Atomic Force Microscopy: A Versatile Tool in Cancer Research
Simple Summary
Abstract
1. Introduction
2. Concise Overview of AFM
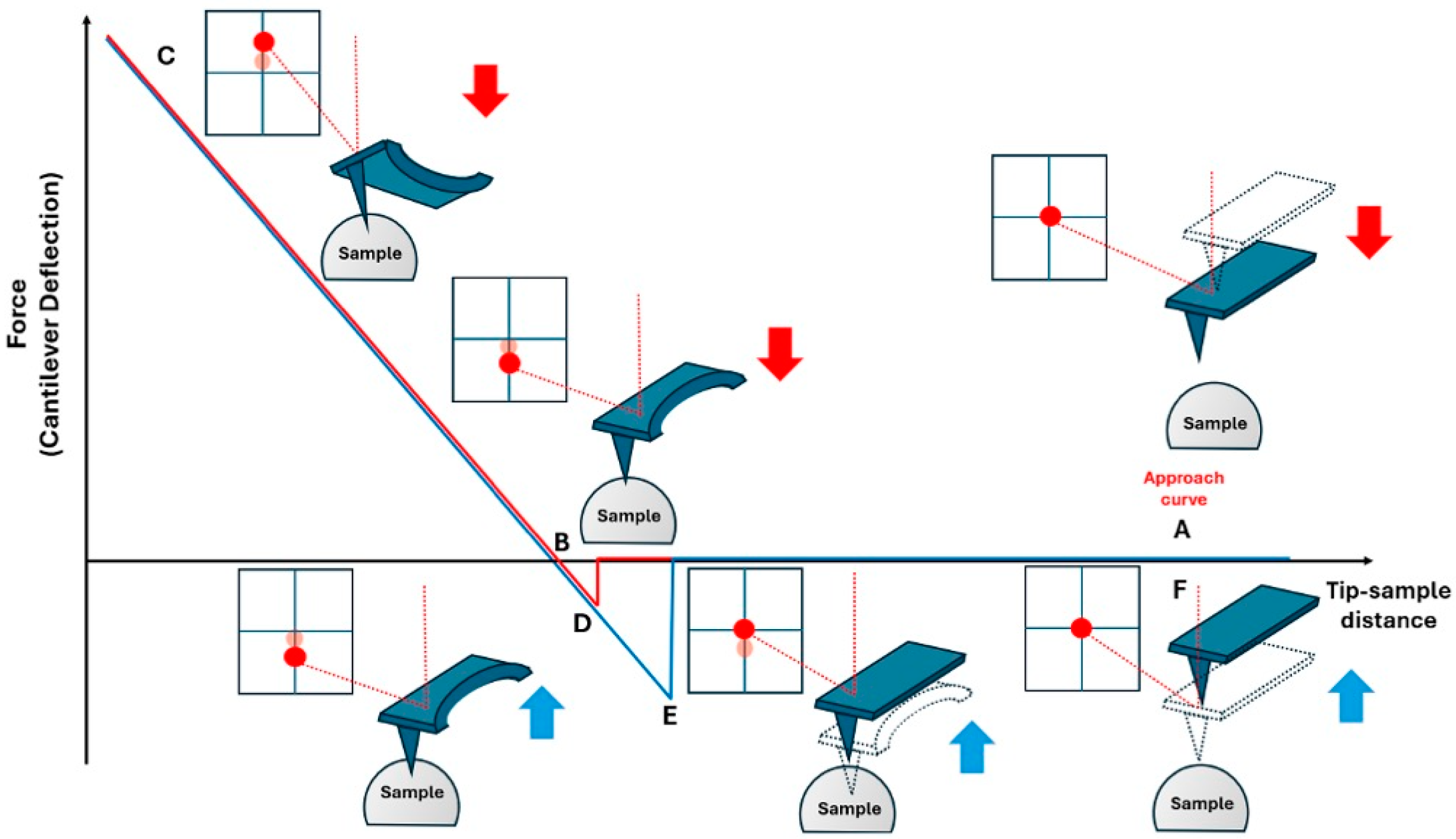
2.1. Principles of Operation
2.2. Topographic Techniques
2.3. Static and Dynamic Force Spectroscopy Characterization
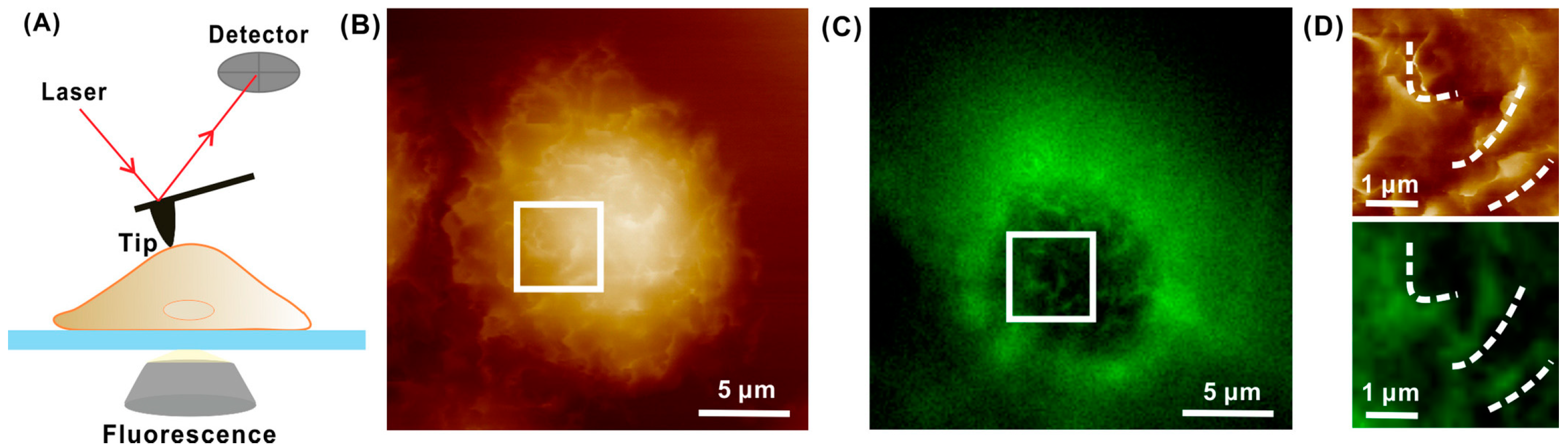
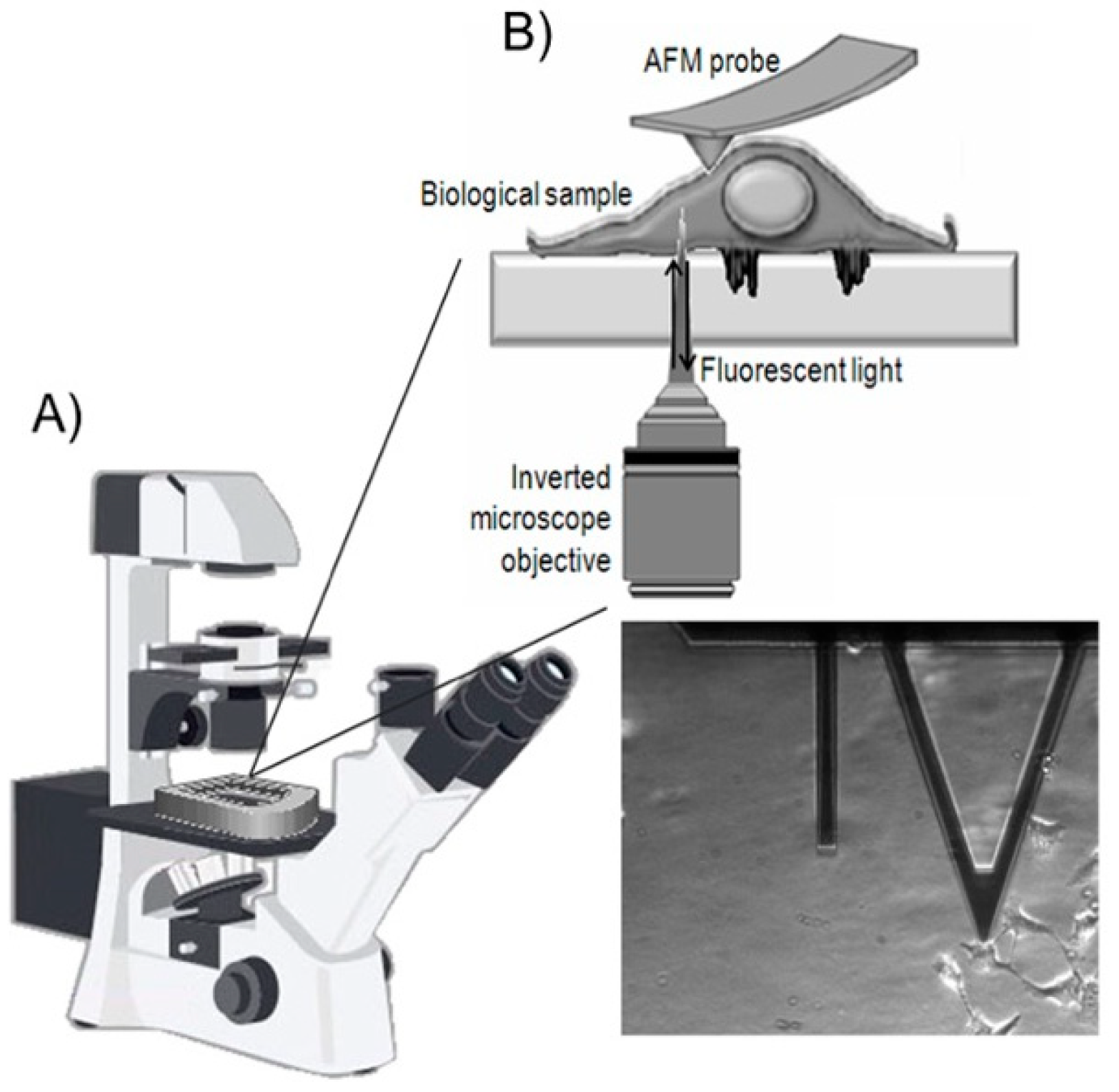
3. AFM Integrated with Optical Technique
4. Current Uses and Importance of AFM in Cancer Biomedicine
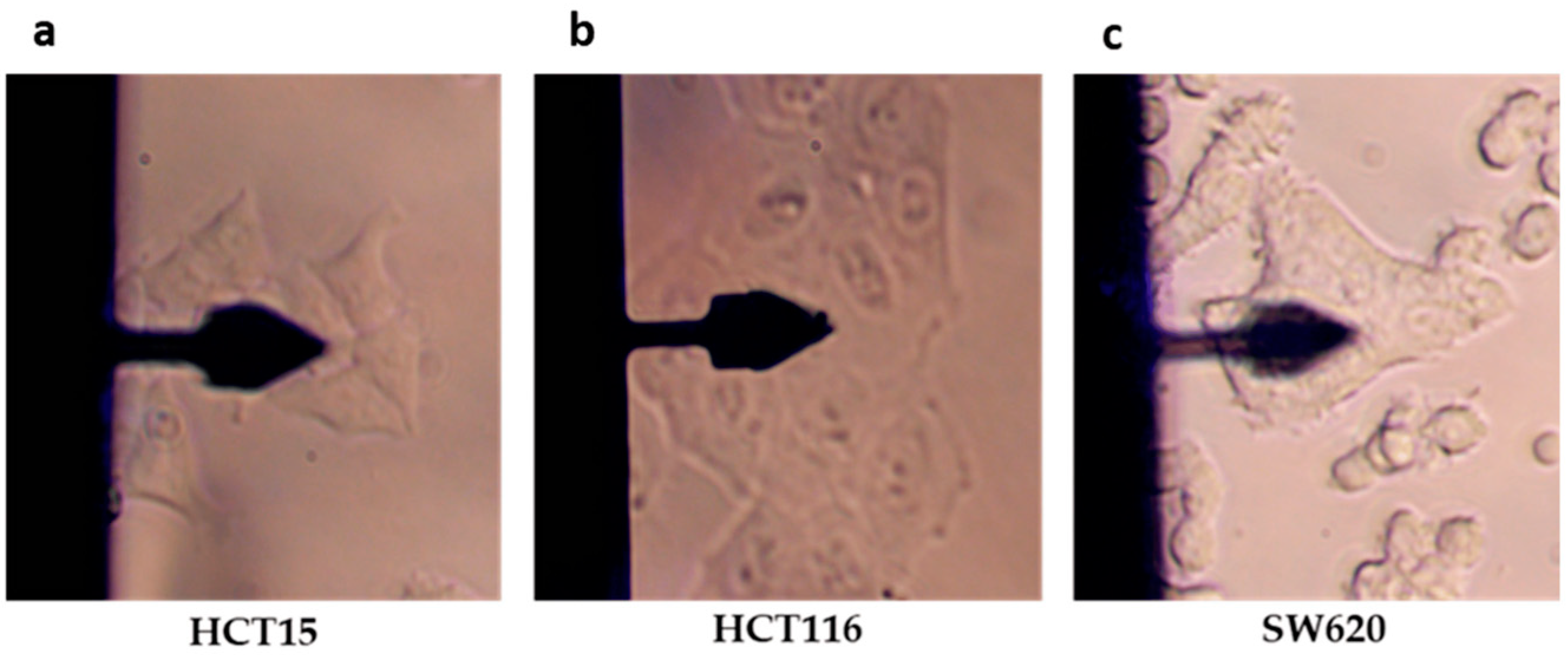
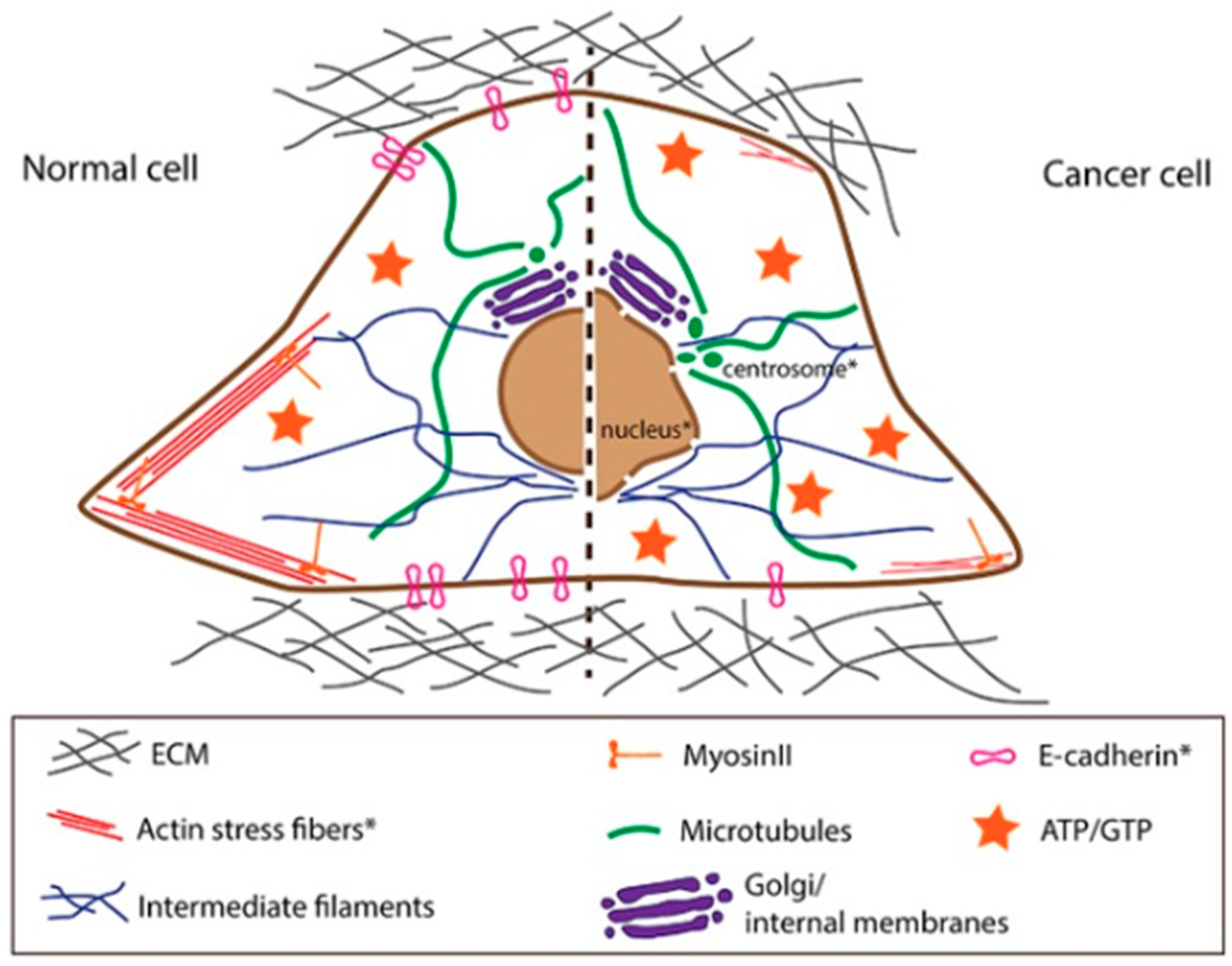
4.1. AFM Use to Investigate Structural and Morphological Changes in Cancer Cells
4.2. The Role of the Tumor Microenvironment
4.3. Nanomechanics and Morphological Characterization of Cellular Exosomes
4.4. The Role of Genes in Cancer Progression
4.5. Evaluation of Drug Formulations Against Cancer Diseases
5. Conclusions and Future Perspectives
Author Contributions
Funding
Conflicts of Interest
References
- Basoli, F.; Giannitelli, S.M.; Gori, M.; Mozetic, P.; Bonfanti, A.; Trombetta, M.; Rainer, A. Biomechanical Characterization at the Cell Scale: Present and Prospects. Front. Physiol. 2018, 9, 1449. [Google Scholar] [CrossRef]
- Ingber, D.E. Mechanobiology and Diseases of Mechanotransduction. Ann. Med. 2003, 35, 564–577. [Google Scholar] [CrossRef] [PubMed]
- Jansen, K.A.; Donato, D.M.; Balcioglu, H.E.; Schmidt, T.; Danen, E.H.J.; Koenderink, G.H. A Guide to Mechanobiology: Where Biology and Physics Meet. Biochim. Biophys. Acta-Mol. Cell Res. 2015, 1853, 3043–3052. [Google Scholar] [CrossRef]
- Ladoux, B.; Mège, R. Mechanobiology of Collective Cell Behaviours. Nat. Rev. Mol. Cell Biol. 2017, 18, 743–757. [Google Scholar] [CrossRef]
- Roca-Cusachs, P.; Conte, V.; Trepat, X. Quantifying Forces in Cell Biology. Nat. Cell Biol. 2017, 19, 742–751. [Google Scholar] [CrossRef] [PubMed]
- Yip, H.Y.K.; Papa, A. Signaling Pathways in Cancer: Therapeutic Targets, Combinatorial Treatments, and New Developments. Cells 2021, 10, 659. [Google Scholar] [CrossRef] [PubMed]
- Janiszewska, M.; Primi, M.C.; Izard, T. Cell Adhesion in Cancer: Beyond the Migration of Single Cells. J. Biol. Chem. 2020, 295, 2495. [Google Scholar] [CrossRef]
- Aseervatham, J. Cytoskeletal Remodeling in Cancer. Biology 2020, 9, 385. [Google Scholar] [CrossRef]
- Pfeifer, C.R.; Irianto, J.; Discher, D.E. Nuclear Mechanics and Cancer Cell Migration. In Cell Migrations: Causes and Functions; Advances in Experimental Medicine and Biology; Springer: Cham, Switzerland, 2019; Volume 1146, pp. 117–130. [Google Scholar] [CrossRef]
- Buruga, K.; Kalathi, J.T. A Facile Synthesis of Halloysite Nanotubes Based Polymer Nanocomposites for Glass Coating Application. J. Alloys Compd. 2018, 735, 1807–1817. [Google Scholar] [CrossRef]
- Wu, P.H.; Aroush, D.R.B.; Asnacios, A.; Chen, W.C.; Dokukin, M.E.; Doss, B.L.; Durand-Smet, P.; Ekpenyong, A.; Guck, J.; Guz, N.V.; et al. A Comparison of Methods to Assess Cell Mechanical Properties. Nat. Methods 2018, 15, 491–498. [Google Scholar] [CrossRef]
- Runel, G.; Lopez-ramirez, N.; Chlasta, J.; Masse, I. Biomechanical Properties of Cancer Cells. Cells 2021, 10, 887. [Google Scholar] [CrossRef]
- Menezes, P.L.; Ingole, S.P.; Nosonovsky, M.; Kailas, S.V.; Lovell, M.R. Tribology for Scientists and Engineers: From Basics to Advanced Concepts; Springer: Berlin/Heidelberg, Germany, 2013; 948p. [Google Scholar] [CrossRef]
- Cascione, M.; De Matteis, V.; Rinaldi, R. Investigation of Cellular Morpho-Mechanical Alteration as a Novel Diagnostic Tool in Nanomedicine. In Proceedings of the 2023 IEEE Nanotechnology Materials and Devices Conference, Paestum, Italy, 22–25 October 2023; pp. 389–394. [Google Scholar] [CrossRef]
- Dvorak, J.A. The Application of Atomic Force Microscopy to the Study of Living Vertebrate Cells in Culture. Methods 2003, 29, 86–96. [Google Scholar] [CrossRef] [PubMed]
- Cascione, M.; De Matteis, V.; Rinaldi, R.; Leporatti, S. Atomic force microscope combined with optical microscopy for cells investigation. Microsc. Res. Tech. 2017, 80, 109. [Google Scholar] [CrossRef] [PubMed]
- Mathur, A.B.; Truskey, G.A.; Reichert, W.M. Atomic Force and Total Internal Reflection Fluorescence Microscopy for the Study of Force Transmission in Endothelial Cells. Biophys. J. 2000, 78, 1725–1735. [Google Scholar] [CrossRef]
- Kassies, R.; Van Der Werf, K.O.; Lenferink, A.; Hunter, C.N.; Olsen, J.D.; Subramaniam, V.; Otto, C. Combined AFM and Confocal Fluorescence Microscope for Applications in Bio-Nanotechnology. J. Microsc. 2005, 217, 109–116. [Google Scholar] [CrossRef]
- Zhou, L.; Cai, M.; Tong, T.; Wang, H. Progress in the Correlative Atomic Force Microscopy and Optical Microscopy. Sensors 2017, 17, 938. [Google Scholar] [CrossRef]
- Liu, L.; Wei, Y.; Liu, J.; Wang, K.; Zhang, J.; Zhang, P.; Zhou, Y.; Li, B. Spatial High Resolution of Actin Filament Organization by PeakForce Atomic Force Microscopy. Cell Prolif. 2020, 53, e12670. [Google Scholar] [CrossRef]
- Skamrahl, M.; Colin-York, H.; Barbieri, L.; Fritzsche, M. Simultaneous Quantification of the Interplay Between Molecular Turnover and Cell Mechanics by AFM–FRAP. Small 2019, 15, e1902202. [Google Scholar] [CrossRef] [PubMed]
- Datta, R.; Heaster, T.M.; Sharick, J.T.; Gillette, A.A.; Skala, M.C.; Biomed, J. Fluorescence Lifetime Imaging Microscopy: Fundamentals and Advances in Instrumentation, Analysis, and Applications. J. Biomed. Opt. 2020, 25, 71203. [Google Scholar] [CrossRef]
- Becerra, N.; Salis, B.; Tedesco, M.; Flores, S.M.; Vena, P.; Raiteri, R. AFM and Fluorescence Microscopy of Single Cells with Simultaneous Mechanical Stimulation via Electrically Stretchable Substrates. Materials 2021, 14, 4131. [Google Scholar] [CrossRef]
- Hohmann, T.; Dehghani, F. The Cytoskeleton—A Complex Interacting Meshwork. Cells 2019, 8, 362. [Google Scholar] [CrossRef] [PubMed]
- De Matteis, V.; Cascione, M.; Toma, C.C.; Pellegrino, P.; Rizzello, L.; Rinaldi, R. Tailoring Cell Morphomechanical Perturbations through Metal Oxide Nanoparticles. Nanoscale Res. Lett. 2019, 14, 109. [Google Scholar] [CrossRef] [PubMed]
- Calzado-Martín, A.; Encinar, M.; Tamayo, J.; Calleja, M.; San Paulo, A. Effect of Actin Organization on the Stiffness of Living Breast Cancer Cells Revealed by Peak-Force Modulation Atomic Force Microscopy. ACS Nano 2016, 10, 3365–3374. [Google Scholar] [CrossRef]
- Monteiro-Reis, S.; Ferreira, J.P.S.; Pires, R.A.; Lobo, J.; Carvalho, J.A.; Reis, R.L.; Jorge, R.N.; Jerónimo, C. Bladder Wall Stiffness after Cystectomy in Bladder Cancer Patients: A Preliminary Study. Cancers 2023, 15, 359. [Google Scholar] [CrossRef]
- Zhao, X.; Zhong, Y.; Ye, T.; Wang, D.; Mao, B. Discrimination Between Cervical Cancer Cells and Normal Cervical Cells Based on Longitudinal Elasticity Using Atomic Force Microscopy. Nanoscale Res. Lett. 2015, 10, 482. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Sun, L.; Okuda, S.; Yamamoto, D.; Nakayama, M.; Oshima, H.; Saito, H.; Kouyama, Y.; Mimori, K.; Ando, T.; et al. Nano-Scale Physical Properties Characteristic to Metastatic Intestinal Cancer Cells Identified by High-Speed Scanning Ion Conductance Microscope. Biomaterials 2022, 280, 121256. [Google Scholar] [CrossRef]
- Kashani, A.S.; Packirisamy, M. Cancer cells optimize elasticity for efficient migration. R. Soc. Open Sci. 2020, 7, 10. [Google Scholar] [CrossRef]
- Hayashi, K.; Iwata, M. Stiffness of cancer cells measured with an AFM indentation method. J. Mech. Behav. Biomed. Mater. 2015, 49, 105–111. [Google Scholar] [CrossRef]
- Kwon, S.; Yang, W.; Moon, D.; Kim, K.S. Comparison of Cancer Cell Elasticity by Cell Type. J. Cancer 2020, 11, 5403–5412. [Google Scholar] [CrossRef]
- Brás, M.M.; Cruz, T.B.; Maia, A.F.; Oliveira, M.J.; Sousa, S.R.; Granja, P.L.; Radmacher, M. Mechanical Properties of Colorectal Cancer Cells Determined by Dynamic Atomic Force Microscopy: A Novel Biomarker. Cancers 2022, 14, 5053. [Google Scholar] [CrossRef]
- Tang, K.; Xin, Y.; Li, K.; Chen, X.; Tan, Y. Cell Cytoskeleton and Stiffness Are Mechanical Indicators of Organotropism in Breast Cancer. Biology 2021, 10, 259. [Google Scholar] [CrossRef] [PubMed]
- Viljoen, A.; Mathelié-Guinlet, M.; Ray, A.; Strohmeyer, N.; Oh, Y.J.; Hinterdorfer, P.; Müller, D.J.; Alsteens, D.; Dufrêne, Y.F. Force spectroscopy of single cells using atomic force microscopy. Nat. Rev. Methods Primers 2021, 1, 63. [Google Scholar] [CrossRef]
- Smolyakov, G.; Thiebot, B.; Campillo, C.; Labdi, S.; Severac, C.; Pelta, J.; Dague, É. Elasticity, Adhesion, and Tether Extrusion on Breast Cancer Cells Provide a Signature of Their Invasive Potential. ACS Appl. Mater. Interfaces 2016, 8, 27426–27431. [Google Scholar] [CrossRef]
- Gaman, A.; Osiac, E.; Rotaru, I.; Taisescu, C. Surface morphology of leukemic cells from chronic myeloid leukemia under atomic force microscopy. Curr. Health Sci. J. 2013, 39, 45–47. [Google Scholar] [PubMed] [PubMed Central]
- Kaul-Ghanekar, R.; Singh, S.; Mamgain, H.; Jalota-Badhwar, A.; Paknikar, K.M.; Chattopadhyay, S. Tumor suppressor protein SMAR1 modulates the roughness of cell surface: Combined AFM and SEM study. BMC Cancer 2009, 9, 350. [Google Scholar] [CrossRef] [PubMed]
- Schierbaum, N.; Rheinlaender, J.; Schäffer, T.E. Viscoelastic properties of normal and cancerous human breast cells are affected differently by contact to adjacent cells. Acta Biomater. 2017, 55, 239–248. [Google Scholar] [CrossRef]
- Alibert, C.; Goud, B.; Manneville, J.-B. Are cancer cells really softer than normal cells? Biol. Cell 2017, 109, 167–189. [Google Scholar] [CrossRef]
- Buckley, S.T.; Medina, C.; Davies, A.M.; Ehrhardt, C. Cytoskeletal re-arrangement in TGF-β1-induced alveolar epithelial-mesenchymal transition studied by atomic force microscopy and high-content analysis. Nanomed. NBM 2012, 8, 355–364. [Google Scholar] [CrossRef]
- Cascione, M.; De Matteis, V.; Toma, C.C.; Leporatti, S. Morphomechanical alterations induced by transforming growth factor-β1 in epithelial breast cancer cells. Cancers 2018, 10, 234. [Google Scholar] [CrossRef]
- Cascione, M.; Leporatti, S.; Dituri, F.; Giannelli, G. Transforming growth factor-β promotes morphomechanical effects involved in epithelial to mesenchymal transition in living hepatocellular carcinoma. Int. J. Mol. Sci. 2019, 20, 108. [Google Scholar] [CrossRef]
- Footer, M.J.; Kerssemakers, J.W.; Theriot, J.A.; Dogterom, M. Direct measurement of force generation by actin filament polymerization using an optical trap. Proc. Nat. Acad. Sci. USA 2007, 104, 2181–2186. [Google Scholar] [CrossRef] [PubMed]
- Humphrey, J.D.; Dufresne, E.R.; Schwartz, M.A. Mechano-transduction and extracellular matrix homeostasis. Nat. Rev. Mol. Cell. Biol. 2014, 15, 802–812. [Google Scholar] [CrossRef]
- Hartmann, B.; Fleischhauer, L.; Nicolau, M.; Jensen, T.H.L.; Taran, F.-A.; Clausen-Schaumann, H.; Reuten, R. Profiling native pulmonary basement membrane stiffness using atomic force microscopy. Nat. Protoc. 2024, 19, 1498–1528. [Google Scholar] [CrossRef] [PubMed]
- Abidine, Y.; Giannetti, A.; Revilloud, J.; Laurent, V.M.; Verdier, C. Viscoelastic Properties in Cancer: From Cells to Spheroids. Cells 2021, 10, 1704. [Google Scholar] [CrossRef] [PubMed]
- Micalet, A.; Upadhyay, A.; Javanmardi, Y.; de Brito, C.G.; Moeendarbary, E.; Cheema, U. Patient-specific colorectal-cancer-associated fibroblasts modulate tumor microenvironment mechanics. iScience 2024, 27, 110060. [Google Scholar] [CrossRef]
- Stylianou, A.; Gkretsi, V.; Stylianopoulos, T. Transforming growth factor-β modulates pancreatic cancer associated fibroblasts cell shape, stiffness and invasion. BBA-Gen. Subj. 2018, 1862, 1537–1546. [Google Scholar] [CrossRef]
- Stylianou, A.; Voutouri, C.; Mpekris, F.; Stylianopoulos, T. Pancreatic Cancer Presents Distinct Nanomechanical Properties During Progression. Ann. Biomed. Eng. 2023, 51, 1602–1615. [Google Scholar] [CrossRef]
- Sapudom, J.; Rubner, S.; Martin, S.; Kurth, T.; Riedel, S.; Mierke, C.T.; Pompe, T. The phenotype of cancer cell invasion controlled by fibril diameter and pore size of 3D collagen networks. Biomaterials 2015, 52, 367–375. [Google Scholar] [CrossRef]
- Shen, Y.; Wang, X.; Lu, J.; Salfenmoser, M.; Wirsik, N.M.; Schleussner, N.; Imle, A.; Freire Valls, A.; Radhakrishnan, P.; Liang, J.; et al. Reduction of Liver Metastasis Stiffness Improves Response to Bevacizumab in Metastatic Colorectal Cancer. Cancer Cell. 2020, 37, 800–817. [Google Scholar] [CrossRef]
- Mahaffey, B.J.; Fowler, Z.P.; Lung, Z.; Dang, V.; Lee, H.; Munoz, M.A.; Goodin, D.A.; Frieboes, H.B.; Williams, B.J.; Johnson, A.M.; et al. The prognostic effect of mechanical, ultrastructural, and ECM signatures in glioblastoma core and rim. APL Bioeng. 2024, 8, 036101. [Google Scholar] [CrossRef]
- Fan, Y.; Sun, Q.; Li, X.; Feng, J.; Ao, Z.; Li, X.; Wang, J. Substrate Stiffness Modulates the Growth, Phenotype, and Chemoresistance of Ovarian Cancer Cells. Front. Cell Dev. Biol. 2021, 9, 718834. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Chen, J.; Zhang, Z.; Xiao, C.; Zeng, H.; Xu, C.; Yang, X.; Li, Z. Modulating tumor mechanics with nanomedicine for cancer therapy. Biomater. Sci. 2023, 11, 4471–4489. [Google Scholar] [CrossRef]
- Kontomaris, S.V.; Stylianou, A.; Georgakopoulos, A.; Malamou, A. 3D AFM Nanomechanical Characterization of Biological Materials. Nanomaterials 2023, 13, 395. [Google Scholar] [CrossRef]
- Najera, J.; Rosenberger, M.R.; Datta, M. Atomic Force Microscopy Methods to Measure Tumor Mechanical Properties. Cancers 2023, 15, 3285. [Google Scholar] [CrossRef] [PubMed]
- Wan, Z.; Gao, X.T.; Dong, Y.; Zhao, Y.X.; Chen, X.T.; Yang, G.D.; Liu, L. Exosome-mediated cell-cell communication in tumor progression. Am. J. Cancer Res. 2018, 8, 1661–1673. [Google Scholar] [PubMed]
- Cheshomi, H.; Matin, M.M. Exosomes and their importance in metastasis, diagnosis, and therapy of colorectal cancer. J. Cell Biochem. 2019, 120, 2671–2686. [Google Scholar] [CrossRef]
- Xu, R.; Rai, A.; Chen, M.; Suwakulsiri, W.; Greening, D.W.; Simpson, R.J. Extracellular vesicles in cancer—Implications for future improvements in cancer care. Nat. Rev. Clin. Oncol. 2018, 15, 617–638. [Google Scholar] [CrossRef]
- Alderton, G. Exosomes drive premetastatic niche formation. Nat Rev. Cancer 2012, 12, 447. [Google Scholar] [CrossRef]
- Azmi, A.S.; Bao, B.; Sarkar, F.H. Exosomes in cancer development, metastasis, and drug resistance: A comprehensive review. Cancer Metastasis Rev. 2013, 32, 623–642. [Google Scholar] [CrossRef]
- Bastos, N.; Ruivo, C.F.; da Silva, S.; Melo, S.A. Exosomes in cancer: Use them or target them? Semin. Cell Dev. Biol. 2018, 78, 13–21. [Google Scholar] [CrossRef]
- Melo, S.A.; Sugimoto, H.; O’Connell, J.T.; Kato, N.; Villanueva, A.; Vidal, A.; Qiu, L.; Vitkin, E.; Perelman, L.T.; Melo, C.A.; et al. Cancer Exosomes Perform Cell-Independent MicroRNA Biogenesis and Promote Tumorigenesis. Cancer Cell 2014, 26, 707–721. [Google Scholar] [CrossRef]
- Sharma, S.; Rasool, H.I.; Palanisamy, V.; Mathisen, C.; Schmidt, M.; Wong, D.T.; Gimzewski, J.K. Structural-Mechanical Characterization of Nanoparticle Exosomes in Human Saliva, Using Correlative AFM, FESEM, and Force Spectroscopy. ACS Nano 2010, 4, 1921–1926. [Google Scholar] [CrossRef] [PubMed]
- Vorselen, D.; Piontek, M.C.; Roos, W.H.; Wuite, G.J.L. Mechanical Characterization of Liposomes and Extracellular Vesicles, a Protocol. Front. Mol. Biosci. 2020, 7, 139. [Google Scholar] [CrossRef] [PubMed]
- Paul, D.; Roy, A.; Nandy, A.; Datta, B.; Borar, P.; Pal, S.K.; Senapati, D.; Rakshit, T. Identification of Biomarker Hyaluronan on Colon Cancer Extracellular Vesicles Using Correlative AFM and Spectroscopy. J. Phys. Chem. Lett. 2020, 11, 5569–5576. [Google Scholar] [CrossRef]
- Ju, T.; Yang, F.; Wang, S.; Wang, J.; Song, Z.; Xu, H.; Wang, Z.; Chen, Y. Nanoscopic characterization of hepatocytes treated with normoxic and hypoxic tumor-derived exosomes. Micron 2022, 158, 103283. [Google Scholar] [CrossRef] [PubMed]
- Dobra, G.; Bukva, M.; Szabo, Z.; Bruszel, B.; Harmati, M.; Gyukity-Sebestyen, E.; Jenei, A.; Szucs, M.; Horvath, P.; Biro, T.; et al. Small Extracellular Vesicles Isolated from Serum May Serve as Signal-Enhancers for the Monitoring of CNS Tumors. Int. J. Mol. Sci. 2020, 21, 5359. [Google Scholar] [CrossRef]
- Yurtsever, A.; Yoshida, T.; Behjat, A.B.; Araki, Y.; Hanayama, R.; Fukuma, T. Structural and mechanical characteristics of exosomes from osteosarcoma cells explored by 3D-atomic force microscopy. Nanoscale 2021, 13, 6661–6677. [Google Scholar] [CrossRef]
- Sharma, S.; LeClaire, M.; Wohlschlegel, J.; Gimzewski, J. Impact of isolation methods on the biophysical heterogeneity of single extracellular vesicles. Sci. Rep. 2020, 10, 13327. [Google Scholar] [CrossRef]
- Senigagliesi, B.; Samperi, G.; Cefarin, N.; Gneo, P.L.; Petrosino, S.; Apollonio, M.; Caponnetto, F.; Sgarra, R.; Collavin, L.; Cesselli, D.; et al. Triple negative breast cancer-derived small extracellular vesicles as modulator of biomechanics in target cells. Nanomed. Nanotechnol. Biol. Med. 2022, 44, 102582. [Google Scholar] [CrossRef]
- Manganelli, V.; Dini, L.; Tacconi, S.; Dinarelli, S.; Capozzi, A.; Riitano, G.; Recalchi, S.; Caglar, T.R.; Fratini, F.; Misasi, R.; et al. Autophagy Promotes Enrichment of Raft Components within Extracellular Vesicles Secreted by Human 2FTGH Cells. Int. J. Mol. Sci. 2024, 25, 6175. [Google Scholar] [CrossRef]
- LeClaire, M.; Gimzewski, J.; Sharma, S. A review of the biomechanical properties of single extracellular vesicles. NanoSelect 2021, 2, 1–15. [Google Scholar] [CrossRef]
- Di Santo, R.; Romanò, S.; Mazzini, A.; Jovanović, S.; Nocca, G.; Campi, G.; Papi, M.; De Spirito, M.; Di Giacinto, F.; Ciasca, G. Recent Advances in the Label-Free Characterization of Exosomes for Cancer Liquid Biopsy: From Scattering and Spectroscopy to Nanoindentation and Nanodevices. Nanomaterials 2021, 11, 1476. [Google Scholar] [CrossRef]
- Spill, S.F.; Reynolds, D.S.; Kamm, R.D.; Zaman, M.H. Impact of the physical microenvironment on tumor progression and metastasis. Curr. Opin. Biotechnol. 2016, 40, 41–48. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Mo, Y.; Gong, Z.; Yang, X.; Yang, M.; Zhang, S.; Xiong, F.; Xiang, B.; Zhou, M.; Liao, Q.; et al. Circular RNAs in human cancer. Mol. Cancer 2017, 16, 25. [Google Scholar] [CrossRef]
- Zhong, Y.; Du, Y.; Yang, X.; Mo, Y.; Fan, C.; Xiong, F.; Ren, D.; Ye, X.; Li, C.; Wang, Y.; et al. Circular RNAs function as ceRNAs to regulate and control human cancer progression. Mol. Cancer 2018, 17, 79. [Google Scholar] [CrossRef] [PubMed]
- Wei, X.; Lu, Y.; Zhang, X.; Chen, M.-L.; Wang, J.-H. Recent advances in single-cell ultra-trace analysis. TrAC Trends Anal. Chem. 2020, 127, 115886, ISSN 0165-9936. [Google Scholar] [CrossRef]
- Gong, Z.; Yang, Q.; Zeng, Z.; Zhang, W.; Li, X.; Zu, X.; Deng, H.; Chen, P.; Liao, Q.; Xiang, B.; et al. An integrative transcriptomic analysis reveals p53 regulated miRNA, mRNA, and lncRNA networks in nasopharyngeal carcinoma. Tumour Biol. 2016, 37, 3683–3695. [Google Scholar] [CrossRef] [PubMed]
- He, B.; Li, W.; Wu, Y.; Wei, F.; Gong, Z.; Bo, H.; Wang, Y.; Li, X.; Xiang, B.; Guo, C.; et al. Epstein–Barr virus-encoded miR-BART6-3p inhibits cancer cell metastasis and invasion by targeting long non-coding RNA LOC553103. Cell Death Dis. 2016, 7, e2353. [Google Scholar] [CrossRef]
- Lee, H.J.; Diaz, M.F.; Price, K.M.; Ozuna, J.A.; Zhang, S.; Sevick-Muraca, E.M.; Hagan, J.P.; Wenzel, P.L. Fluid shear stress activates YAP1 to promote cancer cell motility. Nat. Commun. 2017, 8, 14122. [Google Scholar] [CrossRef]
- Shim, W.C.; Woo, S.; Park, J.W. Nanoscale Force-Mapping-Based Quantification of Low-Abundance Methylated DNA. Nano Lett. 2022, 22, 1324–1330. [Google Scholar] [CrossRef]
- Luo, C.H.; Shi, Y.; Liu, Y.Q.; Liu, Q.; Mao, M.; Luo, M.; Yang, K.-D.; Wang, W.-Y.; Chen, C.; Niu, Q.; et al. High levels of TIMP1 are associated with increased extracellular matrix stiffness in isocitrate dehydrogenase 1-wild type gliomas. Lab. Investig. 2022, 102, 1304–1313. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Xu, G.; Fu, H.; Li, P.; Li, D.; Deng, K.; Gao, W.; Shang, Y.; Wu, M. Membrane-bound transcription factor LRRC4 inhibits glioblastoma cell motility. Int. J. Biol. Macromol. 2023, 246, 125590. [Google Scholar] [CrossRef] [PubMed]
- Koebley, S.R.; Mikheikin, A.; Leslie, K.; Guest, D.; McConnell-Wells, W.; Lehman, J.H.; Al Juhaishi, T.; Zhang, X.; Roberts, C.H.; Picco, L.; et al. Digital Polymerase Chain Reaction Paired with High-Speed Atomic Force Microscopy for Quantitation and Length Analysis of DNA Length Polymorphisms. ACS Nano 2020, 14, 15385–15393. [Google Scholar] [CrossRef]
- Young, K.M.; Xu, C.; Ahkee, K.; Mezencev, R.; Swingle, S.P.; Yu, T.; Paikeday, A.; Kim, C.; McDonald, J.F.; Qiu, P.; et al. Correlating mechanical and gene expression data on the single cell level to investigate metastatic phenotypes. iScience 2023, 26, 106393. [Google Scholar] [CrossRef] [PubMed]
- Purdy, M.P.; Ducharme, M.; Haak, A.J.; Ravix, J.; Tan, Q.; Sicard, D.; Prakash, Y.S.; Tschumperlin, D.J.; Stewart, E.A. YAP/TAZ are Activated by Mechanical and Hormonal Stimuli in Myometrium and Exhibit Increased Baseline Activation in Uterine Fibroids. Reprod. Sci. 2020, 27, 1074–1085. [Google Scholar] [CrossRef]
- Melters, D.P.; Dalal, Y. Nano-Surveillance: Tracking Individual Molecules in a Sea of Chromatin. J. Mol. Biol. 2021, 433, 166720. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.-Y.; Myong, S. Probing steps in DNA transcription using single-molecule methods. J. Biol. Chem. 2021, 297, 101086. [Google Scholar] [CrossRef]
- Yang, Y.; Li, M.; Sun, X.; Zhou, C.; Wang, Y.; Wang, L.; Chen, L.; Liang, Z.; Zhu, L.; Yang, H. The selective cytotoxicity of DSF-Cu attributes to the biomechanical properties and cytoskeleton rearrangements in the normal and cancerous nasopharyngeal epithelial cells. Int. J. Biochem. Cell Biol. 2017, 84, 96–108. [Google Scholar] [CrossRef]
- Ren, J.; Huang, H.; Liu, Y.; Zheng, X.; Zou, Q. An Atomic Force Microscope Study Revealed Two Mechanisms in the Effect of Anticancer Drugs on Rate-Dependent Young’s Modulus of Human Prostate Cancer Cells. PLoS ONE 2015, 10, e0126107. [Google Scholar] [CrossRef]
- Hung, M.S.; Tsai, M.F. Investigating the Influence of Anti-Cancer Drugs on the Mechanics of Cells Using AFM. BioNanoScience 2015, 5, 156–161. [Google Scholar] [CrossRef]
- Rudzińska, M.; Parodi, A.; Maslova, V.D.; Efremov, Y.M.; Gorokhovets, N.V.; Makarov, V.A.; Popkov, V.A.; Golovin, A.V.; Zernii, E.Y.; Zamyatnin, A.A., Jr. Cysteine Cathepsins Inhibition Affects Their Expression and Human Renal Cancer Cell Phenotype. Cancers 2020, 12, 1310. [Google Scholar] [CrossRef]
- Nishio, T.; Shimada, Y.; Yoshikawa, Y.; Kenmotsu, T.; Schiessel, H.; Yoshikawa, K. The Anticancer Drug Daunomycin Directly Affects Gene Expression and DNA Structure. Int. J. Mol. Sci. 2023, 24, 6631. [Google Scholar] [CrossRef]
- Marinello, J.; Delcuratolo, M.; Capranico, G. Anthracyclines as Topoisomerase II Poisons: From Early Studies to New Perspectives. Int. J. Mol. Sci. 2018, 19, 3480. [Google Scholar] [CrossRef] [PubMed]
- Kaczorowska, A.; Lamperska, W.; Frączkowska, K.; Masajada, J.; Drobczyński, S.; Sobas, M.; Wróbel, T.; Chybicka, K.; Tarkowski, R.; Kraszewski, S.; et al. Profound Nanoscale Structural and Biomechanical Changes in DNA Helix upon Treatment with Anthracycline Drugs. Int. J. Mol. Sci. 2020, 21, 4142. [Google Scholar] [CrossRef] [PubMed]
- Sofińska, K.; Cieśla, M.; Barbasz, J.; Wilkosz, N.; Lipiec, E.; Szymoński, M.; Białas, P. Double-strand breaks quantification by statistical length analysis of DNA fragments imaged with AFM. Measurement 2022, 198, 111362. [Google Scholar] [CrossRef]
- Kim, H.; Hoshi, M.; Iijima, M.; Kuroda, S.; Nakamura, C. Development of a universal method for the measurement of binding affinities of antibody drugs towards a living cell based on AFM force spectroscopy. Anal. Methods 2020, 12, 2922–2927. [Google Scholar] [CrossRef] [PubMed]
- Meng, J.; Paetzell, E.; Bogorad, A.; Soboyejo, W.O. Adhesion between peptides/antibodies and breast cancer cells. J. Appl. Phys. 2010, 107, 114301. [Google Scholar] [CrossRef]
- Wang, M.; Ruan, Y.; Xing, X.; Chen, Q.; Peng, Y.; Cai, J. Curcumin induced nanoscale CD44 molecular redistribution and antigen–antibody interaction on HepG2 cell surface. Anal. Chim. Acta 2011, 697, 83–89. [Google Scholar] [CrossRef]
- Zhang, Q.; Shi, Y.; Xu, H.; Zhou, L.; Gao, J.; Jiang, J.; Cai, M.; Shan, Y. Evaluating the efficacy of the anticancer drug cetuximab by atomic force microscopy. RSC Adv. 2018, 8, 21793–21797. [Google Scholar] [CrossRef]
- Li, M.; Liu, L.; Xi, N.; Wang, Y. Applications of Atomic Force Microscopy in Exploring Drug Actions in Lymphoma-Targeted Therapy at the Nanoscale. BioNanoScience 2016, 6, 22–32. [Google Scholar] [CrossRef]
- Raudenska, M.; Kratochvilova, M.; Vicar, T.; Gumulec, J.; Balvan, J.; Polanska, H.; Pribyl, J.; Masarik, M. Cisplatin enhances cell stiffness and decreases invasiveness rate in prostate cancer cells by actin accumulation. Sci. Rep. 2019, 9, 1660. [Google Scholar] [CrossRef]
- Beton, K.; Brożek-Płuska, B. Biochemistry and Nanomechanical Properties of Human Colon Cells upon Simvastatin, Lovastatin, and Mevastatin Supplementations: Raman Imaging and AFM Studies. J. Phys. Chem. B 2022, 126, 7088. [Google Scholar] [CrossRef] [PubMed]
- De Matteis, V.; Cascione, M.; Toma, C.C.; Leporatti, S. Morphomechanical and organelle perturbation induced by silver nanoparticle exposure. J. Nanopart Res. 2018, 20, 273. [Google Scholar] [CrossRef]
- Cascione, M.; De Matteis, V.; Mandriota, G.; Leporatti, S.; Rinaldi, R. Acute Cytotoxic Effects on Morphology and Mechanical Behavior in MCF-7 Induced by TiO2NPs Exposure. Int. J. Mol. Sci. 2019, 20, 3594. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Ly, A.; Das, S.; Sakthivel, T.S.; Barkam, S.; Seal, S. Cerium oxide nanoparticles at the nano-bio interface: Size-dependent cellular uptake. Artif. Cells Nanomed. Biotechnol. 2018, 46, 956–963. [Google Scholar] [CrossRef]
- Lara-Cruz, C.; Jiménez-Salazar, J.; Ramón-Gallegos, E.; Damian-Matsumura, P.; Batina, N. Increasing roughness of the human breast cancer cell membrane through incorporation of gold nanoparticles. Int. J. Nanomed. 2016, 11, 5149–5161. [Google Scholar] [CrossRef] [PubMed]
- Abduljauwad, S.N.; Ahmed, H.U.R. Enhancing cancer cell adhesion with clay nanoparticles for countering metastasis. Sci. Rep. 2019, 9, 5935. [Google Scholar] [CrossRef]
- Miletić, M.; Aškrabić, S.; Rüger, J.; Vasić, B.; Korićanac, L.; Mondol, A.S.; Dellith, J.; Popp, J.; Schiebe, I.W.; Dohčević-Mitr, Z. Combined Raman and AFM detection of changes in HeLa cervical cancer cells induced by CeO2 nanoparticles–molecular and morphological perspectives. Analyst 2020, 145, 3983–3995. [Google Scholar] [CrossRef]
- Al, M.R.K.; Wu, Y.; Ghosh, D.; Do, B.H.; Chen, K.; Dawson, M.R.; Fang, N.; Sulchek, T.A.; El-Sayed, M.A. Nuclear Membrane-Targeted Gold Nanoparticles Inhibit Cancer Cell Migration and Invasion. ACS Nano 2017, 11, 3716–3726. [Google Scholar] [CrossRef]
- Swift, J.; Ivanovska, I.L.; Buxboim, A.; Harada, T.; Dingal, P.C.D.P.; Pinter, J.; Pajerowski, J.D.; Spinler, K.R.; Shin, J.-W.; Tewari, M.; et al. Nuclear Lamin-A Scales with Tissue Stiffness and Enhances Matrix-Directed Differentiation. Science 2013, 341, 1240104. [Google Scholar] [CrossRef]
- Garcia Diosa, J.A.; Gonzalez Orive, A.; Grundmeier, G.; Camargo Amado, R.J.; Keller, A. Morphological Dynamics of Leukemia Cells on TiO2 Nanoparticle Coatings Studied by AFM. Appl. Sci. 2021, 11, 9898. [Google Scholar] [CrossRef]
- Yang, F.; Riedel, R.; del Pino, P.; Pelaz, B.; Said, A.H.; Soliman, M.; Pinnapireddy, S.R.; Feliu, N.; Parak, W.J.; Bakowsky, U.; et al. Real-time, label-free monitoring of cell viability based on cell adhesion measurements with an atomic force microscope. J. Nanobiotechnol. 2017, 15, 23. [Google Scholar] [CrossRef]
- Winkler, R.; Ciria, M.; Ahmad, M.; Plank, H.; Marcuello, C. A Review of the Current State of Magnetic Force Microscopy to Unravel the Magnetic Properties of Nanomaterials Applied in Biological Systems and Future Directions for Quantum Technologies. Nanomaterials 2023, 13, 2585. [Google Scholar] [CrossRef] [PubMed]
- Kazakova, O.; Puttock, R.; Barton, C.; Corte-León, H.; Jaafar, M.; Neu, V.; Asenjo, A. Frontiers of magnetic force microscopy. J. Appl. Phys. 2019, 125, 060901. [Google Scholar] [CrossRef]
- Mamaeva, S.N.; Kononova, I.V.; Ruzhansky, M.; Nikiforov, P.V.; Nikolaev, N.A.; Pavlov, A.N.; Fedorova, N.F.; Huang, J.; Semenova, M.N.; Barashkova, D.V.; et al. Using Scanning Electron Microscopy and Atomic Force Microscopy to Study the Formation of Nanoparticles on Red Blood Cell Surface in Cervical Cancer Patients. Int. J. Biomed. 2020, 10, 70–75. [Google Scholar] [CrossRef]
- Spyratou, E.; Dilvoi, M.; Patatoukas, G.; Platoni, K.; Makropoulou, M.; Efstathopoulos, E. Probing the Effects of Ionizing Radiation on Young’s Modulus of Human Erythrocytes Cytoskeleton using Atomic Force Microscopy. J. Med. Phys. 2019, 4, 113–117. [Google Scholar] [CrossRef]
- Ma, S.; Wu, J.; Liu, Z.; He, R.; Wang, Y.; Liu, L.; Wang, T.; Wang, W. Quantitative characterization of cell physiological state based on dynamical cell mechanics for drug efficacy indication. J. Pharm. Anal. 2023, 13, 388.e402. [Google Scholar] [CrossRef]
- Deng, X.; Xiong, F.; Li, X.; Xiang, B.; Li, Z.; Wu, X.; Guo, C.; Li, X.; Li, Y.; Li, G.; et al. Application of atomic force microscopy in cancer research. J. Nanobiotechnol. 2018, 16, 102. [Google Scholar] [CrossRef]
- Stylianou, A.; Lekka, M.; Stylianopoulos, T. AFM assessing of nanomechanical fingerprints for cancer early diagnosis and classification: From single cell to tissue level. Nanoscale 2018, 10, 20930–20945. [Google Scholar] [CrossRef]
- Medina-Ramirez, I.E.; Macías-Díaz, J.E.; Masuoka-Ito, D.; Zapien, J.A. Holotomography and atomic force microscopy: A powerful combination to enhance cancer, microbiology, and nanotoxicology research. Discov. Nano 2024, 19, 64. [Google Scholar] [CrossRef]





Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Persano, F.; Parodi, A.; Pallaeva, T.; Kolesova, E.; Zamyatnin, A.A., Jr.; Pokrovsky, V.S.; De Matteis, V.; Leporatti, S.; Cascione, M. Atomic Force Microscopy: A Versatile Tool in Cancer Research. Cancers 2025, 17, 858. https://doi.org/10.3390/cancers17050858
Persano F, Parodi A, Pallaeva T, Kolesova E, Zamyatnin AA Jr., Pokrovsky VS, De Matteis V, Leporatti S, Cascione M. Atomic Force Microscopy: A Versatile Tool in Cancer Research. Cancers. 2025; 17(5):858. https://doi.org/10.3390/cancers17050858
Chicago/Turabian StylePersano, Francesca, Alessandro Parodi, Tatiana Pallaeva, Ekaterina Kolesova, Andrey A. Zamyatnin, Jr., Vadim S. Pokrovsky, Valeria De Matteis, Stefano Leporatti, and Mariafrancesca Cascione. 2025. "Atomic Force Microscopy: A Versatile Tool in Cancer Research" Cancers 17, no. 5: 858. https://doi.org/10.3390/cancers17050858
APA StylePersano, F., Parodi, A., Pallaeva, T., Kolesova, E., Zamyatnin, A. A., Jr., Pokrovsky, V. S., De Matteis, V., Leporatti, S., & Cascione, M. (2025). Atomic Force Microscopy: A Versatile Tool in Cancer Research. Cancers, 17(5), 858. https://doi.org/10.3390/cancers17050858











