Semi-Rational Design of Diaminopimelate Dehydrogenase from Symbiobacterium thermophilum Improved Its Activity toward Hydroxypyruvate for D-serine Synthesis
Abstract
:1. Introduction
2. Results and Discussion
2.1. Enzyme Activity Determination of StDAPDH/H227V
2.2. Prediction of Key Residues for Saturation Mutagenesis
2.3. Screening of Positive Single Mutants
2.4. Iterative Saturation Mutagenesis
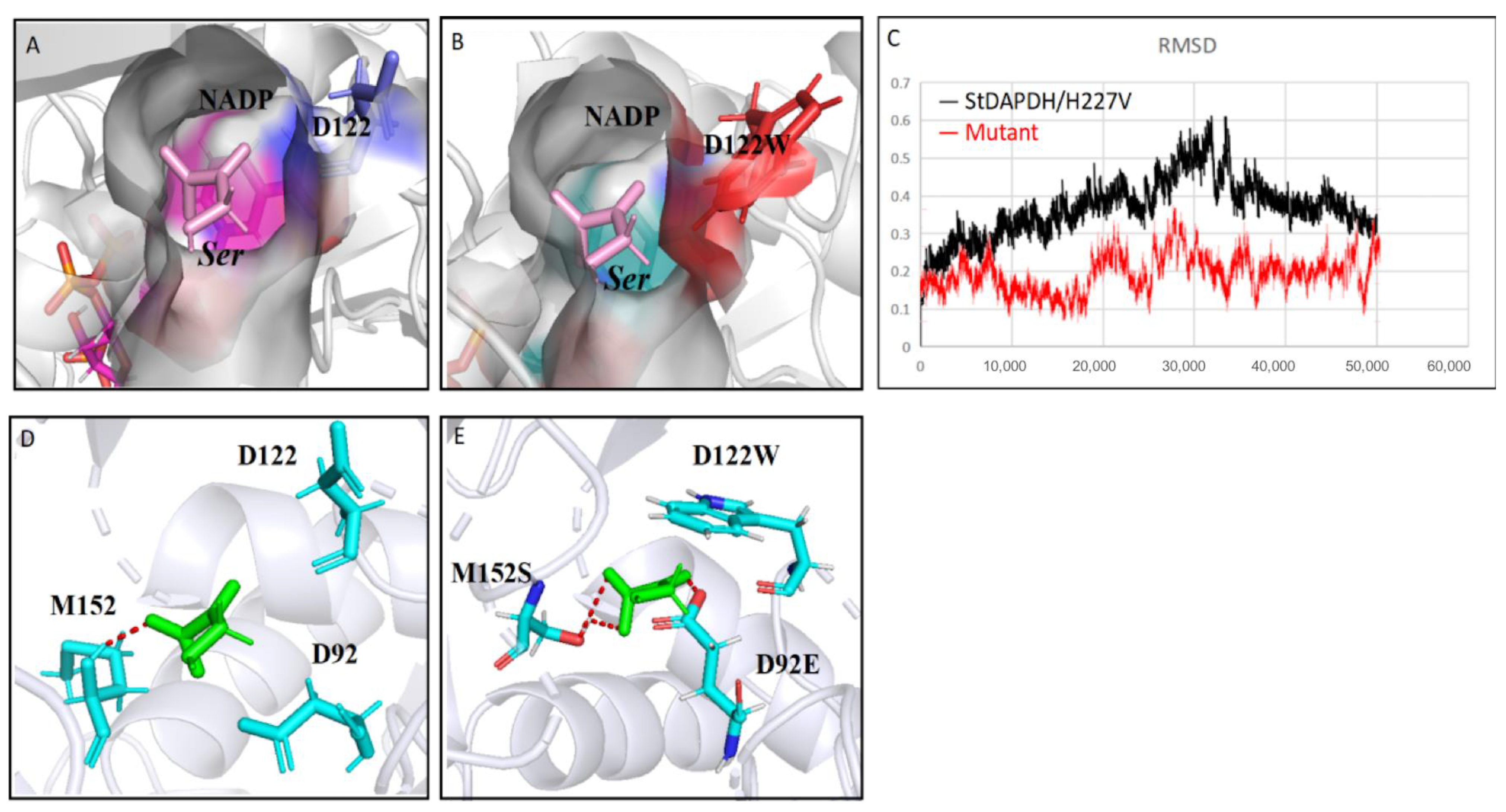
2.5. Role of the Mutation Sites for Enhanced Activity
3. Materials and Methods
3.1. Molecular Modeling and Docking Calculations
3.2. Site-Directed Saturation Mutagenesis
3.3. Protein Expression of StDAPDH Mutants
3.4. Cell Fragmentation
3.5. Mutants Screening
3.6. Purification of Proteins
3.7. Determination of Kinetic Parameters
3.8. Enzymatic Reactions and Detection of L-serine, D-serine, and HPPA
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Genchi, G. An overview on D-amino acids. Amino Acids 2017, 49, 1521–1533. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.Z.; Ma, Q.Y.; Zhu, H.L. Distribution, industrial applications, and enzymatic synthesis of D-amino acids. Appl. Microbiol. Biotechnol. 2015, 99, 3341–3349. [Google Scholar] [CrossRef]
- Bommarius, A.S.; Schwarm, M.; Drauz, K. Biocatalysis to amino acid-based chiral pharmaceuticals—Examples and perspectives. J. Mol. Catal. B Enzym. 1998, 5, 1–11. [Google Scholar] [CrossRef]
- Yang, H.; Zheng, G.; Peng, X. D-Amino acids and D-Tyr-tRNATyr deacylase: Stereospecificity of the translation machine revisited. FEBS Lett. 2003, 552, 95–98. [Google Scholar] [CrossRef] [Green Version]
- Ma, J.S. Unnatural amino acids in drug discovery. Chim. Oggi. 2003, 21, 65–68. [Google Scholar]
- Friedman, M.; Levin, C. Nutritional and medicinal aspects of D amino acids. Amino Acids 2012, 42, 1553–1582. [Google Scholar] [CrossRef]
- Bruckner, H.; Westhauser, T. Chromatographic determination of L- and D-amino acids in plants. Amino Acids 2003, 24, 43–55. [Google Scholar] [CrossRef] [PubMed]
- Wehofsky, N.; Pech, A.; Liebscher, S.; Schmidt, S.; Komeda, H.; Asano, Y.; Bordusa, F. D-amino acid specific proteases and native all-L-proteins: A convenient combination for semisynthesis. Angew. Chem. Int. Ed. 2008, 47, 5456–5460. [Google Scholar] [CrossRef] [PubMed]
- Patil, M.D.; Grogan, G.; Bommarius, A.; Hyungdon, Y. Oxidoreductase-Catalyzed Synthesis of Chiral Amines. ACS Catal. 2018, 8, 10985–11015. [Google Scholar] [CrossRef]
- Misono, H.; Togawa, H.; Yamamoto, T.; Soda, K. Meso-alpha, epsilondiaminopimelate D-dehydrogenase: Distribution and the reaction product. J. Bacteriol. 1979, 137, 22–27. [Google Scholar] [CrossRef] [Green Version]
- Pollegioni, L.; Rosini, E.; Molla, G. Advances in enzymatic synthesis of D-amino acids. Int. J. Mol. Sci. 2020, 21, 3206. [Google Scholar] [CrossRef] [PubMed]
- Bornadel, A.; Bisagni, S.; Pushpanath, A.; Montgomery, S.; Turner, N.; Dominguez, B. Technical considerations for scale-up of imine-reductase-catalyzed reductive amination: A case study. Org. Process Res. Dev. 2019, 23, 1262–1268. [Google Scholar] [CrossRef]
- Gao, X.Z.; Chen, X.; Liu, W.D.; Feng, J.H.; Wu, Q.; Hua, L.; Zhu, D.M. A novel meso-diaminopimelate dehydrogenase from Symbiobacterium thermophilum: Overexpression, characterization, and potential for D-amino acid synthesis. Appl. Environ. Microbiol. 2012, 78, 8595–8600. [Google Scholar] [CrossRef] [Green Version]
- Gao, X.Z.; Huang, F.; Feng, J.H.; Chen, X.; Zhang, H.L.; Wang, Z.X.; Wu, Q.; Zhu, D.M. Engineering the meso-diaminopimelate dehydrogenase from Symbiobacterium thermophilum by site saturation mutagenesis for D-phenylalanine synthesis. Appl. Environ. Microbiol. 2013, 79, 5078–5081. [Google Scholar] [CrossRef] [Green Version]
- Liu, W.; Li, Z.; Huang, C.H.; Guo, R.T.; Zhao, L.M.; Zhang, D.L.; Chen, X.; Wu, Q.; Zhu, D.M. Structural and mutational studies on the unusual substrate specificity of meso-diaminopimelate dehydrogenase from Symbiobacterium thermophilum. Chembiochem 2014, 15, 217–222. [Google Scholar] [CrossRef]
- Cheng, X.K.; Chen, X.; Feng, J.H.; Wu, Q.; Zhu, D.M. Structure-guided engineering of meso-diaminopimelate dehydrogenase for enantioselective reductive amination of sterically bulky 2-keto acids. Catal. Sci. Technol. 2018, 8, 4994–5002. [Google Scholar] [CrossRef]
- Vedha-Peters, K.; Gunawardana, M.; Rozzell, J.D. Creation of a broad-range and highly stereoselective D-amino acid dehydrogenase for the one-step synthesis of D-amino acids. J. Am. Chem. Soc. 2006, 128, 10923–10929. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Akita, H.; Doi, K.; Kawarabayasi, Y. Creation of a thermostable NADP+-dependent D-amino acid dehydrogenase from Ureibacillus thermosphaericus strain A1 meso-diaminopimelate dehydrogenase by site-directed mutagenesis. Biotechnol. Lett. 2012, 34, 1693–1699. [Google Scholar] [CrossRef]
- Hayashi, J.; Seto, T.; Akita, H.; Watanabe, M.; Hoshino, T.; Yoneda, K.; Ohshima, T.; Sakuraba, H. Structure-based engineering of an artificially generated NADP+-dependent D-amino acid dehydrogenase. Appl. Environ. Microbiol. 2017, 83, 1–13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Akita, H.; Suzuki, H.; Doi, K. Efficient synthesis of D-branched-chain amino acids and their labeled compounds with stable isotopes using D-amino acid dehydrogenase. Appl. Microbiol. Biotechnol. 2014, 98, 1135–1143. [Google Scholar] [CrossRef]
- Hanson, R.L.; Davis, B.L.; Goldberg, S.L.; Johnston, R.M.; Parker, W.L.; Tully, T.P.; Montana, M.A.; Patel, R.N. Enzymatic preparation of a D-amino acid from a racemic amino acid or keto acid. Org. Process Res. Dev. 2008, 12, 1119–1129. [Google Scholar] [CrossRef]
- Xu, J.M.; Fu, F.T.; Hu, H.F.; Zheng, Y.G. A high-throughput screening method for amino acid dehydrogenas. Anal. Biochem. 2016, 495, 29–31. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Yao, P.; Gong, R.; Li, J.; Wu, Q.; Zhu, D.M.; Liu, P.; Li, G.Y.; Yao, P.Y.; Lonsdale, R.; et al. Simultaneous engineering of an enzyme’s entrance tunnel and active site: The case of monoamine oxidase MAO-N. Chem. Sci. 2017, 8, 4093–4099. [Google Scholar] [CrossRef] [Green Version]





| Enzyme | Specific Activity (U/mg) | Km (mM) | kcat (s−1) | kcat/Km (s−1mM−1) |
|---|---|---|---|---|
| H227V (control) | 0.022 ± 0.002 | 16.34 ± 0.34 | 1.09 ± 0.02 | 0.066 ± 0.003 |
| D92E | 0.057 ± 0.006 | 12.83 ± 0.29 | 1.47 ± 0.03 | 0.115 ± 0.006 |
| D122W | 0.086 ± 0.007 | 16.97 ± 0.43 | 3.44 ± 0.05 | 0.21 ± 0.005 |
| M152S | 0.049 ± 0.005 | 13.26 ± 0.37 | 1.43 ± 0.02 | 0.14 ± 0.003 |
| N253Y | 0.044 ± 0.003 | 14.23 ± 0.41 | 1.33 ± 0.04 | 0.093 ± 0.005 |
| D92E/D122W | 0.098 ± 0.008 | 12.32 ± 0.24 | 3.45 ± 0.06 | 0.281 ± 0.004 |
| D92E/M152S | 0.065 ± 0.006 | 11.07 ± 0.17 | 1.71 ± 0.03 | 0.154 ± 0.002 |
| D92E/N253Y | 0.054 ± 0.006 | 12.12 ± 0.21 | 1.62 ± 0.02 | 0.133 ± 0.003 |
| D122W/M152S | 0.12 ± 0.01 | 12.04 ± 0.18 | 3.56 ± 0.08 | 0.295 ± 0.005 |
| D122W/N253Y | 0.082 ± 0.008 | 14.82 ± 0.46 | 3.71 ± 0.06 | 0.253 ± 0.004 |
| M152S/N253Y | 0.048 ± 0.05 | 13.18 ± 0.35 | 1.48 ± 0.01 | 0.113 ± 0.002 |
| D92E/D122W/M152S (M3) | 0.19 ± 0.05 | 11.37 ± 0.21 | 3.96 ± 0.07 | 0.348 ± 0.007 |
| D122W/M152S/N253Y | 0.11 ± 0.4 | 11.29 ± 0.14 | 3.34 ± 0.06 | 0.319 ± 0.002 |
| D92E/D122W/M152S/N253Y | 0.15 ± 0.02 | 12.06 ± 0.26 | 3.89 ± 0.05 | 0.321 ± 0.003 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, Z.; Qu, H.; Li, W.; Xu, Y.; Nie, Y. Semi-Rational Design of Diaminopimelate Dehydrogenase from Symbiobacterium thermophilum Improved Its Activity toward Hydroxypyruvate for D-serine Synthesis. Catalysts 2023, 13, 576. https://doi.org/10.3390/catal13030576
Wang Z, Qu H, Li W, Xu Y, Nie Y. Semi-Rational Design of Diaminopimelate Dehydrogenase from Symbiobacterium thermophilum Improved Its Activity toward Hydroxypyruvate for D-serine Synthesis. Catalysts. 2023; 13(3):576. https://doi.org/10.3390/catal13030576
Chicago/Turabian StyleWang, Ziyao, Haojie Qu, Wenqi Li, Yan Xu, and Yao Nie. 2023. "Semi-Rational Design of Diaminopimelate Dehydrogenase from Symbiobacterium thermophilum Improved Its Activity toward Hydroxypyruvate for D-serine Synthesis" Catalysts 13, no. 3: 576. https://doi.org/10.3390/catal13030576








