Abstract
Cholangiocarcinoma (CCA) is a poorly treatable type of cancer and its incidence is dramatically increasing. The lack of understanding of the biology of this tumor has slowed down the identification of novel targets and the development of effective treatments. Based on next generation sequencing profiling, alterations in DNA damage response (DDR)-related genes are paving the way for DDR-targeting strategies in CCA. Based on the notion of synthetic lethality, several DDR-inhibitors (DDRi) have been developed with the aim of accumulating enough DNA damage to induce cell death in tumor cells. Observing that DDRi alone could be insufficient for clinical use in CCA patients, the combination of DNA-damaging regimens with targeted approaches has started to be considered, as evidenced by many emerging clinical trials. Hence, novel therapeutic strategies combining DDRi with patient-specific targeted drugs could be the next level for treating cholangiocarcinoma.
1. Introduction
Most tumor hallmarks described by Hanahan and Weinberg [1,2,3] are directly associated with DNA damage-related alterations in cancer cells. Indeed, the “genome instability and mutation” hallmark is not only one of the leading forces of carcinogenesis, but is also a therapeutic target for inhibition of DNA damage response (DDR) elements such as PARP, ATM, ATR, etc. By increasing genomic instability, DDR inhibitors provide the opportunity to force tumor cells over the edge into apoptosis.
Primary liver cancers, including cholangiocarcinoma (CCA), are the third leading cause of cancer mortality worldwide [4]. In 2020, out of approximately 900,000 patients diagnosed, 830,000 have died. According to a recent report examining twenty-year data on global incidence and trends, both intrahepatic (iCCA) and extrahepatic (eCCA) incidence increased worldwide [5] and this is estimated to continue to grow due to metabolic and infectious etiologic factors in the next years.
Although some advances have been accomplished in the therapeutic approaches of CCA over the years, the survival rate is still not satisfying. The first choice of CCA therapy is resection, and adjuvant treatment is used for unresectable cases [6,7]. If the tumor can be removed successfully, patients can also be administered adjuvant chemotherapy or radiotherapy post-operation to increase complete recovery chances [8,9]. The most common chemotherapeutics used in CCA treatment include: 5-Fluorouracil with Folinic acid [10], Gemcitabine [11], Gemcitabine with Cisplatin [12], Irinotecan [13], or Capecitabine [14]. Interestingly, it has been shown that the tyrosine kinase inhibitor Erlotinib is beneficial in advanced CCA [15], and importantly, radiotherapy ameliorates the survival rate of CCA patients [16,17]. Local treatments are also currently used for CCA including transarterial chemoembolization (TACE), selective internal radiation therapy (SIRT), radiofrequency ablation (RFA), photodynamic therapy (PDT), and brachytherapy (iodine-125 seed implantation) [18,19], thus providing a beneficial extension of survival time. Nevertheless, an increase in survival and quality of life can be achieved by moving towards a more personalized medical approach [20,21]. Although several oncogenic pathways have been identified in CCA, curative therapies have been difficult to develop due to the extreme genetic heterogeneity and drug resistance [9,22,23,24].
Therapy resistance related to tumor heterogeneity is mostly attributed to genetic instability. Since the link between chromosomal abnormalities and cancer was first proposed [25], accumulating evidence has associated numerical and structural chromosomal aberrations to aggressive tumor behavior [26,27,28]. It has been assumed that genetic instability is a fundamental feature of cancer [29] and recent studies have strengthened the suggestion that instability-conferring mutational changes occur early during tumorigenesis [30,31,32,33]. Subsequent genetic instability generates mutations in proto-oncogenes and tumor suppressor genes, triggering tumor progression. The prevalence of genomic instability points to multiple cancer-associated pathways, whose deregulation has been implicated in affecting mitotic chromosome segregation [34,35,36,37].
Genomic instability has long been thought to facilitate treatment resistance by causing heterogeneity at the gene level. However, to maintain the balance between genomic chaos and the acquisition of heterogeneity, cancer cells must constrain chromosome mis-segregation rates within a limited range that maximizes their viability [38,39]. The anti-neoplastic effects of therapies that promote chromosomal instability rest on this premise. For example, widely used anticancer agents such as Taxol, PARP inhibitors and ionizing radiation (IR) are among the strongest inducers of chromosome segregation errors [40,41]. Specifically, exposure to IR promotes the generation of a variety of different lesions within DNA architecture. Radiotherapy-induced DNA damage arises following direct ionization on DNA sugar backbone, or indirectly, by promoting the production of free radicals in cells, which yield oxidative DNA damage and replication stress [42,43]. IR-induced DNA damage includes base modifications (8-OxoG), in addition to crosslinks and DNA ruptures, both at single strand break (SSB) and double strand break (DSB) levels, progressively promoting genomic instability and the activation of DNA damage response patterns in cancer cells [44,45]. DDR activation supports DNA repair and the development of radio-resistance, contributing to therapy failure. Radiotherapy is currently indicated for different CCA patients both in adjuvant regimens and for unresectable. Hence, it is considered rational to target this genetic heterogeneity with DDR inhibitors [46,47,48,49,50,51,52,53]. Using these beneficial tools in combination with patient-specific targeted drugs holds great promise to overcome CCA [54,55,56].
In CCA, DDR inhibitors have a broad area of application likely due to the high aggressivity associated with increased genomic instability. Indeed, both chromosomal or sequence-specific variability represent a fundamental cancer feature that is associated with poor prognosis, metastasis, and therapeutic resistance. It results from errors in chromosome segregation and cell cycle checkpoints during mitosis and leads to both structural and numerical abnormalities. Additionally, genomic instability regulates immune escape by modulating the interaction between tumor and its microenvironment thus driving tumor growth.
In this review we discussed the use of DDR inhibitors in CCA and the combinations of them with other chemotherapeutic or targeted agents (i.e., immunotherapeutic and antifibrotic drugs).
2. Targets and Related Therapies for DDR in Cholangiocarcinoma
Development and progression of CCA have been widely associated with increased DNA damage and genomic instability. Next-generation profiling of CCA as part of the Pan Cancer Analysis of Whole Genome initiative under the management of The International Cancer Genome Consortium (ICGC) project [57,58,59] and United States National Cancer Institutes’ the Cancer Genome Atlas (TCGA) which investigates the genomic alterations in more than 50 different cancers including BTCs [60], revealed the presence of alterations in DDR-related genes, including BRCA1/2, PARP, ATM, ATR, BAP1, ARID1A, RAD51, MLH1, TP53, PALB2, PTEN, FANC, NBN, EMSY and MRE11 [61,62,63,64,65,66,67,68], with the most commonly altered DDR genes being ATM (5%) and BRCA1/2 (4.8%) [62]. According to a very recent study on the genetic determinants of iCCA, the most common oncogenic alterations were IDH1 (20%), ARID1A (20%), TP53 (17%), CDKN2A (15%), BAP1 (15%), FGFR2 (15%), PBRM1 (12%), and KRAS (10%). IDH1/2 mutations were mutually exclusive with FGFR2 fusions (fus) and TP53 (p < 0.0001), KRAS (p = 0.0001), and CDKN2A (p < 0.0001) alterations predicted worse overall survival (OS) [69]. DDR dysregulation is currently considered one of the most relevant intracellular pathways in regulating proliferation, apoptosis and chemoresistance in biliary tract cancers (BTC) [64,70], as germline or somatic mutations in DDR genes were found in the majority of patients (63.5% [67]) and the presence of mutations/alterations in the expression of specific DDR genes can impact on patient response to platinum-based regimens [67,71,72,73,74]. Based on the well-established concept of synthetic lethality (SL), where mutation of two specific genes individually is compatible with viability, but mutation of both leads to cell death, several DDR targeting strategies have been developed [75]. The idea is that targeting one DDR gene in cancer cells that are defective for other DDR genes might be useful, in order to acquire enough DNA damage to trigger cancer cell death [63,76,77]. Thus, targeting DDR genes that are synthetically lethal with a mutation that is common in cancer, can selectively eliminate cancer cells without harming healthy cells. SL, therefore, provides an important means of strategy for the development of cancer-specific therapies [78]. Moreover, pharmacological targeting of DDR strengthens the cytotoxic effects of DNA damaging chemotherapeutics, which still unfortunately represent the main therapeutic regimen for CCA patients, wherein the development of chemoresistance drastically affects the therapeutic outcomes. Hence, different DDR inhibiting molecules are under evaluation both at the preclinical and clinical level for different types of human cancers, including CCA.
2.1. PARP
The PARP (poly (ADP-ribose) polymerase) family of enzymes includes 17 proteins, with PARP1 accounting for 80% of DDR activity [79,80]. PARP1, together with PARP2, function as a DNA damage sensor and signal transducer by recognizing damaged sites in the DNA structure. Upon its binding to DNA breaks, PARP synthetizes negatively charged PAR chains, which are, therefore, attached to target proteins through the PARylation process, a mechanism involved in modifying the chromatin architecture and supporting the recruitment of repair proteins to the sites of damage [81,82]. Once the DNA repair process restores DNA integrity, PARP1 PARylates itself and is released from DNA ruptures [77,83,84]. Interestingly, PARP enzymes participate in regulating not only SSB repair, but also DSB DNA repair through homologous recombination repair (HRR) and non-homologous end joining (NHEJ) mechanisms [85]. PARP1 inhibition leads to persistent SSBs that might result in the disruption of the replication fork, thus generating DSBs [86,87]. In addition, anti-PARP agents preclude autoPARylation and forcefully hold PARP1 on damaged DNA, hampering repair processes and leading to cell death [82,88,89]. Finally, upon the inhibition of PARP-mediated DNA repair, cancer cells are forced to compensate with alternative repair mechanisms, such as NHR, which are more subjected to the introduction of errors and might culminate with massive genomic rearrangements and apoptotic cell death [77,90,91,92]. Hence, using orally deliverable small-molecule inhibitors which target PARP1, PARP2, and PARP3 (PARPi) results in strong anti-cancer effects and to date, several PARP-targeting compounds have been developed. Olaparib (Lynparza) is the first PARPi to be FDA-approved for the management of BRCA mutated ovarian, breast, pancreatic and prostate cancers [93,94,95,96,97,98,99]. Following Olaparib, several alternative PARP inhibiting molecules have been tested in both preclinical and clinical studies, namely, Niraparib, Talazoparib, Rucaparib, Veliparib, all showing overall favorable safety profiles [100,101,102].
Active clinical studies of DDR inhibitors in CCA patients were given in Table 1. To date, disparate PARP inhibiting compounds are under evaluation in CCA, both in preclinical and clinical settings. Preclinical studies have provided promising data about the efficacy of the pharmacological inhibition of PARP in CCA cells, alone or in combination with standard chemotherapy and/or targeted molecules [66,103,104,105]. Current data about the use of PARPi in BTC patients characterized by DDR dysregulation are still open to question, with OS profiles covering from 11 to 65 months [95,103,106,107]. Specifically, CCA patients are presently enrolled in disparate phase I-II trials for the clinical evaluation of the effectiveness of PARPi as anti-CCA agents for monotherapeutic or combination-based approaches [108]. The FDA-approved PARPi Olaparib is under clinical assessment as monotherapy for the determination of overall response rate (ORR) in CCA patients with metastatic disease and alterations in DDR genes [109]. IDH1/2 genetic aberrations occur in 20% of iCCA cases and may be specifically targeted by IDH inhibitors [110]. IDH1/2 mutations have been demonstrated to increase the sensitivity to PARPi [111], therefore, patients with metastatic CCA, harboring IDH1/2 genetic alterations are currently enrolled for the evaluation of ORR and progression free survival (PFS) following Olaparib administration as single-agent therapy [112]. Partly due to the described onset of PARPi-resistance in oncological patients [113], multiple-targeting approaches by using different PARPi-based combinatorial regimens are also under clinical testing. Namely, patients with metastatic and refractory solid malignancies, including bile duct neoplasms, are currently under active recruitment for the evaluation of the therapeutic response to Olaparib administered in combination with other DDR targeting compounds [114,115] and immunotherapy [116,117]. This latter combinatorial approach has revealed, intriguingly, to be a novel potent strategy in the field of anti-tumor therapy, as dysregulated DDR appears to improve immune recognition of cancer cells, by boosting local neoantigen exposure and DAMPs release [118,119,120].
Beside Olaparib, clinical evaluations of other potent PARPi in CCA patients are ongoing, mainly in combinatorial plans with systemic therapies. The single-arm phase I study NCT03337087 aims to estimate tolerability and anti-disease effects of the recently FDA-authorized Rucaparib [107] in combination with different standard chemotherapeutics (Irinotecan, Fluorouracil, Leucovorin) in patients with gastrointestinal tumors, including CCA. Moreover, phase II studies are elucidating the effects of combining Rucaparib-mediated PARP inhibition and immunotherapeutic targeting of PD-1 in CCA patients who already completed a first-line platinum-based therapy for 4–6 months, without any clinical progression [121]. Improving outcomes for clinical PFS and overall survival (OS) are expected. Similar effects were supposed for the phase I evaluation of the potent orally bioavailable PARPi ABT-888 (Veliparib) in CCA patients treated with Gemcitabine and/or Cisplatin [122]. However, to date data are still not available. Finally, investigators plan to assess ORR, together with PFS and OS, following once-daily Niraparib administration in patients with refractory CCA. The recruiting phase recently concluded, and patients who had already received first-line standard therapy have been selected according to the presence of genetic alterations in DDR genes, including BAP1, ATM, ATR, CHK1/2, RAD51 and others [123].
2.2. Wee1
The Wee1 family of Ser/Thr protein kinases includes mainly three members: Wee1, Myt1 (PKMYT1) and Wee1B (Wee2). The first two are implicated in controlling the G2/M transition in somatic cells, being indispensable for regulating the CDK1-Cyclin B1 complex and thus entry and progression of cells into mitosis, whereas Wee2 is involved specifically in the biology of gamete cells, where it regulates the meiotic process [124,125]. Therefore, while Wee1 and Myt1 are expected to work as tumor suppressor genes, they appear to acquire oncogenic properties in malignant cells. In fact, Wee1 and Myt1 have been found overexpressed in a variety of tumors, and a large-scale CRISPR screening revealed that both kinases are essentially required for cancer cell viability [126]. Cancer cells might be dependent on Wee1 family kinases, as their high replication rate needs a powerful network in regulating cell cycle and as p53 is often inactivated in tumors, cancer cells usually rely entirely on the G2/M checkpoint. Moreover, maintaining a sustainable degree of genomic instability in malignant cells appears to require the overexpression of DDR-associated kinases [127,128,129].
Therefore, Wee1 family has been investigated as a potential therapeutic target for human tumors, with a variety of molecules being synthetized and evaluated for anti-cancer activity in several malignancies. Essentially, Wee1/Myt1 inhibiting agents, alone or in combination with chemo- and radiotherapy, function by inducing mitotic catastrophe, as they force G2 transition, also in presence of extensive damaged DNA, and premature entry into mitosis [129]. Until the present time, a plethora of different Wee1 targeting compounds have been preclinically screened and checked in different tumors, including CCA, with encouraging anti-cancer activity both as single agents and in combination with systemic or targeted therapeutics [129,130,131,132]. Among all, Adavosertib (AZD1775, MK1775) and IMP7068 have been chosen for further clinical applications. The orally deliverable small-molecule Adavosertib essentially represents the predominantly studied Wee1 inhibitor, alone or in combinatorial regimens with diverse therapeutics. Intriguingly, phase I-II studies in different malignancies, including CCA, revealed satisfactory outcomes about overall safety and tolerability, along with improving ORR and PFS rates [129,133,134,135,136,137], suggesting that, together with PARP, Wee1 might represent a prominent target to develop in setting up novel therapeutic programs for CCA patients. Interestingly, CCA patients are currently enrolled in the phase II MATCH clinical trial [138], which is actively recruiting patients with refractory BRCA1/2 mutated intrahepatic CCA, to further corroborate data about patients ORR and OS following Adavosertib administration.
2.3. ATR and ATM
In mammalian cells, ATR (ataxia telangiectasia and Rad3-related protein kinase) and ATM (ataxia-telangiectasia mutated protein kinase) represent the main transducers of DNA damage response pathways, coordinating a complex network of cellular processes involved in preserving genomic stability. ATM is predominantly activated upon DSBs, while ATR is implicated in responding to a wider landscape of genotoxic insults [139]. DNA break-points are detected and occupied by replication protein A (RPA), which is then recognized by ATR-interacting protein (ATRIP). ATR kinase is then recruited to DNA breaks, where it is phosphorylated and activated, starting a downstream cascade which induces, among others, the activation of checkpoint kinase 1 (CHK1), a crucial mediator of cell cycle checkpoint regulation [140,141,142]. Conversely, ATM is known to be the prevalent regulator of DSB-induced DNA damage response. By interacting with Nbs1, ATM is recruited to DNA breaks and activated, thus phosphorylating several substrates, including the checkpoint kinase 2 (CHK2) and histone H2AX [143,144,145], which facilitates DNA repair by reducing chromatin density and by originating an epigenetic signal that is recognized by specific DDR proteins [146,147,148,149].
In contrast with PARPi and Wee1, clinical development and application not only of ATR and ATM inhibitors (ATRi, ATMi), but also of the other targets described below, are still quite preliminary. Interestingly, preclinical data suggest that the pharmacological inhibition of ATR and ATM, especially when combined with DNA-damaging agents or PARPi, offers a feasible therapeutic approach to CCA [150,151,152,153]. However, to our knowledge there are no clinical studies enrolling CCA patients for the evaluation of therapeutic efficacy of ATMi. Conversely, the ATP-competitive ATRi Berzosertib (M6620, VX-970) alone or in combination with chemotherapeutics was evaluated in patients with advanced solid malignancies, including CCA, and showed consistent biosafety and tolerability, in addition to providing positive anti-tumor rates in phase I studies [154,155]. Accordingly, the potent oral selective ATRi Ceralasertib (AZD7638), which showed potent antineoplastic properties in several both preclinical and clinical studies (reviewed in [156]), has been selected for clinical use in the treatment of CCA. Multiple-targeting phase II studies are currently evaluating therapeutic advantages of combined administration of Ceralasertib with PARPi for immunotherapy in patients with solid CCA and other malignancies [114,115] who have failed the first-line systemic therapy. As far as it is known, there is no active clinical evaluation of other ATRi in CCA patients, since the phase I evaluation of the ATR kinase inhibitor Elimusertib (BAY 1895344) in BTC and other advanced tumor patients has been suspended due to unacceptable toxicity [157].
2.4. CHK1 and CHK2
As mentioned before, cell cycle checkpoint kinases CHK1 and CHK2 are the major downstream targets of ATR and ATM kinases. Both ATR-CHK1 and ATM-CHK2 signaling cascades converge to the negative regulation of CDC25 family of phosphatases, which crucially manage the phosphorylation status of CDK-Cyclin complexes, being thus key regulators in controlling G1/S and G2/M transitions.
Pharmacological inhibition of CHK1 and CHK2 kinases produces forced entry into mitosis and results in accumulation of DNA damage, ending with cell death [158]. Several specific CHK1/2 inhibiting compounds have been developed thus far. Nevertheless, the application of these compounds in CCA is still quite early and basically poorly explored. A preclinical study suggests that CHK1/2 inhibition by AZD7762 sensitizes CCA cells to radiotherapeutic effects [159]. As far as we know, NCT02124148 is the sole clinical application involving metastatic CCA patients elucidating the efficacy of combining the dual CHK1/2 kinase inhibitor Prexasertib (LY2606368) with standard chemotherapeutics, including cisplatin, showing promising phase I indications about patient response and drug tolerability [160]. Even though available studies provide mainly preliminary information and wider research appears required, current data suggest that targeting DDR at the level of downstream mediators CHK1/2 might represent a reasonable strategy to deepen for the development of novel therapeutic programs for CCA patients.
2.5. Other Targets
As mentioned, several different putative mechanisms can be targeted with the aim of inhibiting DNA damage response pathways. In addition to the most studied proteins such as PARP, Wee1, CHK1/2, ATR and ATM, other druggable targets are currently under evaluation to further the development of molecules with a specific pharmacological activity against DDR, including DNA dependent protein-kinase (DNA-PK) and polo-like kinase 1 (PLK1).
2.5.1. DNA-PK
DNA-PK is a Ser/Thr kinase which belongs to the phosphatidylinositol 3-kinase-related kinase (PIKK) family with a key function during DSBs repair, cell cycle progression and preservation of telomeres [161]. DNA-PK has been shown to be upregulated in BTC, where it is associated with tumor progression [162,163]. The DNA-PK inhibitor NU7026 (LY293646) has been tested for anti-cancer activity in CCA preclinical models. DNA-PK appears to be a negative regulator of DNA repair in CCA, as NU7026 treatment reduces γH2AX levels [164], suggesting that DNA-PK might exert a controversial role in CCA. Nevertheless, the orally deliverable DNA-PK inhibitor Peposertib (M3814) showed good overall tolerability and modest therapeutic response as monotherapy in patients with advanced solid tumors, including CCA [165]. This suggests that focusing on potentiating the efficacy of systemic therapy and developing drug combinations of Peposertib with chemo- and radiation approaches might be rational. Currently, NCT04068194 phase I/II study is actively recruiting patients with locally advanced or metastatic hepatobiliary malignancies to elucidate the effects of Peposertib when combined with immunotherapy (Avelumab) and/or radiotherapy [166].
2.5.2. PLK1
PLK1 is the most studied member of the PLKs family of Ser/Thr protein kinases, which represents a crucial player in managing multiple aspects of cell division, including genomic stability, mitotic entry and exit and cellular response to DNA damaging insults [167,168]. PLK1 has been described as overexpressed and associated with poor prognosis in a plethora of human neoplasms, including CCA [169,170,171]. Indeed, hindering PLK1 by using blocking antibodies, genetic depletion and pharmacological molecules negatively affects cancer cell proliferation and results in the induction of apoptosis [169].
Available preclinical data suggest that the pharmacological blockade of PLK by different selective kinase inhibitors as single agents or in combination with chemo- and radiotherapeutics might offer a promising anti-cancer strategy [172,173,174,175]. Interestingly, PLK inhibition revealed favorable clinical applications for CCA patients. In fact, the phase I study NCT01348347 recently concluded with positive results about overall tolerability, safety and clinical benefits of the oral PLK inhibitor Volasertib in patients with advanced CCA [176,177].

Table 1.
Active clinical studies of DDR inhibitors in CCA patients.
Table 1.
Active clinical studies of DDR inhibitors in CCA patients.
| Target | Treatments | Primary Endpoints | Phase | Study Identifier (ClinicalTrials.gov, accessed on 8 March 2022) |
|---|---|---|---|---|
| PARP | Rucaparib + Irinotecan/5-FU/Leucovorin calcium | MTD; DCR | I/II | NCT03337087 [178] |
| PARP | Niraparib | ORR | II | NCT03207347 [123] |
| PARP | Olaparib | ORR | II | NCT03212274 [112] |
| PARP; ATR | Olaparib + Ceralasertib (AZD6738) | ORR | II | NCT03878095 [114] |
| PARP; PD-L1 | Olaparib + Durvalamab | ORR; DCR | II | NCT03991832 [116] |
| PARP | Olaparib | ORR | II | NCT04042831 [109] |
| PARP; PD-1 | Olaparib + Pembrolizumab | ORR | II | NCT04306367 [117] |
| PARP; ATRATR; PD-L1 | Olaparib + Ceralasertib (AZD6738) Ceralasertib (AZD6738) + Durvalamab | DCR | II | NCT04298021 [115] |
| PARP; PD-1 | Rucaparib + Nivolumab | PFS at 4 months | II | NCT03639935 [121] |
| Wee1 | Adavosertib (AZD1775) | ORR | II | NCT02465060 [138] |
| DNA-PK; PD-L1 | Peposertib + Avelumab | MTD; ORR | I/II | NCT04068194 [166] |
MTD: maximum dose tolerated; DCR: disease control rate; ORR: overall response rate; PFS: progression-free survival.
2.5.3. Evidence and Future Prospects of DDR Inhibition-Based Combination Therapy in CCA
Several both predictive and pharmacodynamic biomarkers have been proposed for pharmacological inhibition of DDR [179]. The presence of mutations in BRCA genes undoubtedly represents the most used criteria to select cancer patients, including those with CCA, for therapeutic regimens which involve DDR inhibitors [139,179]. Recently, a genomic study in a large BTC patients cohort revealed that BRCA mutations are associated with high microsatellite instability and deficient mismatch repair, together with higher median tumor mutation burden, which is broadly known to be related to the patient’s response to immunotherapy [180,181]. BRCA status is defined as a predictive biomarker for therapeutic response to PARPi in several types of cancer, including CCA [139,181]. PARPi have been demonstrated to function as anti-tumor agents also in wild type BRCA cancers [182,183,184], suggesting that screening dysregulations in alternative DDR genes might represent a promising approach to identify other predictive biomarkers for therapeutic response to PARPi and more DDR targeting molecules.
Namely, in iCCA patients, inactivating mutations in BAP1 and amplifications in RAD21 gene provide two novel predictive biomarkers for the clinical efficacy of PARP chemical inhibition [185]. Intriguingly, BAP1 loss can be frequently detected in CCA, and several reports indicate that the chemical inhibition of histone deacetylases (HDAC), which are also involved in DNA repair process, reverses the phenotypic effects of BAP1 inactivation, promoting accumulation of damaged DNA and inducing cancer cell death [186,187,188]. HDAC inhibitors (HDACi) have shown promising preclinical results as anti-CCA molecules [189], with Entinostat undergoing clinical evaluation in combination with immunotherapy [190]. Importantly at epigenetic level, inhibitors of bromodomain and extra-terminal motif (BET) proteins have been recently developed as anticancer agents. This class of inhibitors reversibly binds BET proteins (BRD2, BRD3, BRD4, BRDT) and prevents protein–protein interaction between BET proteins and acetylated histones and transcription factors. BAP1 loss displayed the ability to increase the sensitivity of cancer cells also to BET inhibitors [191], which remarkably result in potent anti-tumor effects in CCA models, especially in combination with PARPi [103]. Hence, BAP1 mutated subgroup of CCA patients might appear as favored candidates for immunoepigenetic combinatorial strategies and for approaches combining different DDRi. Therefore, NGS profiling prior to clinical trial enrolment appears an essential prerequisite to selecting and assigning specific subpopulations of CCA patients to the appropriate therapeutic direction.
As CCA is a highly desmoplastic malignancy, the profound interconnection between cancer cells and tumor microenvironment (TME) along with the inflammation status of CCA should be discussed. Interestingly, inflammation and genomic instability are closely related in a positive feedback loop. Inflammation contributes to carcinogenesis and malignant progression by inducing DNA damage through RONS (reactive oxygen and nitrogen species); DNA damage, in turn, can intensify inflammation [192]. CCAs are classified into two main groups, depending on the type and degree of immune activation. “Hot” CCAs are generally defined by extensive T-cell (CD8+) infiltration, large presence of anti-tumor dendritic cells and NK cells, in addition to the enhanced production of interferon γ and granzyme B and an increased exposure of PD-1 and its ligand. Conversely, “cold” CCAs display low CD8+ infiltration and predominance of immunosuppressive populations, just as in tumor-associated macrophages and myeloid-derived suppressive cells [193]. As mentioned, DDR inhibitors have the potential to convert “cold” tumors to “hot” tumors, enhancing the sensitivity to immune checkpoint blockade, by promoting the exposure of neoantigens on the cancer cell surface and by activating interferon signaling [194]. Thus, selecting CCA patients based on their specific immune landscape might improve therapeutic response to immunotherapy by enhancing the inflammation status through DDR inhibition.
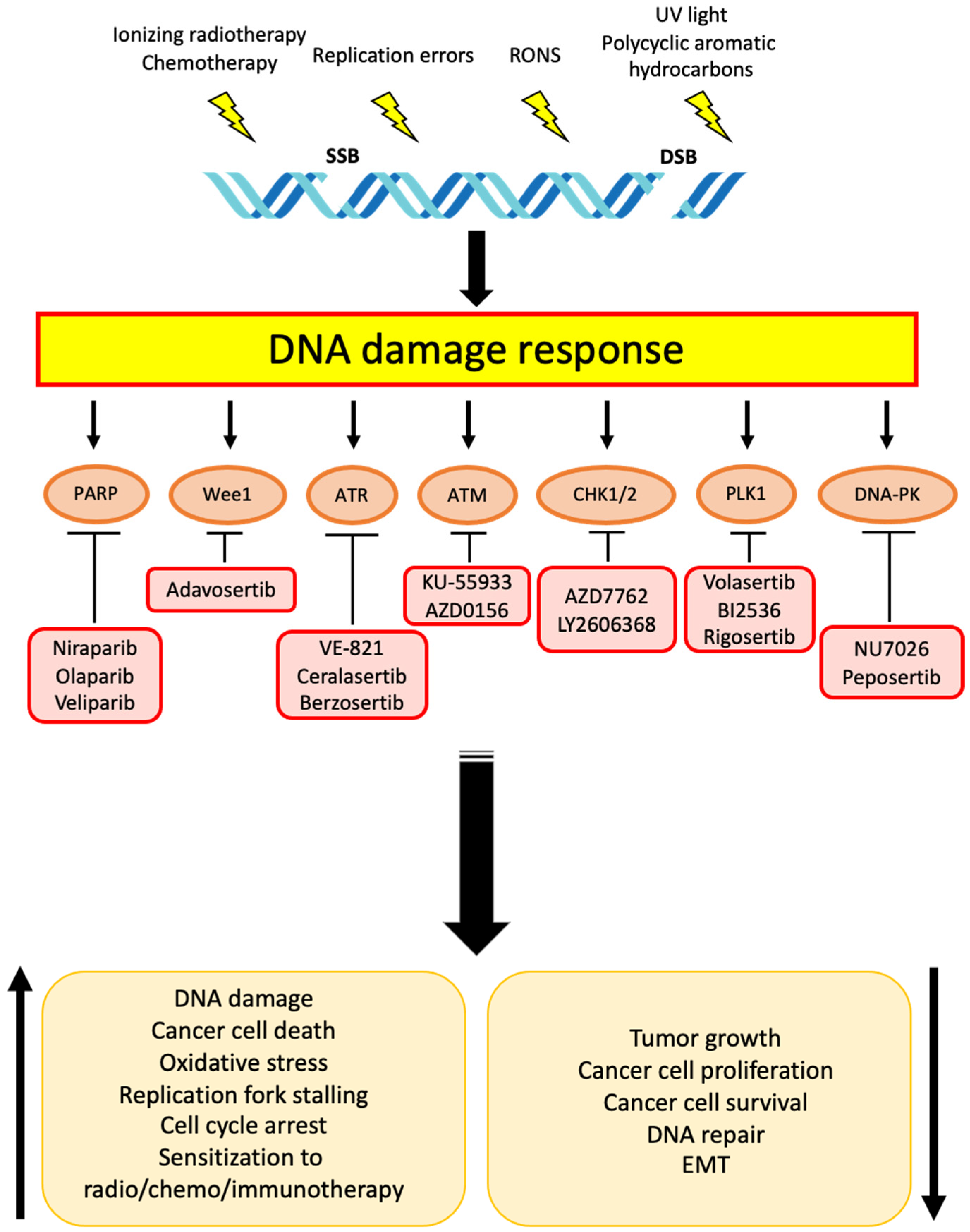
Finally, genotoxic agents have been shown to induce DNA damage in cancer-associated fibroblasts (CAFs), which, in turn, activate the senescence program and reorganize the TME, resulting in reduced sensitivity of cancer cells to radio- and chemotherapy [195,196,197]. Targeting DDR response in such CAFs might not be beneficial, since non-malignant cells harbor a robust machinery for DNA repair and would likely activate alternative ways to compensate for a DDR pathway inhibition. An alternative and more innovative approach might combine DDR inhibitors with antifibrotic drugs which specifically target CAFs by blocking the activity of PDGFRβ, VEGFR3, VEGFA, VEGFC, and EGFR. This strategy has shown promising results as a single-agent in preclinical settings [198], suggesting that TME represents a core element in CCA not only for preserving a pro-tumoral context, but also for providing novel possibilities for drug application. Figure 1 shows the main druggable DDR proteins within specific pharmacological inhibitors and the biological effects that have been observed in CCA cells.
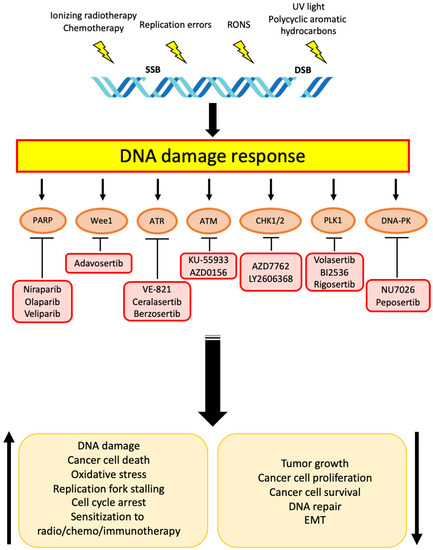
Figure 1.
Schematic representation of DDR inhibitors used in preclinical and clinical studies for CCA therapy. Both exogenous and endogenous factors induce DNA damage (SSBs, DSBs). DNA ruptures activate DNA damage response (DDR) pathways. The main DDR proteins within their specific pharmacological inhibitors and the biological effects that have been observed in CCA cells are shown.
3. Discussion
Alterations in DDR-related genes such as BRCA1/2, PARP, ATM, ATR, BAP1, ARID1A, RAD51, MLH1, TP53 makes CCA an optimal candidate for DDRi treatment, a well-established therapy in clinical application today.
However, the therapy resistance observed against DDRi in aggressive tumor types including iCCA raises the question of how to improve the treatment efficacy. Numerous phase studies show that combinatory therapy approaches using DDRi with other therapy regimens could be the next step in the translational benefit of these inhibitors.
Constituting the 80% of DDR activity in a cell, PARPi basically forms the core of DDR-targeting therapies. As mentioned in the previous section, PARPi (Olaparib, Niraparib, Veliparib, Talazoparib, Rucaparib) showed great promise in clinical studies when combined with conventional chemotherapeutics such as Gemcitabine or Cisplatin, and with immunotherapeutics. Additionally, PARPi were shown to be more effective on tumor cells with specific mutations, such as IDH1/2. Altogether, these properties make PARPi the best possible candidates for further therapeutic developments.
Along with PARPs, Wee1 is considered a useful player to target in cancer cells, since it specifically controls the G2/M checkpoint, and helps malignant cells to maintain a sustainable degree of genomic instability. Wee1 inhibitor Adavosertib was proven useful and thus this small molecule has been included in further clinical studies where it was administered alone or in combinatorial regimens with diverse therapeutics. As a result, it has been shown to be successful in both strategies, and bears hope to develop novel therapeutic approaches for iCCA patients.
Studies performed with the inhibitors developed against ATM/ATR and their downstream elements CHK1/2 remain insufficient to date. Even though there are no current ATMi clinical trials, specific ATRi are currently being evaluated by phase-II studies for therapeutic improvements after combined administration with PARPi or immuno-therapy in patients with solid CCA who have failed the first-line systemic chemotherapy. A similar picture is observed in CHK1/2-targeting studies, being scarce at this moment. However, CHKi appear promising because of their ability to sensitize malignant cells to further therapeutic applications such as radiotherapy or platin-based drug administration.
Even though the most prominent targets are the ones discussed above, other candidates have come into the light in recent years, such as DNA-PK and PLK. Because their inhibitors have exhibited positive results in preliminary studies, they represent a viable target for further research.
However, the success of a treatment is not limited to the development and use of the best inhibitor, but also to the genomic condition of the patient. In order to obtain the best outcome, this needs to be taken into elaborate consideration. Related research has shown that BRCA mutations, which are the most-known alteration in CCA cells in addition to many other solid tumors, may not be the best predictive biomarker for the accurate estimation of DDRi response in CCA patients. Therefore, better biomarker candidates must be identified. Studies have shown that BAP1-inactivating mutations and RAD21 amplifications contribute to the efficacy of PARPi, pointing to the fact that NGS profiling of tumors from patients is the best tool for determining the appropriate therapeutic strategy, especially in targeting tumor subpopulations and decreasing the recurrence possibility. Use of immunotherapy in combination with DDRi to obtain the best response from patients with specific immune characteristics must also be considered. Finally, CAFs that exert powerful effects on TME reorganization should be taken into consideration in designing DDRi therapy. Following a strategy that combines antifibrotic drugs to eliminate these cells represents a rational approach. Future innovative multi-targeted strategies focusing on CCA-intrinsic pathways and TME-extrinsic mediators will likely improve therapeutic efficacy, advancing treatment of this disease.
Author Contributions
Conceptualization, F.M. and C.R.; writing—original draft preparation, Ö.G.G. and G.A.; writing—review and editing, E.U.; supervision, F.M. and C.R.; funding acquisition, F.M. and C.R. All authors have read and agreed to the published version of the manuscript.
Funding
This research was partially funded by Italian Foundation of Cancer Research (Fondazione Italiana per la Ricerca sul Cancro; FIRC) awards (IG23117) given to CR. The APC was funded by AIRC award (IG23117). CR is a member of the European Network for the Study of Cholangiocarcinoma (ENSCCA) and participates in the COST Action EURO-CHOLANGIO-NET granted by the COST Association (CA18122). OGG is STSM recipient of COST Association (CA18122).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Data sharing not applicable. No new data were created or analyzed in this work.
Conflicts of Interest
The authors declare no conflict of interest.
Abbreviations
Cholangiocarcinoma (CCA), intrahepatic CCA (iCCA), extrahepatic CCA (eCCA), biliary tract cancer (BTC), DNA damage response (DDR), synthetic lethality (SL), transarterial chemoembolization (TACE), selective internal radiation therapy (SIRT), radiofrequency ablation (RFA), photodynamic therapy (PDT), ionizing radiation (IR), single strand break (SSB), double strand break (DSB), homologous recombination repair (HRR), non-homologous end joining (NHEJ), non-homologous recombination (NHR), overall response rate (ORR), progression free survival (PFS), overall survival (OS), poly (ADP-ribose) polymerase (PARP), ataxia telangiectasia and Rad3-related protein kinase (ATR), ataxia-telangiectasia mutated protein kinase (ATM), breast cancer gene 1/2 (BRCA1/2), BRCA associated protein 1 (BAP1), isocitrate dehydrogenase 1/2 (IDH1/2), programmed cell death protein 1 (PD-1), checkpoint kinase 1/2 (CHK1/2), protein kinase, membrane associated tyrosine/threonine 1 (PMYT1), cyclin dependent kinase 1 (CDK1), DNA-dependent protein kinase (DNA-PK), polo-like kinase (PLK), histone deacetylase (HDAC), next generation sequencing (NGS), cancer associated fibroblasts (CAFs).
References
- Hanahan, D.; Weinberg, R.A. The Hallmarks of Cancer. Cell 2000, 100, 57–70. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of Cancer: The Next Generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Hanahan, D. Hallmarks of Cancer: New Dimensions. Cancer Discov. 2022, 12, 31–46. [Google Scholar] [CrossRef] [PubMed]
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Florio, A.A.; Ferlay, J.; Znaor, A.; Ruggieri, D.; Alvarez, C.S.; Laversanne, M.; Bray, F.; McGlynn, K.A.; Petrick, J.L. Global Incidence and Trends in Intra- and Extrahepatic Cholangiocarcinoma from 1993 to 2012. Cancer 2020, 126, 2666–2678. [Google Scholar] [CrossRef] [PubMed]
- Benavides, M.; Antón, A.; Gallego, J.; Gómez, M.A.; Jiménez-Gordo, A.; La Casta, A.; Laquente, B.; Macarulla, T.; Rodríguez-Mowbray, J.R.; Maurel, J. Biliary Tract Cancers: SEOM Clinical Guidelines. Clin. Transl. Oncol. 2015, 17, 982–987. [Google Scholar] [CrossRef] [PubMed]
- Nakeeb, A.; Pitt, H.A.; Sohn, T.A.; Coleman, J.; Abrams, R.A.; Piantadosi, S.; Hruban, R.H.; Lillemoe, K.D.; Yeo, C.J.; Cameron, J.L. Cholangiocarcinoma: A Spectrum of Intrahepatic, Perihilar, and Distal Tumors. Ann. Surg. 1996, 224, 463–475. [Google Scholar] [CrossRef]
- Su, C.-H.; Tsay, S.-H.; Wu, C.-C.; Shyr, Y.-M.; King, K.-L.; Lee, C.-H.; Lui, W.-Y.; Liu, T.-J.; P’eng, F.-K. Factors Influencing Postoperative Morbidity, Mortality, and Survival After Resection for Hilar Cholangiocarcinoma. Ann. Surg. 1996, 223, 384–394. [Google Scholar] [CrossRef]
- Razumilava, N.; Gores, G.J. Cholangiocarcinoma. Lancet 2014, 383, 2168–2179. [Google Scholar] [CrossRef]
- Choi, C.W.; Choi, K.; Seo, J.H.; Kim, B.S.; Kim, J.S.; Kim, C.D.; Um, S.H.; Kim, J.S.; Kim, Y.H. Effects of 5-Fluorouracil and Leucovorin in the Treatment of Pancreatic–Biliary Tract Adenocarcinomas. Am. J. Clin. Oncol. 2000, 23, 425–428. [Google Scholar] [CrossRef] [PubMed]
- Park, J.-S.; Oh, S.-Y.; Kim, S.-H.; Kwon, H.-C.; Kim, J.-S.; Jin-Kim, H.; Kim, Y.-H. Single-Agent Gemcitabine in the Treatment of Advanced Biliary Tract Cancers: A Phase II Study. Jpn. J. Clin. Oncol. 2005, 35, 68–73. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Giuliani, F.; Gebbia, V.; Maiello, E.; Borsellino, N.; Bajardi, E.; Colucci, G. Gemcitabine and Cisplatin for Inoperable and/or Metastatic Biliary Tree Carcinomas: A Multicenter Phase II Study of the Gruppo Oncologico Dell’Italia Meridionale (GOIM). Ann. Oncol. 2006, 17, vii73–vii77. [Google Scholar] [CrossRef] [PubMed]
- Bhargava, P.; Jani, C.R.; Savarese, D.M.; O’Donnell, J.L.; Stuart, K.E.; Rocha Lima, C.M. Gemcitabine and Irinotecan in Locally Advanced or Metastatic Biliary Cancer: Preliminary Report. Oncology 2003, 17, 23–26. [Google Scholar] [PubMed]
- Knox, J.J.; Hedley, D.; Oza, A.; Feld, R.; Siu, L.L.; Chen, E.; Nematollahi, M.; Pond, G.R.; Zhang, J.; Moore, M.J. Combining Gemcitabine and Capecitabine in Patients with Advanced Biliary Cancer: A Phase II Trial. J. Clin. Oncol. 2005, 23, 2332–2338. [Google Scholar] [CrossRef] [PubMed]
- Philip, P.A.; Mahoney, M.R.; Allmer, C.; Thomas, J.; Pitot, H.C.; Kim, G.; Donehower, R.C.; Fitch, T.; Picus, J.; Erlichman, C. Phase II Study of Erlotinib in Patients with Advanced Biliary Cancer. J. Clin. Oncol. 2006, 24, 3069–3074. [Google Scholar] [CrossRef]
- Bonet Beltrán, M.; Allal, A.S.; Gich, I.; Solé, J.M.; Carrió, I. Is Adjuvant Radiotherapy Needed after Curative Resection of Extrahepatic Biliary Tract Cancers? A Systematic Review with a Meta-Analysis of Observational Studies. Cancer Treat. Rev. 2012, 38, 111–119. [Google Scholar] [CrossRef]
- Bridgewater, J.A.; Goodman, K.A.; Kalyan, A.; Mulcahy, M.F. Biliary Tract Cancer: Epidemiology, Radiotherapy, and Molecular Profiling. Am Soc. Clin. Oncol. Educ. Book 2016, 35, e194–e203. [Google Scholar] [CrossRef]
- Li, H.; Chen, L.; Zhu, G.-Y.; Yao, X.; Dong, R.; Guo, J.-H. Interventional Treatment for Cholangiocarcinoma. Front. Oncol.. 2021, 11, 671327. [Google Scholar] [CrossRef]
- Renzulli, M.; Ramai, D.; Singh, J.; Sinha, S.; Brandi, N.; Ierardi, A.M.; Albertini, E.; Sacco, R.; Facciorusso, A.; Golfieri, R. Locoregional Treatments in Cholangiocarcinoma and Combined Hepatocellular Cholangiocarcinoma. Cancers 2021, 13, 3336. [Google Scholar] [CrossRef]
- Li, Y.; Song, Y.; Liu, S. The New Insight of Treatment in Cholangiocarcinoma. J. Cancer 2022, 13, 450–464. [Google Scholar] [CrossRef]
- Manne, A.; Woods, E.; Tsung, A.; Mittra, A. Biliary Tract Cancers: Treatment Updates and Future Directions in the Era of Precision Medicine and Immuno-Oncology. Front. Oncol. 2021, 11, 768009. [Google Scholar] [CrossRef] [PubMed]
- Cao, J.; Hu, J.; Liu, S.; Meric-Bernstam, F.; Abdel-Wahab, R.; Xu, J.; Li, Q.; Yan, M.; Feng, Y.; Lin, J.; et al. Intrahepatic Cholangiocarcinoma: Genomic Heterogeneity Between Eastern and Western Patients. JCO Precis. Oncol. 2020, 4, 557–569. [Google Scholar] [CrossRef] [PubMed]
- Lin, Z.; Gong, J.; Zhong, G.; Hu, J.; Cai, D.; Zhao, L.; Zhao, Z. Identification of Mutator-Derived Alternative Splicing Signatures of Genomic Instability for Improving the Clinical Outcome of Cholangiocarcinoma. Front. Oncol. 2021, 11, 666847. [Google Scholar] [CrossRef] [PubMed]
- Brandi, G.; Farioli, A.; Astolfi, A.; Biasco, G.; Tavolari, S. Genetic Heterogeneity in Cholangiocarcinoma: A Major Challenge for Targeted Therapies. Oncotarget 2015, 6, 14744–14753. [Google Scholar] [CrossRef] [PubMed]
- Calkins, G.N. Zur Frage Der Entstehung Maligner Tumoren. By Th. Boveri. Jena, Gustav Fischer. 1914. 64 Pages. Science 1914, 40, 857–859. [Google Scholar] [CrossRef]
- Bakhoum, S.F.; Ngo, B.; Laughney, A.M.; Cavallo, J.-A.; Murphy, C.J.; Ly, P.; Shah, P.; Sriram, R.K.; Watkins, T.B.K.; Taunk, N.K.; et al. Chromosomal Instability Drives Metastasis through a Cytosolic DNA Response. Nature 2018, 553, 467–472. [Google Scholar] [CrossRef]
- Turajlic, S.; Xu, H.; Litchfield, K.; Rowan, A.; Chambers, T.; Lopez, J.I.; Nicol, D.; O’Brien, T.; Larkin, J.; Horswell, S.; et al. Tracking Cancer Evolution Reveals Constrained Routes to Metastases: TRACERx Renal. Cell 2018, 173, 581–594.e12. [Google Scholar] [CrossRef]
- Goh, J.Y.; Feng, M.; Wang, W.; Oguz, G.; Yatim, S.M.J.M.; Lee, P.L.; Bao, Y.; Lim, T.H.; Wang, P.; Tam, W.L.; et al. Chromosome 1q21.3 Amplification Is a Trackable Biomarker and Actionable Target for Breast Cancer Recurrence. Nat. Med. 2017, 23, 1319–1330. [Google Scholar] [CrossRef]
- Cahill, D.P.; Kinzler, K.W.; Vogelstein, B.; Lengauer, C. Genetic Instability and Darwinian Selection in Tumours. Trends Cell Biol. 1999, 9, M57–M60. [Google Scholar] [CrossRef]
- Armaghany, T.; Wilson, J.D.; Chu, Q.; Mills, G. Genetic Alterations in Colorectal Cancer. Gastrointest Cancer Res. 2012, 5, 19–27. [Google Scholar]
- Pikor, L.; Thu, K.; Vucic, E.; Lam, W. The Detection and Implication of Genome Instability in Cancer. Cancer Metastasis Rev. 2013, 32, 341–352. [Google Scholar] [CrossRef] [PubMed]
- Giam, M.; Rancati, G. Aneuploidy and Chromosomal Instability in Cancer: A Jackpot to Chaos. Cell Div. 2015, 10, 3. [Google Scholar] [CrossRef] [PubMed]
- Tubbs, A.; Nussenzweig, A. Endogenous DNA Damage as a Source of Genomic Instability in Cancer. Cell 2017, 168, 644–656. [Google Scholar] [CrossRef] [PubMed]
- Carter, S.L.; Cibulskis, K.; Helman, E.; McKenna, A.; Shen, H.; Zack, T.; Laird, P.W.; Onofrio, R.C.; Winckler, W.; Weir, B.A.; et al. Absolute Quantification of Somatic DNA Alterations in Human Cancer. Nat. Biotechnol 2012, 30, 413–421. [Google Scholar] [CrossRef]
- Bielski, C.M.; Zehir, A.; Penson, A.V.; Donoghue, M.T.A.; Chatila, W.; Armenia, J.; Chang, M.T.; Schram, A.M.; Jonsson, P.; Bandlamudi, C.; et al. Genome Doubling Shapes the Evolution and Prognosis of Advanced Cancers. Nat. Genet. 2018, 50, 1189–1195. [Google Scholar] [CrossRef]
- Taylor, A.M.; Shih, J.; Ha, G.; Gao, G.F.; Zhang, X.; Berger, A.C.; Schumacher, S.E.; Wang, C.; Hu, H.; Liu, J.; et al. Genomic and Functional Approaches to Understanding Cancer Aneuploidy. Cancer Cell 2018, 33, 676–689.e3. [Google Scholar] [CrossRef]
- Thompson, S.L.; Bakhoum, S.F.; Compton, D.A. Mechanisms of Chromosomal Instability. Curr. Biol. 2010, 20, R285–R295. [Google Scholar] [CrossRef]
- Laughney, A.M.; Elizalde, S.; Genovese, G.; Bakhoum, S.F. Dynamics of Tumor Heterogeneity Derived from Clonal Karyotypic Evolution. Cell Rep. 2015, 12, 809–820. [Google Scholar] [CrossRef]
- Burkard, M.E.; Weaver, B.A. Tuning Chromosomal Instability to Optimize Tumor Fitness. Cancer Discov. 2017, 7, 134–136. [Google Scholar] [CrossRef]
- Lee, H.-S.; Lee, N.C.O.; Kouprina, N.; Kim, J.-H.; Kagansky, A.; Bates, S.; Trepel, J.B.; Pommier, Y.; Sackett, D.; Larionov, V. Effects of Anticancer Drugs on Chromosome Instability and New Clinical Implications for Tumor-Suppressing Therapies. Cancer Res. 2016, 76, 902–911. [Google Scholar] [CrossRef]
- Bakhoum, S.F.; Kabeche, L.; Wood, M.D.; Laucius, C.D.; Qu, D.; Laughney, A.M.; Reynolds, G.E.; Louie, R.J.; Phillips, J.; Chan, D.A.; et al. Numerical Chromosomal Instability Mediates Susceptibility to Radiation Treatment. Nat. Commun. 2015, 6, 5990. [Google Scholar] [CrossRef] [PubMed]
- Srinivas, U.S.; Tan, B.W.Q.; Vellayappan, B.A.; Jeyasekharan, A.D. ROS and the DNA Damage Response in Cancer. Redox Biol. 2019, 25, 101084. [Google Scholar] [CrossRef] [PubMed]
- Colin, D.J.; Limagne, E.; Ragot, K.; Lizard, G.; Ghiringhelli, F.; Solary, É.; Chauffert, B.; Latruffe, N.; Delmas, D. The Role of Reactive Oxygen Species and Subsequent DNA-Damage Response in the Emergence of Resistance towards Resveratrol in Colon Cancer Models. Cell Death Dis. 2014, 5, e1533. [Google Scholar] [CrossRef] [PubMed]
- Kim, W.; Lee, S.; Seo, D.; Kim, D.; Kim, K.; Kim, E.; Kang, J.; Seong, K.M.; Youn, H.; Youn, B. Cellular Stress Responses in Radiotherapy. Cells 2019, 8, E1105. [Google Scholar] [CrossRef] [PubMed]
- Huang, R.-X.; Zhou, P.-K. DNA Damage Response Signaling Pathways and Targets for Radiotherapy Sensitization in Cancer. Signal Transduct. Target. Ther. 2020, 5, 60. [Google Scholar] [CrossRef]
- Dai, J.; Jiang, M.; He, K.; Wang, H.; Chen, P.; Guo, H.; Zhao, W.; Lu, H.; He, Y.; Zhou, C. DNA Damage Response and Repair Gene Alterations Increase Tumor Mutational Burden and Promote Poor Prognosis of Advanced Lung Cancer. Front. Oncol. 2021, 11, 708294. [Google Scholar] [CrossRef]
- Jin, J.; Tao, Z.; Cao, J.; Li, T.; Hu, X. DNA Damage Response Inhibitors: An Avenue for TNBC Treatment. Biochim. Biophys. Acta Rev. Cancer 2021, 1875, 188521. [Google Scholar] [CrossRef]
- Minchom, A.; Aversa, C.; Lopez, J. Dancing with the DNA Damage Response: Next-Generation Anti-Cancer Therapeutic Strategies. Ther. Adv. Med. Oncol. 2018, 10, 1758835918786658. [Google Scholar] [CrossRef]
- Lozano, R.; Castro, E.; Aragón, I.M.; Cendón, Y.; Cattrini, C.; López-Casas, P.P.; Olmos, D. Genetic Aberrations in DNA Repair Pathways: A Cornerstone of Precision Oncology in Prostate Cancer. Br. J. Cancer 2021, 124, 552–563. [Google Scholar] [CrossRef]
- Perkhofer, L.; Gout, J.; Roger, E.; de Almeida, F.K.; Wiesmüller, L.; Seufferlein, T.; Kleger, A. DNA Damage Repair as a Target in Pancreatic Cancer: State-of-the-Art and Future Perspectives. Gut 2021, 70, 606–617. [Google Scholar] [CrossRef]
- Knijnenburg, T.A.; Wang, L.; Zimmermann, M.T.; Chambwe, N.; Gao, G.F.; Cherniack, A.D.; Fan, H.; Shen, H.; Way, G.P.; Greene, C.S.; et al. Genomic and Molecular Landscape of DNA Damage Repair Deficiency across The Cancer Genome Atlas. Cell Rep. 2018, 23, 239–254.e6. [Google Scholar] [CrossRef] [PubMed]
- Topatana, W.; Juengpanich, S.; Li, S.; Cao, J.; Hu, J.; Lee, J.; Suliyanto, K.; Ma, D.; Zhang, B.; Chen, M.; et al. Advances in Synthetic Lethality for Cancer Therapy: Cellular Mechanism and Clinical Translation. J. Hematol. Oncol. 2020, 13, 118. [Google Scholar] [CrossRef] [PubMed]
- Gao, A.; Guo, M. Epigenetic Based Synthetic Lethal Strategies in Human Cancers. Biomark. Res. 2020, 8, 44. [Google Scholar] [CrossRef] [PubMed]
- Bekaii-Saab, T.S.; Bridgewater, J.; Normanno, N. Practical Considerations in Screening for Genetic Alterations in Cholangiocarcinoma. Ann. Oncol. 2021, 32, 1111–1126. [Google Scholar] [CrossRef]
- Miller, G.; Socci, N.D.; Dhall, D.; D’Angelica, M.; DeMatteo, R.P.; Allen, P.J.; Singh, B.; Fong, Y.; Blumgart, L.H.; Klimstra, D.S.; et al. Genome Wide Analysis and Clinical Correlation of Chromosomal and Transcriptional Mutations in Cancers of the Biliary Tract. J. Exp. Clin. Cancer Res. 2009, 28, 62. [Google Scholar] [CrossRef]
- Andersen, J.B.; Thorgeirsson, S.S. Genetic Profiling of Intrahepatic Cholangiocarcinoma. Curr. Opin. Gastroenterol. 2012, 28, 266–272. [Google Scholar] [CrossRef]
- International Network of Cancer Genome Projects. Nature 2010, 464, 993–998. [CrossRef]
- Zhang, J.; Baran, J.; Cros, A.; Guberman, J.M.; Haider, S.; Hsu, J.; Liang, Y.; Rivkin, E.; Wang, J.; Whitty, B.; et al. International Cancer Genome Consortium Data Portal—a One-Stop Shop for Cancer Genomics Data. Database 2011, 2011, bar026. [Google Scholar] [CrossRef]
- Cancer Projects | ICGC Data Portal. Available online: https://dcc.icgc.org/projects (accessed on 16 April 2022).
- The Cancer Genome Atlas Program—National Cancer Institute. Available online: https://www.cancer.gov/about-nci/organization/ccg/research/structural-genomics/tcga (accessed on 16 April 2022).
- Sulkowski, P.L.; Corso, C.D.; Robinson, N.D.; Scanlon, S.E.; Purshouse, K.R.; Bai, H.; Liu, Y.; Sundaram, R.K.; Hegan, D.C.; Fons, N.R.; et al. 2-Hydroxyglutarate Produced by Neomorphic IDH Mutations Suppresses Homologous Recombination and Induces PARP Inhibitor Sensitivity. Sci. Transl. Med. 2017, 9, eaal2463. [Google Scholar] [CrossRef]
- Lin, J.; Shi, J.; Guo, H.; Yang, X.; Jiang, Y.; Long, J.; Bai, Y.; Wang, D.; Yang, X.; Wan, X.; et al. Alterations in DNA Damage Repair Genes in Primary Liver Cancer. Clin. Cancer Res. 2019, 25, 4701–4711. [Google Scholar] [CrossRef]
- Ahn, D.H.; Bekaii-Saab, T. Biliary Tract Cancer and Genomic Alterations in Homologous Recombinant Deficiency: Exploiting Synthetic Lethality with PARP Inhibitors. Chin. Clin. Oncol. 2020, 9, 6. [Google Scholar] [CrossRef] [PubMed]
- Lamarca, A.; Barriuso, J.; McNamara, M.G.; Valle, J.W. Biliary Tract Cancer: State of the Art and Potential Role of DNA Damage Repair. Cancer Treat. Rev. 2018, 70, 168–177. [Google Scholar] [CrossRef] [PubMed]
- Lamarca, A.; Kapacee, Z.; Breeze, M.; Bell, C.; Belcher, D.; Staiger, H.; Taylor, C.; McNamara, M.G.; Hubner, R.A.; Valle, J.W. Molecular Profiling in Daily Clinical Practice: Practicalities in Advanced Cholangiocarcinoma and Other Biliary Tract Cancers. J. Clin. Med. 2020, 9, 2854. [Google Scholar] [CrossRef] [PubMed]
- Bezrookove, V.; Patino, J.M.; Nosrati, M.; Desprez, P.-Y.; McAllister, S.; Soroceanu, L.; Baron, A.; Osorio, R.; Kashani-Sabet, M.; Dar, A.A. Niraparib Suppresses Cholangiocarcinoma Tumor Growth by Inducing Oxidative and Replication Stress. Cancers 2021, 13, 4405. [Google Scholar] [CrossRef] [PubMed]
- Chae, H.; Kim, D.; Yoo, C.; Kim, K.-P.; Jeong, J.H.; Chang, H.-M.; Lee, S.S.; Park, D.H.; Song, T.J.; Hwang, S.; et al. Therapeutic Relevance of Targeted Sequencing in Management of Patients with Advanced Biliary Tract Cancer: DNA Damage Repair Gene Mutations as a Predictive Biomarker. Eur J. Cancer 2019, 120, 31–39. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Cao, Y.; Yang, X.; Li, G.; Shi, Y.; Wang, D.; Long, J.; Song, Y.; Mao, J.; Xie, F.; et al. Mutational Spectrum and Precision Oncology for Biliary Tract Carcinoma. Theranostics 2021, 11, 4585–4598. [Google Scholar] [CrossRef]
- Boerner, T.; Drill, E.; Pak, L.M.; Nguyen, B.; Sigel, C.S.; Doussot, A.; Shin, P.; Goldman, D.A.; Gonen, M.; Allen, P.J.; et al. Genetic Determinants of Outcome in Intrahepatic Cholangiocarcinoma. Hepatology 2021, 74, 1429–1444. [Google Scholar] [CrossRef]
- Banales, J.M.; Cardinale, V.; Carpino, G.; Marzioni, M.; Andersen, J.B.; Invernizzi, P.; Lind, G.E.; Folseraas, T.; Forbes, S.J.; Fouassier, L.; et al. Expert Consensus Document: Cholangiocarcinoma: Current Knowledge and Future Perspectives Consensus Statement from the European Network for the Study of Cholangiocarcinoma (ENS-CCA). Nat. Rev. Gastroenterol. Hepatol. 2016, 13, 261–280. [Google Scholar] [CrossRef]
- Kohya, N.; Miyazaki, K.; Matsukura, S.; Yakushiji, H.; Kitajima, Y.; Kitahara, K.; Fukuhara, M.; Nakabeppu, Y.; Sekiguchi, M. Deficient Expression of O(6)-Methylguanine-DNA Methyltransferase Combined with Mismatch-Repair Proteins HMLH1 and HMSH2 Is Related to Poor Prognosis in Human Biliary Tract Carcinoma. Ann. Surg. Oncol. 2002, 9, 371–379. [Google Scholar] [CrossRef]
- Ahn, D.H.; Javle, M.; Ahn, C.W.; Jain, A.; Mikhail, S.; Noonan, A.M.; Ciombor, K.; Wu, C.; Shroff, R.T.; Chen, J.L.; et al. Next-Generation Sequencing Survey of Biliary Tract Cancer Reveals the Association between Tumor Somatic Variants and Chemotherapy Resistance. Cancer 2016, 122, 3657–3666. [Google Scholar] [CrossRef]
- Hwang, I.G.; Jang, J.S.; Do, J.H.; Kang, J.H.; Lee, G.W.; Oh, S.Y.; Kwon, H.C.; Jun, H.J.; Lim, H.Y.; Lee, S.; et al. Different Relation between ERCC1 Overexpression and Treatment Outcomes of Two Platinum Agents in Advanced Biliary Tract Adenocarcinoma Patients. Cancer Chemother Pharm. 2011, 68, 935–944. [Google Scholar] [CrossRef]
- Zhang, M.; Huang, W.-Y.; Andreotti, G.; Gao, Y.-T.; Rashid, A.; Chen, J.; Sakoda, L.C.; Shen, M.-C.; Wang, B.-S.; Chanock, S.; et al. Variants of DNA Repair Genes and the Risk of Biliary Tract Cancers and Stones: A Population-Based Study in China. Cancer Epidemiol. Biomark. Prev. 2008, 17, 2123–2127. [Google Scholar] [CrossRef]
- O’Neil, N.J.; Bailey, M.L.; Hieter, P. Synthetic Lethality and Cancer. Nat. Rev. Genet. 2017, 18, 613–623. [Google Scholar] [CrossRef] [PubMed]
- Ashworth, A. A Synthetic Lethal Therapeutic Approach: Poly(ADP) Ribose Polymerase Inhibitors for the Treatment of Cancers Deficient in DNA Double-Strand Break Repair. J. Clin. Oncol. 2008, 26, 3785–3790. [Google Scholar] [CrossRef] [PubMed]
- Ricci, A.D.; Rizzo, A.; Bonucci, C.; Tober, N.; Palloni, A.; Mollica, V.; Maggio, I.; Deserti, M.; Tavolari, S.; Brandi, G. PARP Inhibitors in Biliary Tract Cancer: A New Kid on the Block? Medicines 2020, 7, 54. [Google Scholar] [CrossRef] [PubMed]
- Kaelin, W.G. The Concept of Synthetic Lethality in the Context of Anticancer Therapy. Nat. Rev. Cancer 2005, 5, 689–698. [Google Scholar] [CrossRef]
- Bai, P. Biology of Poly(ADP-Ribose) Polymerases: The Factotums of Cell Maintenance. Mol. Cell 2015, 58, 947–958. [Google Scholar] [CrossRef]
- Mehta, R.; Wood, A.C.; Yu, J.; Kim, R. Investigational PARP Inhibitors for the Treatment of Biliary Tract Cancer: Spotlight on Preclinical and Clinical Studies. Expert Opin. Investig. Drugs 2021, 30, 451–461. [Google Scholar] [CrossRef]
- Hottiger, M.O.; Hassa, P.O.; Lüscher, B.; Schüler, H.; Koch-Nolte, F. Toward a Unified Nomenclature for Mammalian ADP-Ribosyltransferases. Trends Biochem. Sci. 2010, 35, 208–219. [Google Scholar] [CrossRef]
- Kamaletdinova, T.; Fanaei-Kahrani, Z.; Wang, Z.-Q. The Enigmatic Function of PARP1: From PARylation Activity to PAR Readers. Cells 2019, 8, 1625. [Google Scholar] [CrossRef]
- Amé, J.-C.; Spenlehauer, C.; de Murcia, G. The PARP Superfamily. Bioessays 2004, 26, 882–893. [Google Scholar] [CrossRef] [PubMed]
- Murai, J.; Huang, S.N.; Das, B.B.; Renaud, A.; Zhang, Y.; Doroshow, J.H.; Ji, J.; Takeda, S.; Pommier, Y. Trapping of PARP1 and PARP2 by Clinical PARP Inhibitors. Cancer Res. 2012, 72, 5588–5599. [Google Scholar] [CrossRef] [PubMed]
- Beck, C.; Robert, I.; Reina-San-Martin, B.; Schreiber, V.; Dantzer, F. Poly(ADP-Ribose) Polymerases in Double-Strand Break Repair: Focus on PARP1, PARP2 and PARP3. Exp. Cell Res. 2014, 329, 18–25. [Google Scholar] [CrossRef] [PubMed]
- Alvarez-Gonzalez, R.; Jacobson, M.K. Characterization of Polymers of Adenosine Diphosphate Ribose Generated in Vitro and in Vivo. Biochemistry 1987, 26, 3218–3224. [Google Scholar] [CrossRef] [PubMed]
- Alemasova, E.E.; Lavrik, O.I. Poly(ADP-Ribosyl)Ation by PARP1: Reaction Mechanism and Regulatory Proteins. Nucleic Acids Res. 2019, 47, 3811–3827. [Google Scholar] [CrossRef] [PubMed]
- Brown, J.S.; O’Carrigan, B.; Jackson, S.P.; Yap, T.A. Targeting DNA Repair in Cancer: Beyond PARP Inhibitors. Cancer Discov. 2017, 7, 20–37. [Google Scholar] [CrossRef] [PubMed]
- Ray Chaudhuri, A.; Nussenzweig, A. The Multifaceted Roles of PARP1 in DNA Repair and Chromatin Remodelling. Nat. Rev. Mol. Cell Biol 2017, 18, 610–621. [Google Scholar] [CrossRef]
- Hanzlikova, H.; Caldecott, K.W. Perspectives on PARPs in S Phase. Trends Genet. 2019, 35, 412–422. [Google Scholar] [CrossRef]
- Azarm, K.; Smith, S. Nuclear PARPs and Genome Integrity. Genes Dev. 2020, 34, 285–301. [Google Scholar] [CrossRef]
- Hanzlikova, H.; Kalasova, I.; Demin, A.A.; Pennicott, L.E.; Cihlarova, Z.; Caldecott, K.W. The Importance of Poly(ADP-Ribose) Polymerase as a Sensor of Unligated Okazaki Fragments during DNA Replication. Mol. Cell 2018, 71, 319–331.e3. [Google Scholar] [CrossRef]
- de Bono, J.; Mateo, J.; Fizazi, K.; Saad, F.; Shore, N.; Sandhu, S.; Chi, K.N.; Sartor, O.; Agarwal, N.; Olmos, D.; et al. Olaparib for Metastatic Castration-Resistant Prostate Cancer. N. Engl. J. Med. 2020, 382, 2091–2102. [Google Scholar] [CrossRef] [PubMed]
- Friedlander, M.; Matulonis, U.; Gourley, C.; du Bois, A.; Vergote, I.; Rustin, G.; Scott, C.; Meier, W.; Shapira-Frommer, R.; Safra, T.; et al. Long-Term Efficacy, Tolerability and Overall Survival in Patients with Platinum-Sensitive, Recurrent High-Grade Serous Ovarian Cancer Treated with Maintenance Olaparib Capsules Following Response to Chemotherapy. Br. J. Cancer 2018, 119, 1075–1085. [Google Scholar] [CrossRef]
- Golan, T.; Hammel, P.; Reni, M.; Van Cutsem, E.; Macarulla, T.; Hall, M.J.; Park, J.-O.; Hochhauser, D.; Arnold, D.; Oh, D.-Y.; et al. Maintenance Olaparib for Germline BRCA-Mutated Metastatic Pancreatic Cancer. N. Engl. J. Med. 2019, 381, 317–327. [Google Scholar] [CrossRef] [PubMed]
- Kaufman, B.; Shapira-Frommer, R.; Schmutzler, R.K.; Audeh, M.W.; Friedlander, M.; Balmaña, J.; Mitchell, G.; Fried, G.; Stemmer, S.M.; Hubert, A.; et al. Olaparib Monotherapy in Patients with Advanced Cancer and a Germline BRCA1/2 Mutation. J. Clin. Oncol. 2015, 33, 244–250. [Google Scholar] [CrossRef] [PubMed]
- Moore, K.; Colombo, N.; Scambia, G.; Kim, B.-G.; Oaknin, A.; Friedlander, M.; Lisyanskaya, A.; Floquet, A.; Leary, A.; Sonke, G.S.; et al. Maintenance Olaparib in Patients with Newly Diagnosed Advanced Ovarian Cancer. N. Engl. J. Med. 2018, 379, 2495–2505. [Google Scholar] [CrossRef] [PubMed]
- Pujade-Lauraine, E.; Ledermann, J.A.; Selle, F.; Gebski, V.; Penson, R.T.; Oza, A.M.; Korach, J.; Huzarski, T.; Poveda, A.; Pignata, S.; et al. Olaparib Tablets as Maintenance Therapy in Patients with Platinum-Sensitive, Relapsed Ovarian Cancer and a BRCA1/2 Mutation (SOLO2/ENGOT-Ov21): A Double-Blind, Randomised, Placebo-Controlled, Phase 3 Trial. Lancet Oncol. 2017, 18, 1274–1284. [Google Scholar] [CrossRef]
- Robson, M.E.; Tung, N.; Conte, P.; Im, S.-A.; Senkus, E.; Xu, B.; Masuda, N.; Delaloge, S.; Li, W.; Armstrong, A.; et al. OlympiAD Final Overall Survival and Tolerability Results: Olaparib versus Chemotherapy Treatment of Physician’s Choice in Patients with a Germline BRCA Mutation and HER2-Negative Metastatic Breast Cancer. Ann. Oncol. 2019, 30, 558–566. [Google Scholar] [CrossRef]
- Robson, M.; Im, S.-A.; Senkus, E.; Xu, B.; Domchek, S.M.; Masuda, N.; Delaloge, S.; Li, W.; Tung, N.; Armstrong, A.; et al. Olaparib for Metastatic Breast Cancer in Patients with a Germline BRCA Mutation. N. Engl. J. Med. 2017, 377, 523–533. [Google Scholar] [CrossRef]
- Litton, J.K.; Rugo, H.S.; Ettl, J.; Hurvitz, S.A.; Gonçalves, A.; Lee, K.-H.; Fehrenbacher, L.; Yerushalmi, R.; Mina, L.A.; Martin, M.; et al. Talazoparib in Patients with Advanced Breast Cancer and a Germline BRCA Mutation. N. Engl. J. Med. 2018, 379, 753–763. [Google Scholar] [CrossRef]
- Peyraud, F.; Italiano, A. Combined PARP Inhibition and Immune Checkpoint Therapy in Solid Tumors. Cancers 2020, 12, E1502. [Google Scholar] [CrossRef]
- Fehling, S.C.; Miller, A.L.; Garcia, P.L.; Vance, R.B.; Yoon, K.J. The Combination of BET and PARP Inhibitors Is Synergistic in Models of Cholangiocarcinoma. Cancer Lett. 2020, 468, 48–58. [Google Scholar] [CrossRef]
- Mao, Y.; Huang, X.; Shuang, Z.; Lin, G.; Wang, J.; Duan, F.; Chen, J.; Li, S. PARP Inhibitor Olaparib Sensitizes Cholangiocarcinoma Cells to Radiation. Cancer Med. 2018, 7, 1285–1296. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wild, A.T.; Turcan, S.; Wu, W.H.; Sigel, C.; Klimstra, D.S.; Ma, X.; Gong, Y.; Holland, E.C.; Huse, J.T.; et al. Targeting Therapeutic Vulnerabilities with PARP Inhibition and Radiation in IDH-Mutant Gliomas and Cholangiocarcinomas. Sci. Adv. 2020, 6, eaaz3221. [Google Scholar] [CrossRef] [PubMed]
- Golan, T.; Raitses-Gurevich, M.; Kelley, R.K.; Bocobo, A.G.; Borgida, A.; Shroff, R.T.; Holter, S.; Gallinger, S.; Ahn, D.H.; Aderka, D.; et al. Overall Survival and Clinical Characteristics of BRCA-Associated Cholangiocarcinoma: A Multicenter Retrospective Study. Oncologist 2017, 22, 804–810. [Google Scholar] [CrossRef]
- Xie, Y.; Jiang, Y.; Yang, X.-B.; Wang, A.-Q.; Zheng, Y.-C.; Wan, X.-S.; Sang, X.-T.; Wang, K.; Zhang, D.-D.; Xu, J.-J.; et al. Response of BRCA1-Mutated Gallbladder Cancer to Olaparib: A Case Report. World J. Gastroenterol 2016, 22, 10254–10259. [Google Scholar] [CrossRef] [PubMed]
- Hanna, D.; Chopra, N.; Hochhauser, D.; Khan, K. The Role of PARP Inhibitors in Gastrointestinal Cancers. Crit. Rev. Oncol. Hematol 2022, 171, 103621. [Google Scholar] [CrossRef] [PubMed]
- Academic and Community Cancer Research United. A Phase II Study of Olaparib in Patients with Advanced Biliary Tract Cancer with Aberrant DNA Repair Gene Mutations. 2022. Available online: https://clinicaltrials.gov/ct2/show/NCT04042831 (accessed on 8 March 2022).
- Makawita, S.; Borad, M.J.; Carapeto, F.; Kwong, L.; Bekaii-Saab, T.S.; Murugesan, K.; Ross, J.S.; Danziger, N.; Israel, M.A.; McGregor, K.; et al. IDH1 and IDH2 Driven Intrahepatic Cholangiocarcinoma (IHCC): A Comprehensive Genomic and Immune Profiling Study. J. Clin. Oncol. 2021, 39, 4009. [Google Scholar] [CrossRef]
- IDH-Mutant Tumors Vulnerable to PARP Inhibition. Cancer Discov. 2017, 7, OF4. [CrossRef]
- National Cancer Institute (NCI). A Phase 2 Study of the PARP Inhibitor Olaparib (AZD2281) in IDH1 and IDH2 Mutant Advanced Solid Tumors. 2022. Available online: https://clinicaltrials.gov/ct2/show/NCT03212274 (accessed on 8 March 2022).
- Rose, M.; Burgess, J.T.; O’Byrne, K.; Richard, D.J.; Bolderson, E. PARP Inhibitors: Clinical Relevance, Mechanisms of Action and Tumor Resistance. Front. Cell Dev. Biol. 2020, 8, 564601. [Google Scholar] [CrossRef]
- National Cancer Institute (NCI). A Phase II Study of Olaparib and AZD6738 in Isocitrate Dehydrogenase (IDH) Mutant Solid Tumors. 2022. Available online: https://clinicaltrials.gov/ct2/show/NCT03878095 (accessed on 8 March 2022).
- Oh, D.-Y. DDR-Umbrella Study of DDR (DNA-Damage Response) Targeting Agents in Advanced Biliary Tract Cancer. 2021. Available online: https://clinicaltrials.gov/ct2/show/NCT04298021 (accessed on 8 March 2022).
- University Health Network, Toronto. A Phase II Study of Olaparib and Durvalumab (MEDI 4736) in Patients With IDH-Mutated Solid Tumors. 2021. Available online: https://clinicaltrials.gov/ct2/show/NCT03991832 (accessed on 8 March 2022).
- Georgetown University. A Phase II, Single-Arm Study of Combination Pembrolizumab and Olaparib in the Treatment of Patients With Advanced Cholangiocarcinoma. 2020. Available online: https://clinicaltrials.gov/ct2/show/NCT04306367 (accessed on 8 March 2022).
- Xie, H.; Wang, W.; Qi, W.; Jin, W.; Xia, B. Targeting DNA Repair Response Promotes Immunotherapy in Ovarian Cancer: Rationale and Clinical Application. Front. Immunol. 2021, 12, 661115. [Google Scholar] [CrossRef]
- Jiang, M.; Jia, K.; Wang, L.; Li, W.; Chen, B.; Liu, Y.; Wang, H.; Zhao, S.; He, Y.; Zhou, C. Alterations of DNA Damage Response Pathway: Biomarker and Therapeutic Strategy for Cancer Immunotherapy. Acta Pharm. Sin. B 2021, 11, 2983–2994. [Google Scholar] [CrossRef] [PubMed]
- Brown, J.S.; Sundar, R.; Lopez, J. Combining DNA Damaging Therapeutics with Immunotherapy: More Haste, Less Speed. Br. J. Cancer 2018, 118, 312–324. [Google Scholar] [CrossRef] [PubMed]
- University of Michigan Rogel Cancer Center. Phase II Multi-Center Study of PARP Inhibitor Rucaparib in Combination with Anti-PD-1 Antibody Nivolumab in Patients with Advanced or Metastatic Biliary Tract Cancer Following Platinum Therapy. 2022. Available online: https://clinicaltrials.gov/ct2/show/NCT03639935 (accessed on 8 March 2022).
- National Cancer Institute (NCI). Phase I Study of Veliparib (ABT-888) in Combination with Cisplatin Plus Gemcitabine in Advanced Biliary, Pancreatic, Urothelial, and Non-Small Cell Lung Cancer. 2013. Available online: https://clinicaltrials.gov/ct2/show/NCT01282333 (accessed on 8 March 2022).
- University of Florida. A Phase II Trial of the PARP Inhibitor, Niraparib, in BAP1 and Other DNA Damage Response (DDR) Pathway Deficient Neoplasms (UF-STO-ETI-001). 2021. Available online: https://clinicaltrials.gov/ct2/show/NCT03207347 (accessed on 8 March 2022).
- Schmidt, M.; Rohe, A.; Platzer, C.; Najjar, A.; Erdmann, F.; Sippl, W. Regulation of G2/M Transition by Inhibition of WEE1 and PKMYT1 Kinases. Molecules 2017, 22, E2045. [Google Scholar] [CrossRef] [PubMed]
- Nakanishi, M.; Ando, H.; Watanabe, N.; Kitamura, K.; Ito, K.; Okayama, H.; Miyamoto, T.; Agui, T.; Sasaki, M. Identification and Characterization of Human Wee1B, a New Member of the Wee1 Family of Cdk-Inhibitory Kinases. Genes Cells 2000, 5, 839–847. [Google Scholar] [CrossRef] [PubMed]
- Asquith, C.R.M.; Laitinen, T.; East, M.P. PKMYT1: A Forgotten Member of the WEE1 Family. Nat. Rev. Drug Discov. 2020, 19, 157. [Google Scholar] [CrossRef]
- Liu, Y.; Qi, J.; Dou, Z.; Hu, J.; Lu, L.; Dai, H.; Wang, H.; Yang, W. Systematic Expression Analysis of WEE Family Kinases Reveals the Importance of PKMYT1 in Breast Carcinogenesis. Cell Prolif. 2020, 53, e12741. [Google Scholar] [CrossRef]
- Jeong, D.; Kim, H.; Kim, D.; Ban, S.; Oh, S.; Ji, S.; Kang, D.; Lee, H.; Ahn, T.S.; Kim, H.J.; et al. Protein Kinase, Membrane-associated Tyrosine/Threonine 1 Is Associated with the Progression of Colorectal Cancer. Oncol. Rep. 2018, 39, 2829–2836. [Google Scholar] [CrossRef]
- Ghelli Luserna di Rorà, A.; Cerchione, C.; Martinelli, G.; Simonetti, G. A WEE1 Family Business: Regulation of Mitosis, Cancer Progression, and Therapeutic Target. J. Hematol Oncol. 2020, 13, 126. [Google Scholar] [CrossRef]
- Seo, H.-R.; Nam, A.-R.; Bang, J.-H.; Oh, K.-S.; Kim, J.-M.; Yoon, J.; Kim, T.-Y.; Oh, D.-Y. Inhibition of WEE1 Potentiates Sensitivity to PARP Inhibitor in Biliary Tract Cancer. Cancer Res. Treat. 2022, 54, 541–553. [Google Scholar] [CrossRef]
- Nam, A.-R.; Jin, M.-H.; Bang, J.-H.; Oh, K.-S.; Seo, H.-R.; Oh, D.-Y.; Bang, Y.-J. Inhibition of ATR Increases the Sensitivity to WEE1 Inhibitor in Biliary Tract Cancer. Cancer Res. Treat. 2020, 52, 945–956. [Google Scholar] [CrossRef]
- Jin, M.-H.; Nam, A.-R.; Bang, J.-H.; Oh, K.-S.; Seo, H.-R.; Kim, J.-M.; Yoon, J.; Kim, T.-Y.; Oh, D.-Y. WEE1 Inhibition Reverses Trastuzumab Resistance in HER2-Positive Cancers. Gastric Cancer 2021, 24, 1003–1020. [Google Scholar] [CrossRef] [PubMed]
- Takebe, N.; Naqash, A.R.; O’Sullivan Coyne, G.; Kummar, S.; Do, K.; Bruns, A.; Juwara, L.; Zlott, J.; Rubinstein, L.; Piekarz, R.; et al. Safety, Antitumor Activity, and Biomarker Analysis in a Phase I Trial of the Once-Daily Wee1 Inhibitor Adavosertib (AZD1775) in Patients with Advanced Solid Tumors. Clin. Cancer Res. 2021, 27, 3834–3844. [Google Scholar] [CrossRef] [PubMed]
- Någård, M.; Ah-See, M.-L.; So, K.; Vermunt, M.; Thistlethwaite, F.; Labots, M.; Roxburgh, P.; Ravaud, A.; Campone, M.; Valkenburg-van Iersel, L.; et al. Effect of Food on the Pharmacokinetics of the WEE1 Inhibitor Adavosertib (AZD1775) in Patients with Advanced Solid Tumors. Cancer Chemother. Pharm. 2020, 86, 97–108. [Google Scholar] [CrossRef] [PubMed]
- Lheureux, S.; Cristea, M.C.; Bruce, J.P.; Garg, S.; Cabanero, M.; Mantia-Smaldone, G.; Olawaiye, A.B.; Ellard, S.L.; Weberpals, J.I.; Wahner Hendrickson, A.E.; et al. Adavosertib plus Gemcitabine for Platinum-Resistant or Platinum-Refractory Recurrent Ovarian Cancer: A Double-Blind, Randomised, Placebo-Controlled, Phase 2 Trial. Lancet 2021, 397, 281–292. [Google Scholar] [CrossRef]
- Lu, Y.-L.; Huang, Y.-T.; Wu, M.-H.; Chou, T.-C.; J Wong, R.; Lin, S.-F. Efficacy of Adavosertib Therapy against Anaplastic Thyroid Cancer. Endocr. Relat. Cancer 2021, 28, 311–324. [Google Scholar] [CrossRef]
- Liu, J.F.; Xiong, N.; Campos, S.M.; Wright, A.A.; Krasner, C.; Schumer, S.; Horowitz, N.; Veneris, J.; Tayob, N.; Morrissey, S.; et al. Phase II Study of the WEE1 Inhibitor Adavosertib in Recurrent Uterine Serous Carcinoma. J. Clin. Oncol. 2021, 39, 1531–1539. [Google Scholar] [CrossRef]
- National Cancer Institute (NCI). Molecular Analysis for Therapy Choice (MATCH). 2022. Available online: https://clinicaltrials.gov/ct2/show/NCT02465060 (accessed on 8 March 2022).
- Choi, W.; Lee, E.S. Therapeutic Targeting of DNA Damage Response in Cancer. Int J. Mol. Sci. 2022, 23, 1701. [Google Scholar] [CrossRef]
- Menolfi, D.; Zha, S. ATM, ATR and DNA-PKcs Kinases-the Lessons from the Mouse Models: Inhibition ≠ Deletion. Cell BioSci. 2020, 10, 8. [Google Scholar] [CrossRef]
- Bradbury, A.; Hall, S.; Curtin, N.; Drew, Y. Targeting ATR as Cancer Therapy: A New Era for Synthetic Lethality and Synergistic Combinations? Pharmacol. Ther. 2020, 207, 107450. [Google Scholar] [CrossRef]
- Blackford, A.N.; Jackson, S.P. ATM, ATR, and DNA-PK: The Trinity at the Heart of the DNA Damage Response. Mol. Cell 2017, 66, 801–817. [Google Scholar] [CrossRef]
- You, Z.; Chahwan, C.; Bailis, J.; Hunter, T.; Russell, P. ATM Activation and Its Recruitment to Damaged DNA Require Binding to the C Terminus of Nbs1. Mol. Cell Biol. 2005, 25, 5363–5379. [Google Scholar] [CrossRef] [PubMed]
- Shiloh, Y. ATM and Related Protein Kinases: Safeguarding Genome Integrity. Nat. Rev. Cancer 2003, 3, 155–168. [Google Scholar] [CrossRef] [PubMed]
- Lavin, M.F. Ataxia-Telangiectasia: From a Rare Disorder to a Paradigm for Cell Signalling and Cancer. Nat. Rev. Mol. Cell Biol. 2008, 9, 759–769. [Google Scholar] [CrossRef]
- Chapman, J.R.; Jackson, S.P. Phospho-Dependent Interactions between NBS1 and MDC1 Mediate Chromatin Retention of the MRN Complex at Sites of DNA Damage. EMBO Rep. 2008, 9, 795–801. [Google Scholar] [CrossRef]
- Bouquet, F.; Muller, C.; Salles, B. The Loss of GammaH2AX Signal Is a Marker of DNA Double Strand Breaks Repair Only at Low Levels of DNA Damage. Cell Cycle 2006, 5, 1116–1122. [Google Scholar] [CrossRef]
- Banáth, J.P.; Macphail, S.H.; Olive, P.L. Radiation Sensitivity, H2AX Phosphorylation, and Kinetics of Repair of DNA Strand Breaks in Irradiated Cervical Cancer Cell Lines. Cancer Res. 2004, 64, 7144–7149. [Google Scholar] [CrossRef]
- Xie, A.; Puget, N.; Shim, I.; Odate, S.; Jarzyna, I.; Bassing, C.H.; Alt, F.W.; Scully, R. Control of Sister Chromatid Recombination by Histone H2AX. Mol. Cell 2004, 16, 1017–1025. [Google Scholar] [CrossRef]
- Moolmuang, B.; Ruchirawat, M. The Antiproliferative Effects of Ataxia-Telangiectasia Mutated and ATM- and Rad3-Related Inhibitions and Their Enhancements with the Cytotoxicity of DNA Damaging Agents in Cholangiocarcinoma Cells. J. Pharm. Pharmacol. 2021, 73, 40–51. [Google Scholar] [CrossRef]
- Nam, A.-R.; Yoon, J.; Jin, M.-H.; Bang, J.-H.; Oh, K.-S.; Seo, H.-R.; Kim, J.-M.; Kim, T.-Y.; Oh, D.-Y. ATR Inhibition Amplifies Antitumor Effects of Olaparib in Biliary Tract Cancer. Cancer Lett. 2021, 516, 38–47. [Google Scholar] [CrossRef]
- Nam, A.-R.; Jin, M.H.; Park, J.E.; Bang, J.-H.; Oh, D.-Y.; Bang, Y.-J. Therapeutic Targeting of the DNA Damage Response Using an ATR Inhibitor in Biliary Tract Cancer. Cancer Res. Treat. 2019, 51, 1167–1179. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.-R.; Wu, C.-E.; Yeh, C.-N. ATM Inhibitor Suppresses Gemcitabine-Resistant BTC Growth in a Polymerase θ Deficiency-Dependent Manner. Biomolecules 2020, 10, E1529. [Google Scholar] [CrossRef] [PubMed]
- Yap, T.A.; O’Carrigan, B.; Penney, M.S.; Lim, J.S.; Brown, J.S.; de Miguel Luken, M.J.; Tunariu, N.; Perez-Lopez, R.; Rodrigues, D.N.; Riisnaes, R.; et al. Phase I Trial of First-in-Class ATR Inhibitor M6620 (VX-970) as Monotherapy or in Combination With Carboplatin in Patients With Advanced Solid Tumors. J. Clin. Oncol. 2020, 38, 3195–3204. [Google Scholar] [CrossRef] [PubMed]
- Shapiro, G.I.; Wesolowski, R.; Devoe, C.; Lord, S.; Pollard, J.; Hendriks, B.S.; Falk, M.; Diaz-Padilla, I.; Plummer, R.; Yap, T.A. Phase 1 Study of the ATR Inhibitor Berzosertib in Combination with Cisplatin in Patients with Advanced Solid Tumours. Br. J. Cancer 2021, 125, 520–527. [Google Scholar] [CrossRef] [PubMed]
- Barnieh, F.M.; Loadman, P.M.; Falconer, R.A. Progress towards a Clinically-Successful ATR Inhibitor for Cancer Therapy. Curr Res. Pharmacol. Drug Discov. 2021, 2, 100017. [Google Scholar] [CrossRef] [PubMed]
- Testing the Addition of an Anti-Cancer Drug, BAY 1895344, to the Usual Chemotherapy Treatment (Cisplatin, or Cisplatin and Gemcitabine) for Advanced Solid Tumors with Emphasis on Urothelial Cancer. Available online: https://clinicaltrials.gov/ct2/show/NCT04491942 (accessed on 8 March 2022).
- Reinhardt, H.C.; Yaffe, M.B. Kinases That Control the Cell Cycle in Response to DNA Damage: Chk1, Chk2, and MK2. Curr. Opin. Cell Biol. 2009, 21, 245–255. [Google Scholar] [CrossRef]
- Hematulin, A.; Sagan, D.; Sawanyawisuth, K.; Seubwai, W.; Wongkham, S. Association between Cellular Radiosensitivity and G1/G2 Checkpoint Proficiencies in Human Cholangiocarcinoma Cell Lines. Int J. Oncol. 2014, 45, 1159–1166. [Google Scholar] [CrossRef]
- Moore, K.N.; Hong, D.S.; Patel, M.R.; Pant, S.; Ulahannan, S.V.; Jones, S.; Meric-Bernstam, F.; Wang, J.S.; Aljumaily, R.; Hamilton, E.P.; et al. A Phase 1b Trial of Prexasertib in Combination with Standard-of-Care Agents in Advanced or Metastatic Cancer. Target. Oncol. 2021, 16, 569–589. [Google Scholar] [CrossRef]
- Mohiuddin, I.S.; Kang, M.H. DNA-PK as an Emerging Therapeutic Target in Cancer. Front. Oncol. 2019, 9, 635. [Google Scholar] [CrossRef]
- Yu, Z.; Sui, J.; Ding, Y.; Cao, Z.; Zhou, P.; Wu, D. Expression of DNA-PK in hepato- and cholangio-neoplasms and its significance. Zhonghua Gan Zang Bing Za Zhi 2004, 12, 652–655. [Google Scholar]
- Medová, M.; Medo, M.; Hovhannisyan, L.; Muñoz-Maldonado, C.; Aebersold, D.M.; Zimmer, Y. DNA-PK in Human Malignant Disorders: Mechanisms and Implications for Pharmacological Interventions. Pharmacol. Ther. 2020, 215, 107617. [Google Scholar] [CrossRef]
- Lustri, A.M.; Di Matteo, S.; Fraveto, A.; Costantini, D.; Cantafora, A.; Napoletano, C.; Bragazzi, M.C.; Giuliante, F.; De Rose, A.M.; Berloco, P.B.; et al. TGF-β Signaling Is an Effective Target to Impair Survival and Induce Apoptosis of Human Cholangiocarcinoma Cells: A Study on Human Primary Cell Cultures. PLoS ONE 2017, 12, e0183932. [Google Scholar] [CrossRef]
- van Bussel, M.T.J.; Awada, A.; de Jonge, M.J.A.; Mau-Sørensen, M.; Nielsen, D.; Schöffski, P.; Verheul, H.M.W.; Sarholz, B.; Berghoff, K.; El Bawab, S.; et al. A First-in-Man Phase 1 Study of the DNA-Dependent Protein Kinase Inhibitor Peposertib (Formerly M3814) in Patients with Advanced Solid Tumours. Br. J. Cancer 2021, 124, 728–735. [Google Scholar] [CrossRef]
- National Cancer Institute (NCI). A Phase I/II Study of M3814 and Avelumab in Combination With Hypofractionated Radiation in Patients With Advanced/Metastatic Solid Tumors and Hepatobiliary Malignancies. 2022. Available online: https://clinicaltrials.gov/ct2/show/NCT04068194 (accessed on 8 March 2022).
- Lens, S.M.A.; Voest, E.E.; Medema, R.H. Shared and Separate Functions of Polo-like Kinases and Aurora Kinases in Cancer. Nat. Rev. Cancer 2010, 10, 825–841. [Google Scholar] [CrossRef] [PubMed]
- Mundt, K.E.; Golsteyn, R.M.; Lane, H.A.; Nigg, E.A. On the Regulation and Function of Human Polo-like Kinase 1 (PLK1): Effects of Overexpression on Cell Cycle Progression. Biochem. Biophys. Res. Commun. 1997, 239, 377–385. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Sun, Q.; Wang, X. PLK1, A Potential Target for Cancer Therapy. Transl. Oncol. 2017, 10, 22–32. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Zhang, Y.; Yang, X.; Chen, T.; Han, T. Analysis of Differentially Expressed MRNAs and the Prognosis of Cholangiocarcinoma Based on TCGA Database. Transl. Cancer Res. 2020, 9, 4739–4749. [Google Scholar] [CrossRef] [PubMed]
- Lin, P.; Zhong, X.-Z.; Wang, X.-D.; Li, J.-J.; Zhao, R.-Q.; He, Y.; Jiang, Y.-Q.; Huang, X.-W.; Chen, G.; He, Y.; et al. Survival Analysis of Genome-Wide Profiles Coupled with Connectivity Map Database Mining to Identify Potential Therapeutic Targets for Cholangiocarcinoma. Oncol. Rep. 2018, 40, 3189–3198. [Google Scholar] [CrossRef]
- Fingas, C.D.; Mertens, J.C.; Razumilava, N.; Sydor, S.; Bronk, S.F.; Christensen, J.D.; Rizvi, S.H.; Canbay, A.; Treckmann, J.W.; Paul, A.; et al. Polo-like Kinase 2 Is a Mediator of Hedgehog Survival Signaling in Cholangiocarcinoma. Hepatology 2013, 58, 1362–1374. [Google Scholar] [CrossRef]
- Sydor, S.; Jafoui, S.; Wingerter, L.; Swoboda, S.; Mertens, J.C.; Gerken, G.; Canbay, A.; Paul, A.; Fingas, C.D. Bcl-2 Degradation Is an Additional pro-Apoptotic Effect of Polo-like Kinase Inhibition in Cholangiocarcinoma Cells. World J. Gastroenterol 2017, 23, 4007–4015. [Google Scholar] [CrossRef]
- Thrum, S.; Lorenz, J.; Mössner, J.; Wiedmann, M. Polo-like Kinase 1 Inhibition as a New Therapeutic Modality in Therapy of Cholangiocarcinoma. Anticancer Res. 2011, 31, 3289–3299. [Google Scholar]
- Malacrida, A.; Rigolio, R.; Celio, L.; Damian, S.; Cavaletti, G.; Mazzaferro, V.; Miloso, M. In Vitro Evaluation of Rigosertib Antitumoral and Radiosensitizing Effects against Human Cholangiocarcinoma Cells. Int. J. Mol. Sci. 2021, 22, 8230. [Google Scholar] [CrossRef] [PubMed]
- Nokihara, H.; Yamada, Y.; Fujiwara, Y.; Yamamoto, N.; Wakui, H.; Nakamichi, S.; Kitazono, S.; Inoue, K.; Harada, A.; Taube, T.; et al. Phase I Trial of Volasertib, a Polo-like Kinase Inhibitor, in Japanese Patients with Advanced Solid Tumors. Invest New Drugs 2016, 34, 66–74. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.-C.; Su, W.-C.; Yen, C.-J.; Hsu, C.-H.; Su, W.-P.; Yeh, K.-H.; Lu, Y.-S.; Cheng, A.-L.; Huang, D.C.-L.; Fritsch, H.; et al. A Phase I Study of Two Dosing Schedules of Volasertib (BI 6727), an Intravenous Polo-like Kinase Inhibitor, in Patients with Advanced Solid Malignancies. Br. J. Cancer 2014, 110, 2434–2440. [Google Scholar] [CrossRef] [PubMed]
- Academic and Community Cancer Research United. Phase I Study of Irinotecan Liposome (Nal-IRI), Fluorouracil, Leucovorin and Rucaparib in the Treatment of Select Gastrointestinal Metastatic Malignancies Followed by a Phase Ib of First and Second Line Treatment of Both Unselected and Selected (for BRCA 1/2 and PALB2 Mutations) Patients with Metastatic Adenocarcinoma of the Pancreas Then Followed by a Phase II Study of First Line Treatment of Selected Patients with Metastatic Adenocarcinoma of the Pancreas with Genomic Markers (Signature) of Homologous Recombination Deficiency (HRD). 2022. Available online: https://clinicaltrials.gov/ct2/show/NCT03337087 (accessed on 8 March 2022).
- Cleary, J.M.; Aguirre, A.J.; Shapiro, G.I.; D’Andrea, A.D. Biomarker-Guided Development of DNA Repair Inhibitors. Mol. Cell 2020, 78, 1070–1085. [Google Scholar] [CrossRef] [PubMed]
- Goodman, A.M.; Kato, S.; Bazhenova, L.; Patel, S.P.; Frampton, G.M.; Miller, V.; Stephens, P.J.; Daniels, G.A.; Kurzrock, R. Tumor Mutational Burden as an Independent Predictor of Response to Immunotherapy in Diverse Cancers. Mol. Cancer Ther. 2017, 16, 2598–2608. [Google Scholar] [CrossRef]
- Spizzo, G.; Puccini, A.; Xiu, J.; Goldberg, R.M.; Grothey, A.; Shields, A.F.; Arora, S.P.; Khushman, M.; Salem, M.E.; Battaglin, F.; et al. Molecular Profile of BRCA-Mutated Biliary Tract Cancers. ESMO Open 2020, 5, e000682. [Google Scholar] [CrossRef]
- Pilié, P.G.; Gay, C.M.; Byers, L.A.; O’Connor, M.J.; Yap, T.A. PARP Inhibitors: Extending Benefit Beyond BRCA-Mutant Cancers. Clin. Cancer Res. 2019, 25, 3759–3771. [Google Scholar] [CrossRef]
- Lord, C.J.; Ashworth, A. BRCAness Revisited. Nat. Rev. Cancer 2016, 16, 110–120. [Google Scholar] [CrossRef]
- Bajrami, I.; Frankum, J.R.; Konde, A.; Miller, R.E.; Rehman, F.L.; Brough, R.; Campbell, J.; Sims, D.; Rafiq, R.; Hooper, S.; et al. Genome-Wide Profiling of Genetic Synthetic Lethality Identifies CDK12 as a Novel Determinant of PARP1/2 Inhibitor Sensitivity. Cancer Res. 2014, 74, 287–297. [Google Scholar] [CrossRef]
- Sabbatino, F.; Liguori, L.; Malapelle, U.; Schiavi, F.; Tortora, V.; Conti, V.; Filippelli, A.; Tortora, G.; Ferrone, C.R.; Pepe, S. Case Report: BAP1 Mutation and RAD21 Amplification as Predictive Biomarkers to PARP Inhibitor in Metastatic Intrahepatic Cholangiocarcinoma. Front. Oncol. 2020, 10, 567289. [Google Scholar] [CrossRef]
- Kuznetsov, J.N.; Aguero, T.H.; Owens, D.A.; Kurtenbach, S.; Field, M.G.; Durante, M.A.; Rodriguez, D.A.; King, M.L.; Harbour, J.W. BAP1 Regulates Epigenetic Switch from Pluripotency to Differentiation in Developmental Lineages Giving Rise to BAP1-Mutant Cancers. Sci. Adv. 2019, 5, eaax1738. [Google Scholar] [CrossRef]
- Kuznetsoff, J.N.; Owens, D.A.; Lopez, A.; Rodriguez, D.A.; Chee, N.T.; Kurtenbach, S.; Bilbao, D.; Roberts, E.R.; Volmar, C.-H.; Wahlestedt, C.; et al. Dual Screen for Efficacy and Toxicity Identifies HDAC Inhibitor with Distinctive Activity Spectrum for BAP1-Mutant Uveal Melanoma. Mol. Cancer Res. 2021, 19, 215–222. [Google Scholar] [CrossRef]
- Sacco, J.J.; Kenyani, J.; Butt, Z.; Carter, R.; Chew, H.Y.; Cheeseman, L.P.; Darling, S.; Denny, M.; Urbé, S.; Clague, M.J.; et al. Loss of the Deubiquitylase BAP1 Alters Class I Histone Deacetylase Expression and Sensitivity of Mesothelioma Cells to HDAC Inhibitors. Oncotarget 2015, 6, 13757–13771. [Google Scholar] [CrossRef]
- Pant, K.; Peixoto, E.; Richard, S.; Gradilone, S.A. Role of Histone Deacetylases in Carcinogenesis: Potential Role in Cholangiocarcinoma. Cells 2020, 9, E780. [Google Scholar] [CrossRef]
- Sidney Kimmel Comprehensive Cancer Center at Johns Hopkins. A Phase 2 Clinical Trial of Entinostat in Combination with Nivolumab for Patients with Previously Treated Unresectable or Metastatic Cholangiocarcinoma and Pancreatic Adenocarcinoma. 2021. Available online: https://clinicaltrials.gov/ct2/show/NCT03250273 (accessed on 8 March 2022).
- Xu, Y.-Y.; Ren, Z.-L.; Liu, X.-L.; Zhang, G.-M.; Huang, S.-S.; Shi, W.-H.; Ye, L.-X.; Luo, X.; Liu, S.-W.; Li, Y.-L.; et al. BAP1 Loss Augments Sensitivity to BET Inhibitors in Cancer Cells. Acta Pharmacol. Sin. 2021. [Google Scholar] [CrossRef]
- Kay, J.; Thadhani, E.; Samson, L.; Engelward, B. Inflammation-Induced DNA Damage, Mutations and Cancer. DNA Repair 2019, 83, 102673. [Google Scholar] [CrossRef] [PubMed]
- Loeuillard, E.; Conboy, C.B.; Gores, G.J.; Rizvi, S. Immunobiology of Cholangiocarcinoma. JHEP Rep. 2019, 1, 297–311. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Zhang, Q.; Wang, R.; Li, Y.; Sun, Y.; Yang, L. Targeting DNA Damage Repair for Immune Checkpoint Inhibition: Mechanisms and Potential Clinical Applications. Front. Oncol. 2021, 11, 648687. [Google Scholar] [CrossRef] [PubMed]
- Tommelein, J.; De Vlieghere, E.; Verset, L.; Melsens, E.; Leenders, J.; Descamps, B.; Debucquoy, A.; Vanhove, C.; Pauwels, P.; Gespach, C.P.; et al. Radiotherapy-Activated Cancer-Associated Fibroblasts Promote Tumor Progression through Paracrine IGF1R Activation. Cancer Res. 2018, 78, 659–670. [Google Scholar] [CrossRef]
- Gomez-Sarosi, L.; Sun, Y.; Coleman, I.; Bianchi-Frias, D.; Nelson, P.S. DNA Damage Induces a Secretory Program in the Quiescent TME That Fosters Adverse Cancer Phenotypes. Mol. Cancer Res. 2017, 15, 842–851. [Google Scholar] [CrossRef]
- Banerjee, J.; Mishra, R.; Li, X.; Jackson, R.S.; Sharma, A.; Bhowmick, N.A. A Reciprocal Role of Prostate Cancer on Stromal DNA Damage. Oncogene 2014, 33, 4924–4931. [Google Scholar] [CrossRef] [PubMed]
- Louis, C.; Edeline, J.; Coulouarn, C. Targeting the Tumor Microenvironment in Cholangiocarcinoma: Implications for Therapy. Expert Opin. Ther. Targets 2021, 25, 153–162. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).