Biodiversity in the Rhizosphere of Selected Winter Wheat (Triticum aestivum L.) Cultivars—Genetic and Catabolic Fingerprinting
Abstract
:1. Introduction
2. Materials and Methods
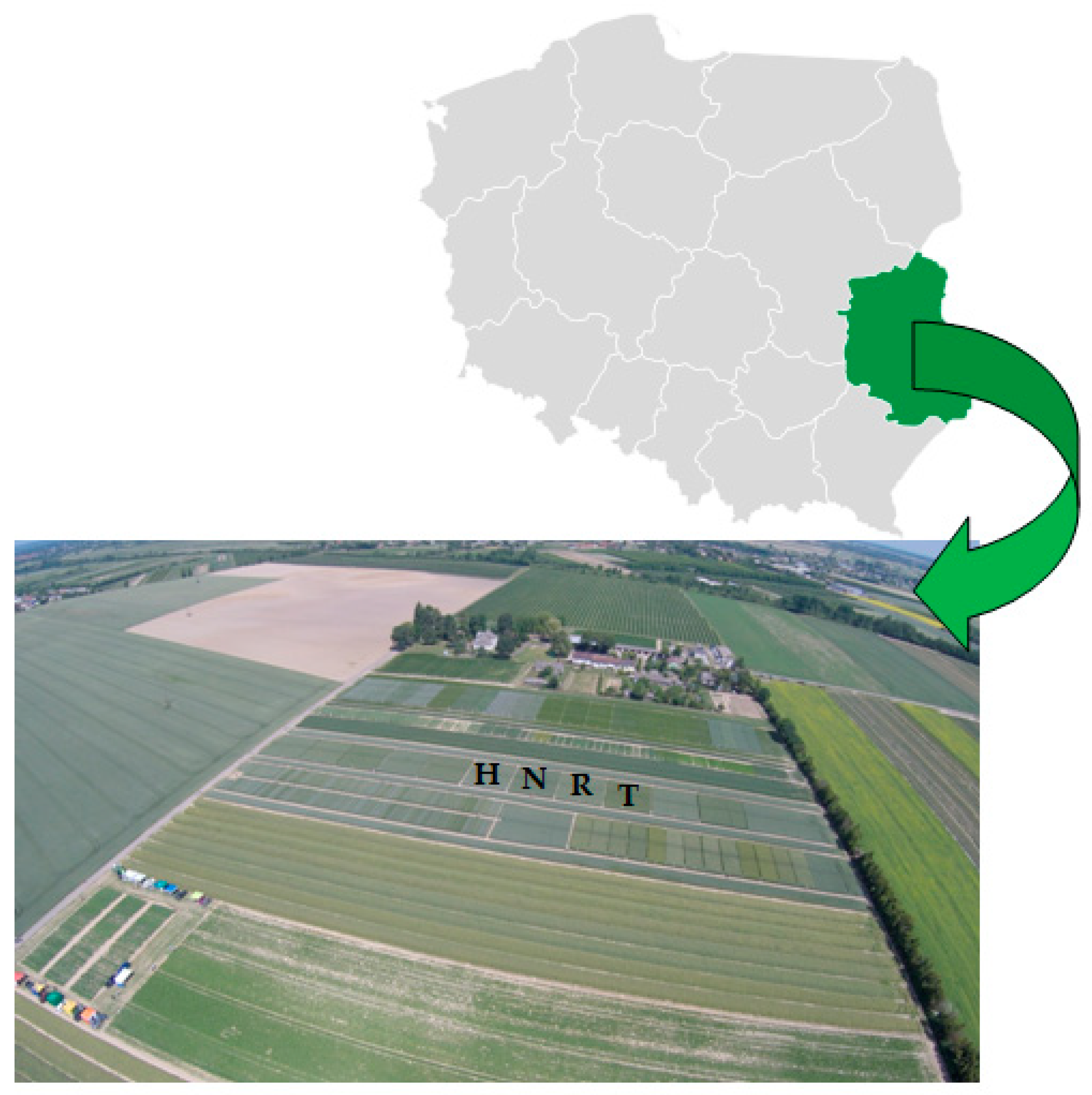
2.1. Soil Sampling and Site Description
2.2. Soil Chemical Properties
2.3. Isolation of DNA and Next Generation Sequencing
2.4. Community Level Physiological Profiling (CLPP) Analysis Using Biolog EcoPlates
2.5. Statistical Analyses
3. Results
3.1. Chemical Properties of Rhizospheric Soils
3.2. Biodiversity in the T. aestivum Rhizosphere—Taxonomic Phylum and Classes Level
3.3. Biodiversity in the T. aestivum Rhizosphere—Taxonomic Genus Level
3.4. Beta-Diversity in the T. aestivum Rhizosphere—Taxonomic Species Level
3.5. Catabolic Activity in the Rhizosphere of Four Selected T. aestivum Cultivars—Community Level Physiological Profiling (CLPP)
3.6. Correlations between Chemical Factors and Dominant Bacterial Phyla Identified in Rhizospheres of Four Wheat Cultivars
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
| Soil before Seeding of Cultivars | pH (H2O) | Eh (mV) | TOC (%) | NH4-N | NO3-N | NO2-N | P-OLSEN |
|---|---|---|---|---|---|---|---|
| mg/kg | |||||||
| Hondia | 5.20 a | 341.24 a | 1.09 a | 1.11 a | 4.06 a | 0.11 a | 3.98 a |
| Nordkap | 6.27 b | 381.88 b | 0.74 b | 0.69 b | 3.07 b | 0.09 a | 3.79 a |
| Rotax | 7.36 c | 347.16 a | 0.76 b | 1.59 c | 4.83 c | 0.26 b | 5.11 b |
| Tytanika | 6.19 b | 399.40 c | 0.60 c | 0.45 d | 4.71 c | 0.08 a | 5.38 b |
| Rhizospheric Soil of Cultivars | Total Reads | Reads Passing Quality Filtering | Remaining Paired-End Assembled Reads | Remaining Filtered Sequences |
|---|---|---|---|---|
| Hondia | 145,364 | 128,179 | 102,543 | 87,161 |
| Nordkap | 245,115 | 213,263 | 168,011 | 123,153 |
| Rotax | 247,580 | 215,385 | 169,774 | 123,304 |
| Tytanika | 237,454 | 211,244 | 166,944 | 117,293 |
| TOTAL | 875,513 | 768,044 | 607,272 | 450,911 |
| Variables | PC1 (69.05%) | PC2 (19.68%) |
| Amines and Amides | −0.590 | −0.169 |
| Aminoacids | −0.839 | −0.051 |
| Carboxylic and acetic acids | −0.959 | 0.275 |
| Hydrocarbons | 0.876 | −0.215 |
| Polymers | 0.489 | −0.782 |
| AWCD | −0.736 | −0.636 |
| Shannon-Wiener | 0.921 | −0.262 |
| Evenness | −0.626 | −0.729 |
| Tytanika | −2.956 | −0.962 |
| Rotax | 2.637 | 0.174 |
| Hondia | −1.045 | 1.817 |
| Nordkap | 1.364 | −1.029 |
| Richness | 0.940 | −0.056 |
| Variables | PC1 (44.69%) | PC2 (27.89%) |
| beta-Methyl-D-Glucoside | 0.209 | 0.872 |
| D-Galactonic Acid gamma-Lactone | −0.199 | −0.742 |
| L-Arginine | −0.900 | 0.113 |
| Pyruvic Acid Methyl Ester | −0.894 | 0.199 |
| D-Xylose | 0.601 | −0.141 |
| D-Galacturonic Acid | −0.905 | −0.301 |
| L-Asparagine | −0.884 | 0.041 |
| Tween 40 | −0.066 | 0.141 |
| i-Erythritol | 0.771 | 0.545 |
| 2-Hydroxy Benzoic Acid | 0.173 | −0.687 |
| L-Phenylalanine | 0.956 | 0.148 |
| Tween 80 | −0.185 | 0.883 |
| D-Mannitol | −0.685 | 0.727 |
| 4-Hydroxy Benzoic Acid | −0.810 | 0.142 |
| L-Serine | −0.585 | 0.563 |
| Alpha-Cyclodextrin | 0.425 | −0.332 |
| N-Acetyl-D-Glucosamine | −0.188 | −0.975 |
| Gamma-Hydroxybutyric Acid | 0.039 | 0.713 |
| L-Threonine | 0.245 | −0.772 |
| Glycogen | 0.239 | −0.753 |
| D-Glucosaminic Acid | 0.181 | −0.961 |
| Itaconic Acid | 0.815 | 0.577 |
| Glycyl-L-Glutamic Acid | −0.288 | 0.177 |
| D-Cellobiose | 0.966 | 0.157 |
| Glucose-1-Phosphate | 0.858 | 0.348 |
| Alpha-Ketobutyric Acid | 0.827 | 0.520 |
| Phenylethylamine | −0.989 | −0.141 |
| Alpha-D-Lactose | 0.782 | −0.141 |
| D,L-alpha-Glycerol Phosphate | −0.844 | −0.167 |
| D-Malic Acid | −0.772 | 0.208 |
| Putrescine | −0.890 | 0.412 |






References
- Roesch, L.F.W.; Fulthorpe, R.R.; Riva, A.; Casella, G.; Hadwin, A.K.M.; Kent, A.D.; Daroub, S.H.; Camargo, F.A.O.; Farmerie, W.G.; Triplett, E.W. Pyrosequencing enumerates and contrasts soil microbial diversity. ISME J. 2007, 1, 283–290. [Google Scholar] [CrossRef] [PubMed]
- Murphy, C.A.; Foster, B.L.; Gao, C. Temporal dynamics in rhizosphere bacterial communities of three perennial grassland species. Agronomy 2016, 6, 17. [Google Scholar] [CrossRef] [Green Version]
- Hassan, M.K.; McInroy, J.A.; Kloepper, J.W. The interactions of rhizodeposits with plant growth-promoting rhizobacteria in the rhizosphere: A review. Agriculture 2019, 9, 142. [Google Scholar] [CrossRef] [Green Version]
- Gałązka, A.; Grzęda, E.; Jończyk, K. Changes of microbial diversity in rhizosphere soils of new quality varieties of winter wheat cultivation in organic farming. Sustainability 2019, 11, 4057. [Google Scholar] [CrossRef] [Green Version]
- Sheirdil, R.A.; Hayat, R.; Zhang, X.X.; Abbasi, N.A.; Ali, S.; Ahmed, M.; Khattak, J.Z.K.; Ahmad, S. Exploring potential soil bacteria for sustainable wheat (Triticum aestivum L.) production. Sustainability 2019, 11, 3361. [Google Scholar] [CrossRef] [Green Version]
- Alawiye, T.T.; Babalola, O.O. Bacterial diversity and community structure in typical plant rhizosphere. Diversity 2019, 11, 179. [Google Scholar] [CrossRef] [Green Version]
- Berg, G.; Smalla, K. Plant species and soil type cooperatively shape the structure and function of microbial communities in the rhizosphere. FEMS Microbiol. Ecol. 2009, 68, 1–13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gregory, P.J. Roots, rhizosphere and soil: The route to a better understanding of soil science? Eur. J. Soil Sci. 2006, 57, 2–12. [Google Scholar] [CrossRef]
- Furtak, K.; Gawryjołek, G.; Gajda, A.M.; Gałązka, A. Effects of maize and winter wheat grown under different cultivation techniques on biological activity of soil. Plant Soil Environ. 2017, 63, 449–454. [Google Scholar] [CrossRef] [Green Version]
- Gałązka, A.; Gawryjołek, K.; Grządziel, J.; Frąc, M.; Księżak, J. Microbial community diversity and the interaction of soil under maize growth in different cultivation techniques. Plant Soil Environ. 2017, 63, 264–270. [Google Scholar] [CrossRef] [Green Version]
- Rosenzweig, N.; Bradeen, J.M.; Tu, Z.J.; McKay, S.J.; Kinkel, L.L. Rhizosphere bacterial communities associated with long-lived perennial prairie plants vary in diversity, composition and structure. Can. J. Microbiol. 2013, 59, 494–502. [Google Scholar] [CrossRef] [Green Version]
- Kowalchuk, G.A.; Buma, D.S.; de Boer, W.; Klinkhamer, P.G.L.; van Veen, J.A. Effects of aboveground plant species composition and diversity on the diversity of soil-borne microorganisms. Antonie Leeuwenhoek 2002, 81, 509–520. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hassan, M.K.; McInroy, J.A.; Jones, J.; Shantharaj, D.; Liles, M.R.; Kloepper, J.W. Pectin-rich amendment enhances soybean growth promotion and nodulation mediated by Bacillus velezensis strains. Plants 2019, 8, 120. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, X.; Zhang, R.; Gao, J.; Wang, X.; Fan, F.; Ma, X.; Yin, H.; Zhang, C.; Feng, K.; Deng, Y. Thirty-one years of rice-rice-green manure rotations shape the rhizosphere microbial community and enrich beneficial bacteria. Soil Biol. Biochem. 2017, 104, 208–217. [Google Scholar] [CrossRef]
- Babalola, O.O. Beneficial bacteria of agricultural importance. Biotechnol. Lett. 2010, 32, 1559–1570. [Google Scholar] [CrossRef] [PubMed]
- Olanrewaju, O.; Glick, B.; Babalola, O. Mechanisms of action of plant growth promoting bacteria. World J. Microbiol. Biotechnol. 2017, 33, 197. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dutta, S.; Surovy, M.Z.; Gupta, D.R.; Mahmud, N.U.; Chanclud, E.; Win, J.; Kamoun, S.; Islam, T. Genomic analyses reveal that biocontrol of wheat blast by Bacillus spp. may be linked with production of antimicrobial compounds and induced systemic resistance in host plants. Figshare 2018. [Google Scholar] [CrossRef]
- Guo, Q.; Li, Y.; Lou, Y.; Shi, M.; Jiang, Y.; Zhou, J.; Sun, Y.; Xue, Q.; Lai, H. Bacillus amyloliquefaciens Ba13 induces plant systemic resistance and improves rhizosphere micro ecology against tomato yellow leaf curlvirus disease. Appl. Soil Ecol. 2019, 137, 154–166. [Google Scholar] [CrossRef]
- Duy, M.; Hoi, N.; Ve, N.; Thuc, L.; Trang, N. Influence of cellulomonas Flavigena, Azospirillum sp. and Pseudomonas sp. on rice growth and yield grown in submerged soil amended in rice straw. Recent Trends PGPR Res. Sustain. Crop Product. 2016, 8, 238–242. [Google Scholar]
- Kumawat, K.; Sharma, P.; Sirari, A.; Singh, I.; Gill, B.; Singh, U.; Saharan, K. Synergism of Pseudomonas aeruginosa (LSE-2) nodule endophyte with Bradyrhizobium sp. (LSBR-3) for improving plant growth, nutrie nt acquisition and soil health in soybean. World J. Microbiol. Biotechnol. 2019, 35, 47. [Google Scholar] [CrossRef]
- Harman, G.E.; Uphoff, N. Symbiotic root-endophytic soil microbes improve crop productivity and provide environmental benefits. Scientifica 2019, 2019, 9106395. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Periera, L.B.; Andrade, G.S.; Meneghin, S.P.; Vicentini, R.; Ottoboni, L.M.M. Prospecting plant-growth prooting bacteria isolated from the rhizosphere sugarcane under drought stress. Curr. Microbiol. 2019, 76, 1345–1354. [Google Scholar] [CrossRef] [PubMed]
- Grządziel, J.; Gałązka, A. Microplot long-term experiment reveals strong soil type influence on bacteria composition and its functional diversity. Appl. Soil Ecol. 2018, 124, 117–123. [Google Scholar] [CrossRef]
- White, R., III; Rivas-Ubach, A.; Borkum, M.; Köberl, M.; Bilbao, A.; Colby, S.; Hoyt, D.; Bingol, K.; Kim, Y.; Wendler, J. The state of rhizospheric science in the era of multi-omics: A practical guide to omics technologies. Rhizosphere 2017, 3, 212–221. [Google Scholar] [CrossRef]
- Yang, R.; Liang, X.; Torrion, J.A.; Walsh, O.S.; O’Brien, K.; Liu, Q. The influence of water and nitrogen availability on the expression of end-use quality parameters of spring wheat. Agronomy 2019, 8, 257. [Google Scholar] [CrossRef] [Green Version]
- Begcy, K.; Weigert, A.; Ogolla Egesa, A.; Dresselhous, T. Compared to Australian cultivars, European summer wheat (Triticum aestivum) overreacts when moderate heat stress is applied as the pollen development stage. Agronomy 2018, 8, 99. [Google Scholar] [CrossRef] [Green Version]
- Shiferaw, B.; Smale, M.; Braun, H.J.; Duveiller, E.; Reynolds, M.; Muricho, G. Crops that feed the world 10. Past successes and future challenges to the role played by wheat in global food security. Food Secur. 2013, 5, 291–317. [Google Scholar] [CrossRef] [Green Version]
- Nelson, A.G.; Frick, B.; Niziol, D.; Clapperton, J.; Spaner, D.; Quideau, S. Spring wheat genotypes differentially alter soil microbial communities and wheat bread making quality in organic and conventional systems. Can. J. Plant Sci. 2011, 91, 485–495. [Google Scholar] [CrossRef]
- Friedrich, T.; Kassam, A. No-till farming and the environment: Do no-till system require more chemicals? Out. Pest Manag. 2012, 8, 153–157. [Google Scholar] [CrossRef]
- Alarcon, R.; Hernandez-Plaza, E.; Navarette, L.; Sanchez, M.J.; Escudero, A.; Hernanz, J.L.; Sanchez-Giron, V.; Sanchez, A.M. Effects of no-tillage and no-inversion tillage on weed community diversity and crop yield over nine years in a Mediterranean cereal-legume cropland. Soil Till. Res. 2018, 179, 54–62. [Google Scholar] [CrossRef]
- Praeg, N.; Pauli, H.; Illmer, P. Microbial diversity in bulk and rhizosphere soil of Ranunculus glacialis along a high alpine altitudinal gradient. Front. Microbiol. 2019, 10, 1429. [Google Scholar] [CrossRef] [PubMed]
- PNR-0431. Chemical, Biological and Agricultural Soil Analyses; Soil Sampling; PKN: Warsaw, Poland, 1997. (In Polish) [Google Scholar]
- Wolińska, A.; Kuźniar, A.; Zielenkiewicz, U.; Banach, A.; Izak, D.; Stępniewska, Z.; Błaszczyk, M. Metagenomic analysis of some potential nitrogen-fixing bacteria in arable soils at different formation processes. Microb. Ecol. 2017, 73, 162–176. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wolińska, A.; Kuźniar, A.; Zielenkiewicz, U.; Izak, D.; Szafranek-Nakonieczna, A.; Banach, A.; Błaszczyk, M. Bacteroidetes as a sensitive indicator of agricultural soil usage revealed by culture-independent approach. Appl. Soil Ecol. 2017, 119, 128–137. [Google Scholar] [CrossRef]
- Wolińska, A.; Kuźniar, A.; Zielenkiewicz, U.; Banach, A.; Błaszczyk, M. Indicators of arable soils fatigue—Bacterial families and genera: A metagenomic approach. Ecol. Ind. 2018, 93, 490–500. [Google Scholar] [CrossRef]
- Thijs, S.; Op De Beeck, M.; Beckers, B.; Truyens, S.; Stevens, V.; Van Hamme, J.D.; Weyens, N.; Vangronsveld, J. Comparative evaluation of four bacteria-specific primer pairs for 16S rRNA gene surveys. Front. Microbiol. 2017, 8, 494. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Kuczyński, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Met. 2010, 7, 335–336. [Google Scholar] [CrossRef] [Green Version]
- DeSantis, T.Z.; Hugenholtz, P.; Larsen, N.; Rojas, M.; Brodie, E.L.; Keller, K.; Dalevi, D.; Hu, P.; Andersen, G.L. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 2006, 72, 5069–5072. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Aronesty, E. ea-Utilis: Command-Line Tools for Processing Biological Sequencing Data. 2011. Available online: http://code.google.com/p/ea-utils (accessed on 4 February 2020).
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef] [Green Version]
- Pohland, B.; Owen, B. Biolog EcoPlates standard methods. TAS Tech. Biul. Biol. 2009, 1, 1–3. [Google Scholar]
- Frac, M.; Oszust, K.; Lipiec, J. Community level physiological profiles (CLPP), characterization and microbial activity of soil amended with dairy sewage sludge. Sensors 2012, 12, 3253–3268. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bucher, R.; Andres, C.; Wedel, M.F.; Entling, M.H.; Nickel, H. Biodiversity in low-intensity pastures, straw meadows, and fallows of a fen area—A multitrophic comparison. Agric. Ecosyst. Environ. 2016, 1, 190–196. [Google Scholar] [CrossRef]
- Wolińska, A.; Frąc, M.; Oszust, K.; Szafranek-Nakonieczna, A.; Zielenkiewicz, U.; Stępniewska, Z. Microbial biodiversity of meadows under different modes of land use: Catabolic and genetic fingerprinting. World J. Microbiol. Biotechnol. 2017, 33, 154. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Swędrzyńska, D.; Grześ, S. Microbiological parameters of soil under sugar beet as a response to the long term-application of different tillage systems. Pol. J. Environ. Stud. 2015, 24, 285–294. [Google Scholar] [CrossRef] [Green Version]
- Wolińska, A.; Rekosz-Burlaga, H.; Goryluk-Salmonowicz, A.; Błaszczyk, M.; Stępniewska, Z. Bacterial abundance and dehydrogenase activity in selected agricultural soils from Lublin region. Pol. J. Environ. Stud. 2015, 24, 2677–2682. [Google Scholar] [CrossRef]
- Wakelin, S.A.; Gupta, V.V.S.R.; Forrester, S.T. Regional and local factors affecting diversity, abundance and activity of free-living, N2-fixing bacteria in Australian agricultural soils. Pedobiologia 2010, 53, 391–399. [Google Scholar] [CrossRef]
- Simon, Z.; Mtei, K.; Gessesse, A.; Ndakidemi, P.A. Isolation and characterization of nitrogen fixing rhizobia from cultivated and uncultivated soils of northern Tanzania. Am. J. Plant Sci. 2014, 5, 4050–4067. [Google Scholar] [CrossRef] [Green Version]
- Romero, I.C.; Jacobson, M.; Fuhrman, J.A.; Fogel, M.; Capone, D.G. Long-term nitrogen and phosphorus fertilization effects on N2 fixation rates and nifH gene community patterns in mangrove sediments. Mar. Ecol. 2012, 33, 117–127. [Google Scholar] [CrossRef]
- Wolińska, A.; Szafranek-Nakonieczna, A.; Banach, A.; Błaszczyk, M.; Stępniewska, Z. The impact of agricultural soil usage on activity and abundance of ammonifying bacteria in selected soils from Poland. SpringerPlus 2016. [Google Scholar] [CrossRef] [Green Version]
- Banach, A.; Wolińska, A.; Błaszczyk, M.; Stępniewska, Z. The influence of soil properties and land use on the phosphate level in soils from Lubelskie region. Pol. J. Agron. 2015, 22, 3–9. [Google Scholar]
- Robben, C.; Fister, S.; Witte, A.K.; Schoder, D.; Rossmanith, P.; Mester, P. Induction of the viable but non-culturable state in bacterial pathogens by household cleaners and inorganic salts. Sci. Rep. 2018, 8, 15132. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Deng, J.; Jing, J.; Zhu, W.; Zhou, Y. Variations in soil bacteria community diversity and structures among different revegetation types in the Baishilazi nature reserve. Front. Microbiol. 2018, 9, 2874. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fierer, N. Embracing the unknown: Disentangling the complexities of the soil microbiome. Nat. Rev. Microbiol. 2017, 15, 579–590. [Google Scholar] [CrossRef] [PubMed]
- Wood, J.L.; Tang, C.; Franks, A.E. Competitive traits are more important than stress-tolerance traits in a cadmium-contaminated rhizosphere: A role for trait theory in microbial ecology. Front. Microbiol. 2018, 9, 121. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Suarez, C.; Ratering, S.; Kramer, I.; Schnell, S. Cellvibrio diazotrophicus sp. nov., a nitrogen-fixing bacteria isolated from the rhizosphere of salt meadow plants and emended description of the genus. Cellvibrio. Int. J. Syst. Evol. Microbiol. 2014, 64, 481–486. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, A.; Kobayashi, Y.O.; Someya, N.; Ikeda, S. Community analysis of root- and tuber-associated bacteria in field-grown potato plants harboring different resistance levels against common scab. Microbes Environ. 2015, 30, 301–309. [Google Scholar] [CrossRef] [Green Version]
- Chung, E.J.; Park, T.S.; Jeon, C.O.; Chung, Y.R. Chitinophaga oryziterrae sp. nov., isolated from the rhizosphere soil of rice (Oryza sativa L.). Int. J. Syst. Evol. Microbiol. 2012, 62, 3030–3035. [Google Scholar] [CrossRef] [Green Version]
- Kavamura, V.N.; Hayat, R.; Clark, I.M.; Rossmann, M.; Mendes, R.; Hirsch, P.R.; Mauchline, T.H. Inorganic nitrogen application affects both taxonomical and predicted functional structure of wheat rhizosphere bacterial communities. Front. Microbiol. 2018, 9, 1074. [Google Scholar] [CrossRef] [Green Version]
- Koberl, M.; Dita, M.; Martinuz, A.; Staver, C.; Berg, G. Members of Gammaproteobacteria as indicator species of healthy banana plants on Fusarium wilt-infested fields in Central America. Sci. Rep. 2017, 7, 45318. [Google Scholar] [CrossRef]
- Koch, I.H.; Gich, F.; Dunfield, P.F.; Overmann, J. Edaphobacter modestus gen. nov., sp. nov., and Edaphobacter aggregans sp. nov., acidobacteria isolated from alpine and forest soils. Int. J. Syst. Evol. Microbiol. 2008, 58, 1114–1122. [Google Scholar] [CrossRef] [Green Version]
- Hong, Y.; Liao, D.; Hu, A.; Wang, H.; Chen, J.; Khan, S.; Su, J.; Li, H. Diversity of endophytic and rhizoplane bacterial communities associated with exotic Spartina alterniflora and native mangrove using Illumina amplicon sequencing. Can. J. Microbiol. 2015, 61, 723–733. [Google Scholar] [CrossRef] [Green Version]
- Beirn, L.A.; Hempfling, J.W.; Schmid, C.J.; Murphy, J.A.; Clarke, B.B.; Crouch, J.O. Differences among soil-inhabiting microbial communities in Poa Anna turf throughout the growing season. Crop Sci. 2017, 57, 262–273. [Google Scholar] [CrossRef]
- Zhang, J.; Wu, D.; Zhang, J.; Liu, Z.; Song, F. Saccharopolyspora shandongensis sp. nov., isolated from wheat-field soil. Int. J. Syst. Evol. Microbiol. 2008, 58, 1094–1099. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rameshkumar, N.; Lang, E.; Tanaka, N. Description of Vogesella oryzae sp. nov., isolated from the rhizosphere of saline tolerant pokkali rice. Syst. Appl. Microbiol. 2016, 39, 20–24. [Google Scholar] [CrossRef] [PubMed]
- Kuźniar, A.; Włodarczyk, K.; Grządziel, J.; Goraj, W.; Gałązka, A.; Wolińska, A. Culture-independent analysis of an endophytic core microbiome in two species of wheat: Triticum aestivum L. (cv. ‘Hondia’) and the first report of microbiota in Triticum spelta L. (cv. ‘Rokosz’). Syst. Appl. Microbiol. 2019. [Google Scholar] [CrossRef]
- Si, P.; Shao, W.; Yu, H.; Yang, X.; Gao, D.; Qiao, X.; Wang, Z.; Wu, G. Rhizosphere microenvironments of eight common deciduous fruit trees were shaped by microbes in Northern China. Front. Microbiol. 2018, 9, 3147. [Google Scholar] [CrossRef]






| T. aestivum Cultivar | pH (H2O) | Eh (mV) | TOC (%) | NH4-N | NO3-N | NO2-N | P-OLSEN |
|---|---|---|---|---|---|---|---|
| mg/kg | |||||||
| Hondia | 3.87 a | 348.73 a | 1.13 a | 1.16 a | 4.01 a | 0.11 a | 4.07 a |
| Nordkap | 6.67 b | 389.60 b | 0.79 b | 0.73 b | 3.11 b | 0.10 a | 3.75 a |
| Rotax | 7.23 c | 350.86 a | 0.71 b | 1.64 c | 4.79 c | 0.24 b | 5.15 b |
| Tytanika | 6.48 b | 403.30 c | 0.64 c | 0.49 d | 4.73 c | 0.09 a | 5.49 b |
| Genera | Triticum aestivum L. Cultivars | |||
|---|---|---|---|---|
| Hondia | Nordkap | Rotax | Tytanika | |
| Aquicella | + | − | − | − |
| Caldithrix | − | + | − | + |
| Cellvibrio | − | − | + | − |
| Chitinophaga | + | − | − | − |
| Chondromyces | − | + | + | + |
| Cuthoniobacter | − | + | + | + |
| Desulfovibrio | − | + | − | − |
| Edaphobacter | + | − | − | + |
| Flavobacterium | − | − | + | − |
| Gemmatimonas | − | − | − | + |
| Janthinobacterium | − | − | + | − |
| Kaistobacter | − | − | + | − |
| Legionella | + | − | − | − |
| Nitrosospira | + | − | − | − |
| Pedobacter | − | − | + | − |
| Pedosphaera | − | + | − | − |
| Rhodoplanes | − | − | − | + |
| Saccharopolyspora | − | − | − | + |
| Segetibacter | + | − | − | − |
| Vogesella | + | + | + | + |
| Cultivar | Average Well Color Development (AWCD) | Shannon-Wiener (H’) | Evenness (E) | Richness (R) |
|---|---|---|---|---|
| Tytanika | 1.300 a ± 0.034 | 3.233 b ± 0.015 | 0.984 a ± 0.006 | 26.67 b ± 0.577 |
| Rotax | 1.267 a ± 0.111 | 3.287 a ± 0.020 | 0.979 b ± 0.005 | 28.87 a ± 0.177 |
| Hondia | 1.266 a ± 0.080 | 3.247 b ± 0.036 | 0.978 b ± 0.007 | 27.67 b ± 0.007 |
| Nordkap | 1.270 a ± 0.128 | 3.300 a ± 0.042 | 0.980 a ± 0.008 | 29.00 a ± 0.126 |
| Factor | Proteobacteria | Bacteroidetes | Firmicutes | Actinobacteria | Acidobacteria | Verrucomicrobia |
|---|---|---|---|---|---|---|
| pH | 0.602 * | −0.574 * | −0.400 | 0.614 * | −0.853 *** | 0.915 *** |
| Eh | −0.378 | −0.977 *** | 0.659 ** | 0.332 | 0.120 | 0.199 |
| TOC | −0.252 | 0.720 ** | 0.166 | −0.806 ** | 0.663 * | −0.687 * |
| NH4-N | 0.692 * | 0.800 ** | −0.912 *** | −0.061 | −0.536 | 0.208 |
| NO3-N | −0.099 | 0.173 | −0.445 | 0.839 *** | −0.362 | −0.116 |
| NO2-N | 0.729 ** | 0.613 * | −0.947 *** | 0.176 | −0.699 * | 0.366 |
| P-Olsen | −0.093 | −0.177 | −0.317 | 0.975 *** | −0.449 | 0.091 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wolińska, A.; Kuźniar, A.; Gałązka, A. Biodiversity in the Rhizosphere of Selected Winter Wheat (Triticum aestivum L.) Cultivars—Genetic and Catabolic Fingerprinting. Agronomy 2020, 10, 953. https://doi.org/10.3390/agronomy10070953
Wolińska A, Kuźniar A, Gałązka A. Biodiversity in the Rhizosphere of Selected Winter Wheat (Triticum aestivum L.) Cultivars—Genetic and Catabolic Fingerprinting. Agronomy. 2020; 10(7):953. https://doi.org/10.3390/agronomy10070953
Chicago/Turabian StyleWolińska, Agnieszka, Agnieszka Kuźniar, and Anna Gałązka. 2020. "Biodiversity in the Rhizosphere of Selected Winter Wheat (Triticum aestivum L.) Cultivars—Genetic and Catabolic Fingerprinting" Agronomy 10, no. 7: 953. https://doi.org/10.3390/agronomy10070953







