Downregulation of LOC441461 Promotes Cell Growth and Motility in Human Gastric Cancer
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Expression Data from TCGA
2.2. Information Gain (IG)
2.3. Survival Analysis
2.4. Gastric Cancer Cell Lines and Culture
2.5. siRNA Transfection
2.6. Subcellular Fractionation and RNA Isolation
2.7. qRT-PCR
2.8. Proliferation Assay
2.9. Colony Formation Assay
2.10. Cell Cycle Assay
2.11. Apoptosis Assay
2.12. Migration and Invasion Assays
2.13. Wound Healing Assay
2.14. RNA-Seq Analysis
2.15. Differentially Expressed Gene Analysis
2.16. Pathway Enrichment Analysis
2.17. Transcription Factor Enrichment Analysis Using the LncRNA-TF Interactome
2.18. Transcription Factor Target Gene Extraction
2.19. Hierarchical Clustering
2.20. Statistical Analysis
3. Results
3.1. Comparison of Gene Expression between Stage I and Stage IV Gastric Cancer
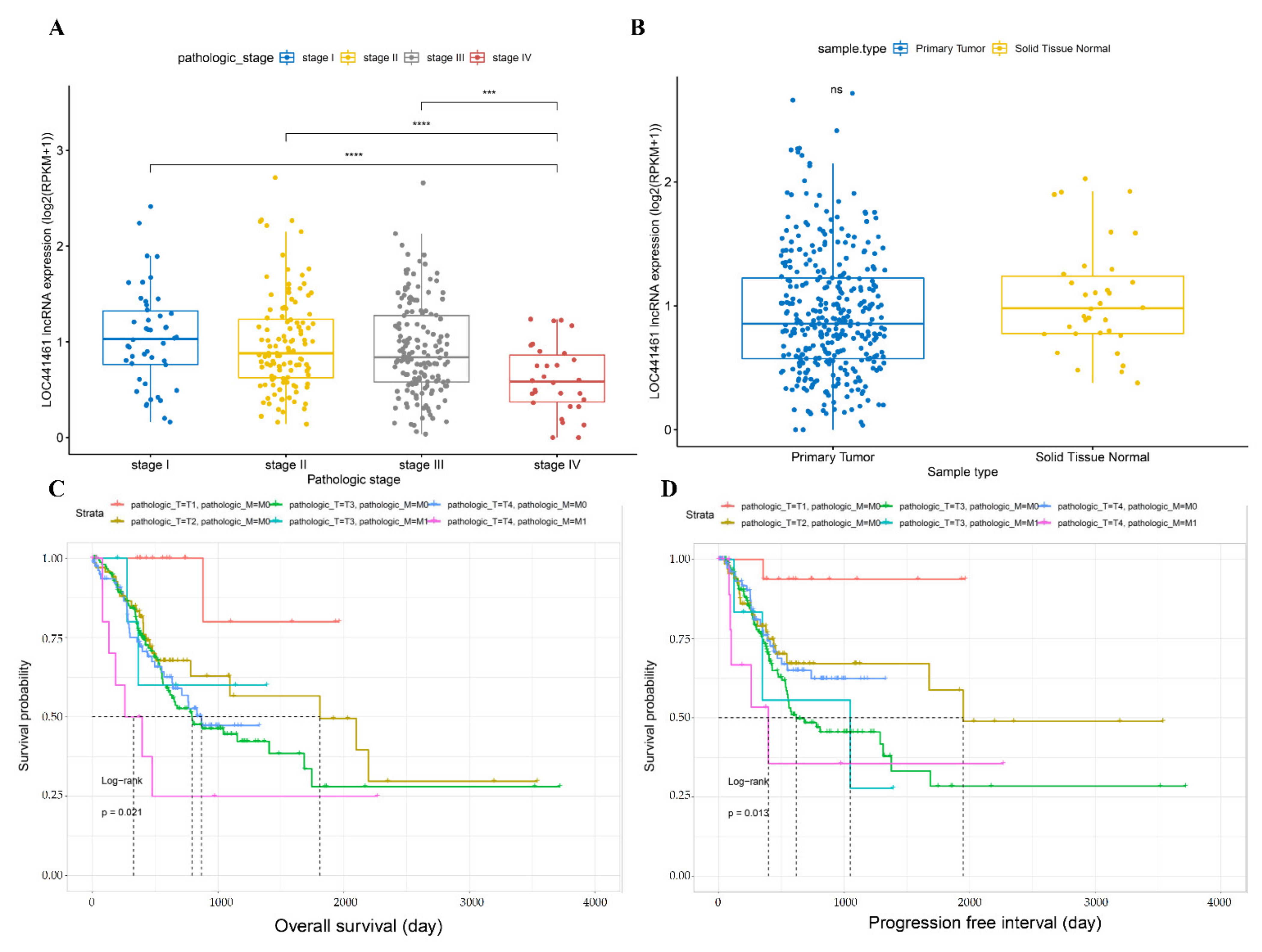
3.2. Comparison of LOC441461 Expression and Prognosis between Different Gastric Cancer Stages
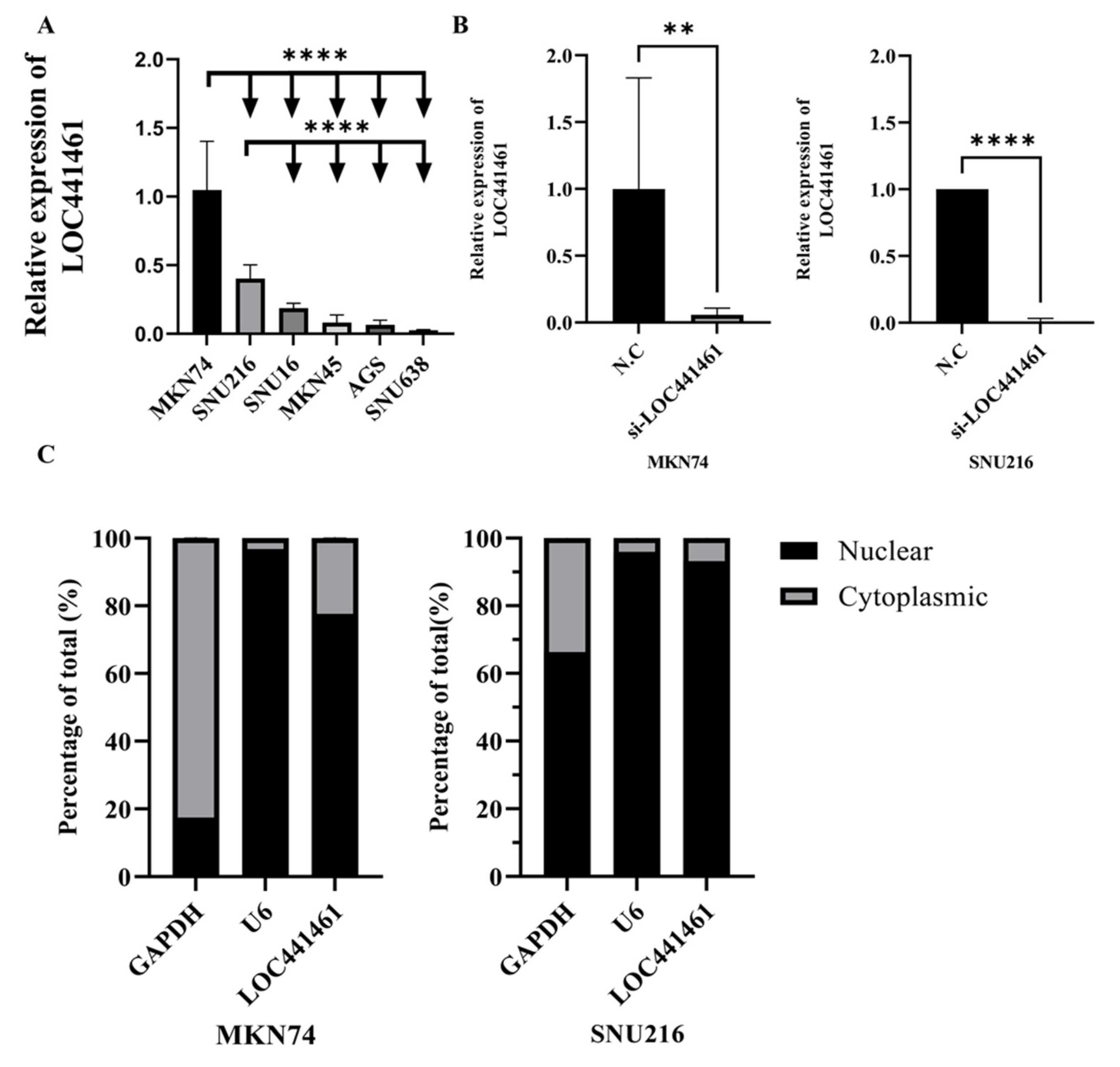
3.3. Selection of Gastric Cancer Cell Lines and si-LOC441461 Treatment
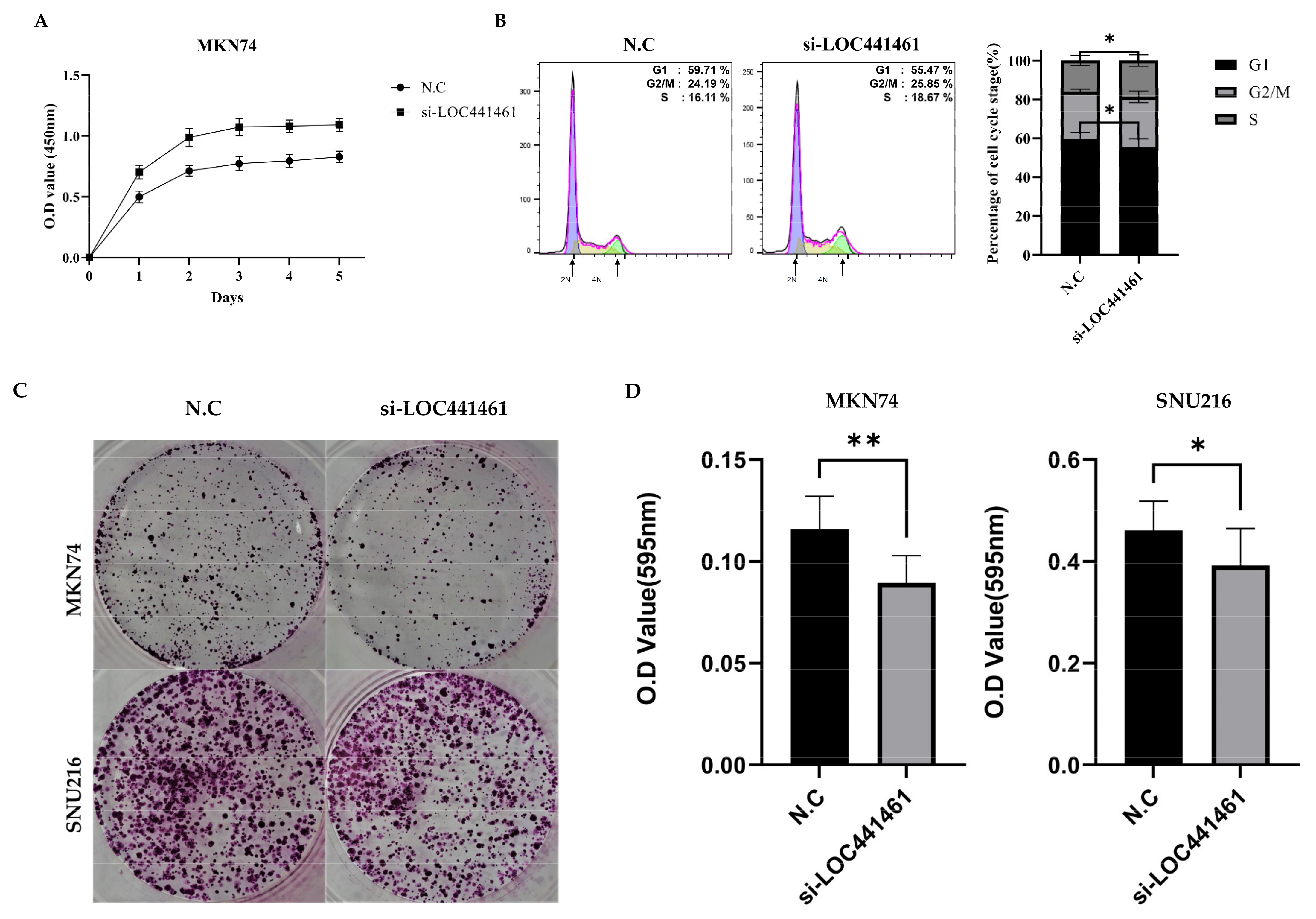
3.4. Downregulation of LOC441461 Induces Proliferation and Accelerates Cell Cycle Progression
3.5. LOC441461 Knockdown Promotes 5-FU-Induced Gastric Cancer Cell Apoptosis
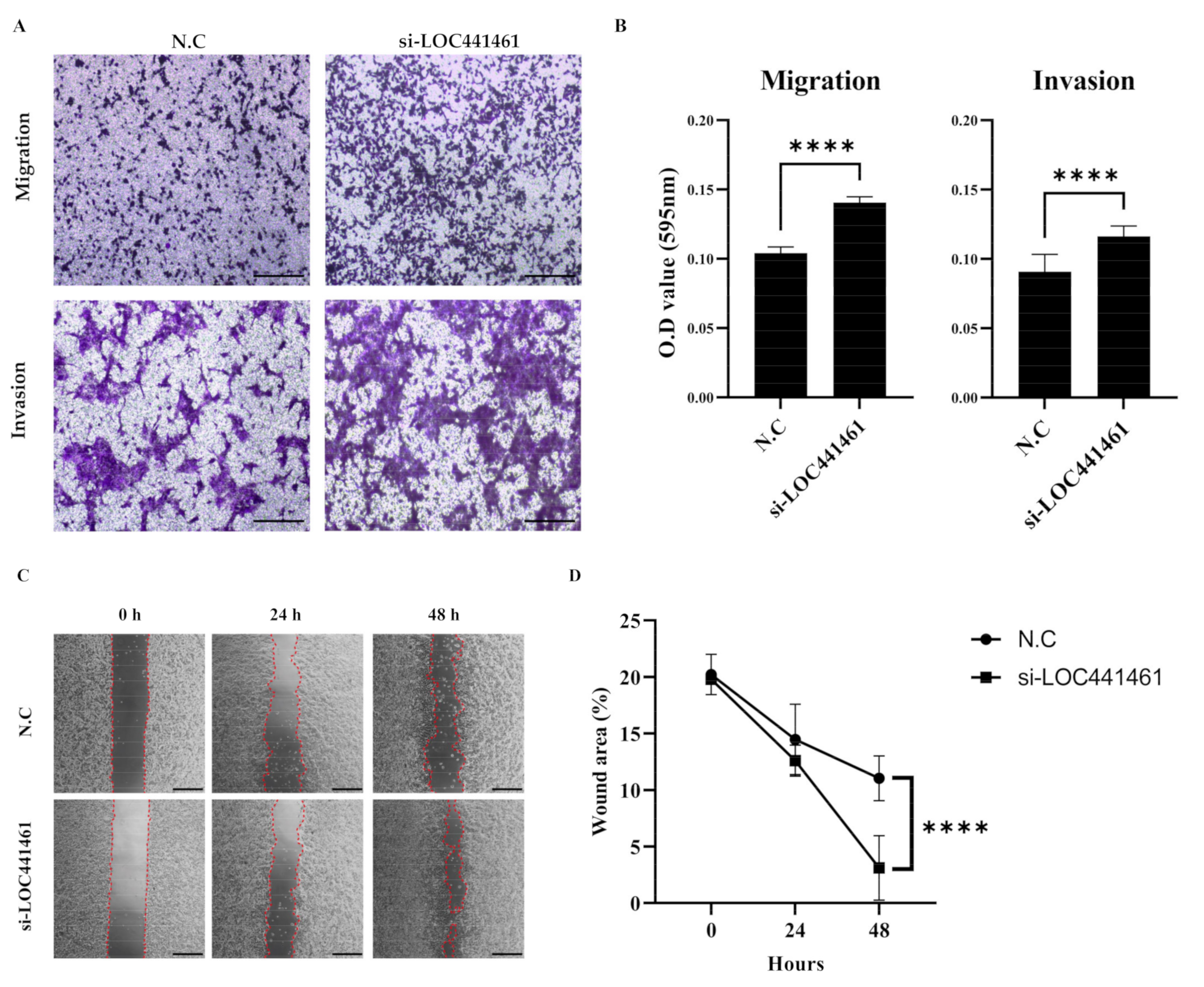
3.6. LOC441461 Knockdown Increases Gastric Cancer Cell Motility and Invasiveness
3.7. Effect of LOC441461 Knockdown on the Transcriptomic Landscape of Gastric Cancer Cells
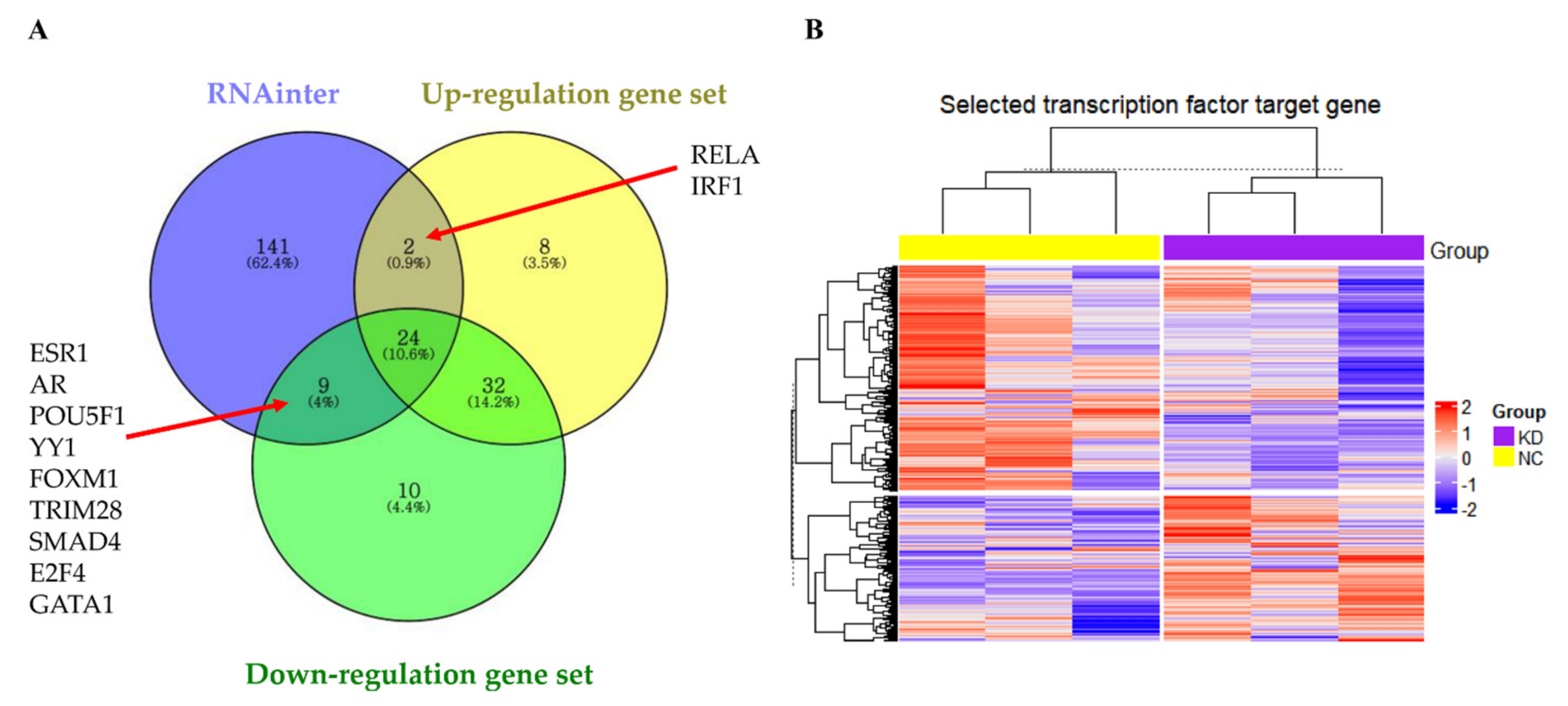
3.8. LOC441461 Changes the Nature of Gastric Cancer Cells by Modulating Transcription Factor Activity
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Smyth, E.C.; Nilsson, M.; Grabsch, H.I.; van Grieken, N.C.; Lordick, F. Gastric cancer. Lancet 2020, 396, 635–648. [Google Scholar] [CrossRef]
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Byrd, D.R.; Carducci, M.; Compton, C.; Fritz, A.; Greene, F. AJCC Cancer Staging Manual; Springer: New York, NY, USA, 2010. [Google Scholar]
- Yao, K.; Uedo, N.; Kamada, T.; Hirasawa, T.; Nagahama, T.; Yoshinaga, S.; Oka, M.; Inoue, K.; Mabe, K.; Yao, T.; et al. Guidelines for endoscopic diagnosis of early gastric cancer. Dig. Endosc. 2020, 32, 663–698. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Gong, Y.; Xu, H. Clinical and pathological staging of gastric cancer: Current perspectives and implications. Eur. J. Surg. Oncol. 2020, 46, e14–e19. [Google Scholar] [CrossRef] [PubMed]
- Roland, N.; Porter, G.; Fish, B.; Makura, Z. Tumour assessment and staging: United Kingdom national multidisciplinary guidelines. J. Laryngol. Otol. 2016, 130, S53–S58. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sano, T.; Coit, D.G.; Kim, H.H.; Roviello, F.; Kassab, P.; Wittekind, C.; Yamamoto, Y.; Ohashi, Y. Proposal of a new stage grouping of gastric cancer for TNM classification: International Gastric Cancer Association staging project. Gastric Cancer 2017, 20, 217–225. [Google Scholar] [CrossRef] [PubMed]
- Wittekind, C. The development of the TNM classification of gastric cancer. Pathol. Int. 2015, 65, 399–403. [Google Scholar] [CrossRef]
- Joshi, S.S.; Badgwell, B.D. Current treatment and recent progress in gastric cancer. CA Cancer J. Clin. 2021, 71, 264–279. [Google Scholar] [CrossRef]
- NCCN. Stomach Patient (NCCN Guidelines); NCCN: Bethesda, MA, USA, 2021. [Google Scholar]
- Yao, R.-W.; Wang, Y.; Chen, L.-L. Cellular functions of long noncoding RNAs. Nat. Cell Biol. 2019, 21, 542–551. [Google Scholar] [CrossRef]
- Zhang, X.; Wang, W.; Zhu, W.; Dong, J.; Cheng, Y.; Yin, Z.; Shen, F. Mechanisms and functions of long non-coding RNAs at multiple regulatory levels. Int. J. Mol. Sci. 2019, 20, 5573. [Google Scholar] [CrossRef]
- Zhu, J.; Fu, H.; Wu, Y.; Zheng, X. Function of lncRNAs and approaches to lncRNA-protein interactions. Sci. China Life Sci. 2013, 56, 876–885. [Google Scholar] [CrossRef] [PubMed]
- Gao, N.; Li, Y.; Li, J.; Gao, Z.; Yang, Z.; Li, Y.; Liu, H.; Fan, T. Long non-coding RNAs: The regulatory mechanisms, research strategies, and future directions in cancers. Front. Oncol. 2020, 10, 2903. [Google Scholar] [CrossRef]
- Gil, N.; Ulitsky, I. Regulation of gene expression by cis-acting long non-coding RNAs. Nat. Rev. Genet. 2020, 21, 102–117. [Google Scholar] [CrossRef] [PubMed]
- Statello, L.; Guo, C.-J.; Chen, L.-L.; Huarte, M. Gene regulation by long non-coding RNAs and its biological functions. Nat. Rev. Mol. Cell Biol. 2021, 22, 96–118. [Google Scholar] [CrossRef]
- Romero-Barrios, N.; Legascue, M.F.; Benhamed, M.; Ariel, F.; Crespi, M. Splicing regulation by long noncoding RNAs. Nucleic Acids Res. 2018, 46, 2169–2184. [Google Scholar] [CrossRef] [PubMed]
- Jiang, M.-C.; Ni, J.-J.; Cui, W.-Y.; Wang, B.-Y.; Zhuo, W. Emerging roles of lncRNA in cancer and therapeutic opportunities. Am. J. Cancer Res. 2019, 9, 1354. [Google Scholar]
- Mathieu, E.-L.; Belhocine, M.; Dao, L.; Puthier, D.; Spicuglia, S. Functions of lncRNA in development and diseases. Med. Sci. 2014, 30, 790–796. [Google Scholar]
- Loewen, G.; Jayawickramarajah, J.; Zhuo, Y.; Shan, B. Functions of lncRNA HOTAIR in lung cancer. J. Hematol. Oncol. 2014, 7, 1–10. [Google Scholar] [CrossRef]
- Ren, S.; Liu, Y.; Xu, W.; Sun, Y.; Lu, J.; Wang, F.; Wei, M.; Shen, J.; Hou, J.; Gao, X. Long noncoding RNA MALAT-1 is a new potential therapeutic target for castration resistant prostate cancer. J. Urol. 2013, 190, 2278–2287. [Google Scholar] [CrossRef]
- Liu, Z.-Q.; He, W.-F.; Wu, Y.-J.; Zhao, S.-L.; Wang, L.; Ouyang, Y.-Y.; Tang, S.-Y. LncRNA SNHG1 promotes EMT process in gastric cancer cells through regulation of the miR-15b/DCLK1/Notch1 axis. BMC Gastroenterol. 2020, 20. [Google Scholar] [CrossRef]
- Cheng, T.; Wang, Y.; Bryant, S.H. FSelector: A Ruby gem for feature selection. Bioinformatics 2012, 28, 2851–2852. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Romanski, P.; Kotthoff, L.; Kotthoff, M.L. Package ‘FSelector’. 2013. Available online: http://cran.r-project.org/web/packages/FSelector/index.html (accessed on 18 November 2021).
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Ge, S.X.; Jung, D.; Yao, R. ShinyGO: A graphical gene-set enrichment tool for animals and plants. Bioinformatics 2020, 36, 2628–2629. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.; Tang, Q.; He, J.; Li, L.; Yang, N.; Yu, S.; Wang, M.; Zhang, Y.; Lin, J.; Cui, T. RNAInter v4. 0: RNA interactome repository with redefined confidence scoring system and improved accessibility. Nucleic Acids Res. 2022, 50, D326–D332. [Google Scholar] [CrossRef]
- Consortium, E.P. An integrated encyclopedia of DNA elements in the human genome. Nature 2012, 489, 57. [Google Scholar] [CrossRef]
- Lachmann, A.; Xu, H.; Krishnan, J.; Berger, S.I.; Mazloom, A.R.; Ma’ayan, A. ChEA: Transcription factor regulation inferred from integrating genome-wide ChIP-X experiments. Bioinformatics 2010, 26, 2438–2444. [Google Scholar] [CrossRef]
- Gu, Z.; Eils, R.; Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 2016, 32, 2847–2849. [Google Scholar] [CrossRef]
- Cui, H.; Jiang, Z.; Zeng, S.; Wu, H.; Zhang, Z.; Guo, X.; Dong, K.; Wang, J.; Shang, L.; Li, L. A new candidate oncogenic lncRNA derived from pseudogene WFDC21P promotes tumor progression in gastric cancer. Cell Death Dis. 2021, 12, 903. [Google Scholar] [CrossRef]
- Zhao, Z.; Liu, H.; Zhou, X.; Fang, D.; Ou, X.; Ye, J.; Peng, J.; Xu, J. Necroptosis-Related lncRNAs: Predicting Prognosis and the Distinction between the Cold and Hot Tumors in Gastric Cancer. J. Oncol. 2021, 2021, 1–16. [Google Scholar] [CrossRef]
- Winkle, M.; El-Daly, S.M.; Fabbri, M.; Calin, G.A. Noncoding RNA therapeutics—Challenges and potential solutions. Nat. Rev. Drug Discov. 2021, 20, 629–651. [Google Scholar] [CrossRef]
- Bridges, M.C.; Daulagala, A.C.; Kourtidis, A. LNCcation: lncRNA localization and function. J. Cell Biol. 2021, 220, e202009045. [Google Scholar] [CrossRef]
- Ma, X.; Wang, G.; Fan, H.; Li, Z.; Chen, W.; Xiao, J.; Ni, P.; Liu, K.; Shen, K.; Wang, Y.; et al. Long noncoding RNA FAM225A promotes the malignant progression of gastric cancer through the miR-326/PADI2 axis. Cell Death Discov. 2022, 8, 20. [Google Scholar] [CrossRef] [PubMed]
- Ghafouri-Fard, S.; Shoorei, H.; Anamag, F.T.; Taheri, M. The role of non-coding RNAs in controlling cell cycle related proteins in cancer cells. Front. Oncol. 2020, 10, 2616. [Google Scholar] [CrossRef] [PubMed]
- Kitagawa, M.; Kitagawa, K.; Kotake, Y.; Niida, H.; Ohhata, T. Cell cycle regulation by long non-coding RNAs. Cell. Mol. Life Sci. 2013, 70, 4785–4794. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.-Y.; Tang, J.-N.; Xie, H.-X.; Du, Y.-A.; Huang, L.; Yu, P.-F.; Cheng, X.-D. 5-Fluorouracil chemotherapy of gastric cancer generates residual cells with properties of cancer stem cells. Int. J. Biol. Sci. 2015, 11, 284. [Google Scholar] [CrossRef]
- Sethy, C.; Kundu, C.N. 5-Fluorouracil (5-FU) resistance and the new strategy to enhance the sensitivity against cancer: Implication of DNA repair inhibition. Biomed. Pharmacother. 2021, 137, 111285. [Google Scholar] [CrossRef]
- Ba, M.-C.; Ba, Z.; Long, H.; Cui, S.-Z.; Gong, Y.-F.; Yan, Z.-F.; Lin, K.-P.; Wu, Y.-B.; Tu, Y.-N. LncRNA AC093818.1 accelerates gastric cancer metastasis by epigenetically promoting PDK1 expression. Cell Death Dis. 2020, 11, 64. [Google Scholar] [CrossRef]
- Bakir, B.; Chiarella, A.M.; Pitarresi, J.R.; Rustgi, A.K. EMT, MET, Plasticity, and Tumor Metastasis. Trends Cell Biol. 2020, 30, 764–776. [Google Scholar] [CrossRef]
- Peyre, L.; Meyer, M.; Hofman, P.; Roux, J. TRAIL receptor-induced features of epithelial-to-mesenchymal transition increase tumour phenotypic heterogeneity: Potential cell survival mechanisms. Br. J. Cancer 2021, 124, 91–101. [Google Scholar] [CrossRef]
- Zhang, H.; Qin, G.; Zhang, C.; Yang, H.; Liu, J.; Hu, H.; Wu, P.; Liu, S.; Yang, L.; Chen, X. TRAIL promotes epithelial-to-mesenchymal transition by inducing PD-L1 expression in esophageal squamous cell carcinomas. J. Exp. Clin. Cancer Res. 2021, 40, 1–16. [Google Scholar] [CrossRef]
- Trivedi, R.; Mishra, D.P. Trailing TRAIL resistance: Novel targets for TRAIL sensitization in cancer cells. Front. Oncol. 2015, 5, 69. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Xu, C.; Kong, X.; Li, X.; Kong, X.; Wang, Y.; Ding, X.; Yang, Q. Trail resistance induces epithelial-mesenchymal transition and enhances invasiveness by suppressing PTEN via miR-221 in breast cancer. PLoS ONE 2014, 9, e99067. [Google Scholar] [CrossRef]
- Hong, D.; Messier, T.L.; Tye, C.E.; Dobson, J.R.; Fritz, A.J.; Sikora, K.R.; Browne, G.; Stein, J.L.; Lian, J.B.; Stein, G.S. Runx1 stabilizes the mammary epithelial cell phenotype and prevents epithelial to mesenchymal transition. Oncotarget 2017, 8, 17610. [Google Scholar] [CrossRef] [PubMed]
- Hong, D. RUNX1 Control of Mammary Epithelial and Breast Cancer Cell Phenotypes; University of Massachusetts Medical School: Worcester, MA, USA, 2017. [Google Scholar]
- Marchese, F.P.; Raimondi, I.; Huarte, M. The multidimensional mechanisms of long noncoding RNA function. Genome Biol. 2017, 18, 206. [Google Scholar] [CrossRef] [PubMed]
- Chanut, A.; Duguet, F.; Marfak, A.; David, A.; Petit, B.; Parrens, M.; Durand-Panteix, S.; Boulin-Deveza, M.; Gachard, N.; Youlyouz-Marfak, I. RelA and RelB cross-talk and function in Epstein–Barr virus transformed B cells. Leukemia 2014, 28, 871–879. [Google Scholar] [CrossRef]
- Ye, T.; Yang, M.; Huang, D.; Wang, X.; Xue, B.; Tian, N.; Xu, X.; Bao, L.; Hu, H.; Lv, T. MicroRNA-7 as a potential therapeutic target for aberrant NF-κB-driven distant metastasis of gastric cancer. J. Exp. Clin. Cancer Res. 2019, 38, 1–18. [Google Scholar] [CrossRef]
- Kimura, T.; Nakayama, K.; Penninger, J.; Kitagawa, M.; Harada, H.; Matsuyama, T.; Tanaka, N.; Kamijo, R.; Vilček, J.; Mak, T.W. Involvement of the IRF-1 transcription factor in antiviral responses to interferons. Science 1994, 264, 1921–1924. [Google Scholar] [CrossRef]
- Nozawa, H.; Oda, E.; Ueda, S.; Tamura, G.; Maesawa, C.; Muto, T.; Taniguchi, T.; Tanaka, N. Functionally inactivating point mutation in the tumor-suppressor IRF-1 gene identified in human gastric cancer. Int. J. Cancer 1998, 77, 522–527. [Google Scholar] [CrossRef]
- Gao, J.; Tian, Y.; Zhang, J. Overexpression of interferon regulatory factor 1 enhances chemosensitivity to 5-fluorouracil in gastric cancer cells. J. Cancer Res. Ther. 2012, 8, 57. [Google Scholar] [CrossRef]
- Jiang, L.; Liu, J.-y.; Shi, Y.; Tang, B.; He, T.; Liu, J.-j.; Fan, J.-y.; Wu, B.; Xu, X.-h.; Zhao, Y.-l. MTMR2 promotes invasion and metastasis of gastric cancer via inactivating IFNγ/STAT1 signaling. J. Exp. Clin. Cancer Res. 2019, 38, 1–16. [Google Scholar] [CrossRef]
- Chang, W.-C.; Huang, S.-F.; Lee, Y.-M.; Lai, H.-C.; Cheng, B.-H.; Cheng, W.-C.; Ho, J.Y.-P.; Jeng, L.-B.; Ma, W.-L. Cholesterol import and steroidogenesis are biosignatures for gastric cancer patient survival. Oncotarget 2017, 8, 692. [Google Scholar] [CrossRef] [PubMed]
- Saigusa, S.; Tanaka, K.; Toiyama, Y.; Yokoe, T.; Okugawa, Y.; Ioue, Y.; Miki, C.; Kusunoki, M. Correlation of CD133, OCT4, and SOX2 in rectal cancer and their association with distant recurrence after chemoradiotherapy. Ann. Surg. Oncol. 2009, 16, 3488–3498. [Google Scholar] [CrossRef] [PubMed]
- Bauer, L.; Langer, R.; Becker, K.; Hapfelmeier, A.; Ott, K.; Novotny, A.; Höfler, H.; Keller, G. Expression profiling of stem cell-related genes in neoadjuvant-treated gastric cancer: A NOTCH2, GSK3B and β-catenin gene signature predicts survival. PLoS ONE 2012, 7, e44566. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Peng, Y.; Zhang, M.; Jiang, Z.; Jiang, Y. TRIM28 activates autophagy and promotes cell proliferation in glioblastoma. OncoTargets Ther. 2019, 12, 397. [Google Scholar] [CrossRef] [PubMed]
- Su, C.; Li, H.; Gao, W. TRIM28 is overexpressed in glioma and associated with tumor progression. OncoTargets Ther. 2018, 11, 6447. [Google Scholar] [CrossRef]
- Li, Y.; Ke, Q.; Shao, Y.; Zhu, G.; Li, Y.; Geng, N.; Jin, F.; Li, F. GATA1 induces epithelial-mesenchymal transition in breast cancer cells through PAK5 oncogenic signaling. Oncotarget 2015, 6, 4345. [Google Scholar] [CrossRef]
- Ouranidis, A.; Vavilis, T.; Mandala, E.; Davidopoulou, C.; Stamoula, E.; Markopoulou, C.K.; Karagianni, A.; Kachrimanis, K. mRNA Therapeutic Modalities Design, Formulation and Manufacturing under Pharma 4.0 Principles. Biomedicines 2021, 10, 50. [Google Scholar] [CrossRef]


| Gene | IG | p |
|---|---|---|
| NOV | 0.1150726 | 9.48 × 10−6 |
| CPA3 | 0.1117299 | 4.84 × 10−5 |
| CARD11 | 0.1049524 | 5.29 ×10−5 |
| ZNF467 | 0.1327168 | 7.48 × 10−5 |
| AK097446 | 0.1205477 | 7.52 × 10−5 |
| LOC441461 | 0.1223392 | 8.80× 10−5 |
| TPSAB1 | 0.1262245 | 0.0001181 |
| TNM | Odds Ratio | CI—Low | CI—High | p |
|---|---|---|---|---|
| T1 vs. T2 | 0.30 | 0.09 | 0.88 | 0.086 |
| T1 vs. T3 | 0.46 | 0.20 | 1.11 | 0.141 |
| T1 vs. T4 | 0.1 | 0.02 | 0.32 | 0.001 * |
| T2 vs. T3 | 1.21 | 0.70 | 2.14 | 0.677 |
| T2 vs. T4 | 0.42 | 0.20 | 0.87 | 0.015 * |
| T3 vs. T4 | 0.38 | 0.21 | 0.67 | 0.002 * |
| N0 vs. N1 | 0.84 | 0.48 | 1.45 | 0.68 |
| N0 vs. N2 | 0.90 | 0.50 | 1.60 | 0.777 |
| N0 vs. N3 | 0.58 | 0.30 | 1.06 | 0.047 |
| N1 vs. N2 | 1.08 | 0.56 | 2.08 | 0.508 |
| N1 vs. N3 | 0.65 | 0.32 | 1.29 | 0.089 |
| N2 vs. N3 | 0.63 | 0.30 | 1.26 | 0.037 |
| M0 vs. M1 | 0.08 | 0.02 | 0.3 | 0.001 * |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, S.-s.; Park, J.; Oh, S.; Kwack, K. Downregulation of LOC441461 Promotes Cell Growth and Motility in Human Gastric Cancer. Cancers 2022, 14, 1149. https://doi.org/10.3390/cancers14051149
Lee S-s, Park J, Oh S, Kwack K. Downregulation of LOC441461 Promotes Cell Growth and Motility in Human Gastric Cancer. Cancers. 2022; 14(5):1149. https://doi.org/10.3390/cancers14051149
Chicago/Turabian StyleLee, Sang-soo, JeongMan Park, Sooyeon Oh, and KyuBum Kwack. 2022. "Downregulation of LOC441461 Promotes Cell Growth and Motility in Human Gastric Cancer" Cancers 14, no. 5: 1149. https://doi.org/10.3390/cancers14051149
APA StyleLee, S.-s., Park, J., Oh, S., & Kwack, K. (2022). Downregulation of LOC441461 Promotes Cell Growth and Motility in Human Gastric Cancer. Cancers, 14(5), 1149. https://doi.org/10.3390/cancers14051149







