The Breast Cancer Protein Co-Expression Landscape
Abstract
:Simple Summary
Abstract
1. Introduction
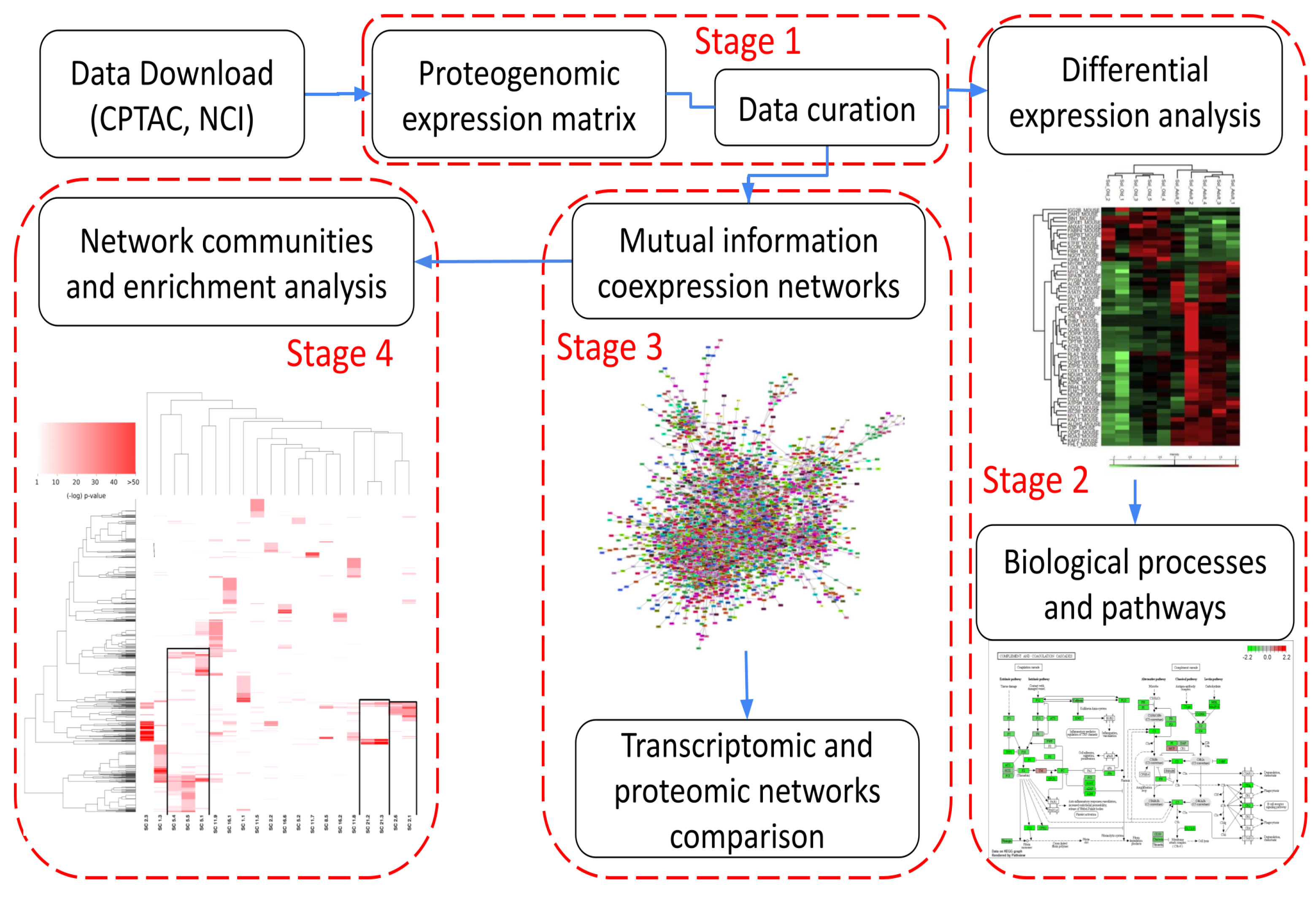
2. Methods
2.1. Stage 1: Data Acquisition, Quality Control, and Data Curation
2.2. Stage 2: Differential Protein Abundance and Pathway Analysis
- Type of process: biological process non-redundant;
- Number of genes per category: 5–2000;
- Fit test: Benjamini and Hochberg (BH);
- Multiple testing correction: false discovery rate (FDR);
- Reference gene list: genome protein-coding.
2.3. Stage 3: Inference and Analysis of Protein Co-Expression Networks
2.4. Stage 4: Network Modularity Analysis and Protein Interaction Networks
3. Results
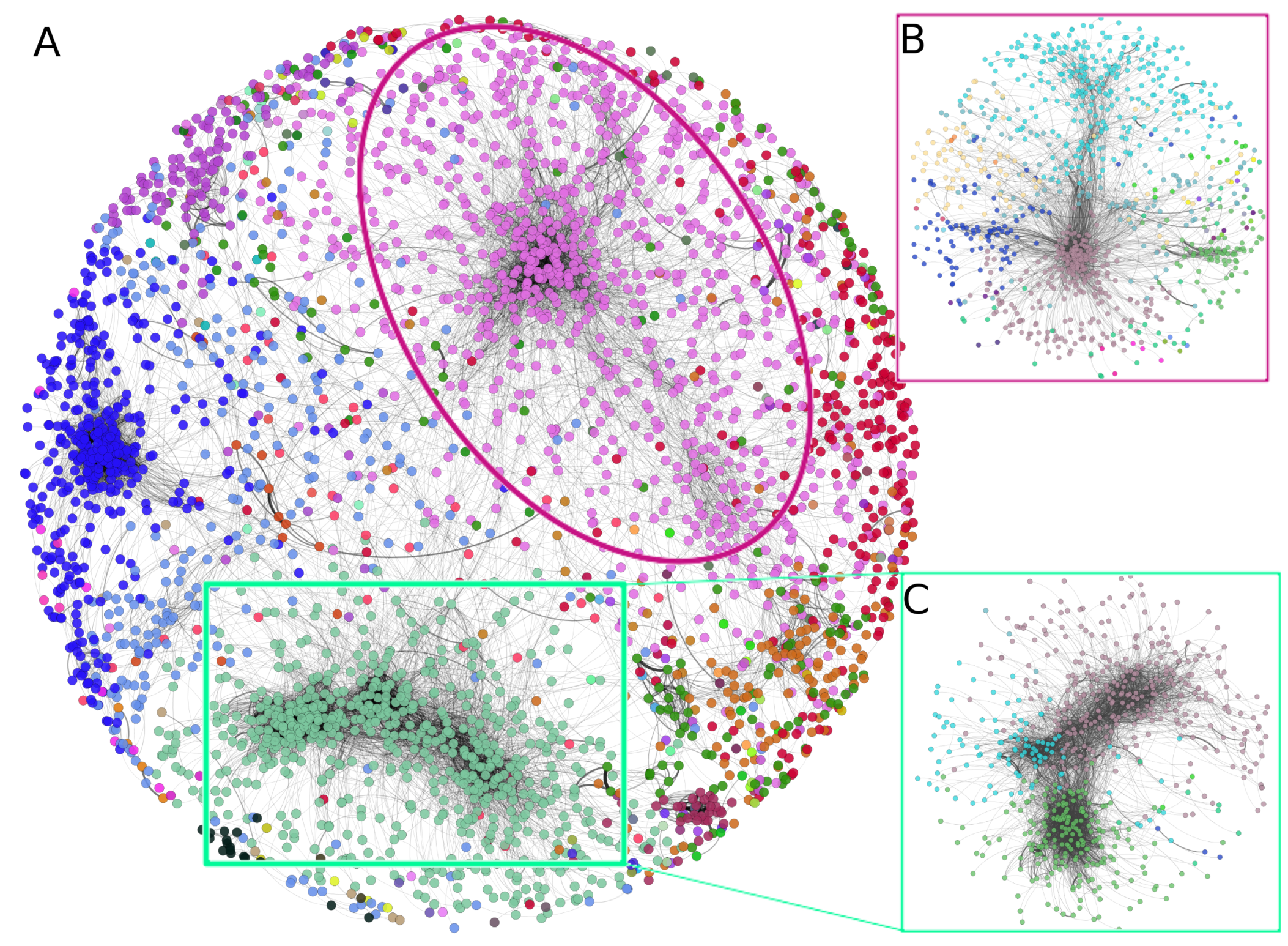
3.1. The Protein Co-Expression Network Shows a Scale-Free Topology
3.2. There Is a Hierarchical Organization in the PCN
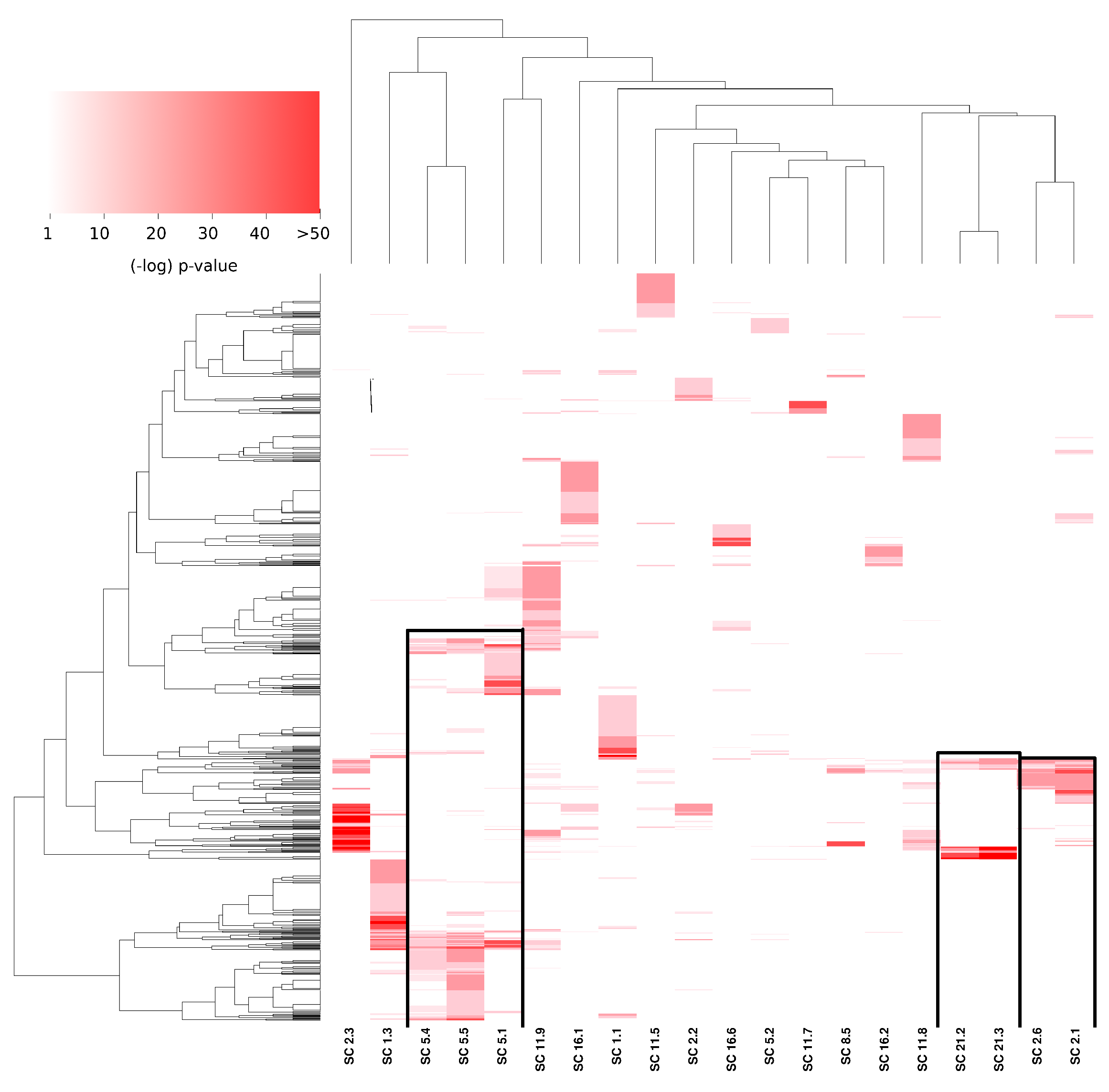
3.3. Subcommunities Are Significantly Associated with Specific Biological Processes
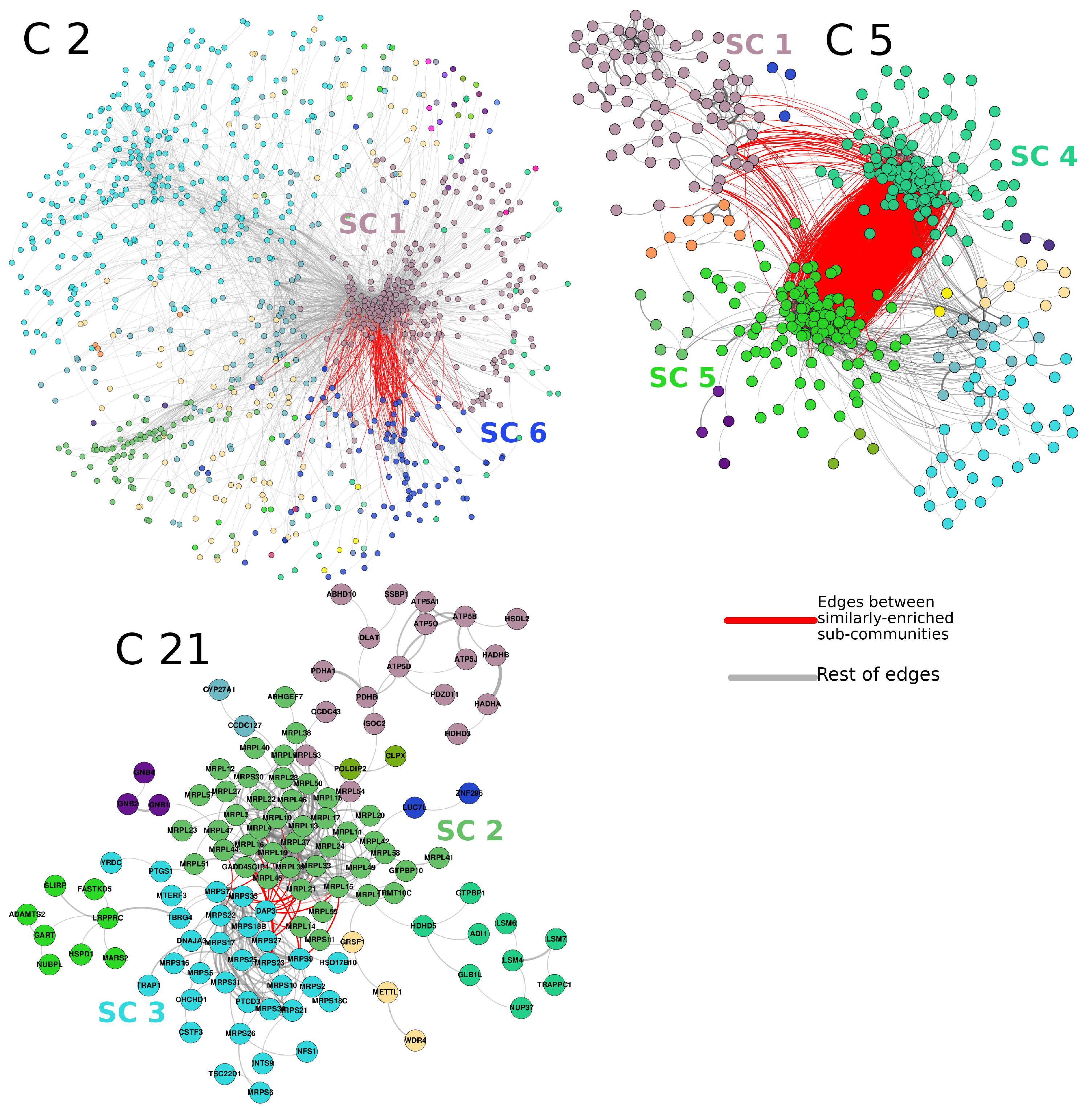
3.4. Subcommunities with Similar Enrichment Are Part of the Same Community, and They Share Several Interactions
3.5. The PCN Is Clustered by Differential Expression Trend
3.6. Protein Co-Expression Is Not Distance-Dependent
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. Ca Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Chong, W.; Zheng, L. Proteomics promises a new era of precision cancer medicine. Signal Transduct. Target. Ther. 2019, 4, 1–2. [Google Scholar]
- Martens, L.; Vizcaíno, J.A. A Golden Age for Working with Public Proteomics Data. Trends Biochem. Sci. 2017, 42, 333–341. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guo, Y.; Xing, Y. Weighted gene co-expression network analysis of pneumocytes under exposure to a carcinogenic dose of chloroprene. Life Sci. 2016, 151, 339–347. [Google Scholar] [CrossRef] [PubMed]
- Brunel, H.; Gallardo-Chacón, J.J.; Buil, A.; Vallverdú, M.; Soria, J.M.; Caminal, P.; Perera, A. Miss: A non-linear methodology based on mutual information for genetic association studies in both population and sib-pairs analysis. Bioinformatics 2010, 26, 1811–1818. [Google Scholar] [CrossRef] [Green Version]
- McShane, E.; Sin, C.; Zauber, H.; Wells, J.N.; Donnelly, N.; Wang, X.; Hou, J.; Chen, W.; Storchova, Z.; Marsh, J.A.; et al. Kinetic Analysis of Protein Stability Reveals Age-Dependent Degradation. Cell 2016, 167, 803–815. [Google Scholar] [CrossRef] [Green Version]
- Goh, W.W.B.; Wong, L. Integrating Networks and Proteomics: Moving Forward. Cell 2016, 34, 951–959. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.; Hou, J.; Tanner, J.J.; Cheng, J. Bioinformatics methods for mass spectrometry-based proteomics data analysis. Int. J. Mol. Sci. 2020, 21, 2873. [Google Scholar] [CrossRef] [Green Version]
- Sinitcyn, P.; Rudolph, J.D.; Cox, J. Computational methods for understanding mass spectrometry—Based shotgun proteomics data. Annu. Rev. Biomed. Data Sci. 2018, 1, 207–234. [Google Scholar] [CrossRef]
- Wickham, H. ggplot2. Wiley Interdiscip. Rev. Comput. Stat. 2011, 3, 180–185. [Google Scholar] [CrossRef]
- Wickham, H.; Francois, R.; Henry, L.; Müller, K. dplyr: A Grammar of Data Manipulation, Version 0.1. 2015. Available online: https://github.com/hadley/dplyr (accessed on 5 May 2022).
- Wickham, H.; Averick, M.; Bryan, J.; Chang, W.; McGowan, L.D.; François, R.; Grolemund, G.; Hayes, A.; Henry, L.; Hester, J.; et al. Welcome to the Tidyverse. J. Open Source Softw. 2019, 4, 1686. [Google Scholar] [CrossRef]
- Smyth, G.K. Limma: Linear Models for Microarray Data. Bioinformatics and Computational Biology Solutions Using R and Bioconductor; Springer: New York, NY, USA, 2005; pp. 397–420. [Google Scholar] [CrossRef] [Green Version]
- Waskom, M.L. Seaborn: Statistical data visualization. J. Open Source Softw. 2021, 6, 3021. [Google Scholar] [CrossRef]
- Hunter, J.D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 2007, 9, 90–95. [Google Scholar] [CrossRef]
- McKinney, W. Pandas: A foundational Python library for data analysis and statistics. Python High Perform. Sci. Comput. 2011, 14, 1–9. [Google Scholar]
- Margolin, A.A.; Nemenman, I.; Basso, K.; Wiggins, C.; Stolovitzky, G.; Dalla Favera, R.; Califano, A. ARACNE: An algorithm for the reconstruction of gene regulatory networks in a mammalian cellular context. BMC Bioinform. 2006, 7, 1–15. [Google Scholar] [CrossRef] [Green Version]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Cline, M.S.; Smoot, M.; Cerami, E.; Kuchinsky, A.; Landys, N.; Workman, C.; Christmas, R.; Avila-Campilo, I.; Creech, M.; Gross, B.; et al. Integration of biological networks and gene expression data using Cytoscape. Nat. Protoc. 2007, 2, 2366–2382. [Google Scholar] [CrossRef] [Green Version]
- de Anda-Jáuregui, G.; Fresno, C.; García-Cortés, D.; Enríquez, J.E.; Hernández-Lemus, E. Intrachromosomal regulation decay in breast cancer. Appl. Math. Nonlinear Sci. 2014, 4, 223–230. [Google Scholar] [CrossRef] [Green Version]
- Su, G.; Kuchinsky, A.; Morris, J.H.; States, D.J.; Meng, F. GLay: Community structure analysis of biological networks. Bioinformatics 2010, 26, 3135–3137. [Google Scholar] [CrossRef] [Green Version]
- Wuchty, S.; Ravasz, E.; Barabási, A.L. The architecture of biological networks. In Complex Systems Science in Biomedicine; Springer: Boston, MA, USA, 2006; pp. 165–181. [Google Scholar]
- Almaas, E.; Barabási, A.L. Power laws in biological networks. In Power Laws, Scale-Free Networks and Genome Biology; Springer: New York, NY, USA, 2006; pp. 1–11. [Google Scholar]
- Santolini, M.; Barabási, A.L. Predicting perturbation patterns from the topology of biological networks. Proc. Natl. Acad. Sci. USA 2018, 115, E6375–E6383. [Google Scholar] [CrossRef] [Green Version]
- Barabási, A.L. Scale-free networks: A decade and beyond. Science 2009, 325, 412–413. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Newman, M.E.; Barabási, A.L.E.; Watts, D.J. The Structure and Dynamics of Networks; Princeton University Press: Princeton, NJ, USA, 2006. [Google Scholar]
- Clauset, A.; Shalizi, C.R.; Newman, M.E. Power-law distributions in empirical data. Siam Rev. 2009, 51, 661–703. [Google Scholar] [CrossRef] [Green Version]
- Alcalá-Corona, S.A.; Velázquez-Caldelas, T.E.; Espinal-Enríquez, J.; Hernández-Lemus, E. Community structure reveals biologically functional modules in mef2c transcriptional regulatory network. Front. Physiol. 2016, 7, 184. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alcalá-Corona, S.A.; de Anda-Jáuregui, G.; Espinal-Enríquez, J.; Hernández-Lemus, E. Network modularity in breast cancer molecular subtypes. Front. Physiol. 2017, 8, 915. [Google Scholar] [CrossRef] [Green Version]
- García-Campos, M.A.; Espinal-Enríquez, J.; Hernández-Lemus, E. Pathway analysis: State of the art. Front. Physiol. 2015, 6, 383. [Google Scholar] [CrossRef] [Green Version]
- García-Cortés, D.; Hernández-Lemus, E.; Espinal-Enríquez, J. Luminal a breast cancer co-expression network: Structural and functional alterations. Front. Genet. 2021, 12, 629475. [Google Scholar] [CrossRef]
- Espinal-Enriquez, J.; Fresno, C.; Anda-Jáuregui, G.; Hernández-Lemus, E. RNA-Seq based genome-wide analysis reveals loss of inter-chromosomal regulation in breast cancer. Sci. Rep. 2017, 7, 1760. [Google Scholar] [CrossRef] [Green Version]
- García-Cortés, D.; de Anda-Jáuregui, G.; Fresno, C.; Hernández-Lemus, E.; Espinal-Enriquez, J. Loss of trans regulation in breast cancer molecular subtypes. bioRxiv 2018. [Google Scholar] [CrossRef]
- de Anda-Jáuregui, G.; Espinal-Enriquez, J.; Hernández-Lemus, E. Spatial organization of the gene regulatory program: An information theoretical approach to breast cancer transcriptomics. Entropy 2019, 21, 195. [Google Scholar] [CrossRef] [Green Version]
- García-Cortés, D.; de Anda-Jáuregui, G.; Fresno, C.; Hernández-Lemus, E.; Espinal-Enríquez, J. Gene co-expression is distance-dependent in breast cancer. Front. Oncol. 2020, 1232. [Google Scholar] [CrossRef]
- González-Espinoza, A.; Zamora, J.; Hernandez-Lemus, E.; Espinal-Enríquez, J. Gene co-expression in breast cancer: A matter of distance. Front. Oncol. 2021, 4743. [Google Scholar] [CrossRef] [PubMed]
- Vella, D.; Zoppis, I.; Mauri, G.; Mauri, P.; Di Silvestre, D. From protein–protein interactions to protein co-expression networks: A new perspective to evaluate large-scale proteomic data. EURASIP J. Bioinform. Syst. Biol. 2017, 2017, 1–16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nishimura, T.; Takadate, T.; Maeda, S.; Suzuki, T.; Minowa, T.; Fukuda, T.; Bando, Y.; Unno, M. Disease-related protein co-expression networks are associated with the prognosis of resectable node-positive pancreatic ductal adenocarcinoma. Res. Sq. 2022; preprint. [Google Scholar] [CrossRef]
- Nishimura, T.; Fujii, K.; Nakamura, H.; Naruki, S.; Sakai, H.; Kimura, H.; Miyazawa, T.; Takagi, M.; Furuya, N.; Marko-Varga, G.; et al. Protein co-expression network-based profiles revealed from laser-microdissected cancerous cells of lung squamous-cell carcinomas. Sci. Rep. 2021, 11, 20209. [Google Scholar] [CrossRef] [PubMed]
- Yamada, K.; Nishimura, T.; Wakiya, M.; Satoh, E.; Fukuda, T.; Amaya, K.; Bando, Y.; Hirano, H.; Ishikawa, T. Protein co-expression networks identified from HOT lesions of ER+ HER2–Ki-67high luminal breast carcinomas. Sci. Rep. 2021, 11, 1705. [Google Scholar] [CrossRef]
- Tretina, K.; Park, E.S.; Maminska, A.; MacMicking, J.D. Interferon-induced guanylate-binding proteins: Guardians of host defense in health and disease. J. Exp. Med. 2019, 216, 482–500. [Google Scholar] [CrossRef] [Green Version]
- Nitta, T.; Nasreen, M.; Seike, T.; Goji, A.; Ohigashi, I.; Miyazaki, T.; Ohta, T.; Kanno, M.; Takahama, Y. IAN family critically regulates survival and development of T lymphocytes. PLoS Biol. 2006, 4, e103. [Google Scholar] [CrossRef] [Green Version]
- Luckheeram, R.V.; Zhou, R.; Verma, A.D.; Xia, B. CD4+ T cells:Differentiation and functions. Clin. Dev. Immunol. 2012, 2012, 925135. [Google Scholar] [CrossRef] [Green Version]
- Patten, L.C.; Belaguli, N.S.; Baek, M.J.; Fagan, S.P.; Awad, S.S.; Berger, D.H. Serum response factor is alternatively spliced in human colon cancer. J. Surg. Res. 2004, 121, 92–100. [Google Scholar] [CrossRef]
- Zamora-Fuentes, J.M.; Hernández-Lemus, E.; Espinal-Enríquez, J. Gene Expression and Co-expression Networks Are Strongly Altered Through Stages in Clear Cell Renal Carcinoma. Front. Genet. 2020, 11, 1232. [Google Scholar] [CrossRef]
- Andonegui-Elguera, S.D.; Zamora-Fuentes, J.M.; Espinal-Enríquez, J.; Hernández-Lemus, E. Loss of Long Distance Co-Expression in Lung Cancer. Front. Genet. 2021, 12, 625741. [Google Scholar] [CrossRef] [PubMed]
- Drago-García, D.; Espinal-Enríquez, J.; Hernández-Lemus, E. Network analysis of EMT and MET micro-RNA regulation in breast cancer. Sci. Rep. 2017, 7, 13534. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- de Anda-Jáuregui, G.; Espinal-Enríquez, J.; Drago-García, D.; Hernández-Lemus, E. Nonredundant, highly connected microRNAs control functionality in breast cancer networks. Int. J. Genom. 2018, 2018, 9585383. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Gómez, C.; Hernández-Lemus, E.; Espinal-Enríquez, J. The Role of Copy Number Variants in Gene Co-Expression Patterns for Luminal B Breast Tumors. Front. Genet. 2022, 13, 806607. [Google Scholar] [CrossRef] [PubMed]
- Ochoa, S.; de Anda-Jáuregui, G.; Hernández-Lemus, E. Multi-omic regulation of the pam50 gene signature in breast cancer molecular subtypes. Front. Oncol. 2020, 10, 845. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of Cancer: The Next Generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [Green Version]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ruhle, M.; Espinal-Enríquez, J.; Hernández-Lemus, E. The Breast Cancer Protein Co-Expression Landscape. Cancers 2022, 14, 2957. https://doi.org/10.3390/cancers14122957
Ruhle M, Espinal-Enríquez J, Hernández-Lemus E. The Breast Cancer Protein Co-Expression Landscape. Cancers. 2022; 14(12):2957. https://doi.org/10.3390/cancers14122957
Chicago/Turabian StyleRuhle, Martín, Jesús Espinal-Enríquez, and Enrique Hernández-Lemus. 2022. "The Breast Cancer Protein Co-Expression Landscape" Cancers 14, no. 12: 2957. https://doi.org/10.3390/cancers14122957
APA StyleRuhle, M., Espinal-Enríquez, J., & Hernández-Lemus, E. (2022). The Breast Cancer Protein Co-Expression Landscape. Cancers, 14(12), 2957. https://doi.org/10.3390/cancers14122957







