Analysis of the Tumor Immune Microenvironment (TIME) in Clear Cell Renal Cell Carcinoma (ccRCC) Reveals an M0 Macrophage-Enriched Subtype: An Exploration of Prognostic and Biological Characteristics of This Immune Phenotype
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Discovery Dataset
2.2. Clustering Based on Immune Cell Subpopulations
2.3. Survival Analysis by Clusters
2.4. Validation Dataset
2.5. Tumor Immune Characterization Using TIDE
2.6. Gene Set Enrichment Analysis (GSEA)
2.7. Multivariable Cox Hazards Analyses
3. Results
3.1. Cohort Description
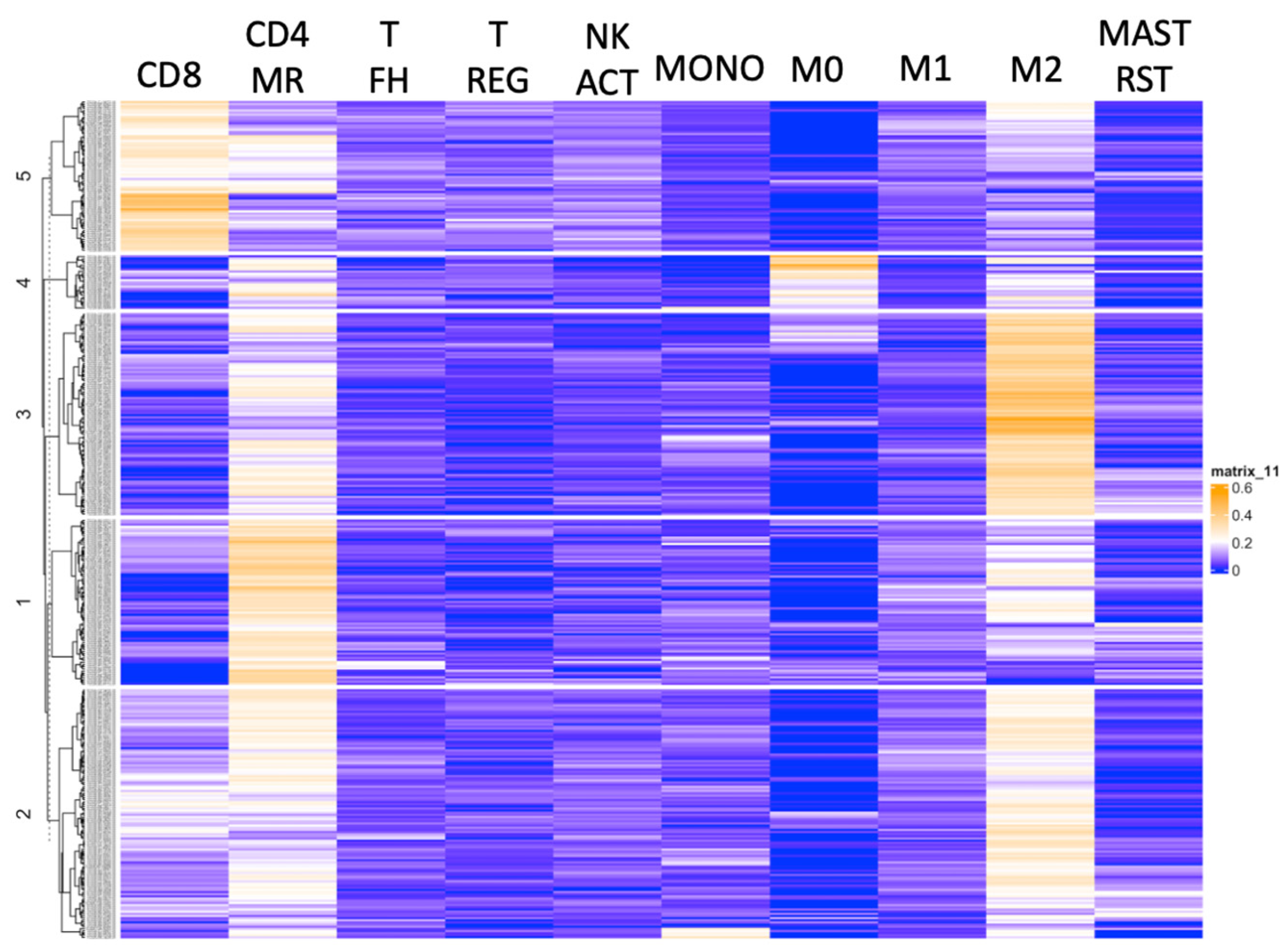
3.2. Hierarchical Clustering Based on the Immune Cell Subsets Identified Macrophage-Enriched Clusters
3.3. M0 Macrophage Enrichment Is Associated with Shorter PFS in Two Separate Cohorts
3.4. M0 Macrophage Enrichment Is Also Associated with Other Pro-Tumorigenic TME Characteristics
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer Statistics, 2021. CA Cancer J. Clin. 2021, 71, 7–33. [Google Scholar] [CrossRef] [PubMed]
- Ljungberg, B.; Campbell, S.C.; Cho, H.Y.; Jacqmin, D.; Lee, J.E.; Weikert, S.; Kiemeney, L.A. The epidemiology of renal cell carcinoma. Eur. Urol. 2011, 60, 615–621. [Google Scholar] [CrossRef] [PubMed]
- Shuch, B.; Amin, A.; Armstrong, A.J.; Eble, J.N.; Ficarra, V.; Lopez-Beltran, A.; Martignoni, G.; Rini, B.I.; Kutikov, A. Understanding pathologic variants of renal cell carcinoma: Distilling therapeutic opportunities from biologic complexity. Eur. Urol. 2015, 67, 85–97. [Google Scholar] [CrossRef] [PubMed]
- Dabestani, S.; Thorstenson, A.; Lindblad, P.; Harmenberg, U.; Ljungberg, B.; Lundstam, S. Renal cell carcinoma recurrences and metastases in primary non-metastatic patients: A population-based study. World J. Urol. 2016, 34, 1081–1086. [Google Scholar] [CrossRef] [PubMed]
- Marconi, L.; Sun, M.; Beisland, C.; Klatte, T.; Ljungberg, B.; Stewart, G.D.; Dabestani, S.; Choueiri, T.K.; Bex, A. Prevalence, Disease-free, and Overall Survival of Contemporary Patients With Renal Cell Carcinoma Eligible for Adjuvant Checkpoint Inhibitor Trials. Clin. Genitourin. Cancer 2021, 19, e92–e99. [Google Scholar] [CrossRef]
- Eggener, S.E.; Yossepowitch, O.; Pettus, J.A.; Snyder, M.E.; Motzer, R.J.; Russo, P. Renal cell carcinoma recurrence after nephrectomy for localized disease: Predicting survival from time of recurrence. J. Clin. Oncol. 2006, 24, 3101–3106. [Google Scholar] [CrossRef]
- Levy, D.A.; Slaton, J.W.; Swanson, D.A.; Dinney, C.P. Stage specific guidelines for surveillance after radical nephrectomy for local renal cell carcinoma. J. Urol. 1998, 159, 1163–1167. [Google Scholar] [CrossRef]
- Şenbabaoğlu, Y.; Gejman, R.S.; Winer, A.G.; Liu, M.; Van Allen, E.M.; de Velasco, G.; Miao, D.; Ostrovnaya, I.; Drill, E.; Luna, A.; et al. Tumor immune microenvironment characterization in clear cell renal cell carcinoma identifies prognostic and immunotherapeutically relevant messenger RNA signatures. Genome Biol. 2016, 17, 231. [Google Scholar] [CrossRef]
- Choueiri, T.K.; Tomczak, P.; Park, S.H.; Venugopal, B.; Ferguson, T.; Chang, Y.-H.; Hajek, J.; Symeonides, S.N.; Lee, J.L.; Sarwar, N.; et al. Adjuvant Pembrolizumab after Nephrectomy in Renal-Cell Carcinoma. N. Engl. J. Med. 2021, 385, 683–694. [Google Scholar] [CrossRef]
- Pal, S.K.; Uzzo, R.; Karam, J.A.; A Master, V.; Donskov, F.; Suarez, C.; Albiges, L.; Rini, B.; Tomita, Y.; Kann, A.G.; et al. Adjuvant atezolizumab versus placebo for patients with renal cell carcinoma at increased risk of recurrence following resection (IMmotion010): A multicentre, randomised, double-blind, phase 3 trial. Lancet 2022, 400, 1103–1116. [Google Scholar] [CrossRef]
- Rappold, P.M.; Silagy, A.W.; Kotecha, R.R.; Hakimi, A.A. Immune checkpoint blockade in renal cell carcinoma. J. Surg. Oncol. 2021, 123, 739–750. [Google Scholar] [CrossRef] [PubMed]
- Denize, T.; Hou, Y.; Pignon, J.C.; Walton, E.; West, D.J.; Freeman, G.J.; Braun, D.A.; Wu, C.J.; Gupta, S.; Motzer, R.J.; et al. Transcriptomic Correlates of Tumor Cell PD-L1 Expression and Response to Nivolumab Monotherapy in Metastatic Clear Cell Renal Cell Carcinoma. Clin. Cancer Res. 2022, 28, 4045–4055. [Google Scholar] [CrossRef] [PubMed]
- Newman, A.M.; Liu, C.L.; Green, M.R.; Gentles, A.J.; Feng, W.; Xu, Y.; Hoang, C.D.; Diehn, M.; Alizadeh, A.A. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 2015, 12, 453–457. [Google Scholar] [CrossRef] [PubMed]
- Ricketts, C.J.; De Cubas, A.A.; Fan, H.; Smith, C.C.; Lang, M.; Reznik, E.; Bowlby, R.; Gibb, E.A.; Akbani, R.; Beroukhim, R.; et al. The Cancer Genome Atlas Comprehensive Molecular Characterization of Renal Cell Carcinoma. Cell Rep. 2018, 23, 313–326.e5. [Google Scholar] [CrossRef]
- Cerami, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012, 2, 401–404. [Google Scholar] [CrossRef]
- Colaprico, A.; Silva, T.C.; Olsen, C.; Garofano, L.; Cava, C.; Garolini, D.; Sabedot, T.S.; Malta, T.M.; Pagnotta, S.M.; Castiglioni, I.; et al. TCGAbiolinks: An R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Res. 2016, 44, e71. [Google Scholar] [CrossRef]
- Silva, T.C.; Colaprico, A.; Olsen, C.; D’Angelo, F.; Bontempi, G.; Ceccarelli, M.; Noushmehr, H. TCGA Workflow: Analyze cancer genomics and epigenomics data using Bioconductor packages. F1000Research 2016, 5, 1542. [Google Scholar] [CrossRef]
- Mounir, M.; Lucchetta, M.; Silva, T.C.; Olsen, C.; Bontempi, G.; Chen, X.; Noushmehr, H.; Colaprico, A.; Papaleo, E. New functionalities in the TCGAbiolinks package for the study and integration of cancer data from GDC and GTEx. PLoS Comput. Biol. 2019, 15, e1006701. [Google Scholar] [CrossRef]
- Chen, B.; Khodadoust, M.S.; Liu, C.L.; Newman, A.M.; Alizadeh, A.A. Profiling Tumor Infiltrating Immune Cells with CIBERSORT. Methods Mol. Biol. 2018, 1711, 243–259. [Google Scholar]
- Braun, D.A.; Hou, Y.; Bakouny, Z.; Ficial, M.; Sant’ Angelo, M.; Forman, J.; Ross-Macdonald, P.; Berger, A.C.; Jegede, O.A.; Elagina, L.; et al. Interplay of somatic alterations and immune infiltration modulates response to PD-1 blockade in advanced clear cell renal cell carcinoma. Nat. Med. 2020, 26, 909–918. [Google Scholar] [CrossRef]
- Gentles, A.J.; Newman, A.M.; Liu, C.L.; Bratman, S.V.; Feng, W.; Kim, D.; Nair, V.S.; Xu, Y.; Khuong, A.; Hoang, C.D.; et al. The prognostic landscape of genes and infiltrating immune cells across human cancers. Nat. Med. 2015, 21, 938–945. [Google Scholar] [CrossRef] [PubMed]
- Gu, Z.; Eils, R.; Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 2016, 32, 2847–2849. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Lichtenberg, T.M.; Hoadley, K.A.; Poisson, L.M.; Lazar, A.J.; Cherniack, A.D.; Kovatich, A.J.; Benz, C.C.; Levine, D.A.; Lee, A.V.; et al. An Integrated TCGA Pan-Cancer Clinical Data Resource to Drive High-Quality Survival Outcome Analytics. Cell 2018, 173, 400–416.e11. [Google Scholar] [CrossRef] [PubMed]
- Therneau, T. A Package for Survival Analysis in S, R package version; Mayo Foundation: Rochester, MN, USA, 2023. [Google Scholar]
- Clark, D.J.; Dhanasekaran, S.M.; Petralia, F.; Pan, J.; Song, X.; Hu, Y.; da Veiga Leprevost, F.; Reva, B.; Lih, T.-S.M.; Chang, H.-Y.; et al. Integrated Proteogenomic Characterization of Clear Cell Renal Cell Carcinoma. Cell 2019, 179, 964–983.e31. [Google Scholar] [CrossRef]
- Jiang, P.; Gu, S.; Pan, D.; Fu, J.; Sahu, A.; Hu, X.; Li, Z.; Traugh, N.; Bu, X.; Li, B.; et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat. Med. 2018, 24, 1550–1558. [Google Scholar] [CrossRef]
- Reyes, A.L.P.; Silva, T.C.; Coetzee, S.G.; Plummer, J.T.; Davis, B.D.; Chen, S.; Hazelett, D.J.; Lawrenson, K.; Berman, B.P.; Gayther, S.A.; et al. GENAVi: A shiny web application for gene expression normalization, analysis and visualization. BMC Genom. 2019, 20, 745. [Google Scholar] [CrossRef]
- Liberzon, A.; Birger, C.; Thorvaldsdóttir, H.; Ghandi, M.; Mesirov, J.P.; Tamayo, P. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 2015, 1, 417–425. [Google Scholar] [CrossRef]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016. [Google Scholar]
- Solinas, G.; Germano, G.; Mantovani, A.; Allavena, P. Tumor-associated macrophages (TAM) as major players of the cancer-related inflammation. J. Leukoc. Biol. 2009, 86, 1065–1073. [Google Scholar] [CrossRef]
- Mantovani, A.; Sica, A. Macrophages, innate immunity and cancer: Balance, tolerance, and diversity. Curr. Opin. Immunol. 2010, 22, 231–237. [Google Scholar] [CrossRef]
- Chevrier, S.; Levine, J.H.; Zanotelli, V.R.T.; Silina, K.; Schulz, D.; Bacac, M.; Ries, C.H.; Ailles, L.; Jewett, M.A.S.; Moch, H.; et al. An Immune Atlas of Clear Cell Renal Cell Carcinoma. Cell 2017, 169, 736–749.e18. [Google Scholar] [CrossRef]
- Zhao, Y.L.; Tian, P.X.; Han, F.; Zheng, J.; Xia, X.X.; Xue, W.J.; Ding, X.M.; Ding, C.G. Comparison of the characteristics of macrophages derived from murine spleen, peritoneal cavity, and bone marrow. J. Zhejiang Univ. Sci. B 2017, 18, 1055–1063. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.; Chen, Z.; Zhong, Q.; Zheng, Z.; Chen, Y.; Shangguan, W.; Zhang, Y.; Yang, J.; Zhu, D.; Xie, W. M2 macrophages secrete CXCL13 to promote renal cell carcinoma migration, invasion, and EMT. Cancer Cell Int. 2021, 21, 677. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Li, H.; Mao, Y.; Wang, X.; Zhang, X.; Yu, X.; Tian, J.; Lei, Z.; Li, C.; Han, Q.; et al. Apoptotic SKOV3 cells stimulate M0 macrophages to differentiate into M2 macrophages and promote the proliferation and migration of ovarian cancer cells by activating the ERK signaling pathway. Int. J. Mol. Med. 2020, 45, 10–22. [Google Scholar] [CrossRef] [PubMed]
- Gabrusiewicz, K.; Rodriguez, B.; Wei, J.; Hashimoto, Y.; Healy, L.M.; Maiti, S.N.; Thomas, G.; Zhou, S.; Wang, Q.; Elakkad, A.; et al. Glioblastoma-infiltrated innate immune cells resemble M0 macrophage phenotype. JCI Insight 2016, 1, e85841. [Google Scholar] [CrossRef]
- Kanno, T.; Kamba, T.; Yamasaki, T.; Shibasaki, N.; Saito, R.; Terada, N.; Toda, Y.; Mikami, Y.; Inoue, T.; Kanematsu, A.; et al. JunB promotes cell invasion and angiogenesis in VHL-defective renal cell carcinoma. Oncogene 2012, 31, 3098–3110. [Google Scholar] [CrossRef]
- Liu, N.; Wang, X.; Wu, H.; Lv, X.; Xie, H.; Guo, Z.; Wang, J.; Dou, G.; Zhang, C.; Sun, M. Computational study of effective matrix metalloproteinase 9 (MMP9) targeting natural inhibitors. Aging 2021, 13, 22867–22882. [Google Scholar] [CrossRef]
- Du, B.; Zhou, Y.; Yi, X.; Zhao, T.; Tang, C.; Shen, T.; Zhou, K.; Wei, H.; Xu, S.; Dong, J.; et al. Identification of Immune-Related Cells and Genes in Tumor Microenvironment of Clear Cell Renal Cell Carcinoma. Front. Oncol. 2020, 10, 1770. [Google Scholar] [CrossRef]
- Hua, X.; Chen, J.; Su, Y.; Liang, C. Identification of an immune-related risk signature for predicting prognosis in clear cell renal cell carcinoma. Aging 2020, 12, 2302–2332. [Google Scholar] [CrossRef]
- Li, R.; Ferdinand, J.R.; Loudon, K.W.; Bowyer, G.S.; Laidlaw, S.; Muyas, F.; Mamanova, L.; Neves, J.B.; Bolt, L.; Fasouli, E.S.; et al. Mapping single-cell transcriptomes in the intra-tumoral and associated territories of kidney cancer. Cancer Cell 2022, 40, 1583–1599.e10. [Google Scholar] [CrossRef]
- Li, X.; Wang, C.Y. From bulk, single-cell to spatial RNA sequencing. Int. J. Oral. Sci. 2021, 13, 36. [Google Scholar] [CrossRef]
- Rappold, P.M.; Vuong, L.; Leibold, J.; Chakiryan, N.H.; Curry, M.; Kuo, F.; Sabio, E.; Jiang, H.; Nixon, B.G.; Liu, M.; et al. A Targetable Myeloid Inflammatory State Governs Disease Recurrence in Clear-Cell Renal Cell Carcinoma. Cancer Discov. 2022, 12, 2308–2329. [Google Scholar] [CrossRef] [PubMed]





| Cluster 1 (n = 77) | Cluster 2 (n = 116) | Cluster 3 (n = 94) | Cluster 4 (n = 25) | Cluster 5 (n = 70) | |
|---|---|---|---|---|---|
| Median Age (years) | 60.9 | 61.8 | 59.4 | 62.4 | 60.2 |
| Gender (M/F, %) | 62/38 | 59/41 | 72/28 | 72/28 | 61/39 |
| Race (W/B/A/NA, %) | 96/4/0/0 | 95/3/1/2 | 93/3/2/2 | 96/0/4/0 | 96/1/3/0 |
| Laterality (R/L, %) | 61/39 | 53/47 | 53/47 | 60/40 | 46/54 |
| Mutation Count (Min, Median, Max) | 1, 48, 553 | 1, 48, 708 | 1, 48, 409 | 1, 48, 89 | 1, 48, 93 |
| Fraction Genome Altered (Mean %) | 15.9 | 13.3 | 16.1 | 23.4 | 15.9 |
| Lab Parameters (↑/↓/WNL/NA %) | |||||
| Serum Calcium | 3/44/23/30 | 1/41/27/32 | 0/49/17/34 | 0/60/16/24 | 1/43/26/30 |
| Hemoglobin | 1/52/32/14 | 2/46/39/14 | 1/45/33/21 | 0/56/36/8 | 0/56/39/6 |
| Platelet Count | 5/10/68/17 | 8/12/66/15 | 3/6/69/21 | 0/20/68/12 | 6/9/77/9 |
| White Blood Cell Count | 34/0/48/0 | 38/3/44/16 | 40/1/36/22 | 32/4/52/12 | 37/1/54/8 |
| Lymph Nodes + (%) | 0 | 8 | 16 | 10 | 23 |
| Grade (%) | |||||
| G1 | 5 | 3 | 0 | 4 | 1 |
| G2 | 57 | 46 | 54 | 32 | 37 |
| G3 | 25 | 41 | 39 | 52 | 46 |
| G4 | 13 | 9 | 5 | 12 | 16 |
| TNM Stage Group (%) | |||||
| I | 60 | 53 | 68 | 40 | 49 |
| II | 12 | 11 | 11 | 12 | 16 |
| III | 27 | 35 | 19 | 48 | 34 |
| IV | 1 | 0 | 1 | 0 | 1 |
| Pathologic T Stage (%) | |||||
| T1, T1a, T1b | 6, 26, 27 | 4, 32, 18 | 7, 38, 22 | 0, 20, 24 | 0, 20, 29 |
| T2, T2a, T2b | 10, 1, 0 | 7, 3, 1 | 11, 0, 0 | 12, 0, 0 | 14, 1, 1 |
| T3, T3a, T3b, T3c | 0, 17, 10, 0 | 1, 24, 8, 2 | 1, 17, 2, 0 | 0, 16, 28, 0 | 0, 21, 11, 0 |
| T4 | 1 | 0 | 1 | 0 | 1 |
| Mutations (% WT, MUT, NA) | |||||
| TP53 | 90/1/9 | 90/4/6 | 87/2/11 | 88/0/12 | 90/0/10 |
| VHL | 44/48/9 | 41/53/6 | 35/54/11 | 52/36/12 | 44/45/10 |
| PBRM1 | 56/35/9 | 62/32/6 | 59/31/11 | 52/36/12 | 74.16.10 |
| SETD2 | 75/16/9 | 87/7/6 | 82/7/11 | 76/12/12 | 84/6/10 |
| TCEB1 | 91/0/9 | 93/1/6 | 88/1/11 | 88/0/12 | 89/1/10 |
| Predicted ICB Response (%) | 27 | 23 | 20 | 4 | 34 |
| Cluster 1 (n = 9) | Cluster 2 (n = 9) | Cluster 3 (n = 25) | Cluster 4 (n = 17) | Cluster 5 (n = 28) | Cluster 6 (n = 11) | |
|---|---|---|---|---|---|---|
| Median Age (years) | 61.1 | 64.1 | 58.6 | 64.2 | 57.6 | 63.7 |
| Gender (M/F, %) | 78/22 | 78/22 | 84/16 | 65/35 | 64/36 | 73/27 |
| Race (W/B/A/NA, %) | 22/0/0/78 | 67/0/0/33 | 48/0/0/52 | 47/0/0/53 | 54/4/4 38 | 64/0/0/36 |
| BMI (Mean) | 25.3 | 28.3 | 33.2 | 32.4 | 31.9 | 30.8 |
| Tumor Site (LP, M, UP, OTH, %) | 11, 22, 33, 33 | 0, 11, 33, 56 | 12, 20, 32, 36 | 6, 35, 24, 35 | 14, 29, 29, 29 | 36, 18, 27, 18 |
| Tumor Size (Mean, cm) | 6.04 | 8.56 | 6.33 | 6.77 | 5.64 | 5.79 |
| Grade (%) | ||||||
| G1 | 0 | 0 | 0 | 24 | 7 | 0 |
| G2 | 89 | 22 | 52 | 35 | 57 | 45 |
| G3 | 11 | 44 | 44 | 41 | 29 | 55 |
| G4 | 0 | 33 | 4 | 0 | 7 | 0 |
| Pathologic Stage (%) | ||||||
| I | 44 | 33 | 40 | 53 | 50 | 55 |
| II | 11 | 11 | 16 | 18 | 7 | 18 |
| III | 22 | 22 | 40 | 24 | 32 | 27 |
| IV | 11 | 33 | 4 | 6 | 11 | 0 |
| Pathologic T Stage (%) | ||||||
| T1, T1a, T1b | 0, 22, 22 | 0, 11, 22 | 0, 20, 20 | 0, 18, 35 | 0, 39, 14 | 0, 36, 27 |
| T2, T2a, T2b | 0, 11, 0 | 0, 11, 11 | 0, 8, 8 | 0, 18, 0, 0 | 0, 4, 4 | 0, 9, 9, |
| T3, T3a, T3b, T3c | 11, 11, 0, 0 | 11, 11, 22, 0 | 4, 36, 4, 0 | 0, 18, 6, 0 | 4, 36, 0, 0 | 0, 18, 0 |
| T4 | 11 | 0 | 0 | 6 | 0 | 0 |
| Lab Parameters (↑/↓/WNL/NA %) | ||||||
| Serum Calcium | 0/0/11/89 | 0/0/56/44 | 0/8/16/76 | 0/12/18/70 | 4/4/18/75 | 0/18/0/82 |
| Hemoglobin (HgB) | 0/11/56/33 | 0/22/56/22 | 0/28/36/36 | 0/39/22/35 | 0/32/32/36 | 9/27/27/36 |
| Platelets (PLT) | 0/0/67/33 | 0/0/78/22 | 0/0/59/41 | 0/0/61/39 | ||
| White Blood Cells (WBC) | 0/0/67/33 | 0/0/78/22 | 0/0/64/36 | 6/0/59/35 | 11/4/50/35 | 0/9/55/36 |
| Laterality (L/R, %) | 78/22 | 56/44 | 40/60 | 29/71 | 46/54 | 73/27 |
| Margins Involved (%) | 22 | 22 | 4 | 0 | 0 | 9 |
| Residual Tumor | ||||||
| R0 | 44 | 67 | 52 | 59 | 43 | 36 |
| R1 | 0 | 0 | 4 | 0 | 0 | 0 |
| R2 | 0 | 0 | 0 | 0 | 0 | 0 |
| Rx | 56 | 33 | 44 | 41 | 57 | 64 |
| Pack Years (Mean) | 22.9 | 18.3 | 34 | 36.5 | 26.3 | 14.7 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Farha, M.; Nallandhighal, S.; Vince, R.; Cotta, B.; Stangl-Kremser, J.; Triner, D.; Morgan, T.M.; Palapattu, G.S.; Cieslik, M.; Vaishampayan, U.; et al. Analysis of the Tumor Immune Microenvironment (TIME) in Clear Cell Renal Cell Carcinoma (ccRCC) Reveals an M0 Macrophage-Enriched Subtype: An Exploration of Prognostic and Biological Characteristics of This Immune Phenotype. Cancers 2023, 15, 5530. https://doi.org/10.3390/cancers15235530
Farha M, Nallandhighal S, Vince R, Cotta B, Stangl-Kremser J, Triner D, Morgan TM, Palapattu GS, Cieslik M, Vaishampayan U, et al. Analysis of the Tumor Immune Microenvironment (TIME) in Clear Cell Renal Cell Carcinoma (ccRCC) Reveals an M0 Macrophage-Enriched Subtype: An Exploration of Prognostic and Biological Characteristics of This Immune Phenotype. Cancers. 2023; 15(23):5530. https://doi.org/10.3390/cancers15235530
Chicago/Turabian StyleFarha, Mark, Srinivas Nallandhighal, Randy Vince, Brittney Cotta, Judith Stangl-Kremser, Daniel Triner, Todd M. Morgan, Ganesh S. Palapattu, Marcin Cieslik, Ulka Vaishampayan, and et al. 2023. "Analysis of the Tumor Immune Microenvironment (TIME) in Clear Cell Renal Cell Carcinoma (ccRCC) Reveals an M0 Macrophage-Enriched Subtype: An Exploration of Prognostic and Biological Characteristics of This Immune Phenotype" Cancers 15, no. 23: 5530. https://doi.org/10.3390/cancers15235530
APA StyleFarha, M., Nallandhighal, S., Vince, R., Cotta, B., Stangl-Kremser, J., Triner, D., Morgan, T. M., Palapattu, G. S., Cieslik, M., Vaishampayan, U., Udager, A. M., & Salami, S. S. (2023). Analysis of the Tumor Immune Microenvironment (TIME) in Clear Cell Renal Cell Carcinoma (ccRCC) Reveals an M0 Macrophage-Enriched Subtype: An Exploration of Prognostic and Biological Characteristics of This Immune Phenotype. Cancers, 15(23), 5530. https://doi.org/10.3390/cancers15235530






