MR Fingerprinting—A Radiogenomic Marker for Diffuse Gliomas
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. MR Fingerprinting Protocol
2.2. Co-Registration
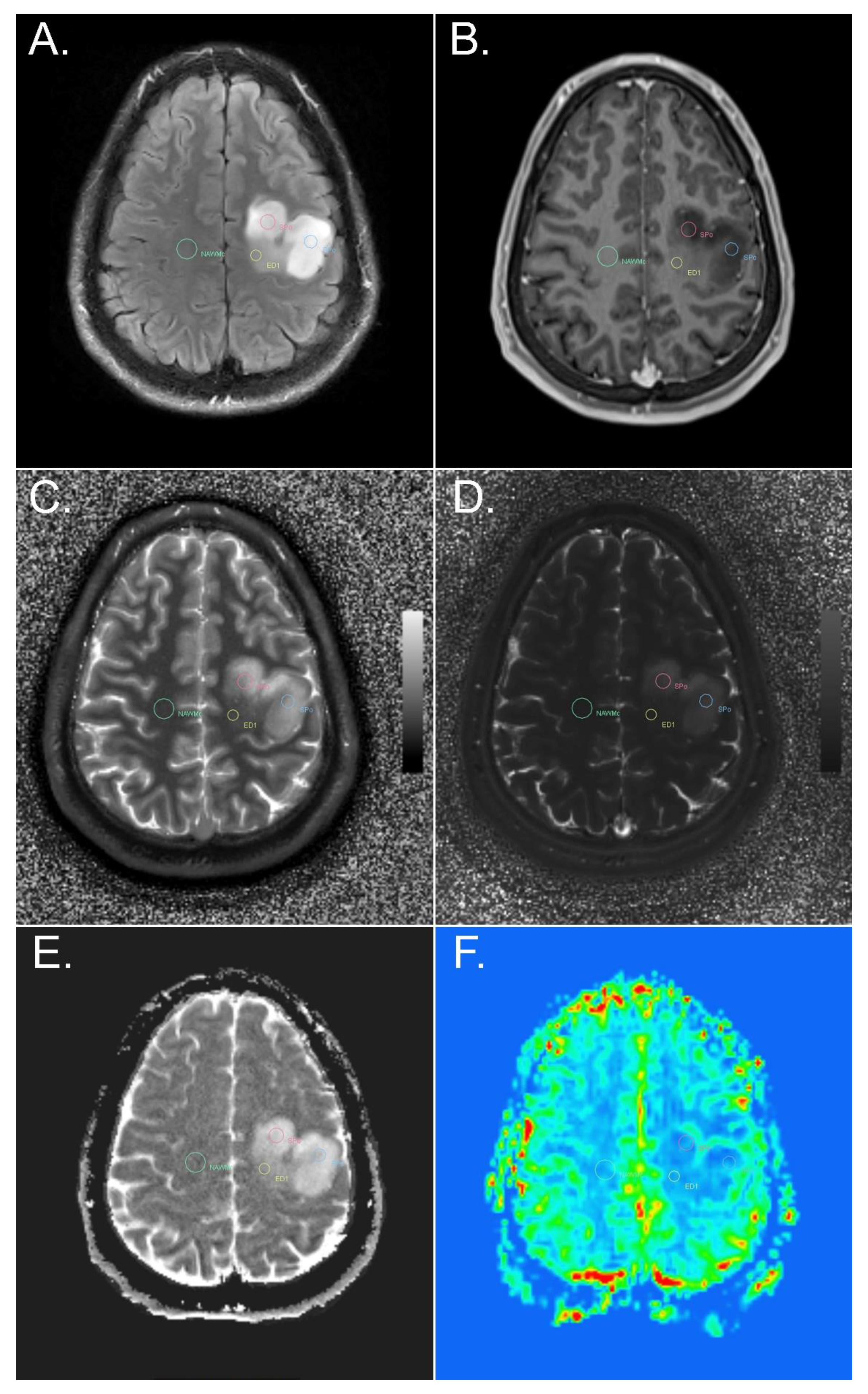
2.3. Region-of-Interest (ROI) Evaluation
2.4. Statistical Analysis
3. Results
3.1. IDH-Mutant versus IDH-Wildtype
3.2. Low Grade Gliomas (LGG) versus High Grade Gliomas (HGG)
3.3. IDH Mutational Status within Different Tumor Grades
3.4. MGMT Methylation Status
3.5. Solid Tumor—NAWM
3.6. Solid Tumor—Peritumoral Edema
3.7. Contrast-Enhancing and Hyperperfused versus Non-Contrast-Enhancing and Non-Hyperperfused Solid Tumor in IDH-Wildtype Gliomas
3.8. Correlation between MRF and Other Advanced MR Methods
3.9. Comparison between MRF and Conventional T1 and T2 Mapping
4. Discussion
Limitations of the Study
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Louis, D.N.; Perry, A.; Wesseling, P.; Brat, D.J.; Cree, I.A.; Figarella-Branger, D.; Hawkins, C.; Ng, H.K.; Pfister, S.M.; Reifenberger, G.; et al. The 2021 WHO Classification of Tumors of the Central Nervous System: A summary. Neuro Oncol. 2021, 23, 1231–1251. [Google Scholar] [CrossRef] [PubMed]
- Cohen, A.L.; Holmen, S.L.; Colman, H. IDH1 and IDH2 mutations in gliomas. Curr. Neurol. Neurosci. Rep. 2013, 13, 345. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Smits, M.; van den Bent, M.J. Imaging Correlates of Adult Glioma Genotypes. Radiology 2017, 284, 316–331. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yan, H.; Parsons, D.W.; Jin, G.; McLendon, R.; Rasheed, B.A.; Yuan, W.; Kos, I.; Batinic-Haberle, I.; Jones, S.; Riggins, G.J.; et al. IDH1 and IDH2 mutations in gliomas. N. Engl. J. Med. 2009, 360, 765–773. [Google Scholar] [CrossRef] [PubMed]
- Sanson, M.; Marie, Y.; Paris, S.; Idbaih, A.; Laffaire, J.; Ducray, F.; El Hallani, S.; Boisselier, B.; Mokhtari, K.; Hoang-Xuan, K.; et al. Isocitrate dehydrogenase 1 codon 132 mutation is an important prognostic biomarker in gliomas. J. Clin. Oncol. 2009, 27, 4150–4154. [Google Scholar] [CrossRef]
- Patel, S.H.; Bansal, A.G.; Young, E.B.; Batchala, P.P.; Patrie, J.T.; Lopes, M.B.; Jain, R.; Fadul, C.E.; Schiff, D. Extent of Surgical Resection in Lower-Grade Gliomas: Differential Impact Based on Molecular Subtype. AJNR Am. J. Neuroradiol. 2019, 40, 1149–1155. [Google Scholar] [CrossRef]
- Van den Bent, M.J.; Brandes, A.A.; Taphoorn, M.J.; Kros, J.M.; Kouwenhoven, M.C.; Delattre, J.Y.; Bernsen, H.J.; Frenay, M.; Tijssen, C.C.; Grisold, W.; et al. Adjuvant procarbazine, lomustine, and vincristine chemotherapy in newly diagnosed anaplastic oligodendroglioma: Long-term follow-up of EORTC brain tumor group study 26951. J. Clin. Oncol. 2013, 31, 344–350. [Google Scholar] [CrossRef]
- Wick, W.; Platten, M.; Meisner, C.; Felsberg, J.; Tabatabai, G.; Simon, M.; Nikkhah, G.; Papsdorf, K.; Steinbach, J.P.; Sabel, M.; et al. Temozolomide chemotherapy alone versus radiotherapy alone for malignant astrocytoma in the elderly: The NOA-08 randomised, phase 3 trial. Lancet Oncol. 2012, 13, 707–715. [Google Scholar] [CrossRef] [Green Version]
- Ellingson, B.M.; Lai, A.; Harris, R.J.; Selfridge, J.M.; Yong, W.H.; Das, K.; Pope, W.B.; Nghiemphu, P.L.; Vinters, H.V.; Liau, L.M.; et al. Probabilistic radiographic atlas of glioblastoma phenotypes. AJNR Am. J. Neuroradiol. 2013, 34, 533–540. [Google Scholar] [CrossRef] [Green Version]
- Yamashita, K.; Hiwatashi, A.; Togao, O.; Kikuchi, K.; Hatae, R.; Yoshimoto, K.; Mizoguchi, M.; Suzuki, S.O.; Yoshiura, T.; Honda, H. MR Imaging-Based Analysis of Glioblastoma Multiforme: Estimation of IDH1 Mutation Status. AJNR Am. J. Neuroradiol. 2016, 37, 58–65. [Google Scholar] [CrossRef] [Green Version]
- Kickingereder, P.; Sahm, F.; Radbruch, A.; Wick, W.; Heiland, S.; Von Deimling, A.; Bendszus, M.; Wiestler, B. IDH mutation status is associated with a distinct hypoxia/angiogenesis transcriptome signature which is non-invasively predictable with rCBV imaging in human glioma. Sci. Rep. 2015, 5, 16238. [Google Scholar] [CrossRef] [PubMed]
- Suh, C.H.; Kim, H.S.; Jung, S.C.; Choi, C.G.; Kim, S.J. Clinically Relevant Imaging Features for MGMT Promoter Methylation in Multiple Glioblastoma Studies: A Systematic Review and Meta-Analysis. AJNR Am. J. Neuroradiol. 2018, 39, 1439–1445. [Google Scholar] [PubMed]
- Cindil, E.; Sendur, H.N.; Cerit, M.N.; Erdogan, N.; Celebi, F.; Dag, N.; Celtikci, E.; Inan, A.; Oner, Y.; Tali, T. Prediction of IDH Mutation Status in High-grade Gliomas Using DWI and High T1-weight DSC-MRI. Acad. Radiol. 2021. [Google Scholar] [CrossRef] [PubMed]
- Choi, C.; Ganji, S.K.; DeBerardinis, R.J.; Hatanpaa, K.J.; Rakheja, D.; Kovacs, Z.; Yang, X.L.; Mashimo, T.; Raisanen, J.M.; Marin-Valencia, I.; et al. 2-hydroxyglutarate detection by magnetic resonance spectroscopy in IDH-mutated patients with gliomas. Nat. Med. 2012, 18, 624–629. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Andronesi, O.C.; Loebel, F.; Bogner, W.; Marjańska, M.; Vander Heiden, M.G.; Iafrate, A.J.; Dietrich, J.; Batchelor, T.T.; Gerstner, E.R.; Kaelin, W.G.; et al. Treatment Response Assessment in IDH-Mutant Glioma Patients by Noninvasive 3D Functional Spectroscopic Mapping of 2-Hydroxyglutarate. Clin. Cancer Res. 2016, 22, 1632–1641. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Andronesi, O.C.; Arrillaga-Romany, I.C.; Ly, K.I.; Bogner, W.; Ratai, E.M.; Reitz, K.; Iafrate, A.J.; Dietrich, J.; Gerstner, E.R.; Chi, A.S.; et al. Pharmacodynamics of mutant-IDH1 inhibitors in glioma patients probed by in vivo 3D MRS imaging of 2-hydroxyglutarate. Nat. Commun. 2018, 9, 1474. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Ke, C.; Liu, J.; Xu, S.; Han, L.; Yang, Y.; Qian, L.; Liu, X.; Zheng, H.; Lv, X.; et al. Diagnostic performance between MR amide proton transfer (APT) and diffusion kurtosis imaging (DKI) in glioma grading and IDH mutation status prediction at 3T. Eur. J. Radiol. 2021, 134, 109466. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.; Zou, T.; Eberhart, C.G.; Villalobos, M.A.; Heo, H.Y.; Zhang, Y.; Wang, Y.; Wang, X.; Yu, H.; Du, Y.; et al. Predicting IDH mutation status in grade II gliomas using amide proton transfer-weighted (APTw) MRI. Magn. Reason. Med. 2017, 78, 1100–1109. [Google Scholar] [CrossRef]
- Thust, S.C.; Heiland, S.; Falini, A.; Jäger, H.R.; Waldman, A.D.; Sundgren, P.C.; Godi, C.; Katsaros, V.K.; Ramos, A.; Bargallo, N.; et al. Glioma imaging in Europe: A survey of 220 centres and recommendations for best clinical practice. Eur. Radiol. 2018, 28, 3306–3317. [Google Scholar] [CrossRef] [Green Version]
- Nakai, K.; Nawashiro, H.; Shima, K.; Kaji, T. An Analysis of T2 Mapping on Brain Tumors. In Brain Edema XV. Acta Neurochirurgica Supplement; Katayama, Y., Maeda, T., Kuroiwa, T., Eds.; Springer: Vienna, Austria, 2013; Volume 118. [Google Scholar] [CrossRef]
- Hattingen, E.; Jurcoane, A.; Daneshvar, K.; Pilatus, U.; Mittelbronn, M.; Steinbach, J.P.; Bähr, O. Quantitative T2 mapping of recurrent glioblastoma under bevacizumab improves monitoring for non-enhancing tumor progression and predicts overall survival. Neuro Oncol. 2013, 15, 1395–1404. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lescher, S.; Jurcoane, A.; Veit, A.; Bähr, O.; Deichmann, R.; Hattingen, E. Quantitative T1 and T2 mapping in recurrent glioblastomas under bevacizumab: Earlier detection of tumor progression compared to conventional MRI. Neuroradiology 2015, 57, 11–20. [Google Scholar] [CrossRef] [PubMed]
- Min, Z.G.; Niu, C.; Rana, N.; Ji, H.M.; Zhang, M. Differentiation of pure vasogenic edema and tumor-infiltrated edema in patients with peritumoral edema by analyzing the relationship of axial and radial diffusivities on 3.0T MRI. Clin. Neurol. Neurosurg. 2013, 115, 1366–1370. [Google Scholar] [CrossRef] [PubMed]
- European Society of Radiology. ESR statement on the stepwise development of imaging biomarkers. Insights Imaging 2013, 4, 147–152. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, D.; Gulani, V.; Seiberlich, N.; Liu, K.; Sunshine, J.L.; Duerk, J.L.; Griswold, M.A. Magnetic resonance fingerprinting. Nature 2013, 495, 187–192. [Google Scholar] [CrossRef] [Green Version]
- Louis, D.N.; Perry, A.; Reifenberger, G.; Von Deimling, A.; Figarella-Branger, D.; Cavenee, W.K.; Ohgaki, H.; Wiestler, O.D.; Kleihues, P.; Ellison, D.W. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: A summary. Acta Neuropathol. 2016, 131, 803–820. [Google Scholar] [CrossRef] [Green Version]
- Saake, M.; Schmidle, A.; Kopp, M.; Hanspach, J.; Hepp, T.; Laun, F.B.; Nagel, A.M.; Dörfler, A.; Uder, M.; Bäuerle, T. MRI Brain Signal Intensity and Relaxation Times in Individuals with Prior Exposure to Gadobutrol. Radiology 2019, 290, 659–668. [Google Scholar] [CrossRef]
- Crawley, P.A.; Henkelman, R.M. A comparison of one-shot and recovery methods in T1 imaging. Magn. Reason. Med. 1988, 7, 23–34. [Google Scholar] [CrossRef]
- Poon, C.S.; Henkelman, R.M. Practical T2 quantitation for clinical applications. J. Magn. Reason. Imaging 1992, 2, 541–553. [Google Scholar] [CrossRef]
- Marques, J.P.; Kober, T.; Krueger, G.; van der Zwaag, W.; Van de Moortele, P.F.; Gruetter, R. MP2RAGE, a self bias-field corrected sequence for improved segmentation and T1-mapping at high field. Neuroimage 2010, 49, 1271–1281. [Google Scholar] [CrossRef] [Green Version]
- Mendlik, T.; Faber, S.C.; Weber, J.; Hohe, J.; Rauch, E.; Reiser, M.; Glaser, C. T2 quantitation of human articular cartilage in a clinical setting at 1.5 T: Implementation and testing of four multiecho pulse sequence designs for validity. Investig. Radiol. 2004, 39, 288–299. [Google Scholar] [CrossRef]
- Jiang, Y.; Ma, D.; Seiberlich, N.; Gulani, V.; Griswold, M.A. MR fingerprinting using fast imaging with steady state precession (FISP) with spiral readout. Magn. Reason. Med. 2015, 74, 1621–1631. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yokota, Y.; Okada, T.; Fushimi, Y.; Yamamoto, A.; Nakajima, S.; Fujimoto, K.; Oshima, S.; Koerzdoerfer, G.; Nittka, M.; Pfeuffer, J.; et al. Acceleration of 2D-MR fingerprinting by reducing the number of echoes with increased in-plane resolution: A volunteer study. Magn. Reson. Mater. Phys. Biol. Med. 2020, 33, 783–791. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Körzdörfer, G.; Cardoso, P.L.; Bär, P.; Kitzer, S.; Bogner, W.; Trattnig, S.; Nittka, M. Data-driven motion detection for MR Fingerprinting. In Proceedings of the ISMRM & SMRT Virtual Conference & Exhibition, 8–14 August 2020; Available online: https://www.ismrm.org/20/program_files/PP25.htm (accessed on 10 December 2021).
- Yu, Z.; Zhao, T.; Assländer, J.; Lattanzi, R.; Sodickson, D.K.; Cloos, M.A. Exploring the sensitivity of magnetic resonance fingerprinting to motion. Magn. Reason. Imaging 2018, 54, 241–248. [Google Scholar] [CrossRef] [PubMed]
- Badve, C.; Yu, A.; Rogers, M.; Ma, D.; Liu, Y.; Schluchter, M.; Sunshine, J.; Griswold, M.; Gulani, V. Simultaneous T1 and T2 Brain Relaxometry in Asymptomatic Volunteers using Magnetic Resonance Fingerprinting. Tomography 2015, 1, 136–144. [Google Scholar] [CrossRef] [PubMed]
- Paulson, E.S.; Schmainda, K.M. Comparison of dynamic susceptibility-weighted contrast-enhanced MR methods: Recommendations for measuring relative cerebral blood volume in brain tumors. Radiology 2008, 249, 601–613. [Google Scholar] [CrossRef] [PubMed]
- Haubold, J.; Demircioglu, A.; Gratz, M.; Glas, M.; Wrede, K.; Sure, U.; Antoch, G.; Keyvani, K.; Nittka, M.; Kannengiesser, S.; et al. Non-invasive tumor decoding and phenotyping of cerebral gliomas utilizing multiparametric (18)F-FET PET-MRI and MR Fingerprinting. Eur. J. Nucl. Med. Mol. Imaging 2020, 47, 1435–1445. [Google Scholar] [CrossRef]
- Badve, C.; Yu, A.; Dastmalchian, S.; Rogers, M.; Ma, D.; Jiang, Y.; Margevicius, S.; Pahwa, S.; Lu, Z.; Schluchter, M.; et al. MR Fingerprinting of Adult Brain Tumors: Initial Experience. AJNR Am. J. Neuroradiol. 2017, 38, 492–499. [Google Scholar] [CrossRef] [Green Version]
- Koeller, K.K.; Rushing, E.J. Oligodendroglioma and its variants: Radiologic-pathologic correlation. Radiographics 2005, 25, 1669–1688. [Google Scholar] [CrossRef]
- Dastmalchian, S.; Kilinc, O.; Onyewadume, L.; Tippareddy, C.; McGivney, D.; Ma, D.; Griswold, M.; Sunshine, J.; Gulani, V.; Barnholtz-Sloan, J.S.; et al. Radiomic analysis of magnetic resonance fingerprinting in adult brain tumors. Eur. J. Nucl. Med. Mol. Imaging 2021, 48, 683–693. [Google Scholar] [CrossRef]
- Hong, E.K.; Choi, S.H.; Shin, D.J.; Jo, S.W.; Yoo, R.E.; Kang, K.M.; Yun, T.J.; Kim, J.H.; Sohn, C.H.; Park, S.H.; et al. Radiogenomics correlation between MR imaging features and major genetic profiles in glioblastoma. Eur. Radiol. 2018, 28, 4350–4361. [Google Scholar] [CrossRef]
- Hoehn-Berlage, M.; Bockhorst, K. Quantitative magnetic resonance imaging of rat brain tumors: In vivo NMR relaxometry for the discrimination of normal and pathological tissues. Technol. Health Care 1994, 2, 247–254. [Google Scholar] [CrossRef] [PubMed]
- Oh, J.; Cha, S.; Aiken, A.H.; Han, E.T.; Crane, J.C.; Stainsby, J.A.; Wright, G.A.; Dillon, W.P.; Nelson, S.J. Quantitative apparent diffusion coefficients and T2 relaxation times in characterizing contrast enhancing brain tumors and regions of peritumoral edema. J. Magn. Reason. Imaging 2005, 21, 701–708. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Ma, D.; Keenan, K.E.; Stupic, K.F.; Gulani, V.; Griswold, M.A. Repeatability of magnetic resonance fingerprinting T1 and T2 estimates assessed using the ISMRM/NIST MRI system phantom. Magn. Reason. Med. 2017, 78, 1452–1457. [Google Scholar] [CrossRef] [PubMed]
- Körzdörfer, G.; Kirsch, R.; Liu, K.; Pfeuffer, J.; Hensel, B.; Jiang, Y.; Ma, D.; Gratz, M.; Bär, P.; Bogner, W.; et al. Reproducibility and Repeatability of MR Fingerprinting Relaxometry in the Human Brain. Radiology 2019, 292, 429–437. [Google Scholar] [CrossRef]
- Ma, D.; Pierre, E.Y.; Jiang, Y.; Schluchter, M.D.; Setsompop, K.; Gulani, V.; Griswold, M.A. Music-based magnetic resonance fingerprinting to improve patient comfort during MRI examinations. Magn. Reson. Med. 2016, 75, 2303–2314. [Google Scholar] [CrossRef] [Green Version]
- Pierre, E.Y.; Ma, D.; Chen, Y.; Badve, C.; Griswold, M.A. Multiscale reconstruction for MR fingerprinting. Magn. Reson. Med. 2016, 75, 2481–2492. [Google Scholar] [CrossRef] [Green Version]
- Assländer, J.; Cloos, M.A.; Knoll, F.; Sodickson, D.K.; Hennig, J.; Lattanzi, R. Low rank alternating direction method of multipliers reconstruction for MR fingerprinting. Magn. Reason. Med. 2018, 79, 83–96. [Google Scholar] [CrossRef]
- Ma, D.; Pierre, E.Y.; McGivney, D.; Mehta, B.; Chen, Y.; Jiang, Y.; Griswold, M. Applications of low rank modeling to fast 3D magnetic resonance fingerprinting (MRF). In Proceedings of the ISMRM, 25th Annual Meeting and Exhibition, Honolulu, HI, USA, 22–27 April 2017. [Google Scholar]
- Kiselev, V.G.; Korzdorfer, G.; Gall, P. Toward Quantification: Microstructure and Magnetic Resonance Fingerprinting. Investig. Radiol. 2021, 56, 1–9. [Google Scholar] [CrossRef]





| Neuropathological Tumor Type (WHO 2016) | Neuropathological Tumor Grade (WHO 2016) | MGMPR Promoter Methylation Status | Age | Gender |
|---|---|---|---|---|
| Diffuse astrocytoma, IDH-mutant | WHO grade II | Methylated | 23 | M |
| Diffuse astrocytoma, IDH-mutant | WHO grade II | Methylated | 33 | M |
| Diffuse astrocytoma, IDH-mutant | WHO grade II | Unmethylated | 54 | F |
| Diffuse astrocytoma, IDH-mutant | WHO grade II | Methylated | 77 | F |
| Diffuse astrocytoma, IDH-mutant | WHO grade II | Methylated | 46 | F |
| Diffuse astrocytoma, IDH-mutant | WHO grade II | Unmethylated | 57 | M |
| Diffuse astrocytoma, IDH-wildtype | WHO grade II | Unmethylated | 27 | M |
| Anaplastic astrocytoma, IDH-mutant | WHO grade III | Unmethylated | 59 | M |
| Anaplastic astrocytoma, IDH-mutant | WHO grade III | Methylated | 29 | M |
| Anaplastic astrocytoma, IDH-mutant | WHO grade III | Methylated | 28 | F |
| Anaplastic astrocytoma, IDH-wildtype | WHO grade III | Unmethylated | 65 | F |
| Glioblastoma, IDH-mutant | WHO grade IV | Methylated | 45 | F |
| Glioblastoma, IDH-wildtype | WHO grade IV | Unmethylated | 47 | F |
| Glioblastoma, IDH-wildtype | WHO grade IV | Unmethylated | 58 | M |
| Glioblastoma, IDH-wildtype | WHO grade IV | Unmethylated | 59 | F |
| Glioblastoma, IDH-wildtype | WHO grade IV | Methylated | 52 | M |
| Glioblastoma, IDH-wildtype | WHO grade IV | Unmethylated | 59 | M |
| Glioblastoma, IDH-wildtype | WHO grade IV | Methylated | 71 | M |
| Glioblastoma, IDH-wildtype | WHO grade IV | Methylated | 61 | M |
| Glioblastoma, IDH-wildtype | WHO grade IV | Methylated | 62 | M |
| Oligoendroglioma, IDH-mutant and 1p/19q-codeleted | WHO grade II | Methylated | 52 | M |
| Oligoendroglioma, IDH-mutant and 1p/19q-codeleted | WHO grade II | Methylated | 38 | F |
| Oligoendroglioma, IDH-mutant and 1p/19q-codeleted | WHO grade II | Methylated | 61 | M |
| Anaplastic oligodendroglioma, IDH-mutant and 1p/19q-codeleted | WHO grade III | Methylated | 51 | M |
| Examination Parameters | 2D ax T2 FLAIR | 2D T2 ax | DWI ax | 3D SWI ax | 3D T1 ax pre | 2D T2 cor | PWI ax | 3D T1 ax Post | 3D FLAIR |
|---|---|---|---|---|---|---|---|---|---|
| TSE + IR | TSE | EPI-SE | GRE | MPRAGE | TSE | SS-EPI | MPRAGE | TSE + IR | |
| Voxel dimensions | 0.9 × 0.9 | 0.8 × 0.6 | 1.8 × 1.8 | 0.9 × 0.9 | 1 × 1 | 0.4 × 0.4 | 1.8 × 1.8 | 1 × 1 | 1 × 1 |
| Matrix size | 256 × 256 | 250 × 384 | 128 × 128 | 256 × 192 | 256 × 256 | 531 × 640 | 128 × 128 | 256 × 256 | 256 × 256 |
| No. slices | 36 | 40 | 30 | 80 | 192 | 56 | 19 | 192 | 176 |
| Field of view [mm2] | 230 | 210 | 230 | 230 | 220 | 230 | 230 | 220 | 250 |
| Slice thickness, mm | 4 | 3 | 5 | 1.75 | 1 | 3 | 5 | 1 | 0.9 |
| TE [ms] | 100 | 88 | 78 | 20 | 3.79 | 115 | 32 | 3.79 | 393 |
| TI [ms] | 2500 | - | - | - | 1100 | - | - | - | 2050 |
| TR [ms] | 9220 | 3490 | 4000 | 28 | 1800 | 4290 | 1400 | 1800 | 7000 |
| TA, [min:s] | 4:38 | 1:25 | 1:38 | 3:52 | 5:44 | 3:40 | 1:17 | 5:44 | 3:39 |
| GRAPPA factor | - | - | 2 | 2 | - | 2 | 2 | - | - |
| BW/pixel, [Hz/pixel] | 170 | 199 | 1502 | 120 | 200 | 176 | 1346 | 200 | 651 |
| FA [°] | 150 | 120 | - | 15 | 12 | 120 | 90 | 12 | T2var |
| Fat saturation | yes | No | Yes | No | No | No | Yes | No | Yes |
| Examination Parameters | 2D ax T2 FLAIR | 2D T2 ax | 3D T1 Sag | 2D Multi-Echo Spin Echo | MRF |
|---|---|---|---|---|---|
| TSE + IR | TSE | MP2RAGE (T1 map) | (T2 map) | ||
| Voxel dimensions [mm2] | 0.6 × 0.6 | 0.7 × 0.7 | 1.0 × 1.0 | 0.7 × 0.7 | 1.0 × 1.0 |
| Matrix size | 384 × 276 | 320 × 240 | 256 × 216 | 320 × 257 | 256 × 256 |
| No. slices | 10 | 23 | 160 | 10 | 10–13 |
| Field of view [mm2] | 230 × 166 | 230 × 170 | 256 × 216 | 230 × 180 | 256 × 256 |
| Slice thickness [mm] | 5.0 | 5.0 | 1.0 | 5.0 | 5.0 |
| TE [ms] | 126 | 118 | 2.98 | 12.6, 25.2, … 201.6 | 2.0 |
| TI [ms] | 2500 | – | 700, 2500 | – | 21.0 |
| TR [ms] | 8500 | 4890 | 5000 | 2100 | 12.14–15.00 (varied by sequence) |
| TA [min:sec] | 3:43 | 1:25 | 8:02 | 3:38 | 3:51–4:51 |
| Acceleration factor | 1 (turbo factor: 19) | 2 | 2 | 3 | 24 (inner k-space), 48 (outer k-space) |
| BW/pixel [Hz/pixel] | 140 | 130 | 240 | 150 | RX-Bandwidth: 400 kHz |
| FA [°] | 180 | 180 | 4, 5 | 180 | 0–74 (varied by sequence) |
| Fat saturation | Yes | – | No | No | no |
| Tumor and Peritumoral Edema | Quantitative Parameter | MRFT1 Mean | MRFT2 Mean |
|---|---|---|---|
| Solid part | T1 mean | 0.913 | |
| <0.001 | |||
| T2 mean | 0.775 | ||
| <0.001 | |||
| ADC mean | 0.697 | 0.813 | |
| <0.001 | <0.001 | ||
| rCBV mean | −0.181 | −0.374 | |
| 0.181 | 0.005 | ||
| Edema ≤ 1 cm adjacent to the solid part | T1 mean | 0.882 | |
| <0.001 | |||
| T2 mean | 0.884 | ||
| <0.001 | |||
| ADC mean | 0.742 | 0.900 | |
| <0.001 | <0.001 | ||
| rCBV mean | −0.174 | −0.223 | |
| 0.376 | 0.254 | ||
| Edema > 1 cm adjacent to the solid part | T1 mean | 0.983 | |
| <0.001 | |||
| T2 mean | 0.983 | ||
| <0.001 | |||
| ADC mean | 0.810 | 0.786 | |
| 0.015 | 0.021 | ||
| rCBV mean | <0.001 | −0.050 | |
| 1.000 | 0.898 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Springer, E.; Cardoso, P.L.; Strasser, B.; Bogner, W.; Preusser, M.; Widhalm, G.; Nittka, M.; Koerzdoerfer, G.; Szomolanyi, P.; Hangel, G.; et al. MR Fingerprinting—A Radiogenomic Marker for Diffuse Gliomas. Cancers 2022, 14, 723. https://doi.org/10.3390/cancers14030723
Springer E, Cardoso PL, Strasser B, Bogner W, Preusser M, Widhalm G, Nittka M, Koerzdoerfer G, Szomolanyi P, Hangel G, et al. MR Fingerprinting—A Radiogenomic Marker for Diffuse Gliomas. Cancers. 2022; 14(3):723. https://doi.org/10.3390/cancers14030723
Chicago/Turabian StyleSpringer, Elisabeth, Pedro Lima Cardoso, Bernhard Strasser, Wolfgang Bogner, Matthias Preusser, Georg Widhalm, Mathias Nittka, Gregor Koerzdoerfer, Pavol Szomolanyi, Gilbert Hangel, and et al. 2022. "MR Fingerprinting—A Radiogenomic Marker for Diffuse Gliomas" Cancers 14, no. 3: 723. https://doi.org/10.3390/cancers14030723








