CircEHD2, CircNETO2 and CircEGLN3 as Diagnostic and Prognostic Biomarkers for Patients with Renal Cell Carcinoma
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Patients
2.2. RNA Isolation, RNase R Treatment, and cel-miR-39 Spike-in
2.3. cDNA Synthesis and Quantitative Real-Time PCR
2.4. Statistics and Data Analysis
3. Results
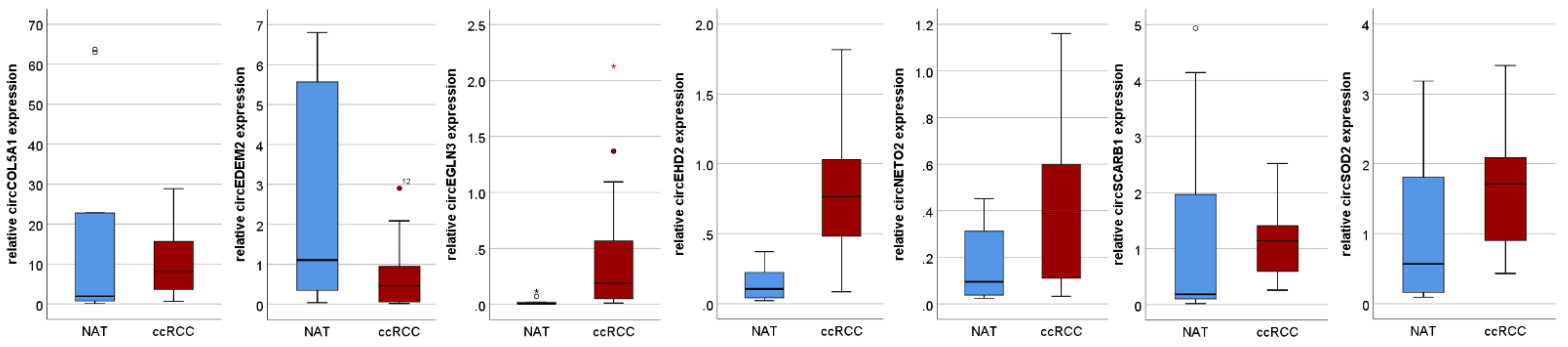
3.1. Discovery Cohort
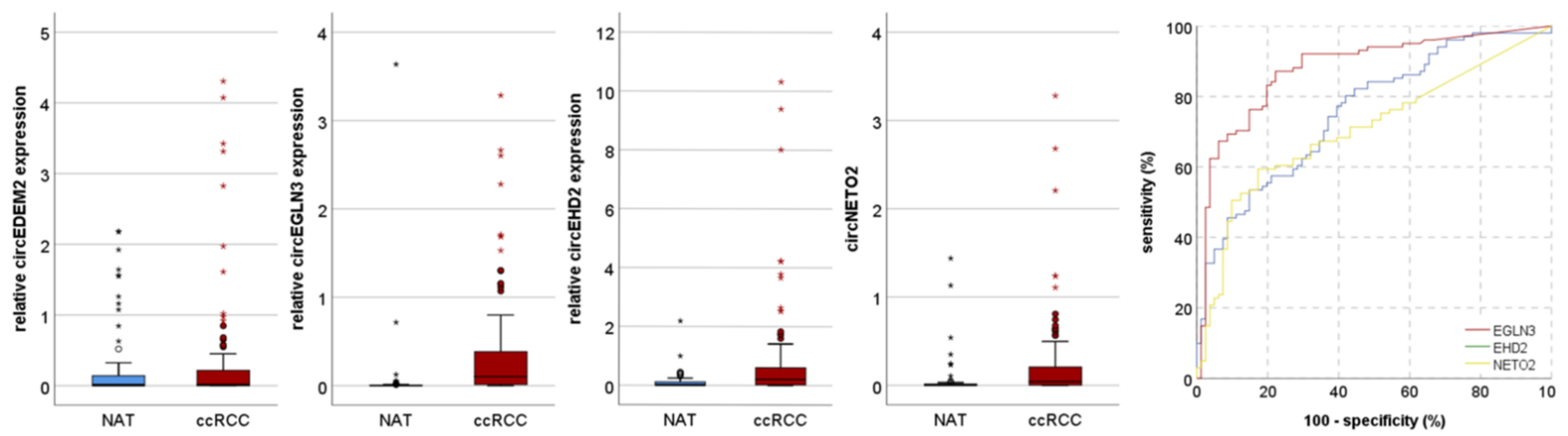
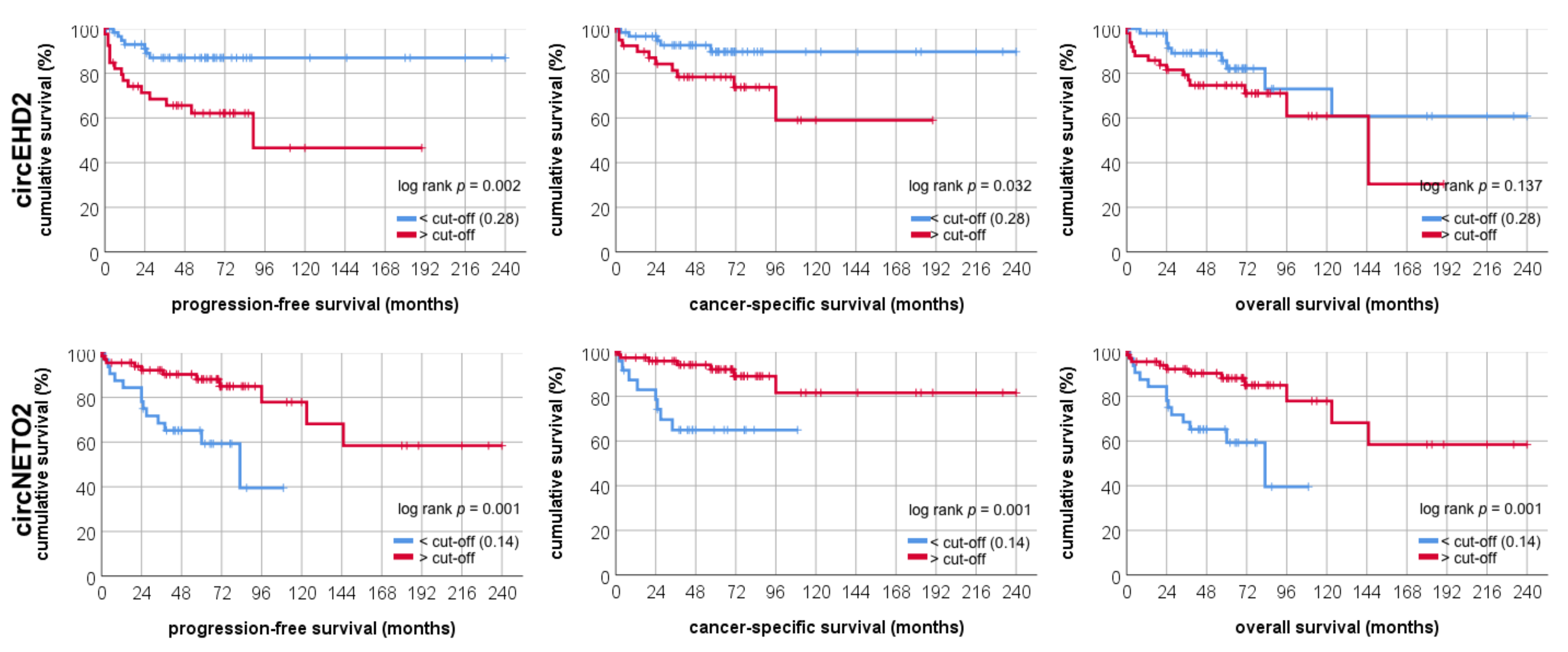
3.2. Validation Cohort
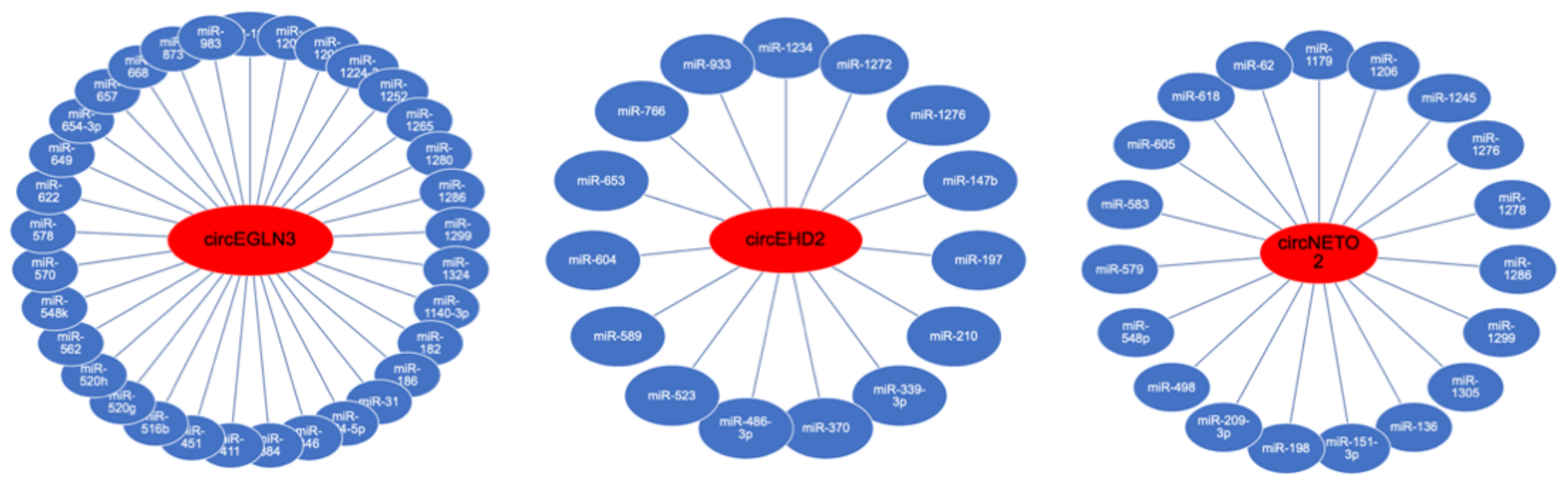
3.3. Analysis of circRNA Interactions
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hofmann, F.; Hwang, E.C.; Lam, T.B.; Bex, A.; Yuan, Y.; Marconi, L.S.; Ljungberg, B. Targeted therapy for metastatic renal cell carcinoma. Cochrane Database Syst. Rev. 2020, 10, CD012796. [Google Scholar] [CrossRef][Green Version]
- Escudier, B.; Porta, C.; Schmidinger, M.; Rioux-Leclercq, N.; Bex, A.; Khoo, V.; Gruenvald, V.; Horwich, A. Renal cell carcinoma: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 2016, 27, v58–v68. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Liu, J.Q.; Chen, M.; Xu, J.; Zhu, D. Circular RNA in cancer development and immune regulation. J. Cell. Mol. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Qi, X.; Liu, L.; Hu, X.; Liu, J.; Yang, J.; Lu, L.; Zhang, Z.; Ma, S.; Li, H.; et al. Emerging Epigenetic Regulation of Circular RNAs in Human Cancer. Mol. Ther. Nucleic Acids 2019, 16, 589–596. [Google Scholar] [CrossRef]
- Zhou, M.; Xiao, M.S.; Li, Z.; Huang, C. New progresses of circular RNA biology: From nuclear export to degradation. RNA Biol. 2020, 1–9. [Google Scholar] [CrossRef]
- Xia, S.; Feng, J.; Lei, L.; Hu, J.; Xia, L.; Wang, J.; Xiang, Y.; Liu, L.; Zhong, S.; Han, L.; et al. Comprehensive characterization of tissue-specific circular RNAs in the human and mouse genomes. Brief. Bioinform. 2017, 18, 984–992. [Google Scholar] [CrossRef]
- Xia, S.; Feng, J.; Chen, K.; Ma, Y.; Gong, J.; Cai, F.; Jin, Y.; Gao, Y.; Xia, L.; Chang, H.; et al. CSCD: A database for cancer-specific circular RNAs. Nucleic Acids Res. 2018, 46, D925–D929. [Google Scholar] [CrossRef]
- Franz, A.; Ralla, B.; Weickmann, S.; Jung, M.; Rochow, H.; Stephan, C.; Erbersdobler, A.; Kilic, E.; Fendler, A.; Jung, K. Circular RNAs in Clear Cell Renal Cell Carcinoma: Their Microarray-Based Identification, Analytical Validation, and Potential Use in a Clinico-Genomic Model to Improve Prognostic Accuracy. Cancers 2019, 11, 1473. [Google Scholar] [CrossRef]
- Lv, Q.; Ma, C.; Li, H.; Tan, X.; Wang, G.; Zhang, Y.; Wang, P. Circular RNA microarray expression profile and potential function of circ0005875 in clear cell renal cell carcinoma. J. Cancer 2020, 11, 7146–7156. [Google Scholar] [CrossRef]
- Zhang, D.; Yang, X.J.; Luo, Q.D.; Fu, D.L.; Li, Z.L.; Zhang, P.; Chong, T. Down-Regulation of Circular RNA_000926 Attenuates Renal Cell Carcinoma Progression through miRNA-411-Dependent CDH2 Inhibition. Am. J. Pathol. 2019, 189, 2469–2486. [Google Scholar] [CrossRef]
- Li, W.; Yang, F.Q.; Sun, C.M.; Huang, J.H.; Zhang, H.M.; Li, X.; Wang, G.C.; Zhang, N.; Che, J.P.; Zhang, W.T.; et al. circPRRC2A promotes angiogenesis and metastasis through epithelial-mesenchymal transition and upregulates TRPM3 in renal cell carcinoma. Theranostics 2020, 10, 4395–4409. [Google Scholar] [CrossRef]
- Lin, L.; Cai, J. Circular RNA circ-EGLN3 promotes renal cell carcinoma proliferation and aggressiveness via miR-1299-mediated IRF7 activation. J. Cell. Biochem. 2020, 121. [Google Scholar] [CrossRef]
- Blondeau, J.J.; Deng, M.; Syring, I.; Schrödter, S.; Schmidt, D.; Perner, S.; Müller, S.C.; Ellinger, J. Identification of novel long non-coding RNAs in clear cell renal cell carcinoma. Clin. Epigenet. 2015, 7, 10. [Google Scholar] [CrossRef]
- Strick, A.; von Hagen, F.; Gundert, L.; Klümper, N.; Tolkach, Y.; Schmidt, D.; Kristiansen, G.; Toma, M.; Ritter, M.; Ellinger, J. The N 6 -methyladenosine (m 6 A) erasers alkylation repair homologue 5 (ALKBH5) and fat mass and obesity-associated protein (FTO) are prognostic biomarkers in patients with clear cell renal carcinoma. BJU Int. 2020, 125, 617–624. [Google Scholar] [CrossRef]
- Ren, H.; Liu, Z.; Liu, S.; Zhou, X.; Wang, H.; Xu, J.; Wang, D.; Yuan, G. Profile and clinical implication of circular RNAs in human papillary thyroid carcinoma. PeerJ 2018, 6. [Google Scholar] [CrossRef]
- Liu, H.; Hu, Y.; Zhuang, B.; Yin, J.; Chen, X.H.; Wang, J.; Li, M.M.; Xu, J.; Wang, X.Y.; Yu, Z.B.; et al. Differential Expression of CircRNAs in Embryonic Heart Tissue Associated with Ventricular Septal Defect. Int. J. Med. Sci. 2018, 15, 703–712. [Google Scholar] [CrossRef]
- Dudekula, D.B.; Panda, A.C.; Grammatikakis, I.; De, S.; Abdelmohsen, K.; Gorospe, M. CircInteractome: A web tool for exploring circular RNAs and their interacting proteins and microRNAs. RNA Biol. 2016, 13, 34–42. [Google Scholar] [CrossRef]
- Budczies, J.; Klauschen, F.; Sinn, B.V.; Győrffy, B.; Schmitt, W.D.; Darb-Esfahani, S.; Denkert, C. Cutoff Finder: A comprehensive and straightforward Web application enabling rapid biomarker cutoff optimization. PLoS ONE 2012, 7, e51862. [Google Scholar] [CrossRef]
- Rochow, H.; Franz, A.; Jung, M.; Weickmann, S.; Ralla, B.; Kilic, E.; Stephan, C.; Fendler, A.; Jung, K. Instability of circular RNAs in clinical tissue samples impairs their reliable expression analysis using RT-qPCR: From the myth of their advantage as biomarkers to reality. Theranostics 2020, 10, 9268–9279. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, Y.; Wang, P.; Fu, X.; Lin, W. Circular RNAs in renal cell carcinoma: Implications for tumorigenesis, diagnosis, and therapy. Mol. Cancer 2020, 19, 1–10. [Google Scholar] [CrossRef]
- Xu, L.; Ma, Y.; Zhang, H.; Lu, Q.J.; Yang, L.; Jiang, G.N.; Liao, W.L. HMGA2 regulates circular RNA ASPH to promote tumor growth in lung adenocarcinoma. Cell Death Dis. 2020, 11, 1–10. [Google Scholar] [CrossRef]
- Sun, J.; Pan, S.; Cui, H.; Li, H. CircRNA SCARB1 Promotes Renal Cell Carcinoma Progression via Mir- 510-5p/SDC3 Axis. Curr. Cancer Drug Targets 2020, 20, 461–470. [Google Scholar] [CrossRef]
- Zhuang, C.; Huang, X.; Yu, J.; Gui, Y. Circular RNA hsa_circ_0075828 promotes bladder cancer cell proliferation through activation of CREB1. BMB Rep. 2020, 53, 82–87. [Google Scholar] [CrossRef]
- Zhao, Z.; Song, J.; Tang, B.; Fang, S.; Zhang, D.; Zheng, L.; Wu, F.; Gao, Y.; Chen, C.; Hu, X.; et al. CircSOD2 induced epigenetic alteration drives hepatocellular carcinoma progression through activating JAK2/STAT3 signaling pathway. J. Exp. Clin. Cancer Res. 2020, 39, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Zhang, S.; Chen, X.; Li, N.; Li, J.; Jia, R.; Pan, Y.; Liang, H. EIF4A3-induced circular RNA MMP9 (circMMP9) acts as a sponge of miR-124 and promotes glioblastoma multiforme cell tumorigenesis. Mol. Cancer 2018, 17, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.; Huang, M.; Xing, L.; Yang, R.; Wang, X.; Jiang, R.; Zhang, L.; Chen, J. The circRNA circSEPT9 mediated by E2F1 and EIF4A3 facilitates the carcinogenesis and development of triple-negative breast cancer. Mol. Cancer 2020, 19, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Méjean, A.; Ravaud, A.; Thezenas, S.; Colas, S.; Beauval, J.B.; Bensalah, K.; Geoffrois, L.; Thiery-Vuillemin, A.; Cormier, L.; Lang, H.; et al. Sunitinib Alone or after Nephrectomy in Metastatic Renal-Cell Carcinoma. N. Engl. J. Med. 2018, 379, 417–427. [Google Scholar] [CrossRef]

| Clinical Parameter | Discovery Cohort | Validation Cohort | ||
|---|---|---|---|---|
| ccRCC (n = 20) | Normal (n = 10) | ccRCC (n = 101) | Normal (n = 81) | |
| Sex | ||||
| male | 15 (75.0%) | 6 (60.0%) | 65 (64.4%) | 59 (72.8%) |
| female | 5 (25.0%) | 4 (40.0%) | 36 (35.6%) | 22 (27.2%) |
| Age | ||||
| mean | 65.4 | 58.7 | 63.7 | 64.1 |
| min-max | 43–78 | 43–78 | 36–89 | 36–89 |
| pT-stage | ||||
| pT1 | 10 (50.0%) | 61 (60.4%) | ||
| pT2 | 0 (0%) | 11 (10.9%) | ||
| pT3 | 9 (45.0%) | 28 (27.7%) | ||
| pT4 | 1 (5.0%) | 1 (1.0%) | ||
| lymph node metastasis | ||||
| cN0/pN0 | 17 (85.0%) | 101 (100%) | ||
| pN1 | 3 (15.0%) | 0 (0%) | ||
| distant metastasis | ||||
| cM0 | 10 (50.0%) | 95 (94.1%) | ||
| cM1 | 10 (50.0%) | 6 (5.9%) | ||
| Grading | ||||
| grade 1 | 1 (5.0%) | 11 (10.9%) | ||
| grade 2 | 12 (60.0%) | 74 (73.3%) | ||
| grade 3 | 3 (15.0%) | 14 (13.9%) | ||
| grade 4 | 4 (20.0%) | 2 (2.0%) | ||
| Parameter | Univariate Analysis | Multivariate Analysis | ||
|---|---|---|---|---|
| p-Value | HR (95% CI) | p-Value | HR (95% CI) | |
| Expression of circRNA | ||||
| Low (<cut-off) | 1.00 | 1.00 | ||
| High circEDEM2 | 0.246 | 1.68 (0.70–4.02) | ||
| High circEGLN3 | 0.492 | 0.70 (0.26–1.92) | ||
| High circEHD2 | 0.005 | 3.64 (1.48–8.92) | 0.009 | 3.58 (1.37–9.38) |
| High circNETO2 | 0.002 | 0.26 (0.11–0.61) | 0.001 | 0.17 (0.60–0.50) |
| Clinicopathological parameters | ||||
| Grading | ||||
| G1/2 | 1.00 | 1.00 | ||
| G3/4 | <0.001 | 5.40 (2.29–12.72) | <0.001 | 9.40 (3.38–26.09) |
| pT-stage | ||||
| pT1/2 | 1.00 | 1.00 | ||
| pT3/4 | 0.002 | 3.76 (1.61–8.75) | 0.009 | 3.32 (1.34–8.20) |
| cM-stage | ||||
| cM0 | 1.00 | 1.00 | ||
| cM1 | 0.002 | 5.79 (1.94–17.25) | <0.001 | 11.91 (3.03–46.82) |
| Parameter | Univariate Analysis | Multivariate Analysis | ||
|---|---|---|---|---|
| p-Value | HR (95% CI) | p-Value | HR (95% CI) | |
| Expression of circRNA | ||||
| Low (<cut-off) | 1.00 | 1.00 | ||
| High circEDEM2 | 0.232 | 1.68 (7.16–3.97) | ||
| High circEGLN3 | 0.883 | 0.938 (0.40–2.20) | ||
| High circEHD2 | 0.005 | 3.64 (1.48–8.92) | 0.042 | 2.67 (1.04–6.85) |
| High circNETO2 | 0.001 | 0.24 (0.10–0.56) | 0.001 | 0.14 (0.05–0.43) |
| Clinicopathological parameters | ||||
| Grading | ||||
| G1/2 | 1.00 | 1.00 | ||
| G3/4 | <0.001 | 5.40 (2.29–12.72) | <0.001 | 11.66 (3.95–34.40) |
| pT-stage | ||||
| pT1/2 | 1.00 | 1.00 | ||
| pT3/4 | 0.002 | 3.76 (1.61–8.75) | 0.024 | 2.87 (1.15–7.17) |
| cM-stage | ||||
| cM0 | 1.00 | 1.00 | ||
| cM1 | 0.002 | 5.78 (1.94–17.25) | <0.001 | 13.82 (3.386–56.39) |
| Parameter | Univariate Analysis | Multivariate Analysis | ||
|---|---|---|---|---|
| p-Value | HR (95% CI) | p-Value | HR (95% CI) | |
| Expression of circRNA | ||||
| Low (<cut-off) | 1.00 | 1.00 | ||
| High circEDEM2 | 0.541 | 1.30 (0.56–3.03) | ||
| High circEGLN3 | 0.677 | 0.81 (0.30–2.21) | ||
| High circEHD2 | 0.019 | 3.07 (1.20–7.85) | 0.008 | 3.91 (1.43–10.67) |
| High circNETO2 | 0.010 | 0.32 (0.13–0.76) | 0.001 | 0.15 (0.05–0.46) |
| Clinicopathological parameters | ||||
| Grading | ||||
| G1/2 | 1.00 | 1.00 | ||
| G3/4 | <0.001 | 5.40 (2.29–12.72) | <0.001 | 9.95 (3.56–27.85) |
| pT-stage | ||||
| pT1/2 | 1.00 | 1.00 | ||
| pT3/4 | 0.002 | 3.76 (1.61–8.75) | 0.007 | 3.52 (1.42–8.70) |
| cM-stage | ||||
| cM0 | 1.00 | 1.00 | ||
| cM1 | 0.002 | 5.79 (1.94–17.25) | <0.001 | 14.03 (3.51–56.13) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Frey, L.; Klümper, N.; Schmidt, D.; Kristiansen, G.; Toma, M.; Ritter, M.; Alajati, A.; Ellinger, J. CircEHD2, CircNETO2 and CircEGLN3 as Diagnostic and Prognostic Biomarkers for Patients with Renal Cell Carcinoma. Cancers 2021, 13, 2177. https://doi.org/10.3390/cancers13092177
Frey L, Klümper N, Schmidt D, Kristiansen G, Toma M, Ritter M, Alajati A, Ellinger J. CircEHD2, CircNETO2 and CircEGLN3 as Diagnostic and Prognostic Biomarkers for Patients with Renal Cell Carcinoma. Cancers. 2021; 13(9):2177. https://doi.org/10.3390/cancers13092177
Chicago/Turabian StyleFrey, Lisa, Niklas Klümper, Doris Schmidt, Glen Kristiansen, Marieta Toma, Manuel Ritter, Abdullah Alajati, and Jörg Ellinger. 2021. "CircEHD2, CircNETO2 and CircEGLN3 as Diagnostic and Prognostic Biomarkers for Patients with Renal Cell Carcinoma" Cancers 13, no. 9: 2177. https://doi.org/10.3390/cancers13092177
APA StyleFrey, L., Klümper, N., Schmidt, D., Kristiansen, G., Toma, M., Ritter, M., Alajati, A., & Ellinger, J. (2021). CircEHD2, CircNETO2 and CircEGLN3 as Diagnostic and Prognostic Biomarkers for Patients with Renal Cell Carcinoma. Cancers, 13(9), 2177. https://doi.org/10.3390/cancers13092177







