Textured-Based Deep Learning in Prostate Cancer Classification with 3T Multiparametric MRI: Comparison with PI-RADS-Based Classification
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Population and MRI Datasets
2.2. Textured-DL Model
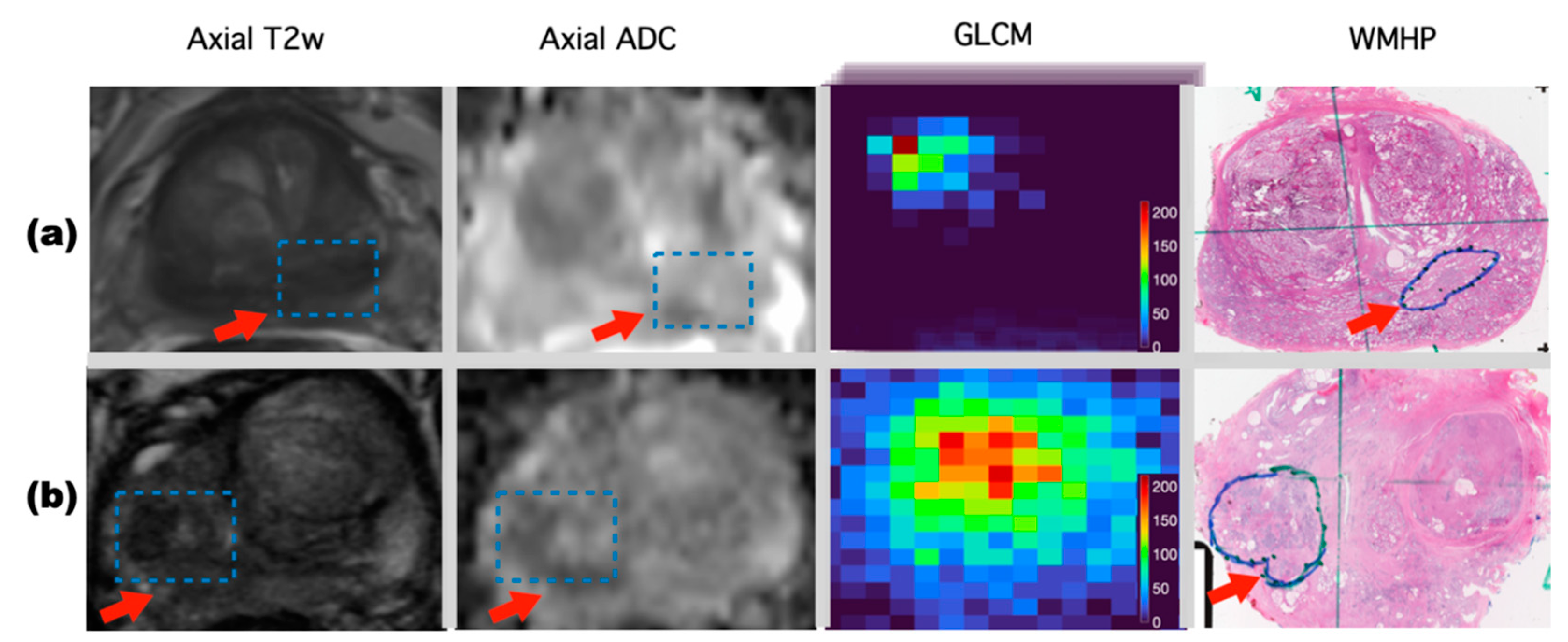
2.2.1. 3D GLCM Extractor
2.2.2. CNN Network
2.3. Model Development and Comparison
2.4. Statistical Analysis
3. Results
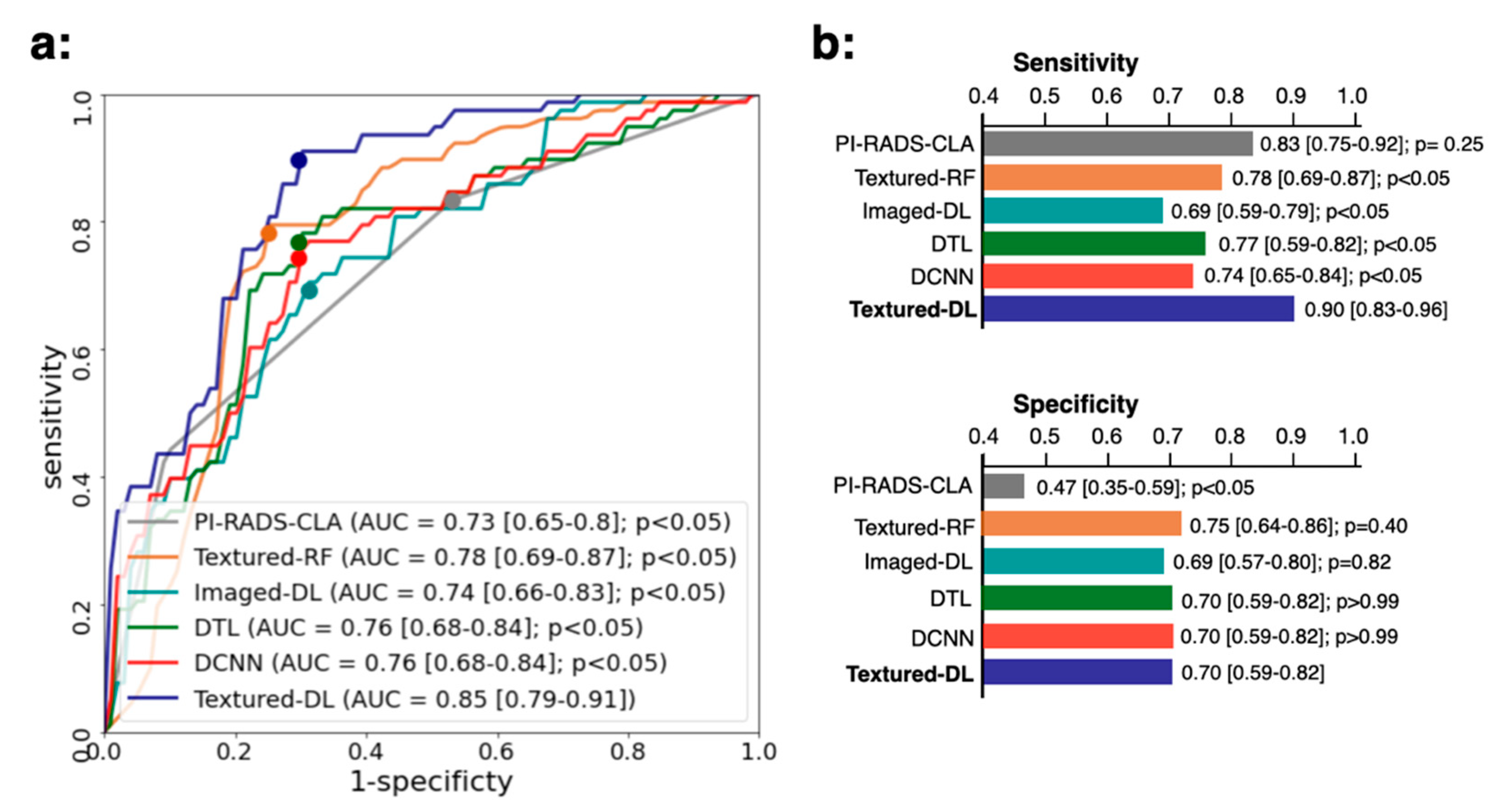
3.1. Model Performance in Comparison with PI-RADS-CLA for All Tumors
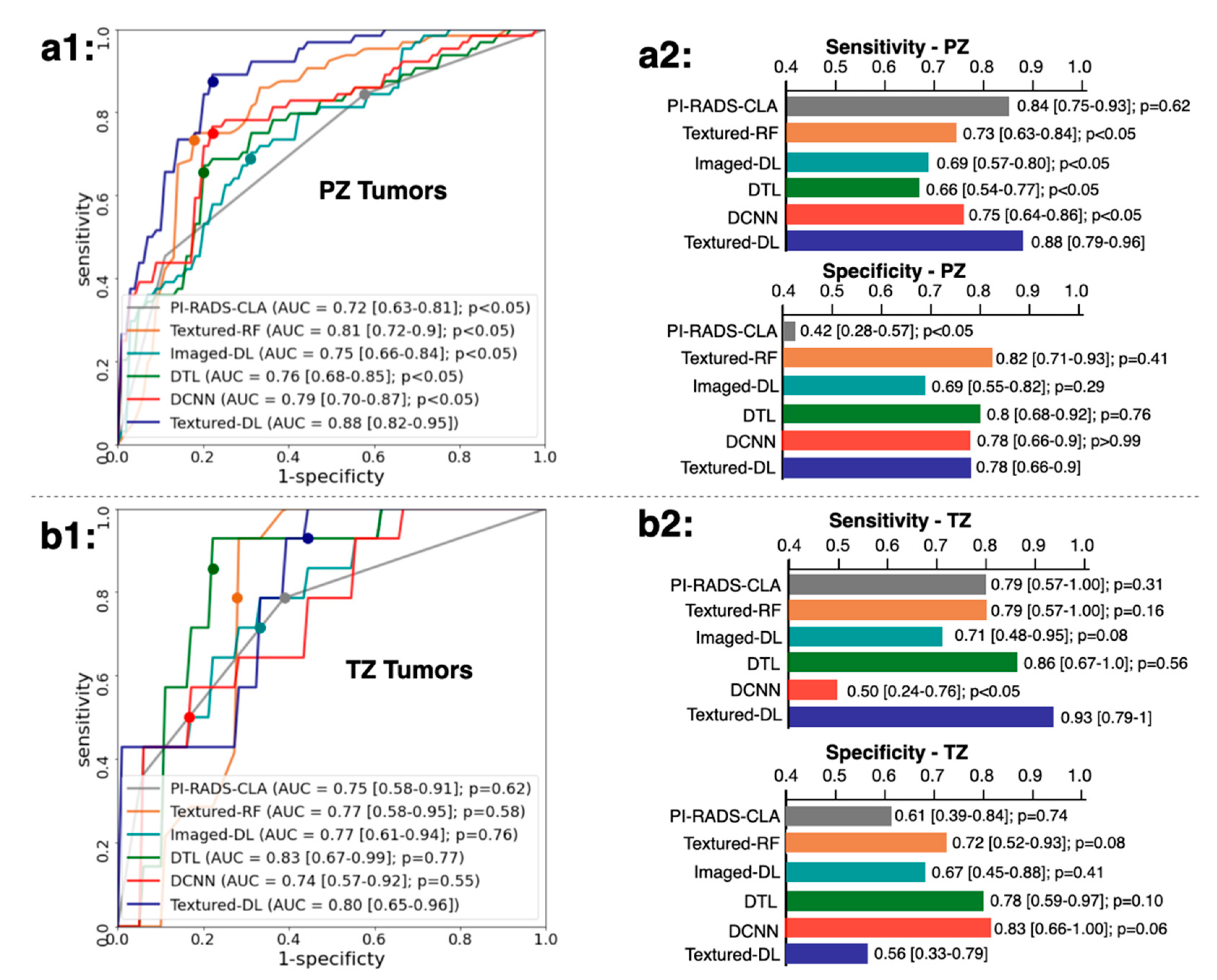
3.2. Classification Performance for Tumors on Different Prostate Zone
3.3. Classification Performance for Solidary and Multi-Focal Tumors
3.4. Classification Performance for Tumors of Different PI-RADS Scores
3.5. Classification Performance for Index Tumors
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Hoeks, C.M.A.; Barentsz, J.O.; Hambrock, T.; Yakar, D.; Somford, D.M.; Heijmink, S.W.T.P.J.; Scheenen, T.W.J.; Vos, P.C.; Huisman, H.; van Oort, I.M.; et al. Prostate Cancer: Multiparametric MR Imaging for Detection, Localization, and Staging. Radiology 2011, 261, 46–66. [Google Scholar] [CrossRef] [PubMed]
- Bjurlin, M.A.; Carroll, P.R.; Eggener, S.; Fulgham, P.F.; Margolis, D.J.; Pinto, P.A.; Rosenkrantz, A.B.; Rubenstein, J.N.; Rukstalis, D.B.; Taneja, S.S.; et al. Update of the standard operating procedure on the use of multiparametric magnetic resonance imaging for the diagnosis, staging and management of prostate cancer. J. Urol. 2020, 203, 706–712. [Google Scholar] [CrossRef] [PubMed]
- Mottet, N.; Bellmunt, J.; Bolla, M.; Briers, E.; Cumberbatch, M.G.; De Santis, M.; Fossati, N.; Gross, T.; Henry, A.M.; Joniau, S.; et al. EAU-ESTRO-SIOG guidelines on prostate cancer. Part 1: Screening, diagnosis, and local treatment with curative intent. Eur. Urol. 2017, 71, 618–629. [Google Scholar] [CrossRef] [PubMed]
- Turkbey, B.; Rosenkrantz, A.B.; Haider, M.A.; Padhani, A.R.; Villeirs, G.; Macura, K.J.; Tempany, C.M.; Choyke, P.L.; Cornud, F.; Margolis, D.J.; et al. Prostate Imaging Reporting and Data System Version 2.1: 2019 Update of Prostate Imaging Reporting and Data System Version 2. Eur. Urol. 2019, 76, 340–351. [Google Scholar] [CrossRef]
- Woo, S.; Suh, C.H.; Kim, S.Y.; Cho, J.Y.; Kim, S.H. Diagnostic performance of prostate imaging reporting and data system version 2 for detection of prostate cancer: A systematic review and diagnostic meta-analysis. Eur. Urol. 2017, 72, 177–188. [Google Scholar] [CrossRef]
- Padhani, A.R.; Weinreb, J.; Rosenkrantz, A.B.; Villeirs, G.; Turkbey, B.; Barentsz, J. Prostate imaging-reporting and data system steering committee: PI-RADS v2 status update and future directions. Eur. Urol. 2019, 75, 385–396. [Google Scholar] [CrossRef] [Green Version]
- Tewes, S.; Mokov, N.; Hartung, D.; Schick, V.; Peters, I.; Schedl, P.; Pertschy, S.; Wacker, F.; Voshage, G.; Hueper, K. Standardized reporting of prostate MRI: Comparison of the prostate imaging reporting and data system (PI-RADS) version 1 and version 2. PLoS ONE 2016, 11, e0162879. [Google Scholar] [CrossRef]
- Costa, D.N.; Pedrosa, I.; Donato, F., Jr.; Roehrborn, C.G.; Rofsky, N.M. MR imaging–transrectal US fusion for targeted prostate biopsies: Implications for diagnosis and clinical management. Radiographics 2015, 35, 696–708. [Google Scholar] [CrossRef] [PubMed]
- Purysko, A.S.; Bittencourt, L.K.; Bullen, J.A.; Mostardeiro, T.R.; Herts, B.R.; Klein, E.A. Accuracy and interobserver agreement for prostate imaging reporting and data system, version 2, for the characterization of lesions identified on multiparametric MRI of the prostate. Am. J. Roentgenol. 2017, 209, 339–349. [Google Scholar] [CrossRef] [PubMed]
- Girometti, R.; Giannarini, G.; Greco, F.; Isola, M.; Cereser, L.; Como, G.; Sioletic, S.; Pizzolitto, S.; Crestani, A.; Ficarra, V.; et al. Interreader agreement of PI-RADS v. 2 in assessing prostate cancer with multiparametric MRI: A study using whole-mount histology as the standard of reference. J. Magn. Reson. Imaging 2019, 49, 546–555. [Google Scholar] [CrossRef] [PubMed]
- Seo, J.W.; Shin, S.-J.; Taik Oh, Y.; Jung, D.C.; Cho, N.H.; Choi, Y.D.; Park, S.Y. PI-RADS version 2: Detection of clinically significant cancer in patients with biopsy gleason score 6 prostate cancer. Am. J. Roentgenol. 2017, 209, W1–W9. [Google Scholar] [CrossRef]
- van der Leest, M.; Cornel, E.; Israël, B.; Hendriks, R.; Padhani, A.R.; Hoogenboom, M.; Zamecnik, P.; Bakker, D.; Setiasti, A.Y.; Veltman, J.; et al. Head-to-head Comparison of Transrectal Ultrasound-guided Prostate Biopsy Versus Multiparametric Prostate Resonance Imaging with Subsequent Magnetic Resonance-guided Biopsy in Biopsy-naïve Men with Elevated Prostate-specific Antigen: A Large Prospective M. Eur. Urol. 2019, 75, 570–578. [Google Scholar] [CrossRef] [Green Version]
- Venderink, W.; van Luijtelaar, A.; Bomers, J.G.R.; van der Leest, M.; de Kaa, C.; Barentsz, J.O.; Sedelaar, J.P.M.; Fütterer, J.J. Results of targeted biopsy in men with magnetic resonance imaging lesions classified equivocal, likely or highly likely to be clinically significant prostate cancer. Eur. Urol. 2018, 73, 353–360. [Google Scholar] [CrossRef] [PubMed]
- Manjunath, B.S.; Ma, W.-Y. Texture features for browsing and retrieval of image data. IEEE Trans. Pattern Anal. Mach. Intell. 1996, 18, 837–842. [Google Scholar] [CrossRef] [Green Version]
- Partio, M.; Cramariuc, B.; Gabbouj, M.; Visa, A. Rock texture retrieval using gray level co-occurrence matrix. In Proceedings of the 5th Nordic Signal Processing Symposium, Hurtigruten, Norway, 4–7 October 2002; Volume 75. [Google Scholar]
- Gatenby, R.A.; Grove, O.; Gillies, R.J. Quantitative imaging in cancer evolution and ecology. Radiology 2013, 269, 8–14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Avanzo, M.; Wei, L.; Stancanello, J.; Vallières, M.; Rao, A.; Morin, O.; Mattonen, S.A.; El Naqa, I. Machine and deep learning methods for radiomics. Med. Phys. 2020, 47, e185–e202. [Google Scholar] [CrossRef]
- Esteva, A.; Kuprel, B.; Novoa, R.A.; Ko, J.; Swetter, S.M.; Blau, H.M.; Thrun, S. Dermatologist-level classification of skin cancer with deep neural networks. Nature 2017, 542, 115–118. [Google Scholar] [CrossRef]
- Liu, Y.; Yang, G.; Mirak, S.A.; Hosseiny, M.; Azadikhah, A.; Zhong, X.; Reiter, R.E.; Lee, Y.; Raman, S.S.; Sung, K. Automatic Prostate Zonal Segmentation Using Fully Convolutional Network With Feature Pyramid Attention. IEEE Access 2019, 7, 163626–163632. [Google Scholar] [CrossRef]
- Liu, H.; Li, H.; Habes, M.; Li, Y.; Boimel, P.; Janopaul-Naylor, J.; Xiao, Y.; Ben-Josef, E.; Fan, Y. Robust Collaborative Clustering of Subjects and Radiomic Features for Cancer Prognosis. IEEE Trans. Biomed. Eng. 2020, 67, 2735–2744. [Google Scholar] [CrossRef]
- Tan, J.; Gao, Y.; Liang, Z.; Cao, W.; Pomeroy, M.J.; Huo, Y.; Li, L.; Barish, M.A.; Abbasi, A.F.; Pickhardt, P.J. 3D-GLCM CNN: A 3-Dimensional Gray-Level Co-Occurrence Matrix-Based CNN Model for Polyp Classification via CT Colonography. IEEE Trans. Med. Imaging 2019, 39, 2013–2024. [Google Scholar] [CrossRef]
- Heinrich, M.P.; Jenkinson, M.; Bhushan, M.; Matin, T.; Gleeson, F.V.; Brady, M.; Schnabel, J.A. MIND: Modality independent neighbourhood descriptor for multi-modal deformable registration. Med. Image Anal. 2012, 16, 1423–1435. [Google Scholar] [CrossRef]
- Paszke, A.; Gross, S.; Chintala, S.; Chanan, G.; Yang, E.; DeVito, Z.; Lin, Z.; Desmaison, A.; Antiga, L.; Lerer, A. Automatic differentiation in pytorch. In Proceedings of the 31st Conference on Neural Information Processing Systems (NIPS 2017), Long Beach, CA, USA, 4–9 December 2017. [Google Scholar]
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. arXiv 2014, arXiv:1412.6980. [Google Scholar]
- Winkel, D.J.; Tong, A.; Lou, B.; Kamen, A.; Comaniciu, D.; Disselhorst, J.A.; Rodriguez-Ruiz, A.; Huisman, H.; Szolar, D.; Shabunin, I.; et al. A Novel Deep Learning Based Computer-Aided Diagnosis System Improves the Accuracy and Efficiency of Radiologists in Reading Biparametric Magnetic Resonance Images of the Prostate: Results of a Multireader, Multicase Study. Investig. Radiol. 2021, 56, 605–613. [Google Scholar] [CrossRef] [PubMed]
- Bonekamp, D.; Kohl, S.; Wiesenfarth, M.; Schelb, P.; Radtke, J.P.; Götz, M.; Kickingereder, P.; Yaqubi, K.; Hitthaler, B.; Gählert, N.; et al. Radiomic machine learning for characterization of prostate lesions with MRI: Comparison to ADC values. Radiology 2018, 289, 128–137. [Google Scholar] [CrossRef] [PubMed]
- der Walt, S.; Schönberger, J.L.; Nunez-Iglesias, J.; Boulogne, F.; Warner, J.D.; Yager, N.; Gouillart, E.; Yu, T. Scikit-image: Image processing in Python. PeerJ 2014, 2, e453. [Google Scholar] [CrossRef] [PubMed]
- Ho, T.K. Random decision forests. In Proceedings of the 3rd International Conference on Document Analysis and Recognition, Montreal, QC, Canada, 14–16 August 1995. [Google Scholar] [CrossRef]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Zhong, X.; Cao, R.; Shakeri, S.; Scalzo, F.; Lee, Y.; Enzmann, D.R.; Wu, H.H.; Raman, S.S.; Sung, K. Deep transfer learning-based prostate cancer classification using 3 Tesla multi-parametric MRI. Abdom. Radiol. 2019, 44, 2030–2039. [Google Scholar] [CrossRef] [PubMed]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Song, Y.; Zhang, Y.-D.; Yan, X.; Liu, H.; Zhou, M.; Hu, B.; Yang, G. Computer-aided diagnosis of prostate cancer using a deep convolutional neural network from multiparametric MRI. J. Magn. Reson. Imaging 2018, 48, 1570–1577. [Google Scholar] [CrossRef]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. arXiv 2014, arXiv:1409.1556. [Google Scholar]
- Weinreb, J.C.; Barentsz, J.O.; Choyke, P.L.; Cornud, F.; Haider, M.A.; Macura, K.J.; Margolis, D.; Schnall, M.D.; Shtern, F.; Tempany, C.M.; et al. PI-RADS prostate imaging–reporting and data system: 2015, version 2. Eur. Urol. 2016, 69, 16–40. [Google Scholar] [CrossRef] [PubMed]
- Agresti, A.; Coull, B.A. Approximate is Better than “Exact” for Interval Estimation of Binomial Proportions. Am. Stat. 1998, 52, 119–126. [Google Scholar] [CrossRef]
- Fluss, R.; Faraggi, D.; Reiser, B. Estimation of the Youden Index and its associated cutoff point. Biom. J. J. Math. Methods Biosci. 2005, 47, 458–472. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- DeLong, E.R.; DeLong, D.M.; Clarke-Pearson, D.L. Comparing the areas under two or more correlated receiver operating characteristic curves: A nonparametric approach. Biometrics 1988, 44, 837–845. [Google Scholar] [CrossRef]
- Eliasziw, M.; Donner, A. Application of the McNemar test to non-independent matched pair data. Stat. Med. 1991, 10, 1981–1991. [Google Scholar] [CrossRef] [PubMed]
- Gaur, S.; Lay, N.; Harmon, S.A.; Doddakashi, S.; Mehralivand, S.; Argun, B.; Barrett, T.; Bednarova, S.; Girometti, R.; Karaarslan, E.; et al. Can computer-aided diagnosis assist in the identification of prostate cancer on prostate MRI? a multi-center, multi-reader investigation. Oncotarget 2018, 9, 33804. [Google Scholar] [CrossRef] [Green Version]
- Zhou, H.; Hallac, R.R.; Yuan, Q.; Ding, Y.; Zhang, Z.; Xie, X.-J.; Francis, F.; Roehrborn, C.G.; Sims, R.D.; Costa, D.N.; et al. Incorporating oxygen-enhanced MRI into multi-parametric assessment of human prostate cancer. Diagnostics 2017, 7, 48. [Google Scholar] [CrossRef] [PubMed] [Green Version]


| Characteristics | Overall | Train/Validation | Test | |
|---|---|---|---|---|
| Patient Number | 402 | 281 | 121 | |
| csPCa lesions/all lesions | 303/466 | 225/324 | 78/142 | |
| Non-csPCa lesions/all lesions | 163/466 | 99/324 | 64/142 | |
| Age (yr.) | 61 (56–66) | 61 (56–67) | 61 (58–66) | |
| Weight (kg.) | 87 (77–95) | 87 (77–96) | 86 (78–94) | |
| PSA (ng/mL) | 8.3 (4.7–8.7) | 8.7 (4.7–8.9) | 7.4 (4.6–8.7) | |
| Tumor Volume (cm3) | 1.1 (0.3–1.2) | 1.1 (0.3–1.2) | 0.9 (0.2–1.1) | |
| GS | False Positive | 80 | 49 | 31 |
| 3 + 3 = 6 | 83 | 50 | 33 | |
| 3 + 4 = 7 | 168 | 126 | 42 | |
| 4 + 3 = 7 | 79 | 57 | 22 | |
| >7 | 56 | 42 | 14 | |
| PI-RADS score | 3 | 119 | 76 | 43 |
| 4 | 197 | 138 | 59 | |
| 5 | 150 | 110 | 40 | |
| Prostate zone | PZ | 364 | 255 | 109 |
| TZ | 99 | 67 | 32 | |
| AFS | 3 | 2 | 1 | |
| Focality | Solitary | 190 | 137 | 53 |
| Multi-focal | 276 | 187 | 89 | |
| Lesion Type | csPCa (%) | Non-csPCa (%) | Method | AUC (95% CI) | Sensitivity (95% CI) | Specificity (95% CI) |
|---|---|---|---|---|---|---|
| PI-RADS score 3 | 13 (30) | 30 (70) | Textured-DL | 0.81 [0.68–0.94] | 0.85 [0.65–1] | 0.73 0.58–0.89] |
| PI-RADS score 4 | 31 (53) | 28 (47) | Textured-DL | 0.84 [0.74–0.94] | 0.87 [0.75–0.99) | 0.71 [0.55–0.88) |
| PI-RADS score 5 | 34 (85) | 6 (15) | Textured-DL | 0.84 [0.66–1] | 0.88 [0.77–0.99] | 0.67 [0.29–1] |
| Tumor Type | csPCa (%) | Non-csPCa (%) | Method | AUC (95% CI) | p-Value | Sensitivity (95% CI) | p-Value | Specificity (95% CI) | p-Value |
|---|---|---|---|---|---|---|---|---|---|
| All index | 77 (84) | 15 (16) | PI-RADS-CLA | 0.65 [0.52–0.78] | 0.32 | 0.83 [0.75–0.91] | 0.25 | 0.27 [0.04–0.49] | 0.18 |
| Textured-DL | 0.73 [0.59–0.88] | 0.90 [0.83–0.96] | 0.47 [0.21–0.72] | ||||||
| Index with PSA < 4 | 10 (77) | 3 (23) | PI-RADS-CLA | 0.72 [0.41–1.00] | 0.06 | 0.7 [0.42–0.98] | 0.32 | 0.67 [0.13–1.00] | >0.99 |
| Textured-DL | 0.87 [0.58–1.00] | 0.9 [0.71–1] | 0.67 [0.13–1.00] | ||||||
| Index with 4 ≤ PSA < 10 | 53 (87) | 8 (13) | PI-RADS-CLA | 0.56 [0.37–0.75] | 0.35 | 0.87 [0.78–0.96] | 0.03 | 0.12 [0.00–0.35] | 0.56 |
| Textured-DL | 0.43 [0.20–0.65] | 0.98 [0.94–1.00] | 0.25 [0.00–0.55] | ||||||
| Index with PSA ≥ 10 | 14 (78) | 4 (22) | PI-RADS-CLA | 0.79 [0.58–0.99] | 0.17 | 0.79 [0.57–1.00] | >0.99 | 0.25 [0.00–0.67] | 0.08 |
| Textured-DL | 0.93 [0.80–1.00] | 0.79 [0.31–0.83] | 1.00 [1.00–1.00] |
| Method | PPV (%) | NPV (%) |
|---|---|---|
| Textured-DL | 58 | 92 |
| PSAD ≥ 0.10 | 29 | 67 |
| PSAD ≥ 0.15 | 35 | 75 |
| PSAD ≥ 0.20 | 25 | 68 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Y.; Zheng, H.; Liang, Z.; Miao, Q.; Brisbane, W.G.; Marks, L.S.; Raman, S.S.; Reiter, R.E.; Yang, G.; Sung, K. Textured-Based Deep Learning in Prostate Cancer Classification with 3T Multiparametric MRI: Comparison with PI-RADS-Based Classification. Diagnostics 2021, 11, 1785. https://doi.org/10.3390/diagnostics11101785
Liu Y, Zheng H, Liang Z, Miao Q, Brisbane WG, Marks LS, Raman SS, Reiter RE, Yang G, Sung K. Textured-Based Deep Learning in Prostate Cancer Classification with 3T Multiparametric MRI: Comparison with PI-RADS-Based Classification. Diagnostics. 2021; 11(10):1785. https://doi.org/10.3390/diagnostics11101785
Chicago/Turabian StyleLiu, Yongkai, Haoxin Zheng, Zhengrong Liang, Qi Miao, Wayne G. Brisbane, Leonard S. Marks, Steven S. Raman, Robert E. Reiter, Guang Yang, and Kyunghyun Sung. 2021. "Textured-Based Deep Learning in Prostate Cancer Classification with 3T Multiparametric MRI: Comparison with PI-RADS-Based Classification" Diagnostics 11, no. 10: 1785. https://doi.org/10.3390/diagnostics11101785








