Mechanism of Eriochloa villosa (Thunb.) Kunth Resistance to Nicosulfuron
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Cultivation of Experimental Materials
2.3. Total RNA Extraction and cDNA Library Preparation
2.4. Unigene Feature Annotations
2.5. Screening and Analysis of Differentially Expressed Genes (DEGs)
2.6. Functional Annotation and Enrichment Analysis of DEGs
2.7. Screening and qRT-PCR Validation of Metabolic Resistance Genes in E. villosa
3. Results
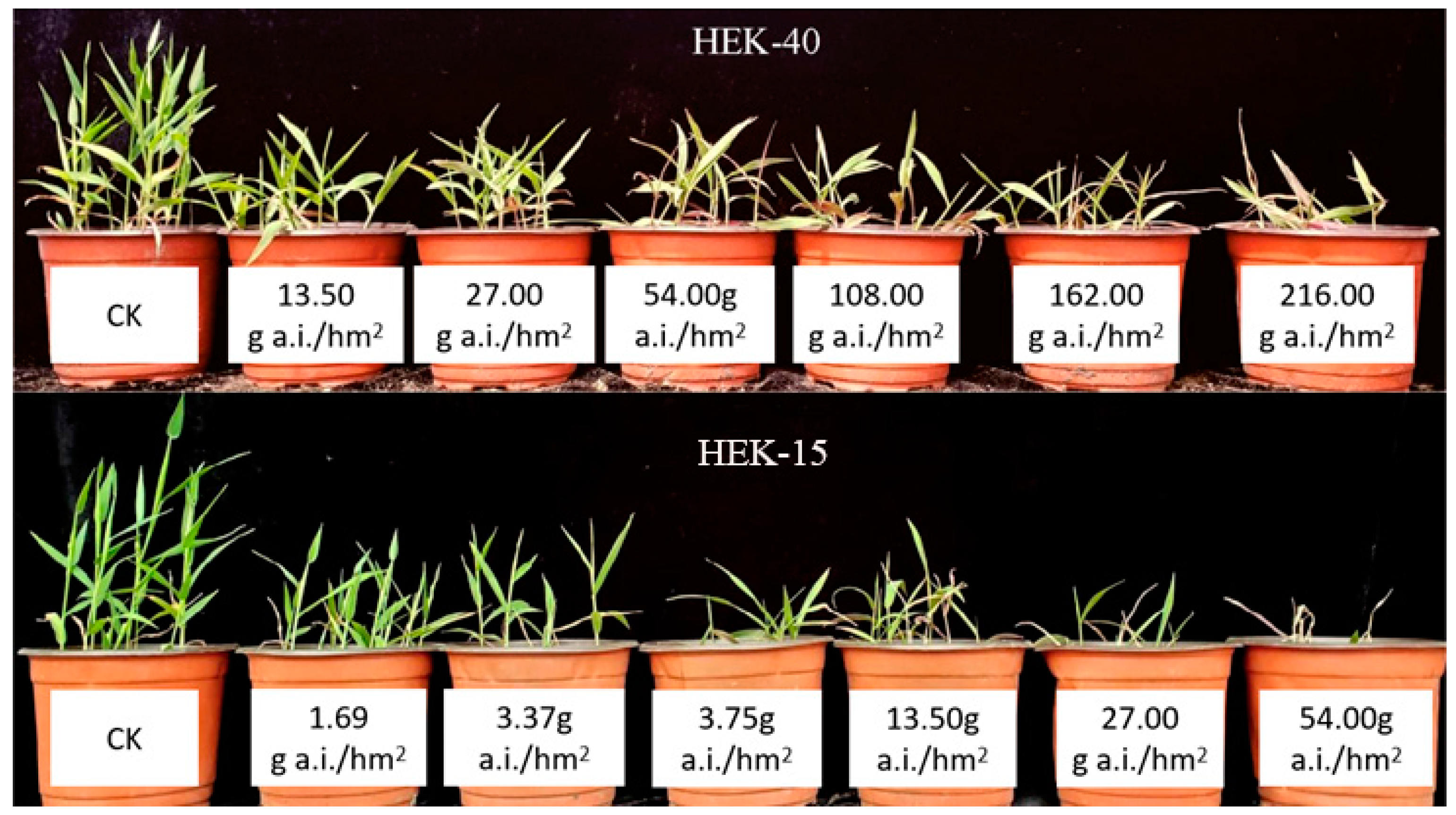
3.1. Analysis of the Resistance of Different E. villosa Populations to Nicosulfuron
3.2. Determining the Quality of E. villosa Seedlings with Different Resistance Levels
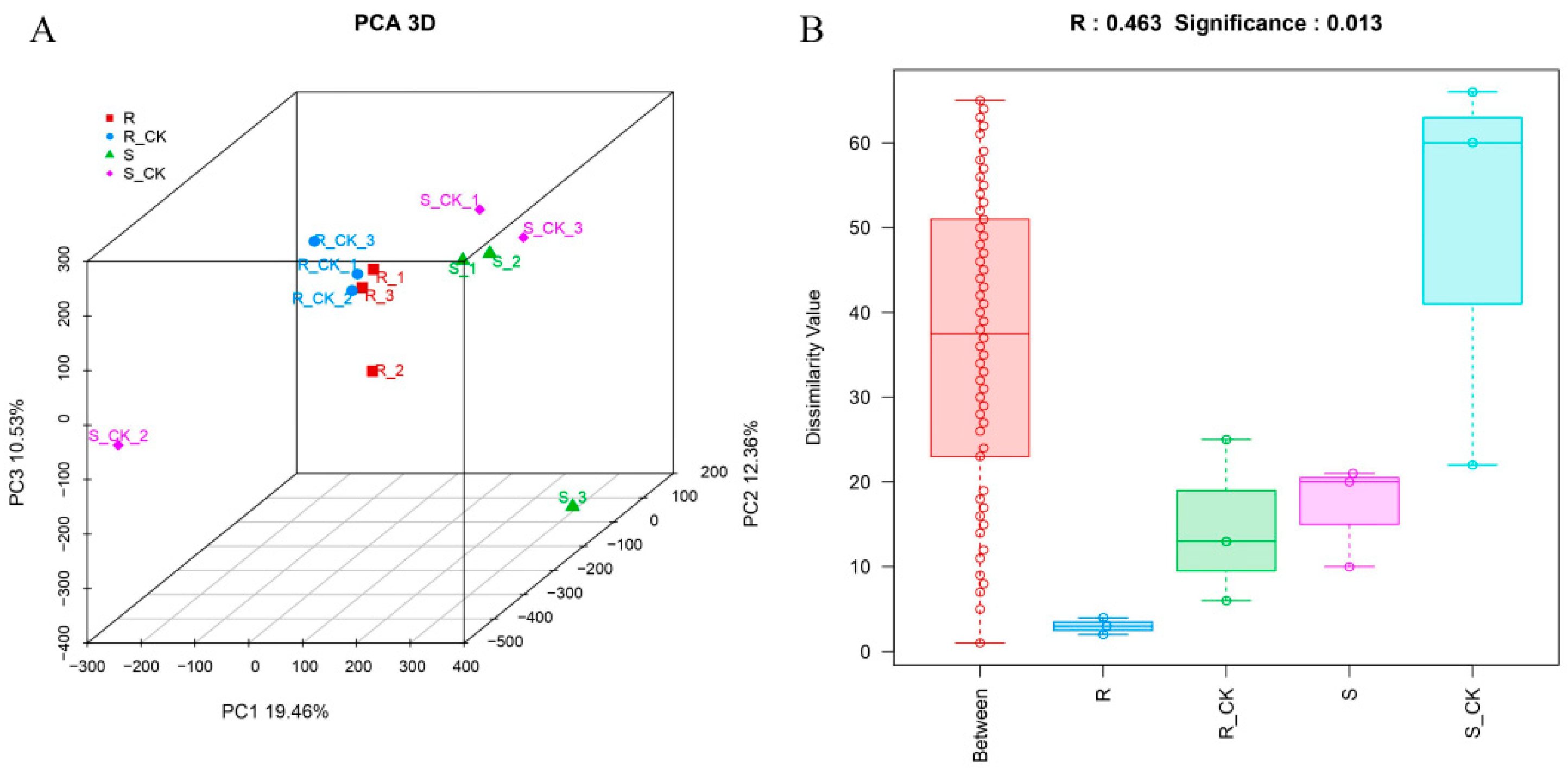
3.3. Principal Component and Dissimilarity Analysis
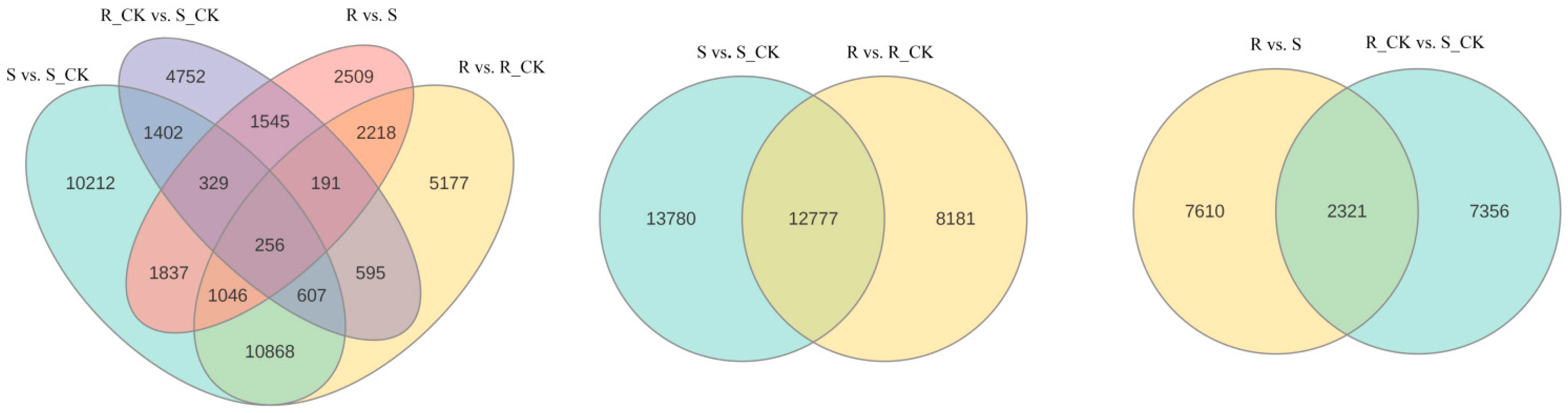
3.4. Differential Expression Analysis of mRNA
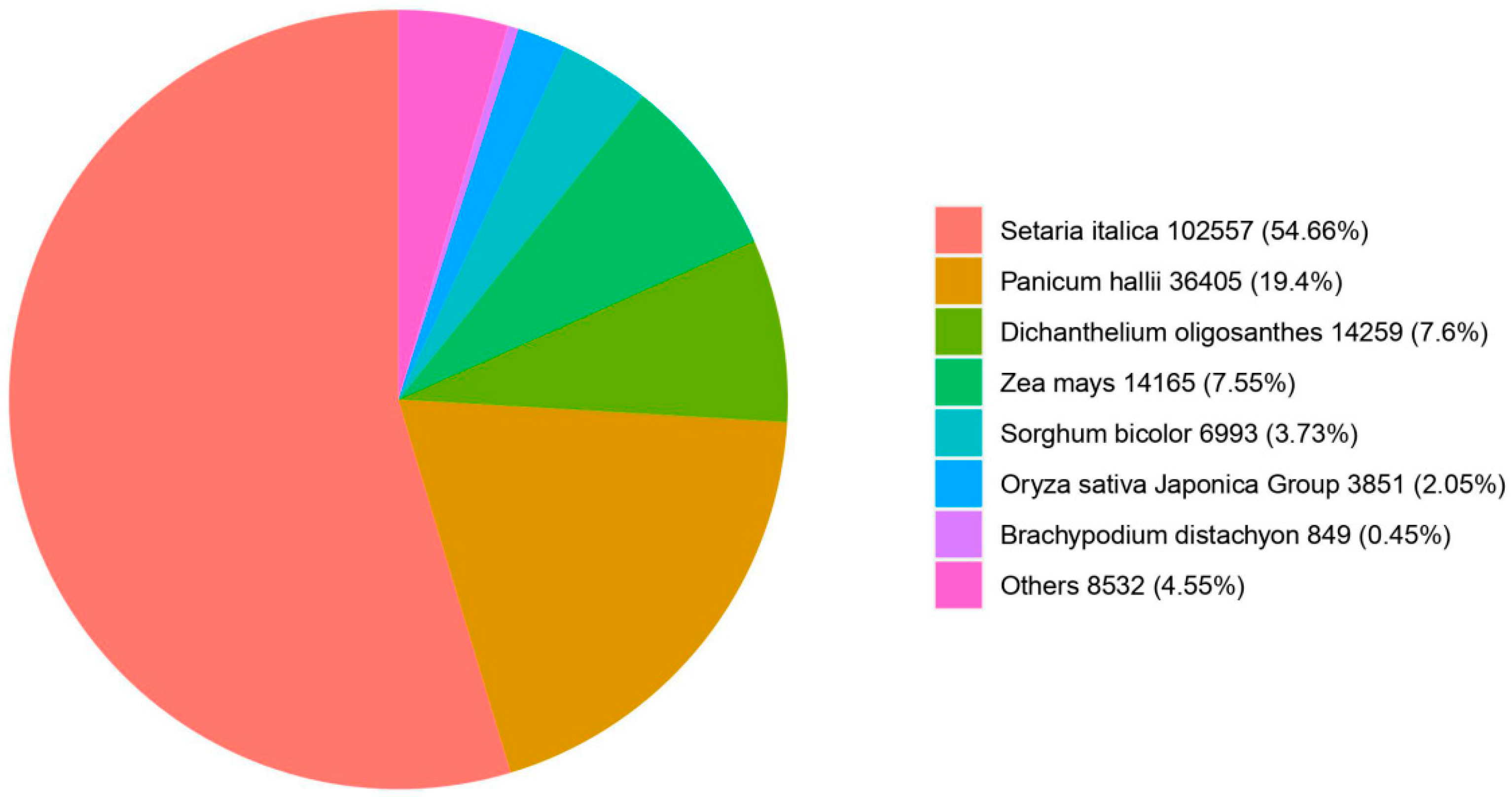
3.5. Homologous Species Comparison of E. villosa Unigenes in the NR Database
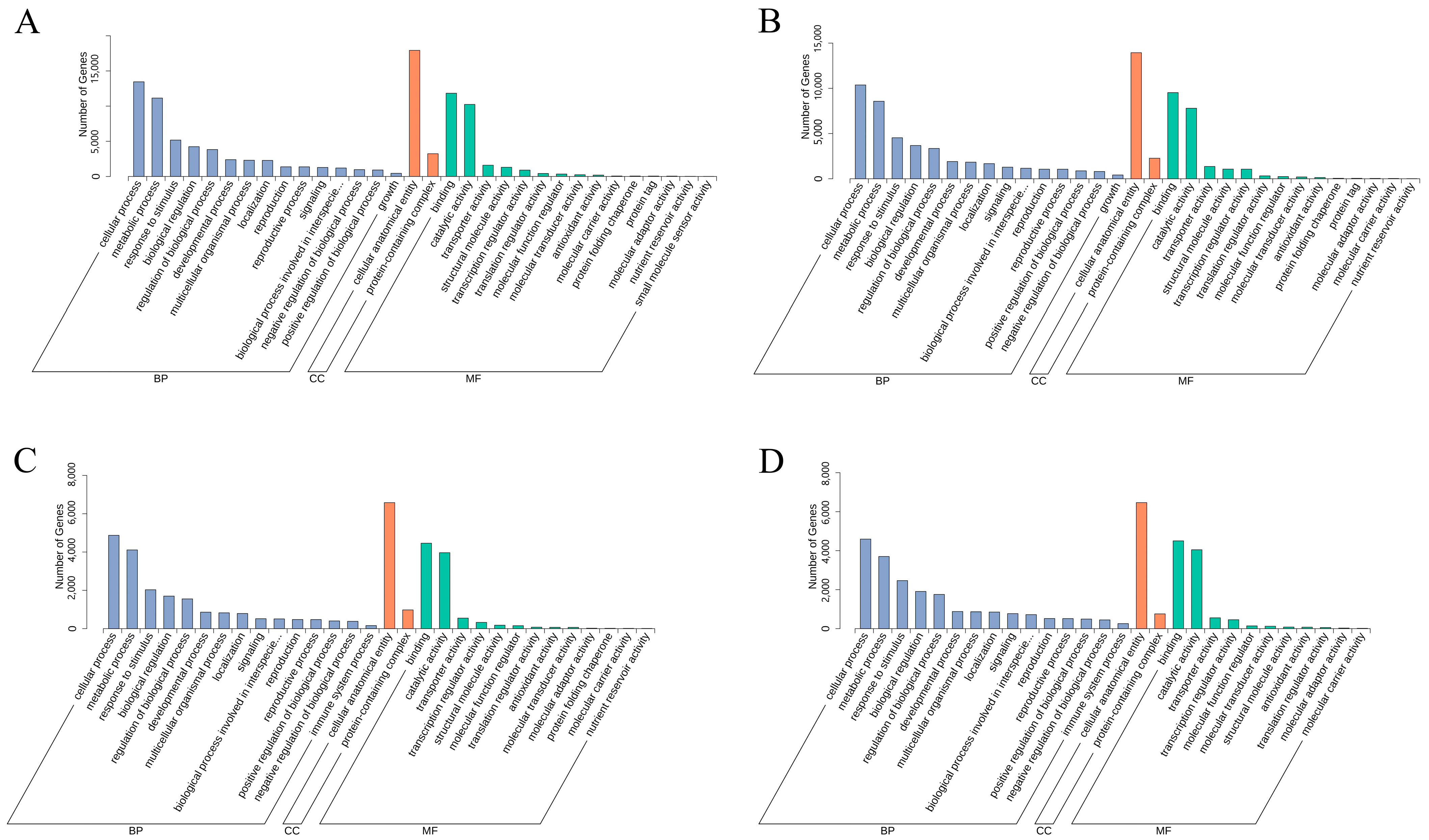
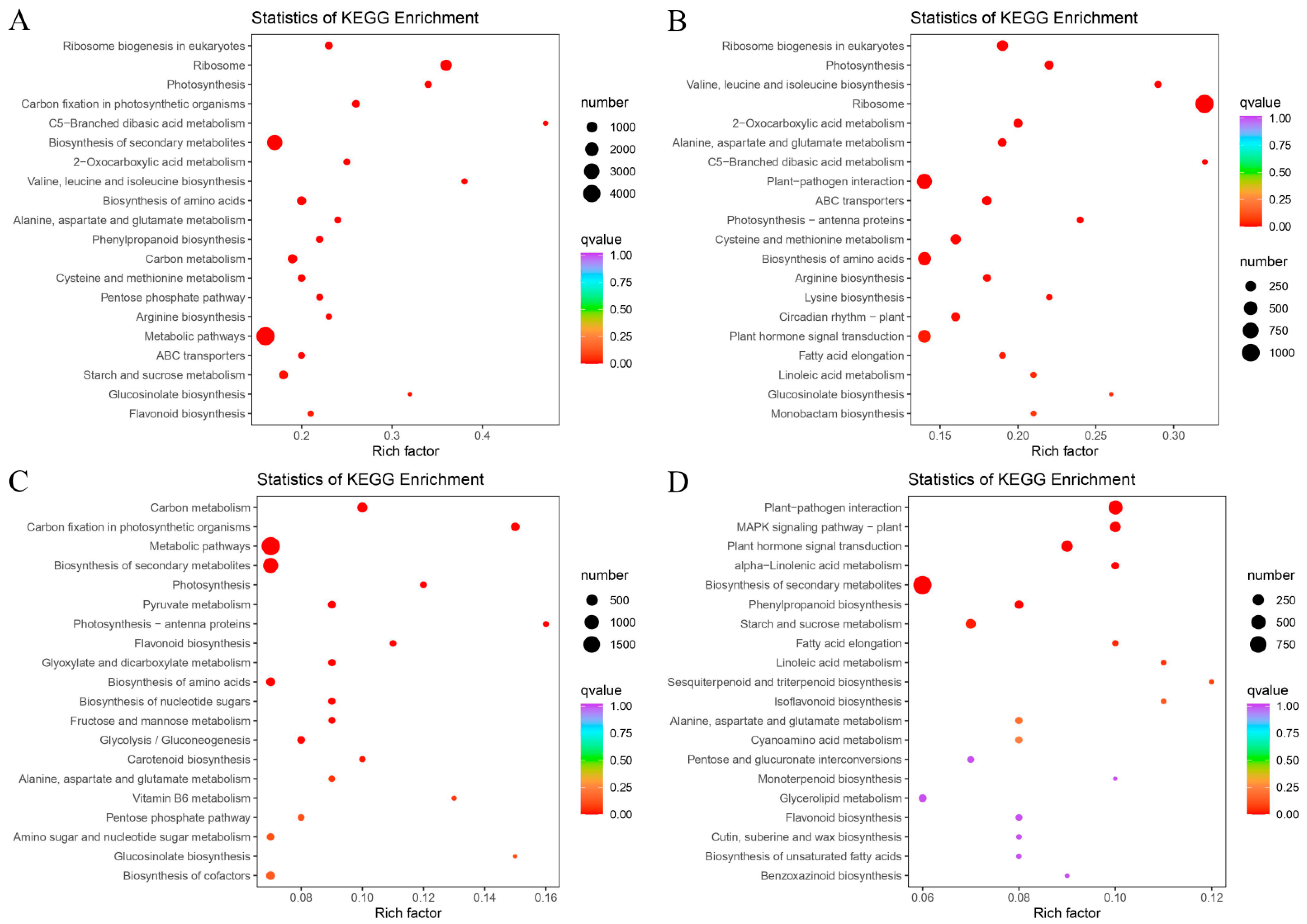
3.6. Functional Annotation and KEGG Enrichment Analysis of E. villosa GO
3.7. Identification of DEGs Involved in the Resistance of E. villosa to Nicotinic Sulfur Metal
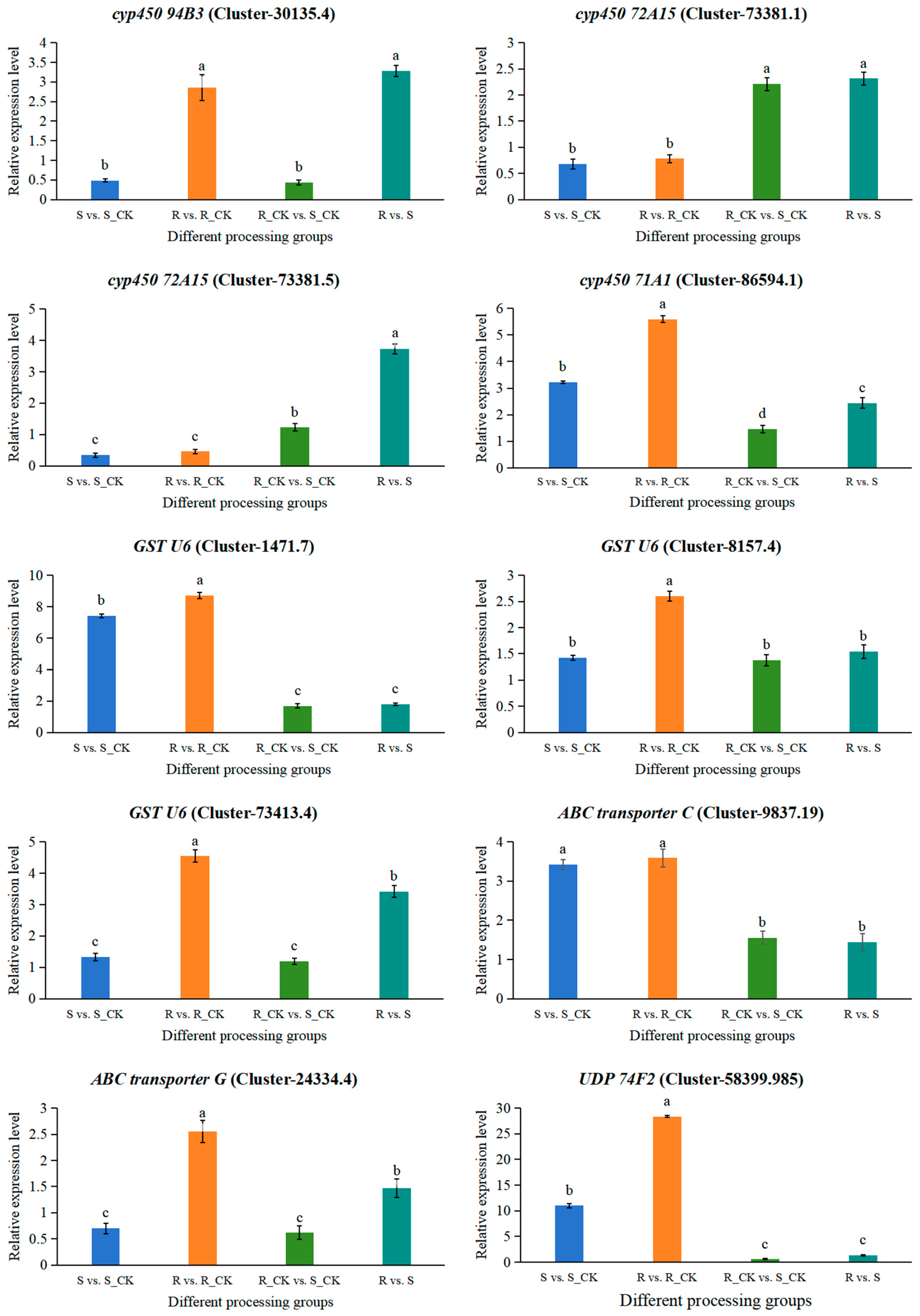
3.8. RT-qPCR Validation of DEGs Involved in the Resistance of E. villosa to Nicosulfuron
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kim, H.; Hwang, I.; Ryu, S.; Han, K.; Kwon, Y. Aryl sulfoxide scaffold useful as herbicide. Appl. Biol. Chem. 2023, 66, 67. [Google Scholar] [CrossRef]
- Hamouzová, K.; Sen, M.K.; Bharati, R.; Košnarová, P.; Chawdhery, M.R.A.; Roy, A.; Soukup, J. Calcium signalling in weeds under herbicide stress: An outlook. Front. Plant Sci. 2023, 14, 1135845. [Google Scholar] [CrossRef] [PubMed]
- Varah, A.; Ahodo, K.; Coutts, S.R.; Hicks, H.L.; Comont, D.; Crook, L.; Hull, R.; Neve, P.; Childs, D.Z.; Freckleton, R.P.; et al. The costs of human-induced evolution in an agricultural system. Nat. Sustain. 2019, 3, 63–71. [Google Scholar] [CrossRef] [PubMed]
- Baek, Y.; Bobadilla, L.K.; Giacomini, D.A.; Montgomery, J.S.; Murphy, B.P.; Tranel, P.J. Evolution of glyphosate-resistant weeds. Rev. Environ. Contam. Toxicol. 2021, 255, 93–128. [Google Scholar] [CrossRef]
- Torra, J.; Cruz, R.A.D. Molecular mechanisms of herbicide resistance in weeds. Genes 2022, 13, 2025. [Google Scholar] [CrossRef]
- Heap, I. The International Survey of Herbicide Resistant Weeds. Available online: https://weedscience.org/Account/FAQ.aspx#cite (accessed on 15 April 2024).
- Jhala, A.J.; Beckie, H.J.; Peters, T.J.; Culpepper, A.S.; Norsworthy, J.K. Interference and management of herbicide-resistant crop volunteers. Weed Sci. 2021, 69, 257–273. [Google Scholar] [CrossRef]
- Scarabel, L.; Panozzo, S.; Loddo, D.; Mathiassen, S.K.; Kristensen, M.; Kudsk, P.; Gitsopoulos, T.; Travlos, I.; Tani, E.; Chachalis, D.; et al. Diversified resistance mechanisms in multi-resistant Lolium spp. in three European countries. Front. Plant Sci. 2020, 11, 608845. [Google Scholar] [CrossRef]
- Llewellyn, R.S.; Lindner, R.K.; Pannell, D.; Powles, S. Herbicide resistance and the adoption of integrated weed management by Western Australia grain growers. Agric. Econ. 2007, 36, 123–130. [Google Scholar] [CrossRef]
- Yanniccari, M.; Gómez-Lobato, M.E.; Istilart, C.; Natalucci, C.; Giménez, D.O.; Castro, A.M. Mechanism of resistance to glyphosate in Lolium perenne from Argentina. Front. Ecol. Evol. 2017, 5. [Google Scholar] [CrossRef]
- Cao, J.; Wu, Q.; Wan, F.; Guo, J.; Wang, R. Reliable and rapid identification of glyphosate-resistance in the invasive weed Amaranthus palmeri in China. Pest Manag. Sci. 2022, 78, 2173–2182. [Google Scholar] [CrossRef]
- McCauley, C.L.; Johnson, W.G.; Young, B.G. Efficacy of halauxifen-methyl on glyphosate-resistant horseweed (Erigeron canadensis). Weed Sci. 2018, 66, 758–763. [Google Scholar] [CrossRef]
- Peterson, M.A.; Collavo, A.; Ovejero, R.; Shivrain, V.; Walsh, M.J. The challenge of herbicide resistance around the world: A current summary. Pest Manag. Sci. 2018, 74, 2246–2259. [Google Scholar] [CrossRef] [PubMed]
- Brown, H.M. Mode of action, crop selectivity, and soil relations of the sulfonylurea herbicides. Pestic. Sci. 1990, 29, 263–281. [Google Scholar] [CrossRef]
- Zhang, C.; Chen, L.; Si, H.; Gao, W.; Liu, P.; Zhang, J. Study on the characteristics and mechanisms of nicosulfuron biodegradation by Bacillus velezensis CF57. J. Basic Microbiol. 2020, 60, 649–658. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Xie, Q.; Song, B.; Riaz, M.; Lal, M.K.; Wang, L.; Lin, X.; Huo, J. Research on phytotoxicity assessment and photosynthetic characteristics of nicosulfuron residues on Beta vulgaris L. J. Environ. Manag. 2024, 353, 120159. [Google Scholar] [CrossRef]
- Cao, Y.; Lan, Y.; Huang, H.; Wei, S.; Li, X.; Sun, Y.; Wang, R.; Huang, Z. Molecular characterization of resistance to nicosulfuron in Setaria viridis. Int. J. Mol. Sci. 2023, 24, 7105. [Google Scholar] [CrossRef]
- Yang, C.; Wang, H.; Duan, Y.; Bei, F.; Jia, S.; Wang, J.; Wang, H.; Liu, W. Enhanced herbicide metabolism and target-site mutations confer multiple resistance to fomesafen and nicosulfuron in Amaranthus retroflexus L. Biology 2023, 12, 592. [Google Scholar] [CrossRef]
- Wang, R.; Sun, Y.; Lan, Y.; Wei, S.; Huang, H.; Li, X.; Huang, Z. ALS gene overexpression and enhanced metabolism conferring Digitaria sanguinalis resistance to nicosulfuron in China. Front. Plant Sci. 2023, 14, 1290600. [Google Scholar] [CrossRef]
- Zhao, B.; Xu, X.; Li, B.; Qi, Z.; Huang, J.; Hu, A.; Wang, G.; Liu, X. Target-site mutation and enhanced metabolism endow resistance to nicosulfuron in a Digitaria sanguinalis population. Pestic. Biochem. Physiol. 2023, 194, 105488. [Google Scholar] [CrossRef]
- Mei, Y.; Si, C.; Liu, M.; Qiu, L.; Zheng, M. Investigation of resistance levels and mechanisms to nicosulfuron conferred by non-target-site mechanisms in large crabgrass (Digitaria sanguinalis L.) from China. Pestic. Biochem. Physiol. 2017, 141, 84–89. [Google Scholar] [CrossRef]
- Hernández, M.; León, R.; Fischer, A.; Gebauer, M.; Galdames, R.; Figueroa, R. Target-site resistance to nicosulfuron in johnsongrass (Sorghum halepense) from Chilean corn fields. Weed Sci. 2015, 63, 631–640. [Google Scholar] [CrossRef]
- Li, Y. Weed Journal of China; China Agriculture Press: Beijing, China, 1998. [Google Scholar]
- Li, W.; Cui, J.; Xu, W.; Shi, S. Effects of Eriochloa villosa (Thunb.) Kunth on the growth and development of spring soybean and its economic threshold in Northeast China. Soybean Sci. 2019, 38, 584–588. [Google Scholar]
- Makleit, P.; Szilágyi, A.; Veres, S. First report of benzoxazinoid compounds in woolly cupgrass (Eriochloa villosa Thunb. Kunth), an invasive plant. Agronomy 2022, 12, 700. [Google Scholar] [CrossRef]
- Follak, S.; Schwarz, M.; Essl, F. First record of Eriochloa villosa (Thunb.) Kunth in Austria and notes on its distribution and agricultural impact in Central Europe. BioInvasions Rec. 2020, 9, 8–16. [Google Scholar] [CrossRef]
- Bunting, J.A.; Sprague, C.L.; Riechers, D.E. Absorption and activity of foramsulfuron in giant foxtail (Setaria faberi) and woolly cupgrass (Eriochloa villosa) with various adjuvants. Weed Sci. 2017, 52, 513–517. [Google Scholar] [CrossRef]
- Szilágyi, A.; Radócz, L.; Hájos, M.T.; Juhász, C.; Kovács, B.; Kovács, G.; Bódi, E.B.; Radványi, C.; Moloi, M.J.; Szőke, L. The impacts of woolly cupgrass on the antioxidative system and growth of a maize hybrid. Plants 2021, 10, 982. [Google Scholar] [CrossRef]
- Han, Y.; Gao, H.; Wang, Y.; Zhang, L.; Jia, J.; Ma, H. Storage time affects the viability, longevity, and germination of Eriochloa villosa (Thunb.) Kunth seeds. Sustainability 2023, 15, 8576. [Google Scholar] [CrossRef]
- Hossein, S.Z.; Hamid, R.M.C.; Zand, E.; Alcántara-de, C.R.; Travlos, I.S.; De, P.R.; Mohammad, T.A. Clodinafop-Propargyl Resistance Genes in Lolium rigidum Guad. Populations Are Associated with Fitness Costs. Agronomy 2018, 8, 106. [Google Scholar] [CrossRef]
- Trapnell, C.; Williams, B.A.; Pertea, G.; Mortazavi, A.; Kwan, G.; van Baren, M.J.; Salzberg, S.L.; Wold, B.J.; Pachter, L. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 2010, 28, 511–515. [Google Scholar] [CrossRef]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef]
- Wang, J.J.; Chen, J.C.; Li, X.J.; Cui, H.L. RNA-Seq transcriptome analysis to identify candidate genes involved in non-target site-based mesosulfuron-methyl resistance in Beckmannia syzigachne. Pestic. Biochem. Physiol. 2021, 171, 104738. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.; Lu, Z.; Huang, H.; Li, W.; Cao, Y.; Wei, S. Target site mutations and cytochrome P450s-involved metabolism confer resistance to nicosulfuron in green foxtail (Setaria viridis). Pestic. Biochem. Physiol. 2021, 179, 104956. [Google Scholar] [CrossRef] [PubMed]
- Dimaano, N.G.; Iwakami, S. Cytochrome P450-mediated herbicide metabolism in plants: Current understanding and prospects. Pest Manag. Sci. 2021, 77, 22–32. [Google Scholar] [CrossRef] [PubMed]
- Zhao, N.; Yan, Y.; Ge, L.; Zhu, B.; Liu, W.; Wang, J. Target site mutations and cytochrome P450s confer resistance to fenoxaprop-P-ethyl and mesosulfuron-methyl in Alopecurus aequalis. Pest Manag. Sci. 2019, 75, 204–214. [Google Scholar] [CrossRef]
- Iwakami, S.; Uchino, A.; Kataoka, Y.; Shibaike, H.; Watanabe, H.; Inamura, T. Cytochrome P450 genes induced by bispyribac-sodium treatment in a multiple-herbicide-resistant biotype of Echinochloa phyllopogon. Pest Manag. Sci. 2014, 70, 549–558. [Google Scholar] [CrossRef]
- Gaines, T.A.; Lorentz, L.; Figge, A.; Herrmann, J.; Maiwald, F.; Ott, M.C.; Han, H.; Busi, R.; Yu, Q.; Powles, S.B.; et al. RNA-Seq transcriptome analysis to identify genes involved in metabolism-based diclofop resistance in Lolium rigidum. Plant J. 2014, 78, 865–876. [Google Scholar] [CrossRef]
- Yan, Q.; Cui, X.; Lin, S.; Gan, S.; Xing, H.; Dou, D. GmCYP82A3, a soybean cytochrome P450 family gene involved in the jasmonic acid and ethylene signaling pathway, enhances plant resistance to biotic and abiotic stresses. PLoS ONE 2016, 11, e0162253. [Google Scholar] [CrossRef]
- Shimabukuro, R.H.; Frear, D.S.; Swanson, H.R.; Walsh, W.C. Glutathione conjugation. An enzymatic basis for atrazine resistance in corn. Plant Physiol. 1971, 47, 10–14. [Google Scholar] [CrossRef]
- Vennapusa, A.R.; Faleco, F.; Vieira, B.; Samuelson, S.; Kruger, G.R.; Werle, R.; Jugulam, M. Prevalence and mechanism of atrazine resistance in waterhemp (Amaranthus tuberculatus) from Nebraska. Weed Sci. 2018, 66, 595–602. [Google Scholar] [CrossRef]
- Brazier, M.; Cole, D.J.; Edwards, R. O-Glucosyltransferase activities toward phenolic natural products and xenobiotics in wheat and herbicide-resistant and herbicide-susceptible black-grass (Alopecurus myosuroides). Phytochemistry 2002, 59, 149–156. [Google Scholar] [CrossRef]
- Pan, L.; Li, J.; Zhang, T.; Zhang, D.; Dong, L.Y. Cross-resistance patterns to acetyl coenzyme a carboxylase (ACCase) inhibitors associated with different ACCase mutations in Beckmannia syzigachne. Weed Res. 2015, 55, 609–620. [Google Scholar] [CrossRef]
- Huang, X.X.; Zhao, S.M.; Zhang, Y.Y.; Li, Y.J.; Shen, H.N.; Li, X.; Hou, B.K. A novel UDP-glycosyltransferase 91C1 confers specific herbicide resistance through detoxification reaction in Arabidopsis. Plant Physiol. Biochem. 2021, 159, 226–233. [Google Scholar] [CrossRef] [PubMed]
- Do, T.H.T.; Martinoia, E.; Lee, Y. Functions of ABC transporters in plant growth and development. Curr. Opin. Plant Biol. 2018, 41, 32–38. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.S.; Tranel, P.J.; Stewart, C.N., Jr. Non-target-site herbicide resistance: A family business. Trends Plant Sci. 2007, 12, 6–13. [Google Scholar] [CrossRef]
- Peng, Y.; Abercrombie, L.L.; Yuan, J.S.; Riggins, C.W.; Sammons, R.D.; Tranel, P.J.; Stewart, C.N., Jr. Characterization of the horseweed (Conyza canadensis) transcriptome using GS-FLX 454 pyrosequencing and its application for expression analysis of candidate non-target herbicide resistance genes. Pest Manag. Sci. 2010, 66, 1053–1062. [Google Scholar] [CrossRef]
- Pan, L.; Yu, Q.; Wang, J.; Han, H.; Mao, L.; Nyporko, A.; Maguza, A.; Fan, L.; Bai, L.; Powles, S. An ABCC-type transporter endowing glyphosate resistance in plants. Proc. Natl. Acad. Sci. USA 2021, 118, e2100136118. [Google Scholar] [CrossRef]
- Zhao, N.; Li, W.; Bai, S.; Guo, W.; Yuan, G.; Wang, F.; Liu, W.; Wang, J. Transcriptome profiling to identify genes involved in mesosulfuron-methyl resistance in Alopecurus aequalis. Front. Plant Sci. 2017, 8, 1391. [Google Scholar] [CrossRef]


| Population | Collection Time | Collection Site | Longitude and Latitude | GR50 g a.i./hm2 | Relational Coefficient R2 | Resistance Index |
|---|---|---|---|---|---|---|
| HEK-40 | 2017.9 | Yilan County, Harbin City | 46°6′50″ N, 130°6′8″ E | 87.69 | 0.9943 | 13.83 |
| HEK-15 | 2017.9 | Beixing Village, Tangyuan County, Jiamusi City | 46°57′46″ N, 130°1′54″ E | 6.34 | 0.9957 | 1.00 |
| Sample | Raw Reads | Clean Reads | Clean Base (G) | Error Rate (%) | Q20 (%) | Q30 (%) | GC Content (%) |
|---|---|---|---|---|---|---|---|
| S_CK1 | 63274590 | 58274224 | 8.74 | 0.03 | 96.54 | 91.31 | 51.68 |
| S_CK2 | 71658798 | 65827016 | 9.87 | 0.03 | 96.75 | 91.87 | 52.81 |
| S_CK3 | 61974484 | 57820668 | 8.67 | 0.03 | 96.77 | 91.88 | 52.35 |
| S_1 | 129772220 | 120361238 | 18.05 | 0.03 | 96.64 | 91.56 | 52.27 |
| S_2 | 88531810 | 79663220 | 11.95 | 0.03 | 96.62 | 91.50 | 51.36 |
| S_3 | 83641322 | 76761782 | 11.51 | 0.03 | 96.61 | 91.48 | 52.25 |
| R_CK1 | 72235082 | 67785044 | 10.17 | 0.03 | 96.55 | 91.33 | 52.13 |
| R_CK2 | 65608280 | 60218848 | 9.03 | 0.03 | 96.63 | 91.56 | 51.57 |
| R_CK3 | 75766050 | 70231348 | 10.53 | 0.03 | 96.5 | 91.25 | 52.1 |
| R_1 | 92895894 | 85161820 | 12.77 | 0.03 | 96.43 | 91.07 | 51.86 |
| R_2 | 74693642 | 68073016 | 10.21 | 0.03 | 96.62 | 91.53 | 51.93 |
| R_3 | 73111380 | 67862266 | 10.18 | 0.03 | 96.72 | 91.80 | 52.03 |
| Level of Gene Expression | Number of DEGs | |||||||
|---|---|---|---|---|---|---|---|---|
| R vs. R_CK | S vs. S_CK | R_CK vs. S_CK | R vs. S | |||||
| Up | Down | Up | Down | Up | Down | Up | Down | |
| >1–5 | 11,169 | 8380 | 12,615 | 12,141 | 2428 | 3343 | 5099 | 3219 |
| >5–10 | 1011 | 240 | 1241 | 386 | 290 | 2999 | 300 | 1124 |
| >10 | 139 | 19 | 140 | 34 | 33 | 584 | 27 | 162 |
| Total | 12,319 | 8639 | 13,996 | 12,561 | 2751 | 6926 | 5426 | 4505 |
| 20,958 | 26,557 | 9677 | 9931 | |||||
| Gene | Annotation | Difference Multiple Log2FC | |||
|---|---|---|---|---|---|
| S vs. S_CK_ | R vs. R_CK_ | R_CK vs. S_CK | R vs. S | ||
| Cluster-73381.1 | CYP 72A15 | −0.85 | −0.31 | 1.04 | 1.54 |
| Cluster-73381.5 | CYP 72A15 | −0.84 | −0.57 | 1.45 | 1.68 |
| Cluster-30135.10 | CYP 94B3 | −1.28 | 1.62 | −0.61 | 2.27 |
| Cluster-30135.3 | CYP 94B3 | −0.55 | 1.70 | −0.30 | 1.94 |
| Cluster-30135.4 | CYP 94B3 | −1.76 | 1.28 | −0.55 | 2.48 |
| Cluster-30135.9 | CYP 94B3 | −1.07 | 1.58 | −0.49 | 2.14 |
| Cluster-86594.1 | CYP 71A1 | 0.22 | 1.76 | 0.21 | 1.73 |
| Cluster-1471.7 | GST U6 | 3.05 | 2.79 | 1.64 | 1.34 |
| Cluster-1471.8 | GST U6 | 2.79 | 3.05 | 1.01 | 1.24 |
| Cluster-73413.4 | GST U6 | 0.89 | 1.63 | 0.38 | 1.09 |
| Cluster-8157.4 | GST U6 | 0.40 | 1.03 | 0.61 | 1.20 |
| Cluster-14424.28 | ABC B4 | 7.00 | 6.04 | 2.65 | 1.68 |
| Cluster-15344.4 | ABC C4 | 2.73 | 2.78 | 1.01 | 1.02 |
| Cluster-14424.33 | ABC B4 | 5.73 | 7.29 | −0.28 | 1.25 |
| Cluster-24334.4 | ABC G25 | −0.98 | 1.65 | −0.46 | 2.15 |
| Cluster-43295.2 | ABC G37 | 0.22 | 1.80 | −0.10 | 1.46 |
| Cluster-74965.22 | ABC G36 | 1.77 | 3.14 | 0.19 | 1.53 |
| Cluster-74965.32 | ABC G40 | 1.65 | 3.18 | 0.29 | 1.78 |
| Cluster-74965.6 | ABC G40 | 1.19 | 2.87 | 0.32 | 1.96 |
| Cluster-9837.12 | ABC C3 | 0.75 | 1.71 | 0.29 | 1.21 |
| Cluster-9837.19 | ABC C3 | 2.52 | 2.99 | 0.85 | 1.29 |
| Cluster-14780.5 | UGT73C1 | −1.68 | 1.52 | −0.28 | 2.90 |
| Cluster-58399.985 | UGT74F2 | 3.53 | 4.94 | 0.18 | 1.57 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guo, J.; Xu, Z.; Jiao, T.; Gao, H.; Wang, Y.; Zhang, L.; Li, M.; Liu, X.; Yan, C.; Han, Y. Mechanism of Eriochloa villosa (Thunb.) Kunth Resistance to Nicosulfuron. Agronomy 2024, 14, 2210. https://doi.org/10.3390/agronomy14102210
Guo J, Xu Z, Jiao T, Gao H, Wang Y, Zhang L, Li M, Liu X, Yan C, Han Y. Mechanism of Eriochloa villosa (Thunb.) Kunth Resistance to Nicosulfuron. Agronomy. 2024; 14(10):2210. https://doi.org/10.3390/agronomy14102210
Chicago/Turabian StyleGuo, Jing, Zeqian Xu, Ting Jiao, Hong Gao, Yuechao Wang, Liguo Zhang, Mukai Li, Xiaomin Liu, Chunxiu Yan, and Yujun Han. 2024. "Mechanism of Eriochloa villosa (Thunb.) Kunth Resistance to Nicosulfuron" Agronomy 14, no. 10: 2210. https://doi.org/10.3390/agronomy14102210






