Development and Yield Traits Indicate That the Constitutive Wound Response Phenotype of Prosystemin Overexpressing Tomato Plants Entails No Fitness Penalty
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Material
2.2. Evaluation of Plants during Vegetative and Reproductive Growth
2.3. Evaluation of Fruit Quality
2.4. Non-Structural Carbohydrate Content in Fruits
2.5. Targeted Fruit Metabolomic Analysis
2.6. Extraction of Total RNA, cDNA Preparation and qPCR Analysis
2.7. Statistical Analysis
3. Results
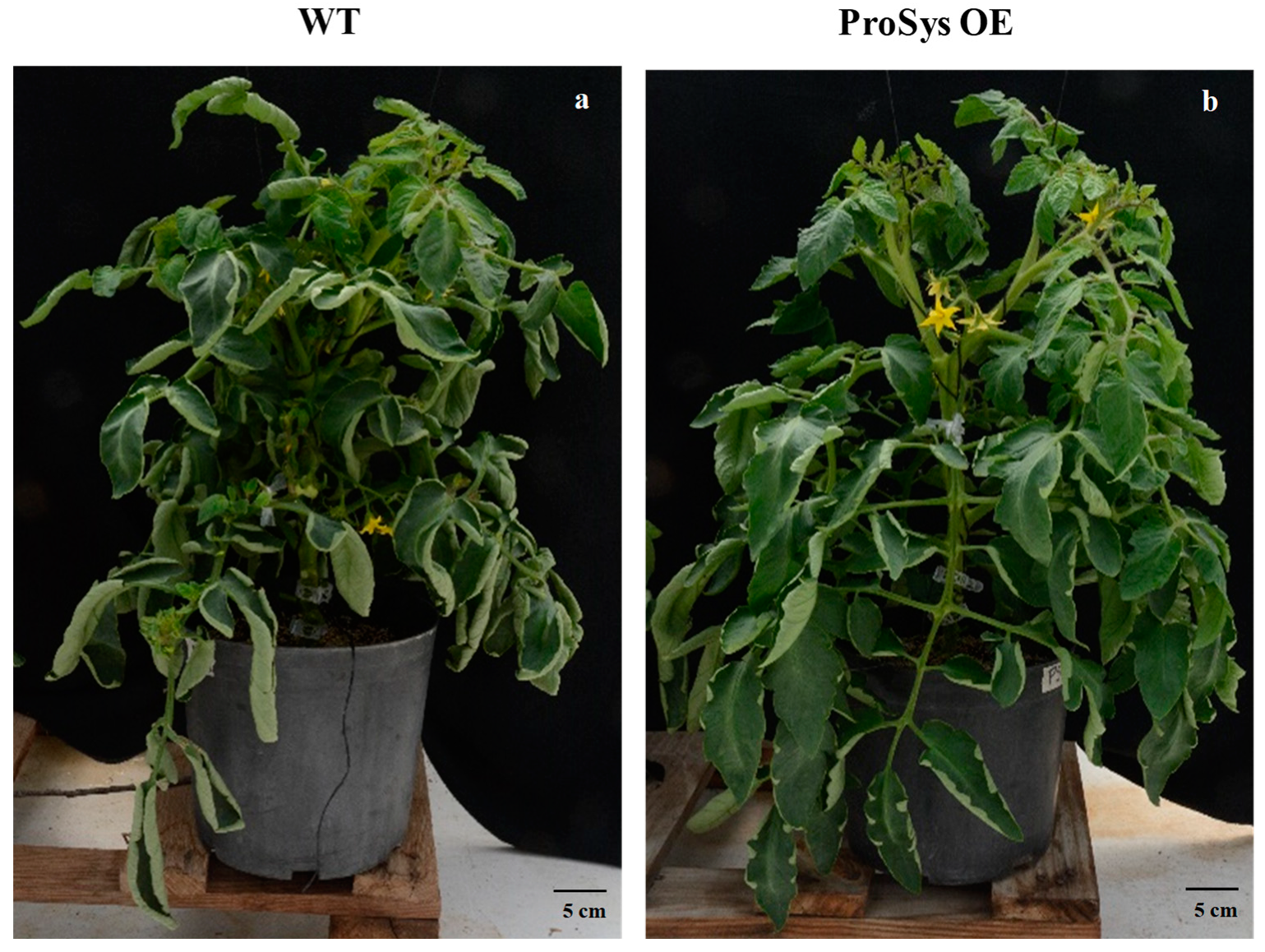
3.1. Plant Growth Rate, Onset of Flowering and Time for Full Fruit Maturation
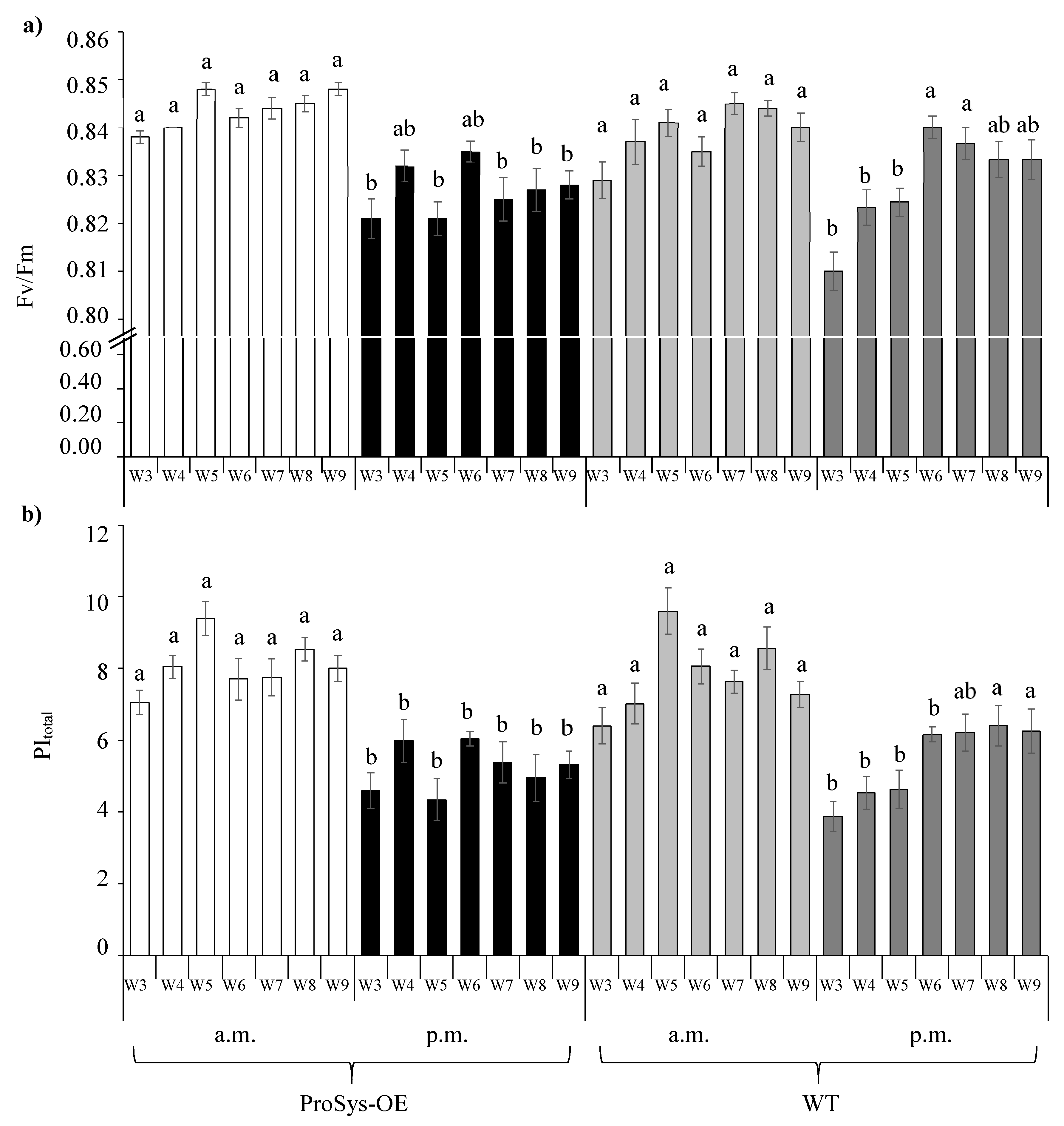
3.2. Photosynthetic Efficiency Parameters
3.3. Fruit Physicochemical Parameters
3.4. Quantitative Gene Expression Profiles
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Pearce, G.; Strydom, D.; Johnson, S.; Ryan, C.A. A polypeptide from tomato leaves induces wound-inducible proteinase inhibitor proteins. Science 1991, 253, 895–897. [Google Scholar] [CrossRef] [PubMed]
- McGurl, B.; Pearce, G.; Orozco-Cardenas, M.; Ryan, C.A. Structure, expression, and antisense inhibition of the systemin precursor gene. Science 1992, 255, 1570–1573. [Google Scholar] [CrossRef] [PubMed]
- McGurl, B.; Orozco-Cardenas, M.; Pearce, G.; Ryan, C.A. Overexpression of the prosystemin gene in transgenic tomato plants generates a systemic signal that constitutively induces proteinase inhibitor synthesis. Proc. Natl. Acad. Sci. USA 1994, 91, 9799–9802. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beloshistov, R.E.; Dreizler, K.; Galiullina, R.A.; Tuzhikov, A.I.; Serebryakova, M.V.; Reichardt, S.; Shaw, J.; Taliansky, M.E.; Pfannstiel, J.; Chichkova, N.V.; et al. Phytaspase-mediated precursor processing and maturation of the wound hormone systemin. New Phytol. 2018, 218, 1167–1178. [Google Scholar] [CrossRef] [PubMed]
- Buonanno, M.; Coppola, M.; Di Lelio, I.; Molisso, D.; Leone, M.; Pennacchio, F.; Langella, E.; Rao, R.; Monti, S.M. Prosystemin, a prohormone that modulates plant defense barriers, is an intrinsically disordered protein. Protein Sci. 2018, 27, 620–632. [Google Scholar] [CrossRef]
- Ryan, C.A.; Pearce, G. Systemins: A functionally defined family of peptide signals that regulate defensive genes in Solanaceae species. Proc. Natl. Acad. Sci. USA 2003, 100, 14577–14580. [Google Scholar] [CrossRef] [Green Version]
- Ryan, C.A.; Moura, D.S. Systemic wound signaling in plants: A new perception. Proc. Natl. Acad. Sci. USA 2002, 99, 6519–6520. [Google Scholar] [CrossRef] [Green Version]
- Howe, G.A.; Schaller, A. Direct defenses in plants and their induction by wounding and insect herbivores. In Induced Plant Resistance against Herbivory; Schaller, A., Ed.; Springer: Heidelberg, Germany, 2008; pp. 7–29. [Google Scholar]
- Wang, L.; Einig, E.; Almeida-Trapp, M.; Albert, M.; Fliegmann, J.; Mithofer, A.; Kalbacher, H.; Felix, G. The systemin receptor SYR1 enhances resistance of tomato against herbivorous insects. Nat. Plants 2018, 4, 152–156. [Google Scholar] [CrossRef]
- Felix, G.; Boller, T. Systemin induces rapid ion fluxes and ethylene biosynthesis in Lycopersicon peruvianum cells. Plant J. 1995, 7, 381–389. [Google Scholar] [CrossRef]
- Moyen, C.; Hammond-Kosack, K.E.; Jones, J.; Knight, M.R.; Johannes, E. Systemin triggers an increase of cytoplasmic calcium in tomato mesophyll cells: Ca2+ mobilization from intra- and extracellular compartments. Plant Cell Environ. 1998, 21, 1101–1111. [Google Scholar] [CrossRef]
- Schaller, A.; Oecking, C. Modulation of plasma membrane H+-ATPase activity differentially activates wound and pathogen defense responses in tomato plants. Plant Cell 1999, 11, 263–272. [Google Scholar]
- Ahmad, F.H.; Wu, X.; Stintzi, A.; Schaller, A.; Schulze, W.X. The systemin signaling cascade as derived from time course analyses of the systemin-responsive phosphoproteome. Mol. Cell Proteom. 2019, 18, 1526–1542. [Google Scholar] [CrossRef] [Green Version]
- Hou, S.; Liu, Z.; Shen, H.; Wu, D. Damage-associated molecular pattern-triggered immunity in plants. Front. Plant Sci. 2019, 10, 646. [Google Scholar] [CrossRef]
- Narváez-Vásquez, J.; Ryan, C.A. The cellular localization of prosystemin: A functional role for phloem parenchyma in systemic wound signaling. Planta 2004, 218, 360–369. [Google Scholar]
- Orozco-Cardenas, M.; McGurl, B.; Ryan, C.A. Expression of an antisense prosystemin gene in tomato plants reduces resistance toward Manduca sexta larvae. Proc. Natl. Acad. Sci. USA 1993, 90, 8273–8276. [Google Scholar] [CrossRef] [Green Version]
- Diaz, J.; ten Have, A.; van Kan, J.A. The role of ethylene and wound signaling in resistance of tomato to Botrytis cinerea. Plant Physiol. 2002, 129, 1341–1351. [Google Scholar] [CrossRef] [Green Version]
- El Oirdi, M.; El Rahman, T.A.; Rigano, L.; El Hadrami, A.; Rodriguez, M.C.; Daayf, F.; Vojnov, A.; Bouarab, K. Botrytis cinerea manipulates the antagonistic effects between immune pathways to promote disease development in tomato. Plant Cell 2011, 23, 2405–2421. [Google Scholar] [CrossRef] [Green Version]
- Coppola, M.; Corrado, G.; Coppola, V.; Cascone, P.; Martinelli, R.; Digilio, M.C.; Pennacchio, F.; Rao, R. Prosystemin overexpression in tomato enhances resistance to different biotic stresses by activating genes of multiple signaling pathways. Plant Mol. Biol. Rep. 2015, 33, 1270–1285. [Google Scholar] [CrossRef] [Green Version]
- Bubici, G.; Carluccio, A.V.; Stavolone, L.; Cillo, F. Prosystemin overexpression induces transcriptional modifications of defense-related and receptor-like kinase genes and reduces the susceptibility to Cucumber mosaic virus and its satellite RNAs in transgenic tomato plants. PLoS ONE 2017, 12, e0171902. [Google Scholar] [CrossRef]
- Corrado, G.; Sasso, R.; Pasquariello, M.; Iodice, L.; Carretta, A.; Cascone, P.; Ariati, L.; Digilio, M.; Guerrieri, E.; Rao, R. Systemin regulates both systemic and volatile signaling in tomato plants. J. Chem. Ecol. 2007, 33, 669–681. [Google Scholar] [CrossRef]
- Coppola, M.; Cascone, P.; Madonna, V.; Di Lelio, I.; Esposito, F.; Avitabile, C.; Romanelli, A.; Guerrieri, E.; Vitiello, A.; Pennacchio, F.; et al. Plant-to-plant communication triggered by systemin primes anti-herbivore resistance in tomato. Sci. Rep. 2017, 7, 15522. [Google Scholar] [CrossRef] [PubMed]
- Dombrowski, J.E. Salt stress activation of wound-related genes in tomato plants. Plant Physiol. 2003, 132, 2098–2107. [Google Scholar] [CrossRef] [Green Version]
- Orsini, F.; Cascone, P.; De Pascale, S.; Barbieri, G.; Corrado, G.; Rao, R.; Maggio, A. Systemin-dependent salinity tolerance in tomato: Evidence of specific convergence of abiotic and biotic stress responses. Physiol. Plant. 2010, 138, 10–21. [Google Scholar] [CrossRef]
- Coppola, M.; Di Lelio, I.; Romanelli, A.; Gualtieri, L.; Molisso, D.; Ruocco, M.; Avitabile, C.; Natale, R.; Cascone, P.; Guerrieri, E.; et al. Tomato plants treated with systemin peptide show enhanced levels of direct and indirect defense associated with increased expression of defense-related genes. Plants 2019, 8, 395. [Google Scholar] [CrossRef] [Green Version]
- Valverde, P.L.; Fornoni, J.; Nunez-Farfan, J. Evolutionary ecology of Datura stramonium: Equal plant fitness benefits of growth and resistance against herbivory. J. Evol. Biol. 2003, 16, 127–137. [Google Scholar] [CrossRef] [PubMed]
- Heidel, A.J.; Clarke, J.D.; Antonovics, J.; Dong, X.N. Fitness costs of mutations affecting the systemic acquired resistance pathway in Arabidopsis thaliana. Genetics 2004, 168, 2197–2206. [Google Scholar] [CrossRef] [Green Version]
- Huot, B.; Yao, J.; Montgomery, B.L.; He, S.Y. Growth-defense tradeoffs in plants: A balancing act to optimize fitness. Mol. Plant 2014, 7, 1267–1287. [Google Scholar] [CrossRef] [Green Version]
- Machado, R.A.R.; Baldwin, I.T.; Erb, M. Herbivory-induced jasmonates constrain plant sugar accumulation and growth by antagonizing gibberellin signaling and not by promoting secondary metabolite production. New Phytol. 2017, 215, 803–812. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Simms, E.L.; Rausher, M.D. Costs and benefits of plant resistance to herbivory. Am. Nat. 1987, 130, 570–581. [Google Scholar] [CrossRef]
- Herms, D.A.; Mattson, W.J. The dilemma of plants: To grow or defend. Q. Rev. Biol. 1992, 67, 283–335. [Google Scholar] [CrossRef] [Green Version]
- Heil, M.; Baldwin, I.T. Fitness costs of induced resistance: Emerging experimental support for a slippery concept. Trends Plant Sci. 2002, 7, 61–67. [Google Scholar] [CrossRef]
- Vos, I.A.; Pieterse, C.M.J.; Van Wees, S.C.M. Costs and benefits of hormone-regulated plant defences. Plant Pathol. 2013, 62, 43–55. [Google Scholar] [CrossRef]
- Cipollini, D.; Walters, D.; Voelckel, C. Costs of resistance in plants: From theory to evidence. In Annual Plant Reviews: Insect-Plant Interactions; Voelckel, C., Jander, G., Eds.; John Wiley & Sons: Hoboken, NJ, USA, 2014; pp. 263–307. [Google Scholar]
- Karasov, T.L.; Chae, E.; Herman, J.J.; Bergelson, J. Mechanisms to mitigate the trade-off between growth and defense. Plant Cell 2017, 29, 666–680. [Google Scholar] [CrossRef] [Green Version]
- Züst, T.; Agrawal, A.A. Trade-offs between plant growth and defense against insect herbivory: An emerging mechanistic synthesis. Annu. Rev. Plant Biol. 2017, 68, 513–534. [Google Scholar] [CrossRef] [Green Version]
- Bowling, S.A.; Guo, A.; Cao, H.; Gordon, A.S.; Klessig, D.F.; Dong, X. A mutation in Arabidopsis that leads to constitutive expression of systemic acquired resistance. Plant Cell 1994, 6, 1845–1857. [Google Scholar]
- Heil, M.; Hilpert, A.; Kaiser, W.; Linsenmair, K.E. Reduced growth and seed set following chemical induction of pathogen defence: Does systemic acquired resistance (SAR) incur allocation costs? J. Ecol. 2000, 88, 645–654. [Google Scholar] [CrossRef]
- Li, X.; Clarke, J.D.; Zhang, Y.; Dong, X. Activation of an EDS1-mediated R-gene pathway in the snc1 mutant leads to constitutive, NPR1- independent pathogen resistance. Mol. Plant Microbe Interact. 2001, 14, 1131–1139. [Google Scholar] [CrossRef] [Green Version]
- Redman, A.M.; Cipollini, D.F.; Schultz, J.C. Fitness costs of jasmonic acid-induced defense in tomato, Lycopersicon esculentum. Oecologia 2001, 126, 380–385. [Google Scholar] [CrossRef]
- Zavala, J.A.; Patankar, A.G.; Gase, K.; Baldwin, I.T. Constitutive and inducible trypsin proteinase inhibitor production incurs large fitness costs in Nicotiana attenuata. Proc. Natl. Acad. Sci. USA 2004, 101, 1607–1612. [Google Scholar] [CrossRef] [Green Version]
- Ballaré, C.L. Light regulation of plant defense. Annu. Rev. Plant Biol. 2014, 65, 335–363. [Google Scholar] [CrossRef]
- Havko, N.E.; Major, I.T.; Jewell, J.B.; Attaran, E.; Browse, J.; Howe, G.A. Control of carbon assimilation and partitioning by jasmonate: An accounting of growth-defense tradeoffs. Plants 2016, 5, 7. [Google Scholar] [CrossRef] [Green Version]
- Campos, M.L.; Yoshida, Y.; Major, I.T.; de Oliveira Ferreira, D.; Weraduwage, S.M.; Froehlich, J.E.; Johnson, B.F.; Kramer, D.M.; Jander, G.; Sharkey, T.D.; et al. Rewiring of jasmonate and phytochrome B signalling uncouples plant growth-defense tradeoffs. Nat. Commun. 2016, 7, 12570. [Google Scholar] [CrossRef] [Green Version]
- Kliebenstein, D.J. False idolatry of the mythical growth versus immunity trade-off in molecular systems plant pathology. Physiol. Mol. Plant Pathol. 2016, 95, 55–59. [Google Scholar] [CrossRef] [Green Version]
- Guo, Q.; Major, I.T.; Howe, G.A. Resolution of growth-defense conflict: Mechanistic insights from jasmonate signaling. Curr. Opin. Plant Biol. 2018, 44, 72–81. [Google Scholar] [CrossRef]
- Ballaré, C.L.; Austin, A.T. Recalculating growth and defense strategies under competition: Key roles of photoreceptors and jasmonates. J. Exp. Bot. 2019, 70, 3425–3434. [Google Scholar] [CrossRef] [PubMed]
- Savchenko, T.V.; Rolletschek, H.; Dehesh, K. Jasmonates-mediated rewiring of central metabolism regulates adaptive responses. Plant Cell Physiol. 2019, 60, 2613–2620. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Milmanda, G.L.; Crocco, C.D.; Reichelt, M.; Mazza, C.A.; Köllner, T.G.; Zhang, T.; Cargnel, M.D.; Lichy, M.Z.; Fiorucci, A.S.; Fankhauser, C.; et al. A light-dependent molecular link between competition cues and defence responses in plants. Nat. Plants 2020, 6, 223–230. [Google Scholar] [CrossRef] [Green Version]
- Corrado, G.; Agrelli, D.; Rocco, M.; Basile, B.; Marra, M.; Rao, R. Systemin-inducible defence against pests is costly in tomato. Biol. Plant. 2011, 55, 305–311. [Google Scholar] [CrossRef]
- Ryan, C.A.; Pearce, G. Systemin: A polypeptide signal for plant defensive genes. Annu. Rev. Cell Dev. Biol. 1998, 14, 1–17. [Google Scholar] [CrossRef] [Green Version]
- Schilmiller, A.L.; Howe, G.A. Systemic signaling in the wound response. Curr. Opin. Plant Biol. 2005, 8, 369–377. [Google Scholar] [CrossRef]
- Metodiev, M.V.; Tsonev, T.D.; Popova, L.P. Effect of jasmonic acid on the stomatal and non-stomatal limitation of leaf photosynthesis in barley leaves. J. Plant Growth Regul. 1996, 15, 75–80. [Google Scholar] [CrossRef]
- Herde, O.; Pena-Cortes, H.; Willmitzer, L.; Fisahn, J. Stomatal responses to jasmonic acid, linolenic acid and abscisic acid in wild-type and ABA-deficient tomato plants. Plant Cell Environ. 1997, 20, 136–141. [Google Scholar] [CrossRef]
- Biber, P.D. Evaluating a chlorophyll content meter on three coastal wetland plant species. J. Agric. Food Environ. Sci. 2007, 24, 1–15. [Google Scholar]
- CIE. CIE 15: Technical Report Colorimetry, 3rd ed.; International Commission on Illumination (CIE): Vienna, Austria, 2004; ISBN 3 901 906 33 9. [Google Scholar]
- 942.15. Acidity (Titratable) of Fruit Products. First Action 1942; AOAC, Association of Official Analytical Chemists: Washington, DC, USA, 1990.
- Wright, D.P.; Scholes, J.D.; Read, D.J. Effects of VA mycorrhizal colonization on photosynthesis and biomass production of Trifolium repens L. Plant Cell Environ. 1998, 21, 209–216. [Google Scholar] [CrossRef]
- Tiessen, A.; Hendriks, J.H.M.; Stitt, M.; Branscheid, A.; Gibon, Y.; Farré, E.M.; Geigenberger, P. Starch synthesis in potato tubers is regulated by post-translational redox modification of ADP-glucose pyrophosphorylase: A novel regulatory mechanism linking starch synthesis to the sucrose supply. Plant Cell 2002, 14, 2191–2213. [Google Scholar] [CrossRef]
- Ramírez-Briones, E.; Rodríguez-Macías, R.; Salcedo-Pérez, E.; Ramírez-Chávez, E.; Molina-Torres, J.; Tiessen, A.; Ordaz-Ortiz, J.; Martínez-Gallardo, N.; Délano-Frier, J.P.; Zañudo-Hernández, J. Seasonal changes in the metabolic profiles and biological activity in leaves of Diospyros digyna and D. rekoi “Zapote” trees. Plants 2019, 8, 449. [Google Scholar] [CrossRef] [Green Version]
- Casarrubias-Castillo, K.; Montero-Vargas, J.M.; Dabdoub-González, N.; Winkler, R.; Martinez-Gallardo, N.A.; Zañudo-Hernández, J.; Avilés-Arnaut, H.; Délano-Frier, J.P. Distinct gene expression and secondary metabolite profiles in suppressor of prosystemin-mediated responses2 (spr2) tomato mutants having impaired mycorrhizal colonization. PeerJ 2020, 8, e8888. [Google Scholar] [CrossRef] [Green Version]
- Mueller, L.A.; Solow, T.H.; Taylor, N.; Skwarecki, B.; Buels, R.; Binns, J.; Lin, C.; Wright, M.H.; Ahrens, R.; Wang, Y.; et al. The SOL genomics network: A comparative resource for Solanaceae biology and beyond. Plant Physiol. 2005, 138, 1310–1317. [Google Scholar] [CrossRef] [Green Version]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Expósito-Rodríguez, M.; Borges, A.A.; Borges-Pérez, A.; Pérez, J.A. Selection of internal control genes for quantitative real-time RT-PCR studies during tomato development process. BMC Plant Biol. 2008, 8, 131. [Google Scholar] [CrossRef] [Green Version]
- Baker, N.R.; Rosenqvist, E. Applications of chlorophyll fluorescence can improve crop production strategies: An examination of future possibilities. J. Exp. Bot. 2004, 55, 1607–1621. [Google Scholar] [CrossRef] [Green Version]
- Çiçek, N.; Arslan, O.; Çulha-Erdal, S.; Eyidoğan, F.; Ekmekçi, Y. Are the photosynthetic performance indexes and the drought factor index satisfactory selection criterion for stress? Fresenius Environ. Bull. 2015, 24, 4190–4198. [Google Scholar]
- Strasser, R.J.; Srivastava, A.; Tsimilli-Michael, M. The fluorescence transient as a tool to characterize and screen photosynthetic samples. In Probing Photosynthesis: Mechanism, Regulation and Adaptation; Yunus, M., Pathre, U., Mohanty, P., Eds.; Taylor and Francis: London, UK, 2000; pp. 443–480. [Google Scholar]
- Tsimilli-Michael, M.; Strasser, R.J. In vivo assessment of stress impact on plants’ vitality: Applications in detecting and evaluating the beneficial role of mycorrhization on host plants. In Mycorrhiza: State of the Art, Genetics and Molecular Biology, Eco-Function, Biotechnology, Eco-Physiology, Structure and Systematics; Varma, A., Ed.; Springer: Berlin, Germany, 2008; pp. 679–703. [Google Scholar]
- Aoun, A.B.; Lechiheb, B.; Benyahya, L.; Ferchichi, A. Evaluation of fruit quality traits of traditional varieties of tomato (Solanum lycopersicum) grown in Tunisia. Afr. J. Food Sci. 2013, 7, 350–354. [Google Scholar] [CrossRef] [Green Version]
- Ryan, C.A. The systemin signaling pathway: Differential activation of plant defensive genes. Biochim. Biophys. Acta 2000, 1477, 112–121. [Google Scholar] [CrossRef]
- Farmer, E.E.; Ryan, C.A. Octadecanoid precursors of jasmonic acid activate the synthesis of wound-inducible protease inhibitors. Plant Cell 1992, 4, 129–134. [Google Scholar] [CrossRef]
- Narváez-Vásquez, J.; Florin-Christensen, J.; Ryan, C.A. Positional specificity of a phospholipase A activity induced by wounding, systemin, and oligosaccharide elicitors in tomato leaves. Plant Cell 1999, 11, 2249–2260. [Google Scholar] [CrossRef] [Green Version]
- Chung, H.S.; Niu, Y.; Browse, J.; Howe, G.A. Top hits in contemporary JAZ: An update on jasmonate signaling. Phytochemistry 2009, 70, 1547–1559. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Seo, H.; Song, J.; Cheong, J.J.; Lee, Y.H.; Lee, Y.W.; Hwang, I.; Lee, J.S.; Choi, Y.D. Jasmonic acid carboxyl methyltransferase: A key enzyme for jasmonate-regulated plant responses. Proc. Natl. Acad. Sci. USA 2001, 98, 4788–4793. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bergey, D.R.; Orozco-Cardenas, M.; de Moura, D.S.; Ryan, C.A. A wound- and systemin-inducible polygalacturonase in tomato leaves. Proc. Natl. Acad. Sci. USA 1999, 96, 1756–1760. [Google Scholar] [CrossRef] [Green Version]
- Vellosillo, T.; Martínez, M.; López, M.A.; Vicente, J.; Cascón, T.; Dolan, L.; Hamberg, M.; Castresana, C. Oxylipins produced by the 9-lip- oxygenase pathway in Arabidopsis regulate lateral root development and defense responses through a specific signaling cascade. Plant Cell 2007, 19, 831–846. [Google Scholar] [CrossRef] [Green Version]
- Vicente, J.; Cascón, T.; Vicedo, B.; García-Agustín, P.; Hamberg, M.; Castresana, C. Role of 9-lipoxygenase and α-dioxygenase oxylipin pathways as modulators of local and systemic defense. Mol. Plant 2012, 5, 914–928. [Google Scholar] [CrossRef] [Green Version]
- Upadhyay, R.K.; Handa, A.K.; Mattoo, A.K. Transcript abundance patterns of 9- and 13-lipoxygenase subfamily gene members in response to abiotic stresses (heat, cold, drought or salt) in tomato (Solanum lycopersicum L.) highlights member-specific dynamics relevant to each stress. Genes 2019, 10, 683. [Google Scholar] [CrossRef] [Green Version]
- Schouten, R.E.; Huijben, T.P.M.; Tijskens, L.M.M.; van Kooten, O. Modelling quality attributes of truss tomatoes: Linking colour and firmness maturity. Postharvest Biol. Technol. 2007, 45, 298–306. [Google Scholar] [CrossRef]
- Li, L.; Zhao, W.; Feng, X.; Chen, L.; Zhang, L.; Zhao, L. Changes in fruit firmness, cell wall composition, and transcriptional profile in the yellow fruit tomato 1 (yft1) mutant. J. Agric. Food Chem. 2019, 67, 463–472. [Google Scholar] [CrossRef]
- White, P.J.; Broadley, M.R. Calcium in plants. Ann. Bot. 2003, 92, 487–511. [Google Scholar] [CrossRef]
- Attaran, E.; Major, I.T.; Cruz, J.A.; Rosa, B.A.; Koo, A.J.; Chen, J.; Kramer, D.M.; He, S.Y.; Howe, G.A. Temporal dynamics of growth and photosynthesis suppression in response to jasmonate signaling. Plant Physiol. 2014, 165, 1302–1314. [Google Scholar] [CrossRef] [Green Version]
- Katsir, L.; Chung, H.S.; Koo, A.J.K.; Howe, G.A. Jasmonate signaling: A conserved mechanism of hormone sensing. Curr. Opin. Plant Biol. 2008, 11, 428–435. [Google Scholar] [CrossRef] [Green Version]
- Ruan, J.; Zhou, Y.; Zhou, M.; Yan, J.; Khurshid, M.; Weng, W.; Cheng, J.; Zhang, K. Jasmonic acid signaling pathway in plants. Int. J. Mol. Sci. 2019, 20, 2479. [Google Scholar] [CrossRef] [Green Version]
- Yang, J.; Duan, G.; Li, C.; Liu, L.; Han, G.; Zhang, Y.; Wang, C. The crosstalks between jasmonic acid and other plant hormone signaling highlight the involvement of jasmonic acid as a core component in plant response to biotic and abiotic stresses. Front. Plant Sci. 2019, 10, 1349. [Google Scholar] [CrossRef] [Green Version]
- Saniewski, M.; Czapski, J. The effect of methyl jasmonate on lycopene and β-carotene accumulation in ripening red tomatoes. Experientia 1983, 39, 1373–1374. [Google Scholar] [CrossRef]
- Saniewski, M.; Nowacki, J.; Czapski, J. The effect of methyl jasmonate on ethylene production and ethylene-forming enzyme activity in tomatoes. J. Plant Physiol. 1987, 129, 175–180. [Google Scholar] [CrossRef]
- Czapski, J.; Saniewski, M. Stimulation of ethylene production and ethylene-forming enzyme activity in fruits of the non-ripening nor and rin tomato mutants by methyl jasmonate. J. Plant Physiol. 1992, 139, 265–268. [Google Scholar] [CrossRef]
- Yilmaz, H.; Yildiz, K.; Muradoǧlu, F. Effect of jasmonic acid on yield and quality of two strawberry cultivars. J. Am. Pomol. Soc. 2003, 57, 32–35. [Google Scholar]
- Concha, C.M.; Figueroa, N.E.; Poblete, L.A.; Oñate, F.A.; Schwab, W.; Figueroa, C.R. Methyl jasmonate treatment induces changes in fruit ripening by modifying the expression of several ripening genes in Fragaria chiloensis fruit. Plant Physiol. Biochem. 2013, 70, 433–444. [Google Scholar] [CrossRef] [PubMed]
- Serrano, M.; Martínez-Esplá, A.; Zapata, P.; Castillo, S.; Martínez-Romero, D.; Guillén, F.; Valverde, J.M.; Valero, D. Effects of methyl jasmonate treatment on fruit quality properties. In Emerging Postharvest Treatment of Fruits and Vegetables; Barman, K., Sharma, S., Siddiqui, M.W., Eds.; Apple Academic Press: Palm Bay, FL, USA, 2018; pp. 85–106. [Google Scholar]
- Hanaka, A.; Nurzyńska-Wierdak, R. Methyl jasmonate—A multifunctional molecule throughout the whole plant life. Acta Sci. Pol. Hortorum Cultus 2019, 18, 237–249. [Google Scholar] [CrossRef]
- Zuñiga, P.E.; Castañeda, Y.; Arrey-Salas, O.; Fuentes, L.; Aburto, F.; Figueroa, C.R. Methyl jasmonate applications from flowering to ripe fruit stages of strawberry (Fragaria × ananassa ‘Camarosa’) reinforce the fruit antioxidant response at post-harvest. Front. Plant Sci. 2020, 11, 538. [Google Scholar] [CrossRef] [PubMed]
- Major, I.T.; Guo, Q.; Zhai, J.; Kapali, G.; Kramer, D.M.; Howe, G.A. A phytochrome B-independent pathway restricts growth at high levels of jasmonate defense. Plant Physiol. 2020, 183, 733–749. [Google Scholar] [CrossRef] [Green Version]
- Chung, H.S.; Koo, A.J.; Gao, X.; Jayanty, S.; Thines, B.; Jones, A.D.; Howe, G.A. Regulation and function of Arabidopsis JASMONATE ZIM domain genes in response to wounding and herbivory. Plant Physiol. 2008, 146, 952–964. [Google Scholar] [CrossRef] [Green Version]
- Narváez-Vásquez, J.; Ryan, C.A. The systemin precursor gene regulates both defensive and developmental genes in Solanum tuberosum. Proc. Natl. Acad. Sci. USA 2002, 99, 15818–15821. [Google Scholar] [CrossRef] [Green Version]
- Pastor, V.; Sanchez-Bel, P.; Gamir, J.; Pozo, M.J.; Flors, V. Accurate and easy method for systemin quantification and examining metabolic changes under different endogenous levels. Plant Methods 2018, 14, 33. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tompa, P. Intrinsically unstructured proteins. Trends Biochem. Sci. 2002, 27, 527–533. [Google Scholar] [CrossRef]
- Oldfield, C.J.; Meng, J.; Yang, J.Y.; Yang, M.Q.; Uversky, V.N.; Dunker, A.K. Flexible nets: Disorder and induced fit in the associations of p53 and 14-3-3 with their partners. BMC Genom. 2008, 9, 1–20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Howton, T.C.; Zhan, Y.A.; Sun, Y.; Mukhtar, M.S. Intrinsically disordered proteins: Controlled chaos or random walk. Int. J. Plant Biol. 2015, 6, 6191. [Google Scholar] [CrossRef] [Green Version]
- Niklas, K.J.; Dunker, A.K.; Yruela, I. The evolutionary origins of cell type diversification and the role of intrinsically disordered proteins. J. Exp. Bot. 2018, 69, 1437–1446. [Google Scholar] [CrossRef]
- Wallmann, A.; Kesten, C. Common functions of disordered proteins across evolutionary distant organisms. Int. J. Mol. Sci. 2020, 21, 2105. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, X.; Jones, W.T.; Rikkerink, E.H.A. GRAS proteins: The versatile roles of intrinsically disordered proteins in plant signaling. Biochem. J. 2012, 442, 1–12. [Google Scholar] [CrossRef] [Green Version]
- Sun, X.; Malhis, N.; Zhao, B.; Xue, B.; Gsponer, J.; Rikkerink, E.H.A. Computational disorder analysis in Ethylene Response Factors uncovers binding motifs critical to their diverse functions. Int. J. Mol. Sci. 2020, 21, 74. [Google Scholar] [CrossRef] [Green Version]
- Gupta, N.; Prasad, V.B.R.; Chattopadhyay, S. LeMYC2 acts as a negative regulator of blue light mediated photomorphogenic growth, and promotes the growth of adult tomato plants. BMC Plant Biol. 2014, 14, 38. [Google Scholar] [CrossRef] [Green Version]
- Ortigosa, A.; Fonseca, S.; Franco-Zorrilla, J.M.; Fernandez-Calvo, P.; Zander, M.; Lewsey, M.G.; García-Casado, G.; Fernández-Barbero, G.; Ecker, J.R.; Solano, R. JA-pathway MYC transcription factors regulate photomorphogenic responses by targeting HY5 gene expression. Plant J. 2020, 102, 138–152. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Yao, Q.; Gao, X.; Jiang, C.; Harberd, N.P.; Fu, X. Shoot-to-root mobile transcription factor HY5 coordinates plant carbon and nitrogen acquisition. Curr. Biol. 2016, 26, 640–646. [Google Scholar] [CrossRef] [Green Version]
- van Gelderen, K.; Kang, C.; Paalman, R.; Keuskamp, D.; Hayes, S.; Pierik, R. Far-red light detection in the shoot regulates lateral root development through the HY5 transcription factor. Plant Cell 2018, 30, 101–116. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ji, Y.; Nuñez Ocaña, D.; Choe, D.; Larsen, D.H.; Marcelis, L.F.M.; Heuvelink, E. Far-red radiation stimulates dry mass partitioning to fruits by increasing fruit sink strength in tomato. New Phytol. 2020, 228, 1914–1925. [Google Scholar] [CrossRef]
- Corrado, G.; Arena, S.; Araujo-Burgos, T.; Coppola, M.; Rocco, M.; Scaloni, A.; Rao, R. The expression of the tomato prosystemin in tobacco induces alterations irrespective of its functional domain. Plant Cell Tiss. Org. 2016, 125, 509–519. [Google Scholar] [CrossRef] [Green Version]
- Tejeda-Sartorius, M.; Martínez de la Vega, O.; Délano-Frier, J.P. Jasmonic acid influences mycorrhizal colonization in tomato plants by modifying the expression of genes involved in carbohydrate partitioning. Physiol. Plant. 2008, 133, 339–353. [Google Scholar] [CrossRef] [PubMed]
- De la Noval-Pons, B.M.; León-Díaz, O.; Martínez-Gallardo, N.A.; Pérez-Ortega, E.; Délano-Frier, J.P. Pattern of β-1, 3-glucanase and chitinase activity in the AMF-systemin interaction in tomato. I. Presymbiotic phase. Cult. Trop. 2017, 38, 36–43. [Google Scholar]
- de la Noval-Pons, B.M.; León-Díaz, O.; Martínez-Gallardo, N.A.; Pérez-Ortega, E.; Délano-Frier, J.P. Activity pattern of β-1,3-glucanases and quitinases in the AMF—systemin interaction in tomato. II Early symbiotic phase. Cult. Trop. 2017, 38, 84–91. [Google Scholar]
- Ruiz-Segoviano, S.A. Efecto de la Sistemina y Otros Péptidos Afines en el Crecimiento de Plantas de Arabidopsis thaliana y Solanum lycopersicum Inoculadas con Rizobacterias Benéficas. Master’s Thesis, Cinvestav-Irapuato, Irapuato, Mexico, 2021. [Google Scholar]



| Genotype | 3 dpt | |||||
|---|---|---|---|---|---|---|
| H 1 | SD | LN | LA | BN | FN | |
| WT | 4.84 ± 0.16 a | 2 0.55 ± 0.02 a | 4.40 ± 0.16 a | 44.3 ± 2 a | - | - |
| ProSys-OE | 5.39 ± 0.2 a | 0.43 ± 0.02 b | 3.0 ± 0.1 b | 20.6 ± 2 b | - | - |
| 18 dpt | ||||||
| WT | 15.6 ± 0.42 b | 0.84 ± 0.02 a | 8.6 ± 0.4 a | 180.4 ± 5 a | - | - |
| ProSys-OE | 17.9 ± 0.31 a | 0.71 ± 0.02 b | 7.4 ± 0.16 b | 108.3 ± 7 b | - | - |
| 33 dpt | ||||||
| WT | 37.6 ± 1.0 b | 1.3 ± 0.02 a | 16.8 ± 0.88 a | 735.7 ± 16 a | 2.44 ± 0.41 a | 12.67 ± 1.63 a |
| ProSys-OE | 40.4 ± 0.82 a | 1.4 ± 0.03 a | 13.3 ± 0.63 b | 698.4 ± 20 a | 2.67 ± 0.61 a | 12.87 ± 0.88 a |
| Genotype | 32 dpt | 39 dpt | 46 dpt | 53 dpt |
|---|---|---|---|---|
| WT | 78.8 ± 1.1 a | 67.6 ± 0.08 b | 73.2 ± 11.3 a | 68.8 ± 1.0 a |
| ProSys-OE | 68.2 ± 1.0 a | 1 73.5 ± 1.5 a | 74.2 ± 1.6 a | 64.0 ± 1.9 b |
| Genotype | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Harvest N° | WT | ProSys OE | |||||||||||||
| Fruit weight (g) | |||||||||||||||
| 1 | 152.3 ± 12.6 a | 177.4 ± 18.0 a | |||||||||||||
| 2 | 94.7 ± 8.1 b | 1 150.3 ± 7.3 a | |||||||||||||
| 3 | 125.9 ± 7.6 b | 145.9 ± 5.5 a | |||||||||||||
| Fruit width (cm) | |||||||||||||||
| 1 | 6.9 ± 0.2 a | 7.1 ± 0.3 a | |||||||||||||
| 2 | 5.9 ± 0.2 b | 6.9 ± 0.2 a | |||||||||||||
| 3 | 6.3 ± 0.1 a | 6.6 ± 0.1 a | |||||||||||||
| Fruit height (cm) | |||||||||||||||
| 1 | 5.9 ± 0.2 a | 6.4 ± 0.3 a | |||||||||||||
| 2 | 5.0 ± 0.1 b | 5.9 ± 0.1 a | |||||||||||||
| 3 | 5.4 ± 0.1 b | 5.8 ± 0.1 a | |||||||||||||
| Number of seeds/fruit size | |||||||||||||||
| WT | ProSys OE | ||||||||||||||
| 2 VL | L | M | S | VS | VL | L | M | S | VS | ||||||
| 3 | 139 ± 7 b | 174 ± 65 a | 72 ± 7 a | 50 ± 1 a | 40 ± 4 a | 173 ± 5 a | 116 ± 8b | 65 ± 5 a | 37 ± 8 b | 19 ± 1 b | |||||
| Total number of fruits | |||||||||||||||
| 1 + 2 + 3 | 75 ± 8 | 80 ± 6 | |||||||||||||
| Color | |||||||||||||||
| L | C | H | L | C | H | ||||||||||
| 1 | 36.2 ± 0.49 a | 28.9 ± 1.3 a | 51.2 ± 0.6 a | 37.5 ± 0.59 a | 30.7 ± 1.0 a | 52.6 ± 0.8 a | |||||||||
| Firmness (N) | |||||||||||||||
| 1 | 0.29 ± 0.3 b | 0.37 ± 0.2 a | |||||||||||||
| Titratable acidity (as % citric acid) | |||||||||||||||
| 2 | 8.7 ± 1.4 b | 11.5 ± 0.5 a | |||||||||||||
| 3 | 9.2 ± 1.5 b | 11.7 ± 0.3 a | |||||||||||||
| Total Soluble Solids (TSS; °Brix) | |||||||||||||||
| 1 | 5.85 ± 0.23 a | 5.64 ± 0.19 b | |||||||||||||
| 2 | 5.12 ± 0.06 b | 5.27 ± 0.03 a | |||||||||||||
| 3 | 5.21 ± 0.1 a | 5.24 ± 0.06 a | |||||||||||||
| Glucose (mg/g) | |||||||||||||||
| 1 | 113.9 ± 1.86 b | 129.2 ± 5.78 a | |||||||||||||
| 2 | 93.7 ± 1.42 b | 103.1 ± 2.63 a | |||||||||||||
| Fructose (mg/g) | |||||||||||||||
| 1 | 95.2 ± 1.36 a | 91.8 ± 3.12 a | |||||||||||||
| 2 | 83.3 ± 1.78 b | 93.8 ± 1.37 a | |||||||||||||
| Sucrose (mg/g) | |||||||||||||||
| 1 | 7.16 ± 0.21 b | 9.9 ± 0.64 a | |||||||||||||
| 2 | 5.45 ± 0.28 b | 6.7 ± 0.32 a | |||||||||||||
| Starch (mg/g) | |||||||||||||||
| 1 | 1.26 ± 0.18 a | 1.7 ± 0.32 a | |||||||||||||
| 2 | 0.63 ± 0.03 b | 0.96 ± 0.10 a | |||||||||||||
| Compound/Genotype | WT | ProSys-OE |
|---|---|---|
| Harvest 1 | ||
| Peak Area | Peak Area | |
| Glyceric acid | 4.67 × 105 ± 88,316 b | 9.84 × 105 ± 1.78 × 105 a 1 |
| Aminomalonic acid | 2.5 × 105 ± 58,383 b | 7.5 × 105 ± 1.9 × 105 a |
| 2-E-butenedioic acid (fumaric acid) | 4.5 × 106 ± 22,736 a | 2.33 × 106 ± 41,209 b |
| 2-Z-butenedioic acid (maleic acid) | 3.93 × 105 ± 52,798 a | 2.13 × 105 ± 33,330 b |
| DL-malic acid | 128.9 × 106 ± 12.1 × 106 a | 78.4 × 106 ± 13.7 × 106 b |
| Oleic acid | 1.06 × 106 ± 198 × 105 a | 5.16 × 105 ± 85,215 b |
| L-glutamine | 213.3 × 106 ± 68.9 × 106 a | 45.3 × 106 ± 7.1 × 106 b |
| Harvest 2 | ||
| Acetic acid | 5.69 × 105 ± 16,6626 a | 1.01 × 105 ± 18,607 b |
| Phosphoric acid | 322.4 × 106 ± 19.9 × 106 a | 157.9 × 106 ± 22.7 × 106 b |
| Glyceric acid | 1.21 × 106 ± 1.07 × 105 a | 6.47 × 105 ± 90,612 b |
| DL-malic acid | 86.1 × 106 ± 5.7 × 106 a | 61.9 × 106 ± 9.5 × 106 b |
| Lactic acid | 1.6 × 105 ± 73,308 b | 4.7 × 105 ± 52,233 a |
| 2-E-butenedioic acid (fumaric acid) | 4.08 × 105 ± 54,794 a | 9.42 × 104 ± 10,999 b |
| 2-Z-butenedioic acid (maleic acid) | 4.18 × 105 ± 89,174 a | 1.14 × 105 ± 15,125 b |
| Butanedioic acid (succinic acid) | 5.9 × 106 ± 7.06 × 105 a | 2.43 × 106 ± 1.8 × 105 b |
| Aminomalonic acid | 3.19 × 105 ± 43,011 b | 6.3 × 105 ± 54,972 a |
| Hexadecanoic acid (palmitic acid) | 61.7 × 106 ± 9.9 × 106 a | 34.3 × 106 ± 2.7 × 106 b |
| Octadecanoic acid (stearic acid) | 32.2 × 106 ± 4.01 × 106 a | 16.9 × 106 ± 1.52 × 106 b |
| Oleic acid | 4.96 × 105 ± 41,024 a | 1.46 × 105 ± 25,905 b |
| Linoleic acid | 2.4 × 106 ± 1.15 × 105 a | 7.8 × 105 ± 57,304 b |
| Glycine | 53.5 × 104 ± 10,788 a | 23.2 × 104± 3775 b |
| L-threonine | 5.45 × 106 ± 1.6 × 106 a | 1.17 × 106 ± 2.84 × 105 b |
| L-serine | 13.1 × 106 ± 2.1 × 106 b | 34.7 × 106 ± 4.1 × 106 a |
| L-phenylalanine | 3.9 × 106 ± 8.6 × 105 b | 7.2 × 106 ± 8.4 × 105 a |
| L-aspartic acid | 32.8 × 106 ± 6.3 × 106 a | 8.9 × 106 ± 1.4 × 106 b |
| L-5-oxo-1-proline | 87.9 × 106 ± 4.7 × 106 a | 70.3 × 107 ± 3.8 × 106 b |
| D-glucuronic acid | 11.9 × 106 ± 1.7 × 106 a | 2.5 × 106 ± 8.7 × 105 b |
| Xylose | 9.05 × 106 ± 6.9 × 105 a | 4.05 × 106 ± 2.7 × 105 b |
| Myo-inositol | 1.34 × 106 ± 1.03 × 105 a | 8.84 × 105 ± 76,541 b |
| Gene/Gene Category | ROOTS | LEAVES | ||||
|---|---|---|---|---|---|---|
| Wound Response | WT | ProSys OE | P/W 2 | WT | ProSys OE | P/W |
| PS | 1 0.006 | 155.7 | 25,950 | 3.8 | 128.5 | 33.8 |
| PLA 2 | 0.51 | 4.2 | 8.2 | ND 3 | ND | - 4 |
| LHA | 4.6 | 7.1 | 1.5 | ND | ND | - |
| LOX D | 1.95 | 31.9 | 16.4 | 0.74 | 1.4 | 1.89 |
| AOS | 0.14 | 22.1 | 157.1 | ND | ND | - |
| JAZ 2 | 70.3 | 41.7 | 0.59 | ND | ND | - |
| JMT | 1.41 | 1.28 | 0.91 | 1.46 | 0.37 | 0.25 |
| PPOD | 2.1 | 6.3 | 3.0 | 0.013 | 0.24 | 18.5 |
| PIN II | 0.11 | 5.85 | 55.2 | 0.045 | 10.2 | 226.7 |
| CYP2 | 58.1 | 95.9 | 1.7 | ND | ND | - |
| SCP | 0.25 | 1.25 | 5.0 | 2.17 | 9.14 | 4.2 |
| AROGP3 | 0.6 | 3.4 | 5.7 | 0.17 | 0.45 | 2.6 |
| 9-LOX | WT | ProSys OE | P/W | WT | ProSys OE | P/W |
| LOX A | 8.8 | 49.8 | 5.7 | ND | ND | - |
| HPL | 0.71 | 1.1 | 1.6 | 2.4 | 6.6 | 2.8 |
| AOS 3 | 2.7 | 37.2 | 13.8 | 0.0 | 0.0 | - |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Luna-Martínez, M.; Martínez-Gallardo, N.; Casarrubias-Castillo, K.; Monti, S.M.; Coppola, M.; Rao, R.; Délano-Frier, J.P. Development and Yield Traits Indicate That the Constitutive Wound Response Phenotype of Prosystemin Overexpressing Tomato Plants Entails No Fitness Penalty. Agronomy 2021, 11, 1148. https://doi.org/10.3390/agronomy11061148
Luna-Martínez M, Martínez-Gallardo N, Casarrubias-Castillo K, Monti SM, Coppola M, Rao R, Délano-Frier JP. Development and Yield Traits Indicate That the Constitutive Wound Response Phenotype of Prosystemin Overexpressing Tomato Plants Entails No Fitness Penalty. Agronomy. 2021; 11(6):1148. https://doi.org/10.3390/agronomy11061148
Chicago/Turabian StyleLuna-Martínez, Mariela, Norma Martínez-Gallardo, Kena Casarrubias-Castillo, Simona M. Monti, Mariangela Coppola, Rosa Rao, and John P. Délano-Frier. 2021. "Development and Yield Traits Indicate That the Constitutive Wound Response Phenotype of Prosystemin Overexpressing Tomato Plants Entails No Fitness Penalty" Agronomy 11, no. 6: 1148. https://doi.org/10.3390/agronomy11061148








