Abstract
The outbreak of Coronavirus Disease 2019 (COVID-19), a severe respiratory disease caused by betacoronavirus SARS-CoV-2, in 2019 that further developed into a pandemic has received an unprecedented response from the scientific community and sparked a general research interest into the biology and ecology of Coronaviridae, a family of positive-sense single-stranded RNA viruses. Aquatic environments, lakes, rivers and ponds, are important habitats for bats and birds, which are hosts for various coronavirus species and strains and which shed viral particles in their feces. It is therefore of high interest to fully explore the role that aquatic environments may play in coronavirus spread, including cross-species transmissions. Besides the respiratory tract, coronaviruses pathogenic to humans can also infect the digestive system and be subsequently defecated. Considering this, it is pivotal to understand whether wastewater can play a role in their dissemination, particularly in areas with poor sanitation. This review provides an overview of the taxonomy, molecular biology, natural reservoirs and pathogenicity of coronaviruses; outlines their potential to survive in aquatic environments and wastewater; and demonstrates their association with aquatic biota, mainly waterfowl. It also calls for further, interdisciplinary research in the field of aquatic virology to explore the potential hotspots of coronaviruses in the aquatic environment and the routes through which they may enter it.
1. Introduction
The outbreak of severe acute respiratory syndrome caused by betacoronavirus SARS-CoV-2 (provisionally known as 2019nCoV) began at the end of 2019 in China and further spread to other countries [1] and across different continents, forcing the World Health Organization (WHO) to first declare a Public Health Emergency of International Concern at the end of January 2020 [2] and to later announce a pandemic of COVID-19 in March 2020 [3]. This is the first time that any coronavirus has sparked such an epidemiological situation, although some other coronaviruses were already known to reveal pathogenicity to humans. SARS-CoV-2 and COVID-19 have received unprecedented research interest encompassing fields of molecular biology [4], mechanism of cell entry [5], diagnostics [6], epidemiological modeling [7], immunology [8], experimental therapies and vaccine development [9,10], clinical medicine [11,12], prejudice and xenophobia [13] and the psychological effects of the pandemic [14,15]. It has also renewed a general interest in coronavirus biology and ecology.
The aquatic environment can be a source of an uncountable number of microorganisms pathogenic to different aquatic and terrestrial animals, as well as humans [16,17,18]. Lakes and rivers are an important habitat for bats and birds (including waterfowl), which represent one of the main reservoirs for various coronaviruses [19,20]. It is established that these animals can shed coronaviral RNA through feces, although it remains to be explored whether this is related to the presence of infectious viral particles [21,22]. Nevertheless, it is of high interest to investigate the association between aquatic environments and this viral group. It would add to the understanding of the role that these ecosystems may potentially play in infections within species as well as in cross-species transmission.
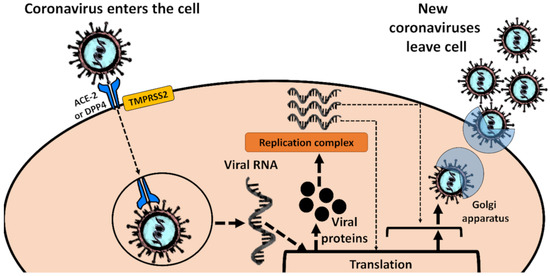
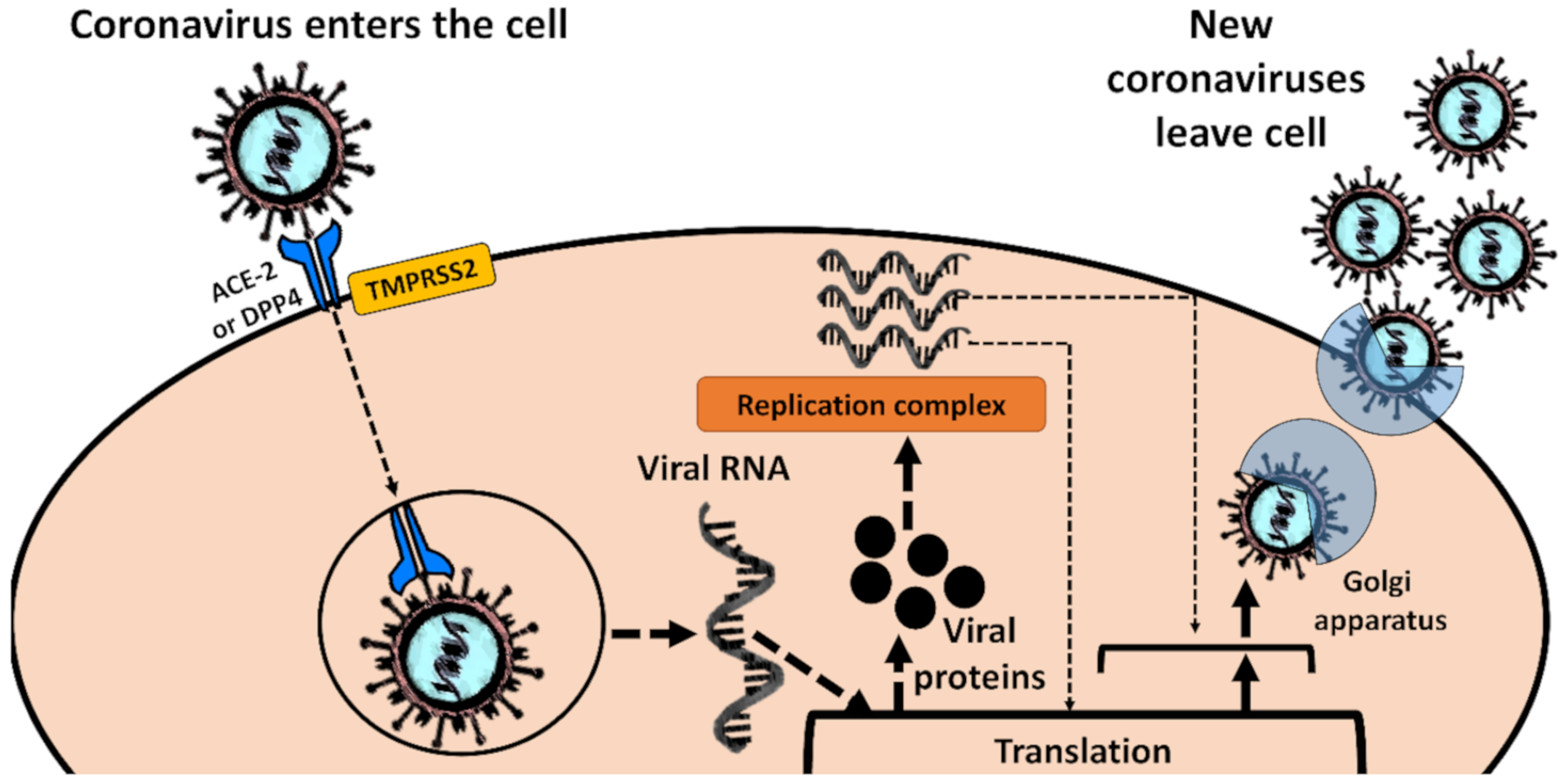
Understanding the role of aquatic ecosystems in this context is also essential from a human health perspective, since untreated water, a well-established source of various pathogens, is used in various areas, particularly those with poor sanitation. It would also help to understand whether aquatic biota could play a potential role as an intermediate host from which humans could contract the coronavirus. Such a process would, however, require the spike glycoprotein that mediates coronavirus entry into cells [23,24] to be first optimized in such a host for binding to human-like angiotensin-converting enzyme 2 (ACE2), alanyl aminopeptidase (CD13), dipeptidyl peptidase 4 (CD26) or other entry receptors through natural selection [5,25].
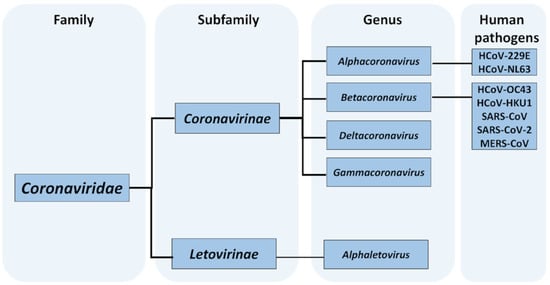
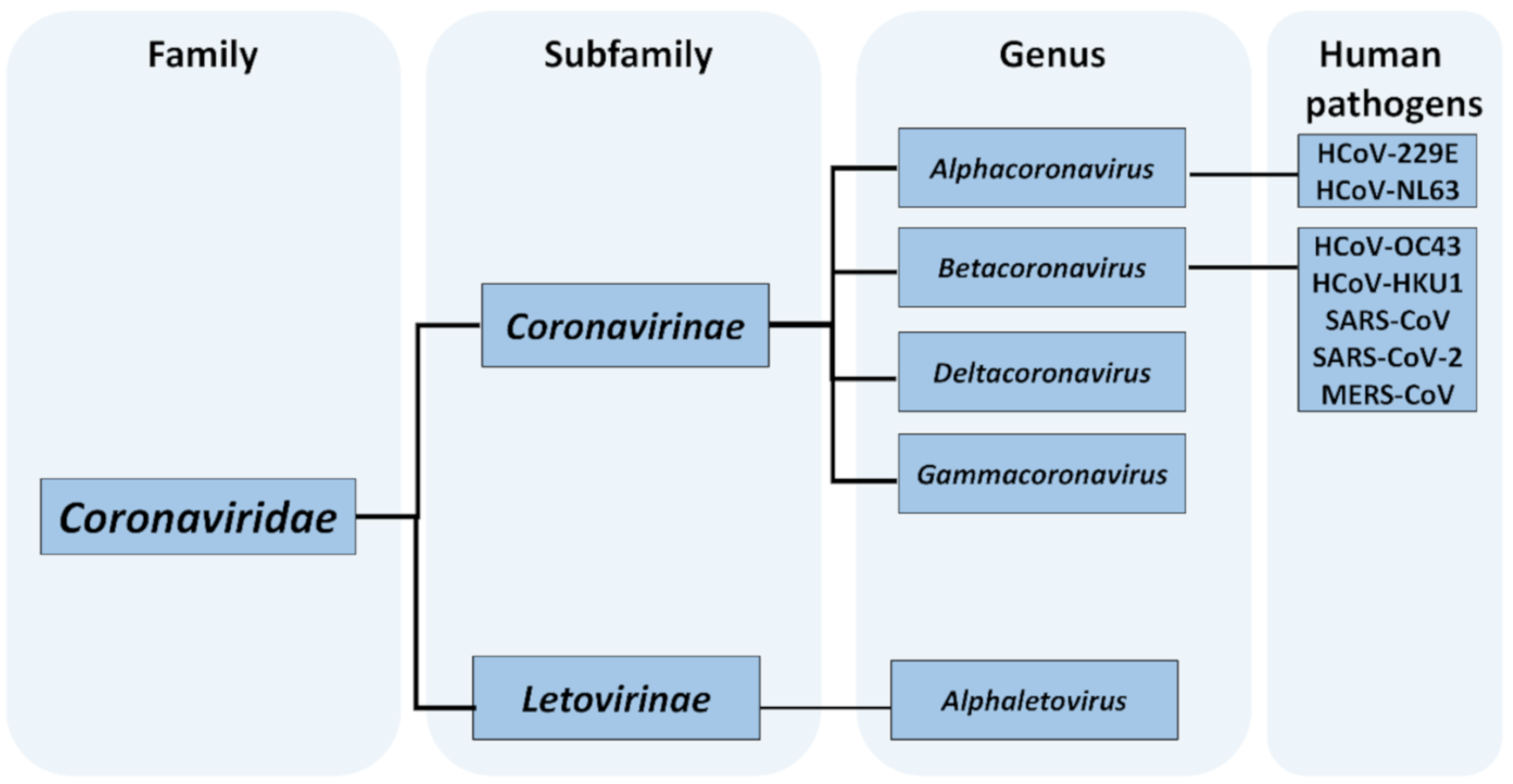
It has been demonstrated that the discharge of inadequately treated sewage effluents is a significant source of enteric viral pathogens in lakes and rivers, including those which are the leading causes of recreational waterborne illnesses [26,27,28]. All seven coronaviruses pathogenic to humans (SARS-CoV, SARS-CoV-2, MERS-CoV, HCoV-229E, HCoV-NL63, HCoV-OC43 and HCoV-HKU1) can infect the respiratory tract and digestive system, and their RNA material can be present in stool [29]. One should note that the detection of the genetic material in fecal material does not necessarily indicate that infectious virions are defecated. However, the presence of cultivable SARS-CoV in stool was already reported, and some preliminary observations, based on a small number of patients [30], suggest that this may also be a case for SARS-CoV-2 [31,32]. How frequently the infectious virus can be present in the human stool and what viral loads can be expected are yet to be explored. Such data would enable understanding whether wastewater, particularly untreated, may serve as a route of their dissemination to the aquatic environment. The understanding of the scale of this process first requires an exploration of the survival of coronaviruses in sewage and its treatment with various methods as well as a monitoring of human pathogenic coronaviruses in untreated and treated wastewater.
In the present review, we provide a brief overview of coronaviruses, their taxonomy, molecular biology, natural reservoirs and pathogenicity; outline their potential to survive in aquatic environments and wastewater; and demonstrate their association with aquatic biota, mainly waterfowl, and other animal species related to aquatic ecosystems. Future research prospects regarding the association between selected coronaviruses and water-related issues are put forward with a call for interdisciplinary research in the field of aquatic virology.
4. Conclusions
Considering that seven strains of coronaviruses are already known to infect humans, some of which can cause severe respiratory disease, and in light of the SARS-CoV-2-caused pandemic with global economic consequences, it is of high interest to further explore all possible routes and intermediate hosts via which further strains pathogenic to humans may emerge. Although the body of work on coronaviruses is extensive, there are numerous knowledge gaps that require further studies. This paper gives an overview of coronaviruses, their survival in the aquatic environment, their association with aquatic biota and their potential to enter aquatic ecosystems via wastewater. Further research is required to explore γ-CoVs and δ-CoVs associated with aquatic birds inhabiting different geographical locations. Considering that waterfowls, as well as bats, can deposit a large mass of droppings directly into the water, it is of high interest to investigate the survival and infectivity of various coronavirus strains related to these animals under realistic conditions of aquatic ecosystems. This is pivotal to understanding whether aquatic environments inhabited by large populations of flock-forming or colony-forming waterfowls, and frequently visited by bats, can represent hotspots of coronaviruses with potential for cross-species transmission. Moreover, there is a need to further explore the understudied Letovirinae subfamily, which is currently represented by only one known species (associated with an amphibian species). It is plausible that marine mammalian species can constitute a significant reservoir for coronaviruses of different genera, but this requires surveillance in wild animals found dead and/or individuals kept in captivity in aquatic parks. Finally, not much is known on the mechanisms of infection of coronaviruses associated with aquatic birds and marine mammals. Any studies identifying potential receptors employed in this process would be valuable to estimate the risks of interspecies transmissions, including those to humans.
Author Contributions
Conceptualization, A.W. and P.R.; writing—Original draft preparation, A.W. and P.R.; supervision, P.R. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Wu, D.; Wu, T.; Liu, Q.; Yang, Z. The SARS-CoV-2 outbreak: What we know. Int. J. Infect. Dis. 2020, 94, 44–48. [Google Scholar] [CrossRef] [PubMed]
- Sohrabi, C.; Alsafi, Z.; O’Neill, N.; Khan, M.; Kerwan, A.; Al-Jabir, A.; Iosifidis, C.; Agha, R. World Health Organization declares global emergency: A review of the 2019 novel coronavirus (COVID-19). Int. J. Surg. 2020, 76, 71–76. [Google Scholar] [CrossRef]
- Cucinotta, D.; Vanelli, M. WHO Declares COVID-19 a Pandemic. Acta Bio Med. Atenei Parm. 2020, 91, 157–160. [Google Scholar] [CrossRef]
- Andersen, K.G.; Rambaut, A.; Lipkin, W.I.; Holmes, E.C.; Garry, R.F. The proximal origin of SARS-CoV-2. Nat. Med. 2020, 26, 450–452. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Kruger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. SARS-CoV-2 Cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 2020, 181, 271–280. [Google Scholar] [CrossRef] [PubMed]
- Ai, T.; Yang, Z.; Hou, H.; Zhan, C.; Chen, C.; Lv, W.; Tao, Q.; Sun, Z.; Xia, L. Correlation of Chest CT and RT-PCR testing in coronavirus disease 2019 (COVID-19) in China: A report of 1014 cases. Radiology 2020, 200642. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.M.; Rui, J.; Wang, Q.P.; Zhao, Z.Y.; Cui, J.A.; Yin, L. A mathematical model for simulating the phase-based transmissibility of a novel coronavirus. Infect. Dis. Poverty 2020, 9, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Wu, D.; Guo, W.; Cao, Y.; Huang, D.; Wang, H.; Wang, T.; Zhang, X.; Chen, H.; Yu, H.; et al. Clinical and immunologic features in severe and moderate Coronavirus Disease 2019. J. Clin. Investig. 2020, 130, 2620–2629. [Google Scholar] [CrossRef]
- Rismanbaf, A. Potential treatments for COVID-19; a narrative literature review. Arch. Acad. Emerg. Med. 2020, 8, e29. [Google Scholar]
- Jin, Y.; Yang, H.; Ji, W.; Wu, W.; Chen, S. Virology, epidemiology, pathogenesis, and control of COVID-19. Viruses 2020, 12, 372. [Google Scholar] [CrossRef]
- Sun, D.; Li, H.; Lu, X.X.; Xiao, H.; Ren, J.; Zhang, F.R.; Liu, Z.S. Clinical features of severe pediatric patients with coronavirus disease 2019 in Wuhan: A single center’s observational study. World J. Pediatr. 2020. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Hu, B.; Hu, C.; Zhu, F.; Liu, X.; Zhang, J.; Wang, B.; Xiang, H.; Cheng, Z.; Xiong, Y.; et al. Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus-infected pneumonia in Wuhan, China. JAMA 2020, 323, 1061–1069. [Google Scholar] [CrossRef] [PubMed]
- Rzymski, P.; Nowicki, M. Preventing COVID-19 prejudice in academia. Science 2020, 368, 1313. [Google Scholar] [CrossRef]
- Li, S.; Wang, Y.; Xue, J.; Zhao, N.; Zhu, T. The impact of COVID-19 epidemic declaration on psychological consequences: A study on active Weibo users. Int. J. Environ. Res. Public Health 2020, 17, 2032. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Pan, R.; Wan, X.; Tan, Y.; Xu, L.; Ho, C.S. Immediate psychological responses and associated factors during the initial stage of the 2019 coronavirus disease (COVID-19) epidemic among the general population in China. Int. J. Environ. Res. Public Health 2020, 17, 1729. [Google Scholar] [CrossRef]
- Klimaszyk, P.; Rzymski, P. The complexity of ecological impacts induced by great cormorants. Hydrobiologia 2016, 771, 13–30. [Google Scholar] [CrossRef]
- Murray, A.G. Using simple models to review the application and implications of different approaches used to simulate transmission of pathogens among aquatic animals. Prev. Vet. Med. 2009, 88, 167–177. [Google Scholar] [CrossRef]
- Oidtmann, B.; Dixon, P.; Way, K.; Joiner, C.; Bayley, A.E. Risk of waterborne virus spread—review of survival of relevant fish and crustacean viruses in the aquatic environment and implications for control measures. Rev. Aquac. 2018, 10, 641–669. [Google Scholar] [CrossRef]
- Fan, Y.; Zhao, K.; Shi, Z.-L.; Zhou, P. Bat coronaviruses in China. Viruses 2019, 11, 210. [Google Scholar] [CrossRef]
- Miłek, J.; Blicharz-Domańska, K. Coronaviruses in avian species—Review with focus on epidemiology and diagnosis in wild birds. J. Vet. Res. 2018, 62, 249–255. [Google Scholar] [CrossRef]
- Dominguez, S.R.; O’Shea, T.J.; Oko, L.M.; Holmes, K.V. Detection of group 1 coronaviruses in bats in North America. Emerg. Infect. Dis. 2007, 13, 1295–1300. [Google Scholar] [CrossRef] [PubMed]
- Chu, D.K.W.; Leung, C.Y.H.; Gilbert, M.; Joyner, P.H.; Ng, E.M.; Tse, T.M.; Guan, Y.; Peiris, J.S.M.; Poon, L.L.M. Avian coronavirus in wild aquatic birds. J. Virol. 2011, 85, 12815–12820. [Google Scholar] [CrossRef] [PubMed]
- Li, F. Structure, Function, and evolution of coronavirus spike proteins. Ann. Rev. Virol. 2016, 3, 237–261. [Google Scholar] [CrossRef] [PubMed]
- Bosch, B.J.; van der Zee, R.; de Haan, C.A.M.; Rottier, P.J.M. The Coronavirus spike protein is a class i virus fusion protein: Structural and Functional characterization of the fusion core complex. J. Virol. 2003, 77, 8801–8811. [Google Scholar] [CrossRef]
- Hofmann, H.; Simmons, G.; Rennekamp, A.J.; Chaipan, C.; Gramberg, T.; Heck, E.; Geier, M.; Wegele, A.; Marzi, A.; Bates, P.; et al. Highly conserved regions within the spike proteins of human coronaviruses 229E and NL63 determine recognition of their respective cellular receptors. J. Virol. 2006, 80, 8639–8652. [Google Scholar] [CrossRef]
- Okoh, A.I.; Sibanda, T.; Gusha, S.S. Inadequately treated wastewater as a source of human enteric viruses in the environment. Int. J. Environ. Res. Public Health 2010, 7, 2620–2637. [Google Scholar] [CrossRef]
- Lenaker, P.L.; Corsi, S.R.; Borchardt, M.A.; Spencer, S.K.; Baldwin, A.K.; Lutz, M.A. Hydrologic, land cover, and seasonal patterns of waterborne pathogens in Great Lakes tributaries. Water Res. 2017, 113, 11–21. [Google Scholar] [CrossRef]
- Eftim, S.E.; Hong, T.; Soller, J.; Boehm, A.; Warren, I.; Ichida, A.; Nappier, S.P. Occurrence of norovirus in raw sewage—A systematic literature review and meta-analysis. Water Res. 2017, 111, 366–374. [Google Scholar] [CrossRef]
- Yang, P.; Wang, X. COVID-19: A new challenge for human beings. Cell. Mol. Immunol. 2020, 17, 555–557. [Google Scholar] [CrossRef]
- Xu, D.; Zhang, Z.; Jin, L.; Chu, F.; Mao, Y.; Wang, H.; Liu, M.; Wang, M.; Zhang, L.; Gao, G.F.; et al. Persistent shedding of viable SARS-CoV in urine and stool of SARS patients during the convalescent phase. Eur. J. Clin. Microbiol. Infect. Dis. 2005, 24, 165–171. [Google Scholar] [CrossRef]
- Wang, W.; Xu, Y.; Gao, R.; Lu, R.; Han, K.; Wu, G.; Tan, W. Detection of SARS-CoV-2 in different types of clinical specimens. JAMA 2020, 323, 1843–1844. [Google Scholar] [CrossRef] [PubMed]
- Xiao, F.; Tang, M.; Zheng, X.; Liu, Y.; Li, X.; Shan, H. Evidence for gastrointestinal infection of SARS-CoV-2. Gastroenterology 2020, 158, 1831–1833. [Google Scholar] [CrossRef] [PubMed]
- ICTV. International Committee on Taxonomy of Viruses. Master Species List 2019, 2. [Google Scholar] [CrossRef]
- Cui, J.; Li, F.; Shi, Z.-L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 2019, 17, 181–192. [Google Scholar] [CrossRef] [PubMed]
- Bukhari, K.; Mulley, G.; Gulyaeva, A.A.; Zhao, L.; Shu, G.; Jiang, J.; Neuman, B.W. Description and initial characterization of metatranscriptomic nidovirus-like genomes from the proposed new family Abyssoviridae, and from a sister group to the Coronavirinae, the proposed genus Alphaletovirus. Virology 2018, 524, 160–171. [Google Scholar] [CrossRef] [PubMed]
- Carstens, E.B. Ratification vote on taxonomic proposals to the International Committee on Taxonomy of Viruses (2009). Arch. Virol. 2010, 155, 133–146. [Google Scholar] [CrossRef]
- Groot, R.J.d.; Ziebuhr, J.; Poon, L.L.; Woo, P.C.; Talbot, P.; Rottier, P.J.M.; Holmes, K.V.; Baric, R.; Perlman, S.; Enjuanes, L.; et al. Revision of the family Coronaviridae. Taxonomic Proposal to the ICTV Executive Committee. 2008. Available online: http://talk.ictvonline.org/files/ictv_official_taxonomy_updates_since_the_8th_report/m/vertebrate-official/default.aspx?pi3174=3 (accessed on 18 May 2020).
- Pellett, P.E.; Mitra, S.; Holland, T.C. Chapter 2—Basics of virology. In Handbook Clinical Neurology; Tselis, A.C., Booss, J., Eds.; Elsevier: New York, NY, USA, 2014; Volume 123, pp. 45–66. [Google Scholar]
- Chen, N.; Zhou, M.; Dong, X.; Qu, J.; Gong, F.; Han, Y.; Qiu, Y.; Wang, J.; Liu, Y.; Wei, Y.; et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: A descriptive study. Lancet 2020, 395, 507–513. [Google Scholar] [CrossRef]
- McBride, R.; van Zyl, M.; Fielding, B.C. The coronavirus nucleocapsid is a multifunctional protein. Viruses 2014, 6, 2991–3018. [Google Scholar] [CrossRef]
- Fehr, A.R.; Perlman, S. Coronaviruses: An overview of their replication and pathogenesis. Methods Mol. Biol. 2015, 1282, 1–23. [Google Scholar] [CrossRef]
- Firth, A.E.; Brierley, I. Non-canonical translation in RNA viruses. J. Gen. Virol. 2012, 93, 1385–1409. [Google Scholar] [CrossRef]
- Lim, Y.X.; Ng, Y.L.; Tam, J.P.; Liu, D.X. Human coronaviruses: A review of virus-host interactions. Diseases 2016, 4, 26. [Google Scholar] [CrossRef] [PubMed]
- Sola, I.; Almazan, F.; Zuniga, S.; Enjuanes, L. Continuous and discontinuous RNA synthesis in coronaviruses. Ann. Rev. Virol. 2015, 2, 265–288. [Google Scholar] [CrossRef] [PubMed]
- Irigoyen, N.; Firth, A.E.; Jones, J.D.; Chung, B.Y.; Siddell, S.G.; Brierley, I. High-resolution analysis of coronavirus gene expression by RNA sequencing and ribosome profiling. PLoS Pathog. 2016, 12, e1005473. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Liu, Q.; Guo, D. Emerging coronaviruses: Genome structure, replication, and pathogenesis. J. Med. Virol. 2020, 92, 418–423. [Google Scholar] [CrossRef]
- Narayanan, K.; Huang, C.; Makino, S. SARS coronavirus accessory proteins. Virus Res. 2008, 133, 113–121. [Google Scholar] [CrossRef]
- Hogue, B.G.; Machamer, C.E. Coronavirus structural proteins and virus assembly. In Nidoviruses; American Society of Microbiology: Washington, DC, USA, 2008. [Google Scholar] [CrossRef]
- Cong, Y.; Kriegenburg, F.; de Haan, C.A.M.; Reggiori, F. Coronavirus nucleocapsid proteins assemble constitutively in high molecular oligomers. Sci. Rep. 2017, 7, 1–10. [Google Scholar] [CrossRef]
- International Committee on Taxonomy of Viruses. Family—Coronaviridae. In Virus Taxonomy; King, A.M.Q., Adams, M.J., Carstens, E.B., Lefkowitz, E.J., Eds.; Elsevier: San Diego, CA, USA, 2012; pp. 806–828. [Google Scholar] [CrossRef]
- Schoeman, D.; Fielding, B.C. Coronavirus envelope protein: Current knowledge. Virol. J. 2019, 16, 69. [Google Scholar] [CrossRef]
- Shang, J.; Wan, Y.; Liu, C.; Yount, B.; Gully, K.; Yang, Y.; Auerbach, A.; Peng, G.; Baric, R.; Li, F. Structure of mouse coronavirus spike protein complexed with receptor reveals mechanism for viral entry. PLoS Pathog. 2020, 16, e1008392. [Google Scholar] [CrossRef]
- Davies, H.A.; Macnaughton, M.R. Comparison of the morphology of three coronaviruses. Arch. Virol. 1979, 59, 25–33. [Google Scholar] [CrossRef]
- Lin, Y.; Yan, X.; Cao, W.; Wang, C.; Feng, J.; Duan, J.; Xie, S. Probing the structure of the SARS coronavirus using scanning electron microscopy. Antivir. Ther. 2004, 9, 287–289. [Google Scholar]
- Masters, P.S. The molecular biology of coronaviruses. Adv. Virus Res. 2006, 66, 193–292. [Google Scholar] [CrossRef] [PubMed]
- De Haan, C.A.M.; de Wit, M.; Kuo, L.; Montalto-Morrison, C.; Haagmans, B.L.; Weiss, S.R.; Masters, P.S.; Rottier, P.J.M. The glycosylation status of the murine hepatitis coronavirus M protein affects the interferogenic capacity of the virus in vitro and its ability to replicate in the liver but not the brain. Virology 2003, 312, 395–406. [Google Scholar] [CrossRef]
- Tatar, G.; Taskin Tok, T. Structures and functions of coronavirus proteins: Molecular modeling of viral nucleoprotein. Int. J. Virol. Infect. Dis. 2017, 2, 001. [Google Scholar]
- Ye, Z.-W.; Yuan, S.; Yuen, K.-S.; Fung, S.-Y.; Chan, C.-P.; Jin, D.-Y. Zoonotic origins of human coronaviruses. Int. J. Biol. Sci. 2020, 16, 1686–1697. [Google Scholar] [CrossRef]
- Corman, V.M.; Muth, D.; Niemeyer, D.; Drosten, C. Hosts and Sources of endemic human coronaviruses. Adv. Virus Res. 2018, 100, 163–188. [Google Scholar] [CrossRef]
- Eboriadou, M.; Haidopoulou, K.; Xanthou, P.; Papa, A. Coronaviruses OC43 and 229E lower respiratory tract co-infections: A clinical report of two cases. Arch. Med. Sci. 2008, 4, 88–90. [Google Scholar]
- Pene, F.; Merlat, A.; Vabret, A.; Rozenberg, F.; Buzyn, A.; Dreyfus, F.; Cariou, A.; Freymuth, F.; Lebon, P. Coronavirus 229E-related pneumonia in immunocompromised patients. Clin. Infect. Dis. 2003, 37, 929–932. [Google Scholar] [CrossRef]
- Walsh, E.E.; Shin, J.H.; Falsey, A.R. Clinical impact of human coronaviruses 229E and OC43 infection in diverse adult populations. J. Infect. Dis. 2013, 208, 1634–1642. [Google Scholar] [CrossRef]
- Gorse, G.J.; O’Connor, T.Z.; Hall, S.L.; Vitale, J.N.; Nichol, K.L. Human coronavirus and acute respiratory illness in older adults with chronic obstructive pulmonary disease. J. Infect. Dis. 2009, 199, 847–857. [Google Scholar] [CrossRef]
- Smuts, H. Human coronavirus NL63 infections in infants hospitalised with acute respiratory tract infections in South Africa. Influ. Respir. Viruses 2008, 2, 135–138. [Google Scholar] [CrossRef]
- Arbour, N.; Day, R.; Newcombe, J.; Talbot, P.J. Neuroinvasion by human respiratory coronaviruses. J. Virol. 2000, 74, 8913–8921. [Google Scholar] [CrossRef] [PubMed]
- Risku, M.; Lappalainen, S.; Rasanen, S.; Vesikari, T. Detection of human coronaviruses in children with acute gastroenteritis. J. Clin. Virol. 2010, 48, 27–30. [Google Scholar] [CrossRef] [PubMed]
- Yeo, C.; Kaushal, S.; Yeo, D. Enteric involvement of coronaviruses: Is faecal-oral transmission of SARS-CoV-2 possible? Lancet Gastroenterol. Hepatol. 2020, 5, 335–337. [Google Scholar] [CrossRef]
- Guan, W.D.; Mok, C.K.P.; Chen, Z.L.; Feng, L.Q.; Li, Z.T.; Huang, J.C.; Ke, C.W.; Deng, X.; Ling, Y.; Wu, S.G.; et al. Characteristics of traveler with Middle East Respiratory Syndrome, China, 2015. Emerg. Infect. Dis. 2015, 21, 2278–2280. [Google Scholar] [CrossRef] [PubMed]
- Gu, J.; Han, B.; Wang, J. COVID-19: Gastrointestinal manifestations and potential fecal-oral transmission. Gastroenterology 2020, 158, 1518–1519. [Google Scholar] [CrossRef] [PubMed]
- Leung, W.K.; To, K.-F.; Chan, P.K.S.; Chan, H.L.Y.; Wu, A.K.L.; Lee, N.; Yuen, K.Y.; Sung, J.J.Y. Enteric involvement of severe acute respiratory syndrome-associated coronavirus infection. Gastroenterology 2003, 125, 1011–1017. [Google Scholar] [CrossRef]
- Ng, S.C.; Tilg, H. COVID-19 and the gastrointestinal tract: More than meets the eye. Gut 2020, 69, 973–974. [Google Scholar] [CrossRef]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Assiri, A.; Al-Tawfiq, J.A.; Al-Rabeeah, A.A.; Al-Rabiah, F.A.; Al-Hajjar, S.; Al-Barrak, A.; Flemban, H.; Al-Nassir, W.N.; Balkhy, H.H.; Al-Hakeem, R.F.; et al. Epidemiological, demographic, and clinical characteristics of 47 cases of Middle East respiratory syndrome coronavirus disease from Saudi Arabia: A descriptive study. Lancet Infect. Dis. 2013, 13, 752–761. [Google Scholar] [CrossRef]
- Lai, K.N.; Tsang, K.W.; Seto, W.H.; Ooi, C.G. Clinical, laboratory, and radiologic manifestation of SARS. Curr. Infect. Dis. Rep. 2004, 6, 213–219. [Google Scholar] [CrossRef]
- Lechien, J.R.; Chiesa-Estomba, C.M.; De Siati, D.R.; Horoi, M.; Le Bon, S.D.; Rodriguez, A.; Dequanter, D.; Blecic, S.; El Afia, F.; Distinguin, L.; et al. Olfactory and gustatory dysfunctions as a clinical presentation of mild-to-moderate forms of the coronavirus disease (COVID-19): A multicenter European study. Eur. Arch. Oto Rhino Laryngol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Poon, L.L.; Guan, Y.; Nicholls, J.M.; Yuen, K.Y.; Peiris, J.S. The aetiology, origins, and diagnosis of severe acute respiratory syndrome. Lancet. Infect. Dis. 2004, 4, 663–671. [Google Scholar] [CrossRef]
- WHO. Summary of Probable SARS Cases with Onset of Illness from 1 November 2002 to 31 July 2003. Available online: https://www.who.int/csr/sars/country/table2004_04_21/en/ (accessed on 18 May 2020).
- File, T.M., Jr.; Tsang, K.W. Severe acute respiratory syndrome: Pertinent clinical characteristics and therapy. Treat. Respir. Med. 2005, 4, 95–106. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Lim, P.L.; Kurup, A.; Gopalakrishna, G.; Chan, K.P.; Wong, C.W.; Ng, L.C.; Se-Thoe, S.Y.; Oon, L.; Bai, X.; Stanton, L.W.; et al. Laboratory-acquired Severe Acute Respiratory Syndrome. N. Engl. J. Med. 2004, 350, 1740–1745. [Google Scholar] [CrossRef] [PubMed]
- Zaki, A.M.; van Boheemen, S.; Bestebroer, T.M.; Osterhaus, A.D.; Fouchier, R.A. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N. Engl. J. Med. 2012, 367, 1814–1820. [Google Scholar] [CrossRef]
- Memish, Z.A.; Perlman, S.; Van Kerkhove, M.D.; Zumla, A. Middle East Respiratory Syndrome. Lancet 2020, 395, 1063–1077. [Google Scholar] [CrossRef]
- Bleibtreu, A.; Bertine, M.; Bertin, C.; Houhou-Fidouh, N.; Visseaux, B. Focus on Middle East respiratory syndrome coronavirus (MERS-CoV). Med. Mal. Infect. 2020, 50, 243–251. [Google Scholar] [CrossRef]
- Gorbalenya, A.E.; Baker, S.C.; Baric, R.S.; de Groot, R.J.; Drosten, C.; Gulyaeva, A.A.; Haagmans, B.L.; Lauber, C.; Leontovich, A.M.; Neuman, B.W.; et al. The species severe acute respiratory syndrome-related coronavirus: Classifying 2019-nCoV and naming it SARS-CoV-2. Nat. Microbiol. 2020, 5, 536–544. [Google Scholar] [CrossRef]
- Dong, E.; Du, H.; Gardner, L. An interactive web-based dashboard to track COVID-19 in real time. Lancet Infect. Dis. 2020, 20, 533–534. [Google Scholar] [CrossRef]
- Lu, G.; Wang, Q.; Gao, G.F. Bat-to-human: Spike features determining ‘host jump’ of coronaviruses SARS-CoV, MERS-CoV, and beyond. Trends Microbiol. 2015, 23, 468–478. [Google Scholar] [CrossRef]
- Menachery, V.D.; Yount, B.L.; Debbink, K.; Agnihothram, S.; Gralinski, L.E.; Plante, J.A.; Graham, R.L.; Scobey, T.; Ge, X.-Y.; Donaldson, E.F.; et al. A SARS-like cluster of circulating bat coronaviruses shows potential for human emergence. Nat. Med. 2015, 21, 1508–1513. [Google Scholar] [CrossRef]
- Reinke, L.M.; Spiegel, M.; Plegge, T.; Hartleib, A.; Nehlmeier, I.; Gierer, S.; Hoffmann, M.; Hofmann-Winkler, H.; Winkler, M.; Pohlmann, S. Different residues in the SARS-CoV spike protein determine cleavage and activation by the host cell protease TMPRSS2. PLoS ONE 2017, 12, e0179177. [Google Scholar] [CrossRef]
- Hofmann, H.; Pyrc, K.; van der Hoek, L.; Geier, M.; Berkhout, B.; Pöhlmann, S. Human coronavirus NL63 employs the severe acute respiratory syndrome coronavirus receptor for cellular entry. Proc. Nat. Acad. Sci. USA 2005, 102, 7988–7993. [Google Scholar] [CrossRef] [PubMed]
- Milewska, A.; Zarebski, M.; Nowak, P.; Stozek, K.; Potempa, J.; Pyrc, K. Human coronavirus NL63 utilizes heparan sulfate proteoglycans for attachment to target cells. J. Virol. 2014, 88, 13221–13230. [Google Scholar] [CrossRef] [PubMed]
- Lang, J.; Yang, N.; Deng, J.; Liu, K.; Yang, P.; Zhang, G.; Jiang, C. Inhibition of SARS pseudovirus cell entry by lactoferrin binding to heparan sulfate proteoglycans. PLoS ONE 2011, 6, e23710. [Google Scholar] [CrossRef] [PubMed]
- Widagdo, W.; Okba, N.M.A.; Li, W.; de Jong, A.; de Swart, R.L.; Begeman, L.; van den Brand, J.M.A.; Bosch, B.-J.; Haagmans, B.L. Species-specific colocalization of Middle East Respiratory Syndrome coronavirus attachment and entry receptors. J. Virol. 2019, 93, e00107–e00119. [Google Scholar] [CrossRef]
- Yeager, C.L.; Ashmun, R.A.; Williams, R.K.; Cardellichio, C.B.; Shapiro, L.H.; Look, A.T.; Holmes, K.V. Human aminopeptidase N is a receptor for human coronavirus 229E. Nature 1992, 357, 420–422. [Google Scholar] [CrossRef] [PubMed]
- Collins, A.R. HLA class I antigen serves as a receptor for human coronavirus OC43. Immunol. Investig. 1993, 22, 95–103. [Google Scholar] [CrossRef]
- Krempl, C.; Schultze, B.; Herrler, G. Analysis of cellular receptors for human coronavirus OC43. Adv. Exp. Med. Biol. 1995, 380, 371–374. [Google Scholar] [CrossRef]
- Huang, X.; Dong, W.; Milewska, A.; Golda, A.; Qi, Y.; Zhu, Q.K.; Marasco, W.A.; Baric, R.S.; Sims, A.C.; Pyrc, K.; et al. Human coronavirus HKU1 spike protein uses O-acetylated sialic acid as an attachment receptor determinant and employs hemagglutinin-esterase protein as a receptor-destroying enzyme. J. Virol. 2015, 89, 7202–7213. [Google Scholar] [CrossRef]
- Chan, C.M.; Lau, S.K.P.; Woo, P.C.Y.; Tse, H.; Zheng, B.-J.; Chen, L.; Huang, J.-D.; Yuen, K.-Y. Identification of major histocompatibility complex class I C molecule as an attachment factor that facilitates coronavirus HKU1 Spike-Mediated Infection. J. Virol. 2009, 83, 1026–1035. [Google Scholar] [CrossRef] [PubMed]
- Shirato, K.; Kawase, M.; Matsuyama, S. Middle East Respiratory syndrome coronavirus infection mediated by the transmembrane serine protease TMPRSS2. J. Virol. 2013, 87, 12552–12561. [Google Scholar] [CrossRef] [PubMed]
- Glowacka, I.; Bertram, S.; Müller, M.A.; Allen, P.; Soilleux, E.; Pfefferle, S.; Steffen, I.; Tsegaye, T.S.; He, Y.; Gnirss, K.; et al. Evidence that TMPRSS2 Activates the severe acute respiratory syndrome coronavirus spike protein for membrane fusion and reduces viral control by the humoral immune response. J. Virol. 2011, 85, 4122–4134. [Google Scholar] [CrossRef]
- Bertram, S.; Dijkman, R.; Habjan, M.; Heurich, A.; Gierer, S.; Glowacka, I.; Welsch, K.; Winkler, M.; Schneider, H.; Hofmann-Winkler, H.; et al. TMPRSS2 activates the human coronavirus 229E for cathepsin-independent host cell entry and is expressed in viral target cells in the respiratory epithelium. J. Virol. 2013, 87, 6150–6160. [Google Scholar] [CrossRef]
- Shirato, K.; Kawase, M.; Matsuyama, S. Wild-type human coronaviruses prefer cell-surface TMPRSS2 to endosomal cathepsins for cell entry. Virology 2018, 517, 9–15. [Google Scholar] [CrossRef] [PubMed]
- Milewska, A.; Nowak, P.; Owczarek, K.; Szczepanski, A.; Zarebski, M.; Hoang, A.; Berniak, K.; Wojarski, J.; Zeglen, S.; Baster, Z.; et al. Entry of human coronavirus NL63 into the cell. J. Virol. 2018, 92, e01917–e01933. [Google Scholar] [CrossRef]
- Spiegel, M.; Schneider, K.; Weber, F.; Weidmann, M.; Hufert, F.T. Interaction of severe acute respiratory syndrome-associated coronavirus with dendritic cells. J. Gen. Virol. 2006, 87, 1953–1960. [Google Scholar] [CrossRef]
- Kaye, M. SARS-associated coronavirus replication in cell lines. Emerg. Infect. Dis. 2006, 12, 128–133. [Google Scholar] [CrossRef]
- Song, Z.; Xu, Y.; Bao, L.; Zhang, L.; Yu, P.; Qu, Y.; Zhu, H.; Zhao, W.; Han, Y.; Qin, C. From SARS to MERS, thrusting coronaviruses into the spotlight. Viruses 2019, 11, 59. [Google Scholar] [CrossRef]
- Mesel-Lemoine, M.; Millet, J.; Vidalain, P.O.; Law, H.; Vabret, A.; Lorin, V.; Escriou, N.; Albert, M.L.; Nal, B.; Tangy, F. A human coronavirus responsible for the common cold massively kills dendritic cells but not monocytes. J. Virol. 2012, 86, 7577–7587. [Google Scholar] [CrossRef]
- Graham, R.L.; Baric, R.S. Recombination, reservoirs, and the modular spike: Mechanisms of coronavirus cross-species transmission. J. Virol. 2010, 84, 3134–3146. [Google Scholar] [CrossRef]
- Woo, P.C.; Lau, S.K.; Lam, C.S.; Tsang, A.K.; Hui, S.W.; Fan, R.Y.; Martelli, P.; Yuen, K.Y. Discovery of a novel bottlenose dolphin coronavirus reveals a distinct species of marine mammal coronavirus in Gammacoronavirus. J. Virol. 2014, 88, 1318–1331. [Google Scholar] [CrossRef]
- Anthony, S.J.; Johnson, C.K.; Greig, D.J.; Kramer, S.; Che, X.; Wells, H.; Hicks, A.L.; Joly, D.O.; Wolfe, N.D.; Daszak, P.; et al. Global patterns in coronavirus diversity. Virus Evolut. 2017, 3, vex012. [Google Scholar] [CrossRef]
- Tang, X.C.; Zhang, J.X.; Zhang, S.Y.; Wang, P.; Fan, X.H.; Li, L.F.; Li, G.; Dong, B.Q.; Liu, W.; Cheung, C.L.; et al. Prevalence and genetic diversity of coronaviruses in bats from China. J. Virol. 2006, 80, 7481–7490. [Google Scholar] [CrossRef] [PubMed]
- Mildenstein, T.; Tanshi, I.; Racey, P.A. Exploitation of bats for bushmeat and medicine. In Bats in the Anthropocene: Conservation Bats in a Changing World; Voigt, C.C., Kingston, T., Eds.; Springer International Publishing: Cham, Switzerland, 2016; pp. 325–375. [Google Scholar] [CrossRef]
- Markotter, W.; Geldenhuys, M.; Jansen van Vuren, P.; Kemp, A.; Mortlock, M.; Mudakikwa, A.; Nel, L.; Nziza, J.; Paweska, J.; Weyer, J. Paramyxo- and coronaviruses in Rwandan bats. Trop. Med. Infect. Dis. 2019, 4, 99. [Google Scholar] [CrossRef] [PubMed]
- Memish, Z.A.; Mishra, N.; Olival, K.J.; Fagbo, S.F.; Kapoor, V.; Epstein, J.H.; Alhakeem, R.; Durosinloun, A.; Al Asmari, M.; Islam, A.; et al. Middle East respiratory syndrome coronavirus in bats, Saudi Arabia. Emerg. Infect. Dis. 2013, 19, 1819–1823. [Google Scholar] [CrossRef] [PubMed]
- Lau, S.K.P.; Zhang, L.; Luk, H.K.H.; Xiong, L.; Peng, X.; Li, K.S.M.; He, X.; Zhao, P.S.; Fan, R.Y.Y.; Wong, A.C.P.; et al. Receptor usage of a novel bat lineage C betacoronavirus reveals evolution of Middle East respiratory syndrome-related coronavirus spike proteins for human dipeptidyl peptidase 4 binding. J. Infect. Dis. 2018, 218, 197–207. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.; Zeng, L.-P.; Yang, X.-L.; Ge, X.-Y.; Zhang, W.; Li, B.; Xie, J.-Z.; Shen, X.-R.; Zhang, Y.-Z.; Wang, N.; et al. Discovery of a rich gene pool of bat SARS-related coronaviruses provides new insights into the origin of SARS coronavirus. PLoS Pathog. 2017, 13, e1006698. [Google Scholar] [CrossRef]
- Zhou, P.; Yang, X.-L.; Wang, X.-G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.-R.; Zhu, Y.; Li, B.; Huang, C.-L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef]
- Chu, D.K.W.; Poon, L.L.M.; Gomaa, M.M.; Shehata, M.M.; Perera, R.A.P.M.; Abu Zeid, D.; El Rifay, A.S.; Siu, L.Y.; Guan, Y.; Webby, R.J.; et al. MERS coronaviruses in dromedary camels, Egypt. Emerg. Infect. Dis. 2014, 20, 1049–1053. [Google Scholar] [CrossRef]
- Corman, V.M.; Kallies, R.; Philipps, H.; Göpner, G.; Müller, M.A.; Eckerle, I.; Brünink, S.; Drosten, C.; Drexler, J.F. Characterization of a novel betacoronavirus related to Middle East respiratory syndrome coronavirus in European hedgehogs. J. Virol. 2014, 88, 717–724. [Google Scholar] [CrossRef] [PubMed]
- Guan, Y.; Zheng, B.J.; He, Y.Q.; Liu, X.L.; Zhuang, Z.X.; Cheung, C.L.; Luo, S.W.; Li, P.H.; Zhang, L.J.; Guan, Y.J.; et al. Isolation and characterization of viruses related to the SARS coronavirus from animals in southern China. Science 2003, 302, 276–278. [Google Scholar] [CrossRef]
- Lam, T.T.-Y.; Shum, M.H.-H.; Zhu, H.-C.; Tong, Y.-G.; Ni, X.-B.; Liao, Y.-S.; Wei, W.; Cheung, W.Y.-M.; Li, W.-J.; Li, L.-F.; et al. Identifying SARS-CoV-2 related coronaviruses in Malayan pangolins. Nature 2020. [Google Scholar] [CrossRef] [PubMed]
- Tao, Y.; Shi, M.; Chommanard, C.; Queen, K.; Zhang, J.; Markotter, W.; Kuzmin, I.V.; Holmes, E.C.; Tong, S. Surveillance of Bat coronaviruses in Kenya identifies relatives of human coronaviruses NL63 and 229E and their recombination history. J. Virol. 2017, 91, e01916–e01953. [Google Scholar] [CrossRef] [PubMed]
- Sabir, J.S.; Lam, T.T.; Ahmed, M.M.; Li, L.; Shen, Y.; Abo-Aba, S.E.; Qureshi, M.I.; Abu-Zeid, M.; Zhang, Y.; Khiyami, M.A.; et al. Co-circulation of three camel coronavirus species and recombination of MERS-CoVs in Saudi Arabia. Science 2016, 351, 81–84. [Google Scholar] [CrossRef] [PubMed]
- Corman, V.M.; Eckerle, I.; Memish, Z.A.; Liljander, A.M.; Dijkman, R.; Jonsdottir, H.; Juma Ngeiywa, K.J.Z.; Kamau, E.; Younan, M.; Al Masri, M.; et al. Link of a ubiquitous human coronavirus to dromedary camels. Proc. Natl. Acad. Sci. USA 2016, 113, 9864–9869. [Google Scholar] [CrossRef]
- Lau, S.K.P.; Woo, P.C.Y.; Li, K.S.M.; Tsang, A.K.L.; Fan, R.Y.Y.; Luk, H.K.H.; Cai, J.-P.; Chan, K.-H.; Zheng, B.-J.; Wang, M.; et al. Discovery of a novel coronavirus, China Rattus coronavirus HKU24, from Norway rats supports the murine origin of betacoronavirus 1 and has implications for the ancestor of betacoronavirus lineage A. J. Virol. 2015, 89, 3076–3092. [Google Scholar] [CrossRef]
- Woo, P.C.Y.; Huang, Y.; Lau, S.K.P.; Yuen, K.-Y. Coronavirus genomics and bioinformatics analysis. Viruses 2010, 2, 1804–1820. [Google Scholar] [CrossRef]
- Wang, W.; Lin, X.-D.; Guo, W.-P.; Zhou, R.-H.; Wang, M.-R.; Wang, C.-Q.; Ge, S.; Mei, S.-H.; Li, M.-H.; Shi, M.; et al. Discovery, diversity and evolution of novel coronaviruses sampled from rodents in China. Virology 2015, 474, 19–27. [Google Scholar] [CrossRef]
- Wang, W.; Lin, X.-D.; Liao, Y.; Guan, X.-Q.; Guo, W.-P.; Xing, J.-G.; Holmes, E.C.; Zhang, Y.-Z. Discovery of a Highly Divergent coronavirus in the Asian house shrew from china illuminates the origin of the alphacoronaviruses. J. Virol. 2017, 91, e00717–e00764. [Google Scholar] [CrossRef]
- Lin, S.Y.; Chen, H.W. Infectious Bronchitis Virus Variants: Molecular Analysis and Pathogenicity Investigation. Int. J. Mol. Sci. 2017, 18, 2030. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Chen, J.; Chen, J.; Kong, X.; Shao, Y.; Han, Z.; Feng, L.; Cai, X.; Gu, S.; Liu, M. Isolation of avian infectious bronchitis coronavirus from domestic peafowl (Pavo cristatus) and teal (Anas). J. Gen. Virol. 2005, 86, 719–725. [Google Scholar] [CrossRef] [PubMed]
- Circella, E.; Camarda, A.; Martella, V.; Bruni, G.; Lavazza, A.; Buonavoglia, C. Coronavirus associated with an enteric syndrome on a quail farm. Avian Pathol. 2007, 36, 251–258. [Google Scholar] [CrossRef]
- Liais, E.; Croville, G.; Mariette, J.; Delverdier, M.; Lucas, M.N.; Klopp, C.; Lluch, J.; Donnadieu, C.; Guy, J.S.; Corrand, L.; et al. Novel avian coronavirus and fulminating disease in guinea fowl, France. Emerg. Infect. Dis. 2014, 20, 105–108. [Google Scholar] [CrossRef] [PubMed]
- Lin, T.L.; Loa, C.C.; Tsai, S.C.; Wu, C.C.; Bryan, T.A.; Thacker, H.L.; Hooper, T.; Schrader, D. Characterization of turkey coronavirus from turkey poults with acute enteritis. Vet. Microbiol. 2002, 84, 179–186. [Google Scholar] [CrossRef]
- Hughes, L.A.; Savage, C.; Naylor, C.; Bennett, M.; Chantrey, J.; Jones, R. Genetically diverse coronaviruses in wild bird populations of northern England. Emerg. Infect. Dis. 2009, 15, 1091–1094. [Google Scholar] [CrossRef]
- Felippe, P.A.; da Silva, L.H.; Santos, M.M.; Spilki, F.R.; Arns, C.W. Genetic diversity of avian infectious bronchitis virus isolated from domestic chicken flocks and coronaviruses from feral pigeons in Brazil between 2003 and 2009. Avian Dis. 2010, 54, 1191–1196. [Google Scholar] [CrossRef]
- Woo, P.C.Y.; Lau, S.K.P.; Lam, C.S.F.; Lau, C.C.Y.; Tsang, A.K.L.; Lau, J.H.N.; Bai, R.; Teng, J.L.L.; Tsang, C.C.C.; Wang, M.; et al. Discovery of seven novel mammalian and avian coronaviruses in the genus deltacoronavirus supports bat coronaviruses as the gene source of alphacoronavirus and betacoronavirus and avian coronaviruses as the gene source of gammacoronavirus and deltacoronavirus. J. Virol. 2012, 86, 3995–4008. [Google Scholar] [CrossRef]
- Boniotti, M.B.; Papetti, A.; Lavazza, A.; Alborali, G.; Sozzi, E.; Chiapponi, C.; Faccini, S.; Bonilauri, P.; Cordioli, P.; Marthaler, D. Porcine epidemic diarrhea virus and discovery of a recombinant swine enteric coronavirus, Italy. Emerg. Infect. Dis. 2016, 22, 83–87. [Google Scholar] [CrossRef]
- Hou, Y.; Yue, X.; Cai, X.; Wang, S.; Liu, Y.; Yuan, C.; Cui, L.; Hua, X.; Yang, Z. Complete genome of transmissible gastroenteritis virus AYU strain isolated in Shanghai, China. J. Virol. 2012, 86, 11935. [Google Scholar] [CrossRef][Green Version]
- Chasey, D.; Cartwright, S.F. Virus-like particles associated with porcine epidemic diarrhoea. Res. Vet. Sci. 1978, 25, 255–256. [Google Scholar] [CrossRef]
- Zhang, M.-J.; Liu, D.-J.; Liu, X.-L.; Ge, X.-Y.; Jongkaewwattana, A.; He, Q.-G.; Luo, R. Genomic characterization and pathogenicity of porcine deltacoronavirus strain CHN-HG-2017 from China. Arch. Virol. 2019, 164, 413–425. [Google Scholar] [CrossRef] [PubMed]
- Vogel, L.; Van der Lubben, M.; te Lintelo, E.G.; Bekker, C.P.J.; Geerts, T.; Schuijff, L.S.; Grinwis, G.C.M.; Egberink, H.F.; Rottier, P.J.M. Pathogenic characteristics of persistent feline enteric coronavirus infection in cats. Vet. Res. 2010, 41, 71. [Google Scholar] [CrossRef] [PubMed]
- Wise, A.G.; Kiupel, M.; Maes, R.K. Molecular characterization of a novel coronavirus associated with epizootic catarrhal enteritis (ECE) in ferrets. Virology 2006, 349, 164–174. [Google Scholar] [CrossRef] [PubMed]
- Pratelli, A. Genetic evolution of canine coronavirus and recent advances in prophylaxis. Vet. Res. 2006, 37, 191–200. [Google Scholar] [CrossRef][Green Version]
- Lau, S.K.; Woo, P.C.; Yip, C.C.; Fan, R.Y.; Huang, Y.; Wang, M.; Guo, R.; Lam, C.S.; Tsang, A.K.; Lai, K.K.; et al. Isolation and characterization of a novel Betacoronavirus subgroup A coronavirus, rabbit coronavirus HKU14, from domestic rabbits. J. Virol. 2012, 86, 5481–5496. [Google Scholar] [CrossRef]
- Kampf, G.; Todt, D.; Pfaender, S.; Steinmann, E. Persistence of coronaviruses on inanimate surfaces and their inactivation with biocidal agents. J. Hosp. Infect. 2020, 104, 246–251. [Google Scholar] [CrossRef]
- Chin, A.W.H.; Chu, J.T.S.; Perera, M.R.A.; Hui, K.P.Y.; Yen, H.-L.; Chan, M.C.W.; Peiris, M.; Poon, L.L.M. Stability of SARS-CoV-2 in different environmental conditions. Lancet Microbe 2020. [Google Scholar] [CrossRef]
- Van Doremalen, N.; Bushmaker, T.; Morris, D.H.; Holbrook, M.G.; Gamble, A.; Williamson, B.N.; Tamin, A.; Harcourt, J.L.; Thornburg, N.J.; Gerber, S.I.; et al. Aerosol and surface stability of SARS-CoV-2 as compared with SARS-CoV-1. N. Engl. J. Med. 2020, 382, 1564–1567. [Google Scholar] [CrossRef]
- Prussin, A.J.; Schwake, D.O.; Lin, K.; Gallagher, D.L.; Buttling, L.; Marr, L.C. Survival of the enveloped virus Phi6 in droplets as a function of relative humidity, absolute humidity, and temperature. Appl. Environ. Microbiol. 2018, 84, e00551-18. [Google Scholar] [CrossRef]
- Hasija, M.; Li, L.; Rahman, N.; Ausar, S.F. Forced degradation studies: An essential tool for the formulation development of vaccines. Vaccin. Dev. Ther. 2013, 3, 11–33. [Google Scholar] [CrossRef][Green Version]
- Sharma, S.; Gray, D.K.; Read, J.S.; O’Reilly, C.M.; Schneider, P.; Qudrat, A.; Gries, C.; Stefanoff, S.; Hampton, S.E.; Hook, S.; et al. A global database of lake surface temperatures collected by in situ and satellite methods from 1985–2009. Sci. Data 2015, 2, 150008. [Google Scholar] [CrossRef] [PubMed]
- Padisák, J.; Reynolds, C.S. Shallow lakes: The absolute, the relative, the functional and the pragmatic. Hydrobiologia 2003, 506, 1–11. [Google Scholar] [CrossRef]
- Darnell, M.E.R.; Subbarao, K.; Feinstone, S.M.; Taylor, D.R. Inactivation of the coronavirus that induces severe acute respiratory syndrome, SARS-CoV. J. Virol. Methods 2004, 121, 85–91. [Google Scholar] [CrossRef]
- Pratelli, A. Canine coronavirus inactivation with physical and chemical agents. Vet. J. 2008, 177, 71–79. [Google Scholar] [CrossRef]
- Perdiz, D.; Grof, P.; Mezzina, M.; Nikaido, O.; Moustacchi, E.; Sage, E. Distribution and repair of bipyrimidine photoproducts in solar UV-irradiated mammalian cells. Possible role of Dewar photoproducts in solar mutagenesis. J. Biol. Chem. 2000, 275, 26732–26742. [Google Scholar] [CrossRef]
- Hicke, J.A.; Slusser, J.; Lantz, K.; Pascual, F.G. Trends and interannual variability in surface UVB radiation over 8 to 11 years observed across the United States. J. Geophys. Res. Atmos. 2008, 113. [Google Scholar] [CrossRef]
- Grigalavicius, M.; Moan, J.; Dahlback, A.; Juzeniene, A. Daily, seasonal, and latitudinal variations in solar ultraviolet A and B radiation in relation to vitamin D production and risk for skin cancer. Int. J. Dermatol. 2016, 55, e23–e28. [Google Scholar] [CrossRef]
- Murray, A.; Jackson, G.A. Viral dynamics: A model of the effects of size, shape, motion and abundance of single-celled planktonic organisms and other particles. Mar. Ecol. Prog. Ser. 1992, 89, 103–116. [Google Scholar] [CrossRef]
- Feichtmayer, J.; Deng, L.; Griebler, C. Antagonistic microbial interactions: Contributions and potential applications for controlling pathogens in the aquatic systems. Front. Microbiol. 2017, 8, 2192. [Google Scholar] [CrossRef]
- Casanova, L.; Rutala, W.A.; Weber, D.J.; Sobsey, M.D. Survival of surrogate coronaviruses in water. Water Res. 2009, 43, 1893–1898. [Google Scholar] [CrossRef] [PubMed]
- Gundy, P.M.; Gerba, C.P.; Pepper, I.L. Survival of coronaviruses in water and wastewater. Food Environ. Virol. 2009, 1, 10. [Google Scholar] [CrossRef]
- Wang, X.-W.; Li, J.-S.; Jin, M.; Zhen, B.; Kong, Q.-X.; Song, N.; Xiao, W.-J.; Yin, J.; Wei, W.; Wang, G.-J.; et al. Study on the resistance of severe acute respiratory syndrome-associated coronavirus. J. Virol. Methods 2005, 126, 171–177. [Google Scholar] [CrossRef] [PubMed]
- Macnaughton, M.R. Occurrence and frequency of coronavirus infections in humans as determined by enzyme-linked immunosorbent assay. Infect. Immun. 1982, 38, 419–423. [Google Scholar] [CrossRef]
- Rabenau, H.F.; Cinatl, J.; Morgenstern, B.; Bauer, G.; Preiser, W.; Doerr, H.W. Stability and inactivation of SARS coronavirus. Med. Microbiol. Immunol. 2005, 194, 1–6. [Google Scholar] [CrossRef]
- Contu, F.; Elsener, B.; Böhni, H. Characterization of implant materials in fetal bovine serum and sodium sulfate by electrochemical impedance spectroscopy. I. Mechanically polished samples. J. Biomed. Mater. Res. 2002, 62, 412–421. [Google Scholar] [CrossRef]
- Raunkjær, K.; Hvitved-Jacobsen, T.; Nielsen, P.H. Measurement of pools of protein, carbohydrate and lipid in domestic wastewater. Water Res. 1994, 28, 251–262. [Google Scholar] [CrossRef]
- Wang, X.W.; Li, J.; Guo, T.; Zhen, B.; Kong, Q.; Yi, B.; Li, Z.; Song, N.; Jin, M.; Xiao, W.; et al. Concentration and detection of SARS coronavirus in sewage from Xiao Tang Shan hospital and the 309th Hospital of the Chinese People’s Liberation Army. Water Sci. Technol. 2005, 52, 213–221. [Google Scholar] [CrossRef]
- Wang, X.W.; Li, J.S.; Guo, T.K.; Zhen, B.; Kong, Q.X.; Yi, B.; Li, Z.; Song, N.; Jin, M.; Xiao, W.J.; et al. Detection of RNA of SARS coronavirus in hospital sewage. Chin. J. Prev. Med. 2004, 38, 257–260. [Google Scholar]
- Goyal, S.M.; Chander, Y.; Yezli, S.; Otter, J.A. Evaluating the virucidal efficacy of hydrogen peroxide vapour. J. Hosp. Infect. 2014, 86, 255–259. [Google Scholar] [CrossRef]
- Zhang, H.; Tang, W.; Chen, Y.; Yin, W. Disinfection threatens aquatic ecosystems. Science 2020, 368, 146–147. [Google Scholar] [CrossRef] [PubMed]
- Duan, S.-M.; Zhao, X.-S.; Wen, R.-F.; Huang, J.-J.; Pi, G.-H.; Zhang, S.-X.; Han, J.; Bi, S.-L.; Ruan, L.; Dong, X.-P. Stability of SARS coronavirus in human specimens and environment and its sensitivity to heating and UV irradiation. Biomed. Environ. Sci. 2003, 16, 246–255. [Google Scholar] [PubMed]
- Salvarina, I.; Gravier, D.; Rothhaupt, K.-O. Seasonal bat activity related to insect emergence at three temperate lakes. Ecol. Evol. 2018, 8, 3738–3750. [Google Scholar] [CrossRef]
- Fukui, D.; Murakami, M.; Nakano, S.; Aoi, T. Effect of emergent aquatic insects on bat foraging in a riparian forest. J. Anim. Ecol. 2006, 75, 1252–1258. [Google Scholar] [CrossRef] [PubMed]
- Chamings, A.; Nelson, T.M.; Vibin, J.; Wille, M.; Klaassen, M.; Alexandersen, S. Detection and characterisation of coronaviruses in migratory and non-migratory Australian wild birds. Sci. Rep. 2018, 8, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Muradrasoli, S.; Bálint, A.; Wahlgren, J.; Waldenström, J.; Belák, S.; Blomberg, J.; Olsen, B. Prevalence and phylogeny of coronaviruses in wild birds from the Bering Strait area (Beringia). PLoS ONE 2010, 5, e13640. [Google Scholar] [CrossRef] [PubMed]
- Barbosa, C.M.; Durigon, E.L.; Thomazelli, L.M.; Ometto, T.; Marcatti, R.; Nardi, M.S.; de Aguiar, D.M.; Pinho, J.B.; Petry, M.V.; Neto, I.S.; et al. Divergent coronaviruses detected in wild birds in Brazil, including a central park in São Paulo. Braz. J. Microbiol. 2019, 50, 547–556. [Google Scholar] [CrossRef]
- Verdugo, C.; Pinto, A.; Ariyama, N.; Moroni, M.; Hernandez, C. Molecular identification of avian viruses in neotropic cormorants (Phalacrocorax brasilianus) in Chile. J. Wildl. Dis. 2018, 55, 105–112. [Google Scholar] [CrossRef]
- Kim, H.R.; Oem, J.K. Surveillance of avian coronaviruses in wild bird populations of Korea. J. Wildl. Dis. 2014, 50, 964–968. [Google Scholar] [CrossRef]
- Wille, M.; Muradrasoli, S.; Nilsson, A.; Järhult, J.D. High prevalence and putative lineage maintenance of avian coronaviruses in Scandinavian waterfowl. PLoS ONE 2016, 11, e0150198. [Google Scholar] [CrossRef]
- Hepojoki, S.; Lindh, E.; Vapalahti, O.; Huovilainen, A. Prevalence and genetic diversity of coronaviruses in wild birds, Finland. Infect. Ecol. Epidemiol. 2017, 7, 1408360. [Google Scholar] [CrossRef] [PubMed]
- Jonassen, C.M.; Kofstad, T.; Larsen, I.-L.; Løvland, A.; Handeland, K.; Follestad, A.; Lillehaug, A. Molecular identification and characterization of novel coronaviruses infecting graylag geese (Anser anser), feral pigeons (Columbia livia) and mallards (Anas platyrhynchos). J. Gen. Virol. 2005, 86, 1597–1607. [Google Scholar] [CrossRef] [PubMed]
- Honkavuori, K.S.; Briese, T.; Krauss, S.; Sanchez, M.D.; Jain, K.; Hutchison, S.K.; Webster, R.G.; Lipkin, W.I. Novel coronavirus and astrovirus in Delaware Bay shorebirds. PLoS ONE 2014, 9, e93395. [Google Scholar] [CrossRef] [PubMed]
- Jordan, B.J.; Hilt, D.A.; Poulson, R.; Stallknecht, D.E.; Jackwood, M.W. Identification of avian coronavirus in wild aquatic birds of the central and eastern USA. J. Wildl. Dis. 2015, 51, 218–221. [Google Scholar] [CrossRef] [PubMed]
- Paim, F.C.; Bowman, A.S.; Miller, L.; Feehan, B.J.; Marthaler, D.; Saif, L.J.; Vlasova, A.N. Epidemiology of deltacoronaviruses (δ-CoV) and gammacoronaviruses (γ-CoV) in wild birds in the United States. Viruses 2019, 11, 897. [Google Scholar] [CrossRef] [PubMed]
- Rzymski, P.; Słodkowicz-Kowalska, A.; Klimaszyk, P.; Solarczyk, P.; Poniedziałek, B. Screening of protozoan and microsporidian parasites in feces of great cormorant (Phalacrocorax carbo). Environ. Sci. Pollut. Res. 2017, 24, 9813–9819. [Google Scholar] [CrossRef]
- Klimaszyk, P. May a cormorant colony be a source of coliform and chemical pollution in a lake? Oceanol. Hydrobiol. Stud. 2012, 41, 67–73. [Google Scholar] [CrossRef]
- Klimaszyk, P.; Brzeg, A.; Rzymski, P.; Piotrowicz, R. Black spots for aquatic and terrestrial ecosystems: Impact of a perennial cormorant colony on the environment. Sci. Total Environ. 2015, 517, 222–231. [Google Scholar] [CrossRef]
- Dessborn, L.; Hessel, R.; Elmberg, J. Geese as vectors of nitrogen and phosphorus to freshwater systems. Inl. Waters 2016, 6, 111–122. [Google Scholar] [CrossRef]
- Green, H.C.; Dick, L.K.; Gilpin, B.; Samadpour, M.; Field, K.G. Genetic markers for rapid PCR-based identification of gull, Canada goose, duck, and chicken fecal contamination in water. Appl. Environ. Microbiol. 2012, 78, 503–510. [Google Scholar] [CrossRef]
- Devane, M.L.; Robson, B.; Nourozi, F.; Scholes, P.; Gilpin, B.J. A PCR marker for detection in surface waters of faecal pollution derived from ducks. Water Res. 2007, 41, 3553–3560. [Google Scholar] [CrossRef] [PubMed]
- Papineau, A.; Berhane, Y.; Wylie, T.; Wylie, K.; Sharpe, S.; Lung, O. Genome Organization of Canada Goose Coronavirus, A Novel Species Identified in a Mass Die-off of Canada Geese. Sci. Rep. 2019, 9, s41598-s019. [Google Scholar] [CrossRef] [PubMed]
- Woo, P.C.Y.; Lau, S.K.P.; Lam, C.S.F.; Lai, K.K.Y.; Huang, Y.; Lee, P.; Luk, G.S.M.; Dyrting, K.C.; Chan, K.-H.; Yuen, K.-Y. Comparative analysis of complete genome sequences of three avian coronaviruses reveals a novel group 3c coronavirus. J. Virol. 2009, 83, 908–917. [Google Scholar] [CrossRef] [PubMed]
- Lau, S.K.P.; Wong, E.Y.M.; Tsang, C.C. Discovery and sequence analysis of four deltacoronaviruses from birds in the Middle East reveal interspecies jumping with recombination as a potential mechanism for avian-to-avian and avian-to-mammalian transmission. J. Virol. 2018, 92. [Google Scholar] [CrossRef]
- Chen, G.-Q.; Zhuang, Q.-Y.; Wang, K.-C.; Liu, S.; Shao, J.-Z.; Jiang, W.-M.; Hou, G.-Y.; Li, J.-P.; Yu, J.-M.; Li, Y.-P.; et al. Identification and Survey of a Novel Avian Coronavirus in Ducks. PLoS ONE 2013, 8, e72918. [Google Scholar] [CrossRef]
- Mihindukulasuriya, K.A.; Wu, G.; St Leger, J.; Nordhausen, R.W.; Wang, D. Identification of a novel coronavirus from a beluga whale by using a panviral microarray. J. Virol. 2008, 82, 5084–5088. [Google Scholar] [CrossRef]
- Nollens, H.H.; Wellehan, J.F.; Archer, L.; Lowenstine, L.J.; Gulland, F.M. Detection of a respiratory coronavirus from tissues archived during a pneumonia epizootic in free-ranging Pacific harbor seals Phoca vitulina richardsii. Dis. Aquat. Organ. 2010, 90, 113–120. [Google Scholar] [CrossRef]
- Chou, C.-F.; Loh, C.B.; Foo, Y.K.; Shen, S.; Fielding, B.C.; Tan, T.H.P.; Khan, S.; Wang, Y.; Lim, S.G.; Hong, W.; et al. ACE2 orthologues in non-mammalian vertebrates (Danio, Gallus, Fugu, Tetraodon and Xenopus). Gene 2006, 377, 46–55. [Google Scholar] [CrossRef]
- Winter, C.; Schwegmann-Weßels, C.; Cavanagh, D.; Neumann, U.; Herrler, G. Sialic acid is a receptor determinant for infection of cells by avian Infectious bronchitis virus. J. Gen. Virol. 2006, 87, 1209–1216. [Google Scholar] [CrossRef]
- Wickramasinghe, I.N.A.; de Vries, R.P.; Gröne, A.; de Haan, C.A.M.; Verheije, M.H. Binding of avian coronavirus spike proteins to host factors reflects virus tropism and pathogenicity. J. Virol. 2011, 85, 8903–8912. [Google Scholar] [CrossRef]
- Chu, V.C.; McElroy, L.J.; Chu, V.; Bauman, B.E.; Whittaker, G.R. The avian coronavirus infectious bronchitis virus undergoes direct low-pH-dependent fusion activation during entry into host cells. J. Virol. 2006, 80, 3180–3188. [Google Scholar] [CrossRef] [PubMed]
- Mordecai, G.J.; Miller, K.M.; Di Cicco, E.; Schulze, A.D.; Kaukinen, K.H.; Ming, T.J.; Li, S.; Tabata, A.; Teffer, A.; Patterson, D.A.; et al. Endangered wild salmon infected by newly discovered viruses. eLife 2019, 8, e47615. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).