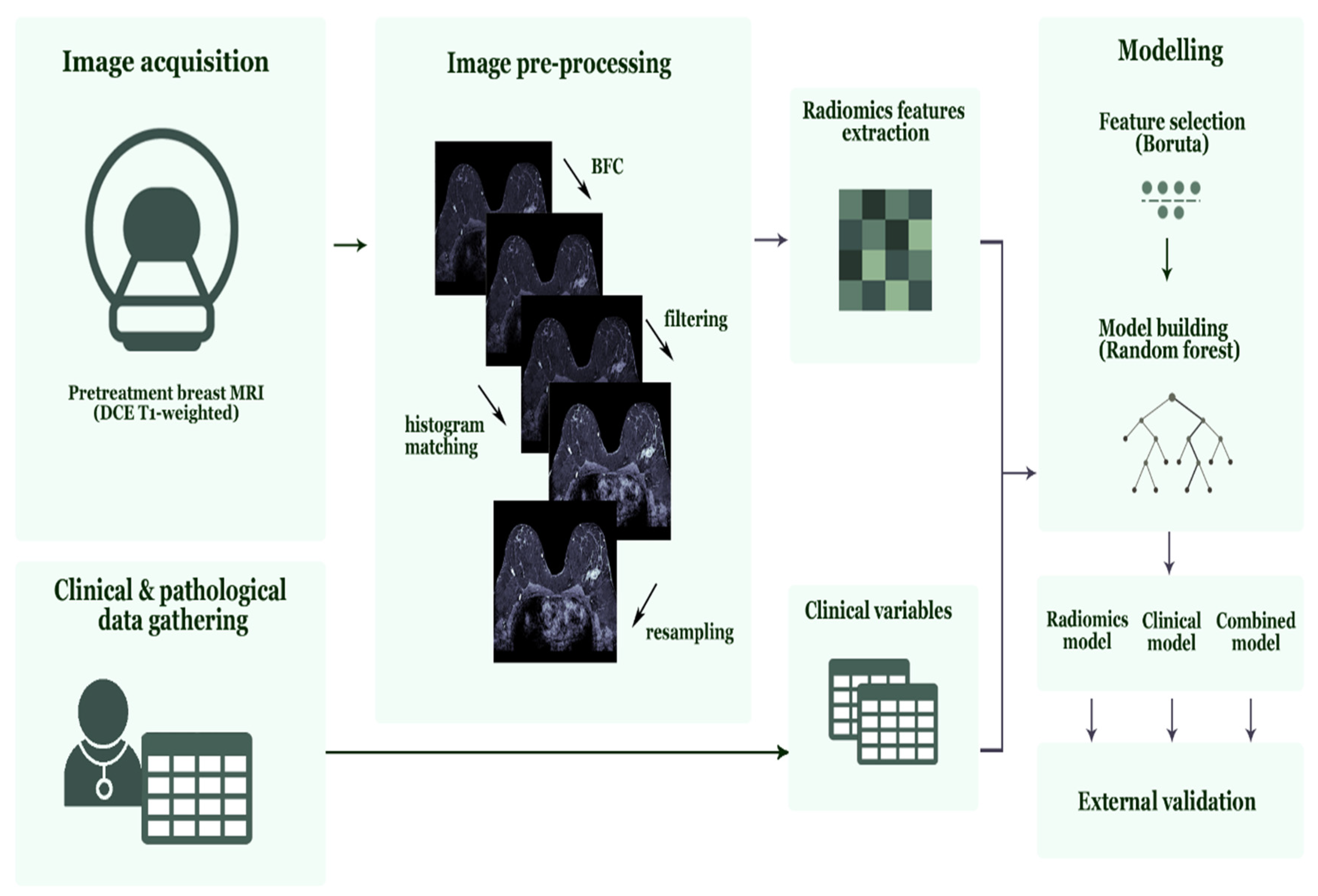
2.6. Image Pre-Processing and Feature Selection
2.7. Feature Selection and Radiomics Model Development
2.8. Clinical and Combined Model Development
2.9. Statistical Analysis
3. Results
3.1. Patients Demographics
3.2. Radiomics Models—Feature Selection and Model Performance
3.3. Clinical Models—Feature Selection and Model Performance
3.4. Combined Models—Feature Selection and Model Performance
3.5. RQS and TRIPOD Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Vugts, G.; Maaskant-Braat, A.J.; Nieuwenhuijzen, G.A.; Roumen, R.M.; Luiten, E.J.; Voogd, A.C. Patterns of Care in the Administration of Neo-adjuvant Chemotherapy for Breast Cancer. A Population-Based Study. Breast J. 2016, 22, 316–321. [Google Scholar] [CrossRef] [PubMed]
- Murphy, B.L.; Day, C.N.; Hoskin, T.L.; Habermann, E.B.; Boughey, J.C. Neoadjuvant Chemotherapy Use in Breast Cancer is Greatest in Excellent Responders: Triple-Negative and HER2+ Subtypes. Ann. Surg. Oncol. 2018, 25, 2241–2248. [Google Scholar] [CrossRef] [PubMed]
- Spronk, P.E.R.; de Ligt, K.M.; van Bommel, A.C.M.; Vrancken Peeters, M.T.F.D.; Siesling, S.; Smorenburg, C.H. Current decisions on neoadjuvant chemotherapy for early breast cancer: Experts’ experiences in the Netherlands. Patient Educ. Couns. 2018, 101, 2111–2115. [Google Scholar] [CrossRef]
- Loibl, S.; Denkert, C.; von Minckwitz, G. Neoadjuvant treatment of breast cancer—Clinical and research perspective. Breast 2015, 24, S73–S77. [Google Scholar] [CrossRef] [PubMed]
- Prevos, R.; Smidt, M.L.; Tjan-Heijnen, V.C.; van Goethem, M.; Beets-Tan, R.G.; Wildberger, J.E.; Lobbes, M.B.I. Pre-treatment differences and early response monitoring of neoadjuvant chemotherapy in breast cancer patients using magnetic resonance imaging: A systematic review. Eur. Radiol. 2012, 22, 2607–2616. [Google Scholar] [CrossRef] [PubMed]
- Lobbes, M.B.; Prevos, R.; Smidt, M.; Tjan-Heijnen, V.C.G.; van Goethem, M.; R Schipper, R.; Beets-Tan, R.G.; Wildberger, J.E. The role of magnetic resonance imaging in assessing residual disease and pathologic complete response in breast cancer patients receiving neoadjuvant chemotherapy: A systematic review. Insights Imaging 2013, 4, 163–175. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Weber, J.J.; Jochelson, M.S.; Eaton, A.; Zabor, E.C.; Barrio, A.V.; Gemignani, M.L.; Pilewskie, M.; Van Zee, K.J.; Morrow, M.; El-Tamer, M. MRI and Prediction of Pathologic Complete Response in the Breast and Axilla after Neoadjuvant Chemotherapy for Breast Cancer: MRI and Pathologic Complete Response. J. Am. Coll. Surg. 2017, 225, 740–746. [Google Scholar] [CrossRef] [PubMed]
- Bouzon, A.; Acea, B.; Soler, R.; Iglesias, A.; Santiago, P.; Mosquera, J.; Calvo, L.; Seoane-Pillado, T.; García, A. Diagnostic accuracy of MRI to evaluate tumour response and residual tumour size after neoadjuvant chemotherapy in breast cancer patients. Radiol. Oncol. 2016, 50, 73–79. [Google Scholar] [CrossRef] [PubMed]
- van der Noordaa, M.E.M.; van Duijnhoven, F.H.; Loo, C.E.; van Werkhoven, E.; van de Vijver, K.K.; Wiersma, T.; Winter-Warnars, H.A.O.; Sonke, G.S.; Vrancken Peeters, M.T.F.D. Identifying pathologic complete response of the breast after neoadjuvant systemic therapy with ultrasound guided biopsy to eventually omit surgery: Study design and feasibility of the MICRA trial (Minimally Invasive Complete Response Assessment). Breast 2018, 40, 76–81. [Google Scholar] [CrossRef] [PubMed]
- Heil, J.; Sinn, P.; Richter, H.; Pfob, A.; Schaefgen, B.; Hennigs, A.; Riedel, F.; Thomas, B.; Thill, M.; Hahn, M.; et al. RESPONDER—Diagnosis of pathological complete response by vacuum-assisted biopsy after neoadjuvant chemotherapy in breast Cancer—A multicenter, confirmative, one-armed, intra-individually-controlled, open, diagnostic trial. BMC Cancer 2018, 18, 851. [Google Scholar] [CrossRef] [PubMed]
- van Loevezijn, A.A.; van der Noordaa, M.E.; van Werkhoven, E.D.; Loo, C.E.; Winter-Warnars, G.A.; Wiersma, T.; van de Vijver, K.K.; Groen, E.J.; Blanken-Peeters, C.F.; Zonneveld, B.J.; et al. Minimally Invasive Complete Response Assessment of the Breast After Neoadjuvant Systemic Therapy for Early Breast Cancer (MICRA trial): Interim Analysis of a Multicenter Observational Cohort Study. Ann. Surg. Oncol. 2020. [Google Scholar] [CrossRef]
- van Griethuysen, J.J.M.; Fedorov, A.; Parmar, C.; Hosny, A.; Aucoin, N.; Narayan, V.; Beets-Tan, R.G.H.; Fillion-Robin, J.-C.; Pieper, S.; Aerts, H.J.W.L. Computational Radiomics System to Decode the Radiographic Phenotype. Cancer Res. 2017, 77, e104–e107. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ibrahim, A.; Primakov, S.; Beuque, M.; Woodruff, H.C.; Halilaj, I.; Wu, G.; Refaee, T.; Granzier, R.; Widaatalla, Y.; Hustinx, R.; et al. Radiomics for precision medicine: Current challenges, future prospects, and the proposal of a new framework. Methods 2021, 188, 20–29. [Google Scholar] [CrossRef] [PubMed]
- Lambin, P.; Leijenaar, R.T.H.; Deist, T.M.; Peerlings, J.; de Jong, E.E.C.; van Timmeren, J.; Sanduleanu, S.; Larue, R.T.H.M.; Even, A.J.G.; Jochems, J.; et al. Radiomics: The bridge between medical imaging and personalized medicine. Nat. Rev. Clin. Oncol. 2017, 14, 749–762. [Google Scholar] [CrossRef]
- Larue, R.T.; Defraene, G.; De Ruysscher, D.; Lambin, P.; van Elmpt, W. Quantitative radiomics studies for tissue characterization: A review of technology and methodological procedures. Br. J. Radiol. 2017, 90, 20160665. [Google Scholar] [CrossRef]
- Wu, G.; Ibrahim, A.; Halilaj, I.; Leijenaar, R.T.; Rogers, W.; Gietema, H.A.; Hendriks, L.E.; Lambin, P.; Woodruff, H.C. The Emerging Role of Radiomics in COPD and Lung Cancer. Respiration 2020, 99, 99–107. [Google Scholar]
- Ahmed, A.; Gibbs, P.; Pickles, M.; Turnbull, L. Texture analysis in assessment and prediction of chemotherapy response in breast cancer. J. Magn. Reson. Imaging 2013, 38, 89–101. [Google Scholar] [CrossRef]
- Parikh, J.; Selmi, M.; Charles-Edwards, G.; Glendenning, J.; Ganeshan, B.; Verma, H.; Mansi, J.; Harries, M.; Tutt, A.; Goh, V. Changes in primary breast cancer heterogeneity may augment midtreatment MR imaging assessment of response to neoadjuvant chemotherapy. Radiology 2014, 272, 100–112. [Google Scholar] [CrossRef]
- Teruel, J.R.; Heldahl, M.G.; Goa, P.E.; Pickles, M.; Lundgren, S.; Bathen, T.F.; Gibbs, P. Dynamic contrast-enhanced MRI texture analysis for pretreatment prediction of clinical and pathological response to neoadjuvant chemotherapy in patients with locally advanced breast cancer. NMR Biomed. 2014, 27, 887–896. [Google Scholar] [CrossRef]
- Chamming’s, F.; Ueno, Y.; Ferre, R.; Kao, E.; Jannot, A.-S.; Chong, J.; Omeroglu, A.; Mesurolle, B.; Reinhold, C.; Gallix, B. Features from Computerized Texture Analysis of Breast Cancers at Pretreatment MR Imaging Are Associated with Response to Neoadjuvant Chemotherapy. Radiology 2018, 286, 412–420. [Google Scholar] [CrossRef] [Green Version]
- Yoon, H.J.; Kim, Y.; Chung, J.; Kim, B.S. Predicting neo-adjuvant chemotherapy response and progression-free survival of locally advanced breast cancer using textural features of intratumoral heterogeneity on F-18 FDG PET/CT and diffusion-weighted MR imaging. Breast J. 2019, 25, 373–380. [Google Scholar] [CrossRef] [PubMed]
- Baessler, B.; Weiss, K.; Pinto Dos Santos, D. Robustness and Reproducibility of Radiomics in Magnetic Resonance Imaging: A Phantom Study. Investig. Radiol. 2019, 54, 221–228. [Google Scholar] [CrossRef] [PubMed]
- Mackin, D.; Fave, X.; Zhang, L.; Fried, D.; Yang, J.; Taylor, B.; Rodriguez-Rivera, E.; Dodge, C.; Jones, A.K.; Court, L. Measuring CT scanner variability of radiomics features. Investig. Radiol. 2015, 50, 757. [Google Scholar] [CrossRef]
- Rai, R.; Holloway, L.C.; Brink, C.; Field, M.; Christiansen, R.L.; Sun, Y.; Barton, M.B.; Liney, G.P. Multicenter evaluation of MRI-based radiomic features: A phantom study. Med. Phys. 2020, 47, 3054–3063. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, A.; Refaee, T.; Primakov, S.; Barufaldi, B.; Acciavatti, R.J.; Granzier, R.W.Y.; Hustinx, R.; Mottaghy, F.M.; Woodruff, H.C.; Wildberger, J.E.; et al. The Effects of In-Plane Spatial Resolution on CT-Based Radiomic Features’ Stability with and without ComBat Harmonization. Cancers 2021, 13, 1848. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, A.; Refaee, T.; Leijenaar, R.T.H.; Primakov, S.; Hustinx, R.; Mottaghy, F.M.; Woodruff, H.C.; Maidment, A.D.A.; Lambin, P. The application of a workflow integrating the variable reproducibility and harmonizability of radiomic features on a phantom dataset. PLoS ONE 2021, 16, e0251147. [Google Scholar] [CrossRef] [PubMed]
- Dreher, C.; Kuder, T.A.; Konig, F.; Mlynarska-Bujny, A.; Tenconi, C.; Paech, D.; Schlemmer, H.-P.; Ladd, M.E.; Bickelhaupt, S. Radiomics in diffusion data: A test-retest, inter- and intra-reader DWI phantom study. Clin. Radiol. 2020, 75, 798.e13–798.e22. [Google Scholar] [CrossRef]
- Peerlings, J.; Woodruff, H.C.; Winfield, J.M.; Ibrahim, A.; Van Beers, B.E.; Heerschap, A.; Jackson, A.; Wildberger, J.E.; Mottaghy, F.M.; DeSouza, N.M.; et al. Stability of radiomics features in apparent diffusion coefficient maps from a multi-centre test-retest trial. Sci. Rep. 2019, 9, 4800. [Google Scholar] [CrossRef] [Green Version]
- Shur, J.; Blackledge, M.; D′Arcy, J.; Collins, D.J.; O’Leach, M.; Koh, D.-M. MRI texture feature repeatability and image acquisition factor robustness, a phantom study and in silico study. Eur. Radiol. Exp. 2021, 5, 1–11. [Google Scholar] [CrossRef]
- Ogston, K.N.; Miller, I.D.; Payne, S.; Hutcheon, A.W.; Sarkar, T.K.; Smith, I.; Schofield, A.; Heys, S.D. A new histological grading system to assess response of breast cancers to primary chemotherapy: Prognostic significance and survival. Breast 2003, 12, 320–327. [Google Scholar] [CrossRef]
- Pinder, S.E.; Provenzano, E.; Earl, H.; Ellis, I.O. Laboratory handling and histology reporting of breast specimens from patients who have received neoadjuvant chemotherapy. Histopathology 2007, 50, 409–417. [Google Scholar] [CrossRef] [PubMed]
- Fan, M.; Wu, G.; Cheng, H.; Zhang, J.; Shao, G.; Li, L. Radiomic analysis of DCE-MRI for prediction of response to neoadjuvant chemotherapy in breast cancer patients. Eur. J. Radiol. 2017, 94, 140–147. [Google Scholar] [CrossRef] [PubMed]
- Braman, N.; Etesami, M.; Prasanna, P.; Dubchuk, C.; Gilmore, H.; Tiwari, P.; Plecha, D.; Madabhushi, A. Intratumoral and peritumoral radiomics for the pretreatment prediction of pathological complete response to neoadjuvant chemotherapy based on breast DCE-MRI. Breast Cancer Res. 2017, 19, 57. [Google Scholar] [CrossRef] [PubMed]
- Granzier, R.W.Y.; Verbakel, N.M.H.; Ibrahim, A.; van Timmermen, J.E.; van Nijnatten, T.J.A.; Leijenaar, R.T.H.; Lobbes, M.B.I.; Smidt, M.L.; Woodruff, H.C. MRI-based radiomics in breast cancer: Feature robustness with respect to inter-observer segmentation variability. Sci. Rep. 2020, 10, 14163. [Google Scholar] [CrossRef]
- Nyul, L.G.; Udupa, J.K.; Zhang, X. New variants of a method of MRI scale standardization. IEEE Trans. Med. Imaging 2000, 19, 143–150. [Google Scholar] [CrossRef]
- Hoebel, K.V.; Patel, J.B.; Beers, A.L.; Chang, K.; Singh, P.; Brown, J.M.; Pinho, M.C.; Batchelor, T.T.; Gerstner, E.R.; Rosen, B.R.; et al. Radiomics Repeatability Pitfalls in a Scan-Rescan MRI Study of Glioblastoma. Radiol. Artif. Intell. 2021, 3, e190199. [Google Scholar] [CrossRef] [PubMed]
- Ligero, M.; Jordi-Ollero, O.; Bernatowicz, K.; Garcia-Ruiz, A.; Delgado-Muñoz, E.; Leiva, D.; Mast, R.; Suarez, C.; Sala-Llonch, R.; Calvo, N.; et al. Minimizing acquisition-related radiomics variability by image resampling and batch effect correction to allow for large-scale data analysis. Eur. Radiol. 2021, 31, 1460–1470. [Google Scholar] [CrossRef]
- Moradmand, H.; Aghamiri, S.M.R.; Ghaderi, R. Impact of image preprocessing methods on reproducibility of radiomic features in multimodal magnetic resonance imaging in glioblastoma. J. Appl. Clin. Med. Phys. 2020, 21, 179–190. [Google Scholar] [CrossRef]
- Sun, X.; Shi, L.; Luo, Y.; Yang, W.; Li, H.; Liang, P.; Li, K.; Mok, V.C.T.; Chu, W.C.W.; Wang, D. Histogram-based normalization technique on human brain magnetic resonance images from different acquisitions. Biomed. Eng. Online 2015, 14, 73. [Google Scholar] [CrossRef] [Green Version]
- Senthilkumaran, N.; Thimmiaraja, J. Histogram Equalization for Image Enhancement Using MRI Brain Images. In Proceedings of the 2014 World Congress on Computing and Communication Technologies, Trichirappalli, India, 27 February–1 March 2014; pp. 80–83. [Google Scholar]
- Duron, L.; Balvay, D.; Vande Perre, S.; Bouchouicha, A.; Savatovsky, J.; Sadik, J.-C.; Thomassin-Naggara, I.; Fournier, L.; Lecler, A. Gray-level discretization impacts reproducible MRI radiomics texture features. PLoS ONE 2019, 14, e0213459. [Google Scholar]
- Kursa, M.B.; Rudnicki, W.R. Feature Selection with the Boruta Package. J. Stat. Softw. 2010, 36, 1–13. [Google Scholar] [CrossRef] [Green Version]
- Kursa, M.B.; Jankowski, A.; Rudnicki, W.R. Boruta—A System for Feature Selection. Fund. Inform. 2010, 101, 271–286. [Google Scholar] [CrossRef]
- Svetnik, V.; Liaw, A.; Tong, C.; Culberson, J.C.; Sheridan, R.P.; Feuston, B.P. Random forest: A classification and regression tool for compound classification and QSAR modeling. J. Chem. Inf. Comput. Sci. 2003, 43, 1947–1958. [Google Scholar] [CrossRef]
- Racine, J.S. RStudio: A platform-independent IDE for R and Sweave. J. Appl. Econom. 2012, 27, 167–172. [Google Scholar] [CrossRef]
- Robin, X.; Turck, N.; Hainard, A.; Tiberti, N.; Lisacek, F.; Sanchez, J.-C.; Müller, M. pROC: An open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinform. 2011, 12, 77. [Google Scholar] [CrossRef] [PubMed]
- Moons, K.G.; Altman, D.G.; Reitsma, J.B.; Ioannidis, J.P.A.; Macaskill, P.; Steyerberg, E.W.; Vickers, A.J.; Ransohoff, D.F.; Collins, G.S. Transparent Reporting of a multivariable prediction model for Individual Prognosis or Diagnosis (TRIPOD): Explanation and elaboration. Ann. Intern. Med. 2015, 162, W1–W73. [Google Scholar] [CrossRef] [Green Version]
- Collins, G.S.; Reitsma, J.B.; Altman, D.G.; Moons, K.G. Transparent reporting of a multivariable prediction model for individual prognosis or diagnosis (TRIPOD): The TRIPOD Statement. BMC Med. 2015, 13, 1. [Google Scholar] [CrossRef] [Green Version]
- Bianchini, L.; Botta, F.; Origgi, D.; Rizzo, S.; Mariani, M.; Summers, P.; García-Polo, P.; Cremonesi, M.; Lascialfari, A. PETER PHAN: An MRI phantom for the optimisation of radiomic studies of the female pelvis. Phys. Med. 2020, 71, 71–81. [Google Scholar] [CrossRef]
- Schwier, M.; van Griethuysen, J.; Vangel, M.G.; Pieper, S.; Peled, S.; Tempany, C.; Aerts, H.J.W.L.; Kikinis, R.; Fennessy, F.M.; Fedorov, A. Repeatability of Multiparametric Prostate MRI Radiomics Features. Sci. Rep. 2019, 9, 9441. [Google Scholar] [CrossRef]
- Fiset, S.; Welch, M.L.; Weiss, J.; Pintilie, M.; Conway, J.L.; Milosevic, M.; Fyles, A.; Traverso, A.; Jaffray, D.; Metser, U.; et al. Repeatability and reproducibility of MRI-based radiomic features in cervical cancer. Radiother. Oncol. 2019, 135, 107–114. [Google Scholar] [CrossRef]
- Scalco, E.; Belfatto, A.; Mastropietro, A.; Rancati, T.; Avuzzi, B.; Messina, A.; Valdagni, R.; Rizzo, G. T2w-MRI signal normalization affects radiomics features reproducibility. Med. Phys. 2020, 47, 1680–1691. [Google Scholar] [CrossRef] [PubMed]
- Choudhery, S.; Gomez-Cardona, D.; Favazza, C.P.; Hoskin, T.L.; Haddad, T.C.; Goetz, M.P.; Boughey, J.C. MRI Radiomics for Assessment of Molecular Subtype, Pathological Complete Response, and Residual Cancer Burden in Breast Cancer Patients Treated With Neoadjuvant Chemotherapy. Acad. Radiol. 2020, in press. [Google Scholar] [CrossRef] [PubMed]
- Henderson, S.; Purdie, C.; Michie, C.; Evans, A.; Lerski, R.; Johnston, M.; Vinnicombe, S.; Thompson, A.M. Interim heterogeneity changes measured using entropy texture features on T2-weighted MRI at 3.0 T are associated with pathological response to neoadjuvant chemotherapy in primary breast cancer. Eur. Radiol. 2017, 27, 4602–4611. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cain, E.H.; Saha, A.; Harowicz, M.R.; Marks, J.R.; Marcom, P.K.; Mazurowski, M.A. Multivariate machine learning models for prediction of pathologic response to neoadjuvant therapy in breast cancer using MRI features: A study using an independent validation set. Breast Cancer Res. Treat. 2019, 173, 455–463. [Google Scholar] [CrossRef] [PubMed]
- Xiong, Q.; Zhou, X.; Liu, Z.; Lei, C.; Yang, C.; Zhang, L.; Zhu, T.; Zhuang, X.; Liang, C.; Tian, L.; et al. Multiparametric MRI-based radiomics analysis for prediction of breast cancers insensitive to neoadjuvant chemotherapy. Clin. Transl. Oncol. 2020, 22, 50–59. [Google Scholar] [CrossRef]
- Liu, Z.; Li, Z.; Qu, J.; Zhang, R.; Zhou, X.; Li, L.; Sun, K.; Tang, Z.; Jiang, H.; Li, H.; et al. Radiomics of multi-parametric MRI for pretreatment prediction of pathological complete response to neoadjuvant chemotherapy in breast cancer: A multicenter study. Clin. Cancer Res. 2019, 25, 3538–3547. [Google Scholar] [CrossRef] [Green Version]

| Hospital | Scanner | Total MRI Exam No. | Group | No. of Tumors for Specific Scanning Parameters | Pixel Spacing | Acquisition Matrix (n) | Slice Thickness (mm) | TR/TE (ms) (n) | Spacing between Slices | Flip Angle |
|---|---|---|---|---|---|---|---|---|---|---|
| MUMC+ | Philips 1.5T (Ingenia) | 124 | a | 44 | (0.97, 0.97) | 340 × 340 | 1 | 3.4/7.5 3.5/7.6 | 1 | 10° |
| b | 66 | (0.95, 0.95) | 378 × 314 (28) 380 × 318 (23) 380 × 316 (18) | 1 | 3.2/7.1 3.4/7.5 3.5/7.6 | 1 | 10° | |||
| c | 9 | (0.80, 0.80) | 344 × 344 | 1 | 3.4/7.5 | 1 | 10° | |||
| d | 3 | (0.92, 0.92) | 400 × 333 (2) 398 × 331 (1) | 1 | 3.5/7.6 3.4/7.5 | 1 | 10° | |||
| e | 1 | (0.88, 0.88) | 384 × 368 | 1 | 3.4/7.5 | 1 | 10° | |||
| f | 1 | (0.85, 0.85) | 384 × 278 | 1 | 2.9/6.5 | 1 | 10° | |||
| Philips 1.5T (Intera) | 28 | a | 25 | (0.97, 0.97) | 340 × 337 | 1 | 3.4/7.4-7.6 | 1 | 10° | |
| b | 1 | (0.99, 0.99) | 376 × 376 | 1 | 3.4/7.4 | 1 | 10° | |||
| c | 1 | (0.95, 0.95) | 364 × 364 | 1 | 3.4/7.5 | 1 | 10° | |||
| d | 1 | (0.85, 0.85) | 368 × 368 | 1 | 3.4/7.4 | 1 | 10° | |||
| ZMC | Philips 1.5T (Achieva) | 123 | a | 94 | (0.97, 0.97) | 340 × 338 | 2 | 3.4/6.9–7.0 | 1 | 12° |
| b | 28 | (0.96, 0.96) | 372 × 368 (15) 372 × 370 (13) | 2 | 3.4/6.9–7.0 | 1 | 12° | |||
| c | 1 | (0.90, 0.90) | 392 × 388 | 2 | 3.4/6.9 | 1 | 12° | |||
| Siemens 3.0T (Skyra) | 39 | a | 39 | (0.69, 0.69) | 288 × 288 | 2 | 1.2/4.0 | unknown | 10° | |
| Siemens 1.5T (Avanto_fit) | 6 | a | 6 | (0.89, 0.89) | 224 × 202 | 2 | 2.4/6.1 | unknown | 10° |
| Characteristics | MUMC+ | ZMC | p-Value |
|---|---|---|---|
| Number of patients | 129 | 161 | - |
| Patient Age (years) (mean; range) | 51 (28–73) | 52 (28–79) | 0.378 |
| Number of tumors | 152 | 168 | - |
| Clinical tumor stage (%) | 0.007 | ||
| T1 | 29 (19.1) | 16 (9.5) | |
| T2 | 99 (65.1) | 103 (61.3) | |
| T3 | 20 (13.2) | 37 (22.0) | |
| T4 | 4 (2.6) | 12 (7.2) | |
| Clinical nodal stage (%) | <0.001 | ||
| N0 | 88 (57.9) | 59 (35.1) | |
| N1 | 44 (29.0) | 87 (51.8) | |
| N2 | 9 (5.9) | 12 (7.1) | |
| N3 | 11 (7.2) | 7 (4.2) | |
| Unknown | 0 (0.0) | 3 (1.8) | |
| Clinical tumor grade (%) | 0.003 | ||
| 1 | 8 (5.3) | 22 (13.1) | |
| 2 | 70 (46.1) | 84 (50.0) | |
| 3 | 68 (44.7) | 62 (36.9) | |
| Unknown | 6 (3.9) | 0 (0.0) | |
| Tumor histology (%) | 0.009 | ||
| Invasive ductal carcinoma | 136 (89.5) | 134 (79.8) | |
| Invasive lobular carcinoma | 10 (6.6) | 14 (8.3) | |
| Invasive mixed ductal/lobular carcinoma | 0 (0.0) | 9 (5.4) | |
| Other invasive carcinoma | 6 (3.9) | 11 (6.5) | |
| Cancer Subtype (%) | 0.921 | ||
| HR+ and HER2− | 80 (52.6) | 82 (48.8) | |
| HR+ and HER2+ | 22 (14.5) | 26 (15.5) | |
| HR− and HER2+ | 19 (12.5) | 22 (13.1) | |
| Triple-negative | 31 (20.4) | 38 (22.6) | |
| Response to NAC (%) | 0.331 | ||
| pCR | 53 (34.9) | 49 (29.2) | |
| Non-pCR | 99 (65.1) | 119 (70.8) |
| Characteristics | MUMC+ | ZMC | ||||
|---|---|---|---|---|---|---|
| Non-pCR | pCR | p-Value | Non-pCR | pCR | p-Value | |
| Number of tumors | 99 | 53 | - | 119 | 49 | - |
| Patient Age (years) (mean; range) | 52 (32–72) | 51 (28–73) | 0.600 | 53 (31–79) | 52 (28–73) | 0.538 |
| Clinical tumor stage (%) | 0.019 * | 0.023 | ||||
| T1 | 12 (12.1) | 17 (32.1) | 6 (5.0) | 10 (20.4) | ||
| T2 | 68 (68.7) | 31 (58.5) | 76 (63.9) | 27 (55.1) | ||
| T3 | 16 (16.2) | 4 (7.5) | 28 (23.5) | 9 (18.4) | ||
| T4 | 3 (3.0) | 1 (1.9) | 9 (7.6) | 3 (6.1) | ||
| Clinical nodal stage (%) | 0.943 | 0.526 | ||||
| N0 | 56 (56.6) | 32 (60.3) | 39 (32.8) | 20 (40.8) | ||
| N1 | 29 (29.3) | 15 (28.3) | 62 (52.1) | 25 (51.0) | ||
| N2 | 6 (6.1) | 3 (5.7) | 11 (9.2) | 1 (2.0) | ||
| N3 | 8 (8.1) | 3 (5.7) | 5 (4.2) | 2 (4.1) | ||
| Unknown | 0 (0.0) | 0 (0.0) | 2 (1.7) | 1 (2.0) | ||
| Clinical tumor grade (%) | <0.001 * | 0.002 | ||||
| 1 | 8 (8.1) | 0 (0.0) | 19 (15.9) | 3 (6.1) | ||
| 2 | 58 (58.6) | 12 (22.7) | 66 (55.5) | 18 (36.7) | ||
| 3 | 32 (32.3) | 36 (67.9) | 34 (28.6) | 28 (57.2) | ||
| Unknown | 1 (1.0) | 5 (9.4) | 0 (0.0) | 0 (0.0) | ||
| Tumor histology (%) | 0.913 | 0.030 | ||||
| Invasive ductal carcinoma | 89 (89.9) | 47 (88.7) | 91 (76.5) | 43 (87.8) | ||
| Invasive lobular carcinoma | 6 (6.1) | 4 (7.5) | 13 (10.9) | 1 (2.0) | ||
| Invasive mixed ductal/lobular carcinoma | 0 (0.0) | 0 (0.0) | 9 (7.6) | 0 (0.0) | ||
| Other invasive carcinoma | 4 (4.0) | 2 (3.8) | 6 (5.0) | 5 (10.2) | ||
| Cancer Subtype (%) | <0.001 * | <0.001 | ||||
| HR+ and HER2− | 64 (64.6) | 16 (30.2) | 75 (63.0) | 7 (14.3) | ||
| HR+ and HER2+ | 15 (15.2) | 7 (13.2) | 14 (11.8) | 12 (24.5) | ||
| HR− and HER2+ | 6 (6.1) | 13 (24.5) | 5 (4.2) | 17 (34.7) | ||
| Triple-negative | 14 (14.1) | 17 (32.1) | 25 (21.0) | 13 (26.5) | ||
| Strategy 1 | Strategy 2 | Strategy 3 | |
|---|---|---|---|
| A (Radiomics) | O_glszm_GrayLevelVariance | W.LHH_firstorder_Kurtosis | O_shape_Sphericity |
| W.HLL_firstorder_Mean | W.LLH_glszm_GrayLevelNon- Uniformity | ||
| W.HLL_glcm_Imc1 | W.LLH_glszm_ZoneEntropy | ||
| W.HLH_glcm_InverseVariance | W.HHL_glcm_Imc1 | ||
| W.LLL_ngtdm_Complexity | W.HHH_glrlm_RunEntropy | ||
| W.LLL_glcm_DifferenceVariance | |||
| B (Clinical) | Age | cT | Age |
| cT | cN | cT | |
| Tumor grade | Tumor grade | Tumor grade | |
| Breast cancer subtype | Breast cancer subtype | Breast cancer subtype | |
| C (Combined) | Tumor grade | Tumor grade | cT |
| Breast cancer subtype | Breast cancer subtype | Tumor grade | |
| O_shape_Sphericity | W.LHL_firstorder_kurtosis | Breast cancer subtype | |
| O_firstorder_Mean | W.HHL_gldm_DependenceVariance | O_shape_Sphericity | |
| W.HLL_glcm_Imc2 | W.LLH_glszm _SmallAreaLowGrayLevelEmphasis | ||
| W.HLL_glszm_ZoneEntropy | |||
| W.HLH_glcm_InverseVariance |
| A (Radiomics) | Strategy 1 | Strategy 2 | Strategy 3 | ||||||
| Training MUMC+ | Validation ZMC | Training ZMC | Validation MUMC+ | Training 70% Mixed | Validation 30% Mixed | ||||
| Train | Test | Train | Test | Train | Test | ||||
| Area under the ROC | 0.71 | 0.78 | 0.55 | 0.64 | 0.67 | 0.52 | 0.60 | 0.65 | 0.50 |
| 95% CI | 0.59–0.82 | 0.63–0.92 | 0.46–0.65 | 0.54–0.75 | 0.49–0.84 | 0.42–0.62 | 0.49–0.71 | 0.51–0.80 | 0.37–0.64 |
| Sensitivity (%) | 53 | 59 | 73 | 44 | 60 | 28 | 38 | 48 | 24 |
| Specificity (%) | 89 | 79 | 36 | 75 | 72 | 62 | 92 | 77 | 88 |
| PPV (%) | 70 | 63 | 32 | 42 | 47 | 28 | 69 | 48 | 47 |
| NPV (%) | 79 | 76 | 77 | 77 | 81 | 62 | 75 | 77 | 72 |
| B (Clinical) | Strategy 1 | Strategy 2 | Strategy 3 | ||||||
| Training MUMC+ | Validation ZMC | Training ZMC | Validation MUMC+ | Training 70% Mixed | Validation 30% Mixed | ||||
| Train | Test | Train | Test | Train | Test | ||||
| Area under the ROC | 0.79 | 0.81 | 0.71 | 0.81 | 0.84 | 0.77 | 0.75 | 0.86 | 0.72 |
| 95% CI | 0.71–0.87 | 0.68–0.95 | 0.62–0.79 | 0.73–0.89 | 0.72–0.96 | 0.70–0.85 | 0.68–0.83 | 0.77–0.95 | 0.61–0.83 |
| Sensitivity (%) | 54 | 86 | 45 | 54 | 71 | 47 | 52 | 71 | 41 |
| Specificity (%) | 87 | 64 | 74 | 85 | 86 | 85 | 77 | 84 | 78 |
| PPV (%) | 69 | 57 | 42 | 59 | 67 | 63 | 52 | 68 | 46 |
| NPV (%) | 78 | 89 | 77 | 82 | 88 | 75 | 77 | 86 | 75 |
| C (Combined) | Strategy 1 | Strategy 2 | Strategy 3 | ||||||
| Training MUMC+ | Validation ZMC | Training ZMC | Validation MUMC+ | Training 70% Mixed | Validation 30% Mixed | ||||
| Train | Test | Train | Test | Train | Test | ||||
| Area under the ROC | 0.82 | 0.83 | 0.73 | 0.79 | 0.86 | 0.69 | 0.79 | 0.86 | 0.71 |
| 95% CI | 0.74–0.90 | 0.70–0.97 | 0.65–0.81 | 0.71–0.88 | 0.74–0.98 | 0.61–0.78 | 0.73–0.86 | 0.76–0.96 | 0.60–0.81 |
| Sensitivity (%) | 53 | 67 | 51 | 51 | 71 | 51 | 52 | 71 | 38 |
| Specificity (%) | 88 | 88 | 82 | 87 | 82 | 67 | 85 | 89 | 83 |
| PPV (%) | 69 | 77 | 53 | 62 | 63 | 45 | 61 | 75 | 50 |
| NPV (%) | 78 | 82 | 80 | 81 | 88 | 72 | 79 | 87 | 75 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Granzier, R.W.Y.; Ibrahim, A.; Primakov, S.P.; Samiei, S.; van Nijnatten, T.J.A.; de Boer, M.; Heuts, E.M.; Hulsmans, F.-J.; Chatterjee, A.; Lambin, P.; et al. MRI-Based Radiomics Analysis for the Pretreatment Prediction of Pathologic Complete Tumor Response to Neoadjuvant Systemic Therapy in Breast Cancer Patients: A Multicenter Study. Cancers 2021, 13, 2447. https://doi.org/10.3390/cancers13102447
Granzier RWY, Ibrahim A, Primakov SP, Samiei S, van Nijnatten TJA, de Boer M, Heuts EM, Hulsmans F-J, Chatterjee A, Lambin P, et al. MRI-Based Radiomics Analysis for the Pretreatment Prediction of Pathologic Complete Tumor Response to Neoadjuvant Systemic Therapy in Breast Cancer Patients: A Multicenter Study. Cancers. 2021; 13(10):2447. https://doi.org/10.3390/cancers13102447
Chicago/Turabian StyleGranzier, Renée W. Y., Abdalla Ibrahim, Sergey P. Primakov, Sanaz Samiei, Thiemo J. A. van Nijnatten, Maaike de Boer, Esther M. Heuts, Frans-Jan Hulsmans, Avishek Chatterjee, Philippe Lambin, and et al. 2021. "MRI-Based Radiomics Analysis for the Pretreatment Prediction of Pathologic Complete Tumor Response to Neoadjuvant Systemic Therapy in Breast Cancer Patients: A Multicenter Study" Cancers 13, no. 10: 2447. https://doi.org/10.3390/cancers13102447








