Implementation of Next Generation Sequencing-Based Liquid Biopsy for Clinical Molecular Diagnostics in Non-Small Cell Lung Cancer (NSCLC) Patients
Abstract
:1. Introduction
2. Materials and Methods
2.1. Patients and Plasma Sample Collection
2.2. Extraction of cfDNA and Assessment of Its Quantity and Quality
2.3. cfDNA Sequencing with the AVENIO ctDNA Expanded Kit
2.4. NGS Data Analysis
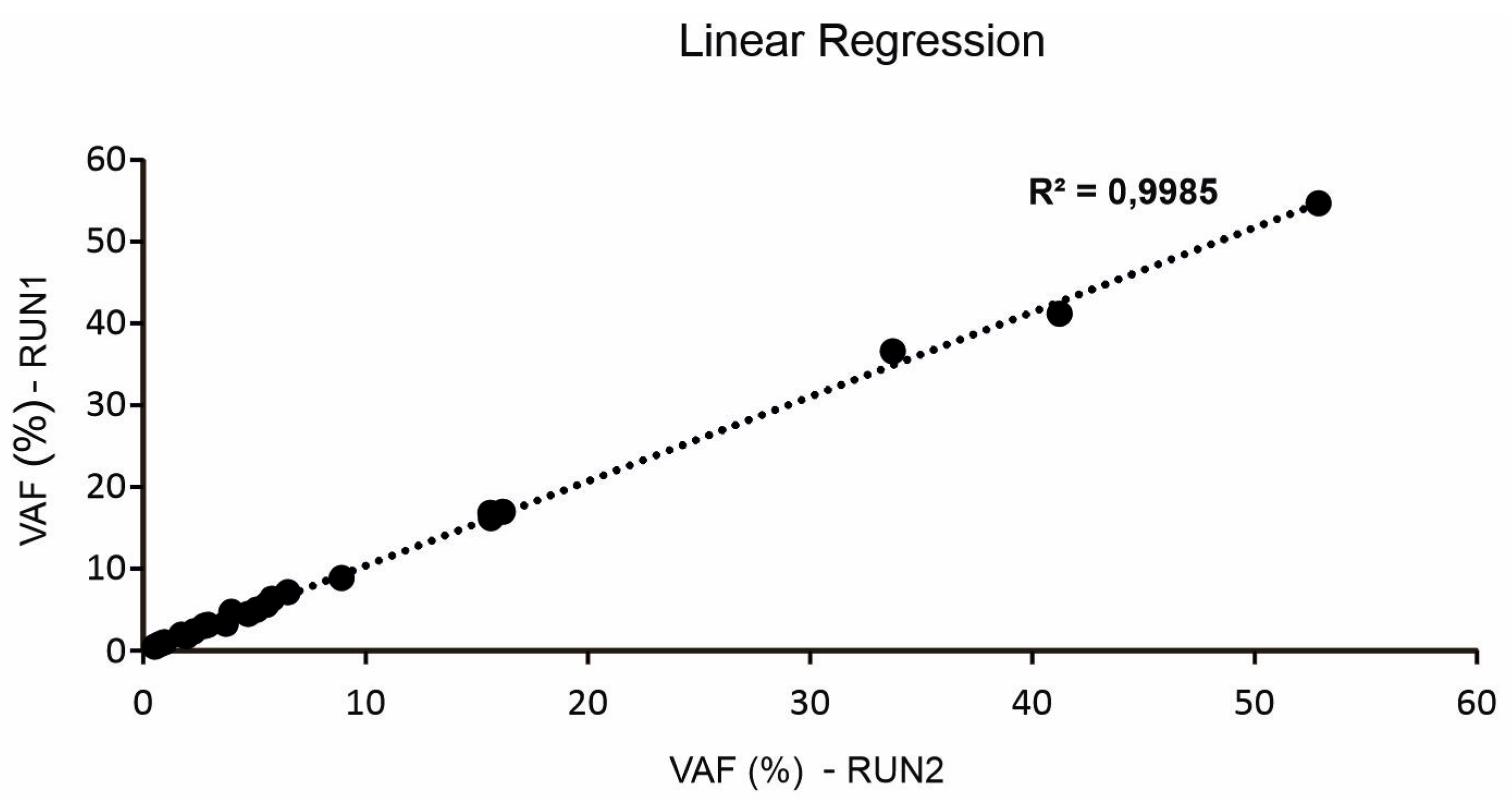
2.5. Inter-Run Assay Reproducibility of cfDNA NGS Analysis
2.6. Tissue Genetic Analysis with the AVENIO Tumor Tissue Expanded Kit
2.7. Statistical Analysis and Data Analysis
3. Results
3.1. Panel Overview
3.2. Technical Assessment of cfDNA NGS Analysis
3.2.1. Assessment of Sensitivity and Limit of Detection (LOD)
3.2.2. Inter-Run Assay Reproducibility of cfDNA NGS Analysis
3.2.3. Comparison of Plasma versus Tissue NGS Analysis
3.3. Study Population
3.4. Data Collection and Interpretation
3.4.1. Assessment of ctDNA Quantity and Quality
3.4.2. Sequencing Performance
3.4.3. Gene Variant Detection
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Meric-Bernstam, F.; Mills, G.B. Overcoming implementation challenges of personalized cancer therapy. Nat. Rev. Clin. Oncol. 2012, 9, 542–548. [Google Scholar] [CrossRef]
- McGranahan, N.; Swanton, C. Clonal Heterogeneity and Tumor Evolution: Past, Present, and the Future. Cell 2017, 168, 613–628. [Google Scholar] [CrossRef] [Green Version]
- Pereira, B.; Chen, C.T.; Goyal, L.; Walmsley, C.; Pinto, C.J.; Baiev, I.; Allen, R.; Henderson, L.; Saha, S.; Taylor, M.S.; et al. Cell-free DNA captures tumor heterogeneity and driver alterations in rapid autopsies with pre-treated metastatic cancer. Nat. Commun. 2021, 12, 3199. [Google Scholar] [CrossRef]
- Indraccolo, S. Evolving use of liquid biopsy in non-small-cell-lung cancer patients. Int. J. Biol. Markers 2020, 35, 23–25. [Google Scholar] [CrossRef] [PubMed]
- Esposito Abate, R.; Frezzetti, D.; Maiello, M.R.; Gallo, M.; Camerlingo, R.; De Luca, A.; De Cecio, R.; Morabito, A.; Normanno, N. Next Generation Sequencing-Based Profiling of Cell Free DNA in Patients with Advanced Non-Small Cell Lung Cancer: Advantages and Pitfalls. Cancers 2020, 12, 3804. [Google Scholar] [CrossRef] [PubMed]
- Li, B.T.; Janku, F.; Jung, B.; Hou, C.; Madwani, K.; Alden, R.; Razavi, P.; Reis-Filho, J.S.; Shen, R.; Isbell, J.M.; et al. Ultra-deep next-generation sequencing of plasma cell-free DNA in patients with advanced lung cancers: Results from the Actionable Genome Consortium. Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 2019, 30, 597–603. [Google Scholar] [CrossRef]
- Pugh, T.J. Circulating Tumour DNA for Detecting Minimal Residual Disease in Multiple Myeloma. Semin. Hematol. 2018, 55, 38–40. [Google Scholar] [CrossRef] [PubMed]
- Thakral, D.; Das, N.; Basnal, A.; Gupta, R. Cell-free DNA for genomic profiling and minimal residual disease monitoring in Myeloma- are we there yet? Am. J. Blood Res. 2020, 10, 26–45. [Google Scholar]
- Aguirre-Ruiz, P.; Ariceta, B.; Viguria, M.C.; Zudaire, M.T.; Blasco-Iturri, Z.; Arnedo, P.; Aguilera-Diaz, A.; Jauregui, A.; Mañú, A.; Prosper, F.; et al. Assessment of Minimal Residual Disease by Next Generation Sequencing in Peripheral Blood as a Complementary Tool for Personalized Transplant Monitoring in Myeloid Neoplasms. J. Clin. Med. 2020, 9, 3818. [Google Scholar] [CrossRef] [PubMed]
- Zulato, E.; Attili, I.; Pavan, A.; Nardo, G.; Del Bianco, P.; Verza, M.; Pasqualini, L.; Pasello, G.; Fassan, M.; Bragadin, A.B.; et al. Early assessment of KRAS mutation in cfDNA correlates with risk of progression and death in advanced non-small-cell lung cancer. Br. J. Cancer 2020, 123, 81–91. [Google Scholar] [CrossRef]
- Mosele, F.; Remon, J.; Mateo, J.; Westphalen, C.B.; Barlesi, F.; Lolkema, M.P.; Normanno, N.; Scarpa, A.; Robson, M.; Meric-Bernstam, F.; et al. Recommendations for the use of next-generation sequencing (NGS) for patients with metastatic cancers: A report from the ESMO Precision Medicine Working Group. Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 2020, 31, 1491–1505. [Google Scholar] [CrossRef]
- Fan, H.C.; Blumenfeld, Y.J.; Chitkara, U.; Hudgins, L.; Quake, S.R. Noninvasive diagnosis of fetal aneuploidy by shotgun sequencing DNA from maternal blood. Proc. Natl. Acad. Sci. USA 2008, 105, 16266–16271. [Google Scholar] [CrossRef] [Green Version]
- Jennings, L.J.; Arcila, M.E.; Corless, C.; Kamel-Reid, S.; Lubin, I.M.; Pfeifer, J.; Temple-Smolkin, R.L.; Voelkerding, K.V.; Nikiforova, M.N. Guidelines for Validation of Next-Generation Sequencing-Based Oncology Panels: A Joint Consensus Recommendation of the Association for Molecular Pathology and College of American Pathologists. J. Mol. Diagn. 2017, 19, 341–365. [Google Scholar] [CrossRef] [Green Version]
- Verma, S.; Moore, M.W.; Ringler, R.; Ghosal, A.; Horvath, K.; Naef, T.; Anvari, S.; Cotter, P.D.; Gunn, S. Analytical performance evaluation of a commercial next generation sequencing liquid biopsy platform using plasma ctDNA, reference standards, and synthetic serial dilution samples derived from normal plasma. BMC Cancer 2020, 20, 945. [Google Scholar] [CrossRef] [PubMed]
- Spence, T.; Perera, S.; Weiss, J.; Grenier, S.; Ranich, L.; Shepherd, F.; Stockley, T.L. Clinical implementation of circulating tumour DNA testing for EGFR T790M for detection of treatment resistance in non-small cell lung cancer. J. Clin. Pathol. 2021, 74, 91–97. [Google Scholar] [CrossRef] [PubMed]
- Cortiula, F.; Pasello, G.; Follador, A.; Nardo, G.; Polo, V.; Scquizzato, E.; Del Conte, A.; Miorin, M.; Giovanis, P.; D’Urso, A.; et al. A Multi-Center, Real-Life Experience on Liquid Biopsy Practice for EGFR Testing in Non-Small Cell Lung Cancer (NSCLC) Patients. Diagnostics 2020, 10, 765. [Google Scholar] [CrossRef]
- Papadopoulou, E.; Tsoulos, N.; Tsantikidi, K.; Metaxa-Mariatou, V.; Stamou, P.E.; Kladi-Skandali, A.; Kapeni, E.; Tsaousi, G.; Pentheroudakis, G.; Petrakis, D.; et al. Clinical feasibility of NGS liquid biopsy analysis in NSCLC patients. PLoS ONE 2019, 14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hu, Y.; Ulrich, B.C.; Supplee, J.; Kuang, Y.; Lizotte, P.H.; Feeney, N.B.; Guibert, N.M.; Awad, M.M.; Wong, K.-K.; Jänne, P.A.; et al. False-Positive Plasma Genotyping Due to Clonal Hematopoiesis. Clin. Cancer Res. 2018, 24, 4437–4443. [Google Scholar] [CrossRef] [Green Version]
- Razavi, P.; Li, B.T.; Brown, D.N.; Jung, B.; Hubbell, E.; Shen, R.; Abida, W.; Juluru, K.; De Bruijn, I.; Hou, C.; et al. High-intensity sequencing reveals the sources of plasma circulating cell-free DNA variants. Nat. Med. 2019, 25, 1928–1937. [Google Scholar] [CrossRef] [PubMed]
- Alborelli, I.; Generali, D.; Jermann, P.; Cappelletti, M.R.; Ferrero, G.; Scaggiante, B.; Bortul, M.; Zanconati, F.; Nicolet, S.; Haegele, J.; et al. Cell-free DNA analysis in healthy individuals by next-generation sequencing: A proof of concept and technical validation study. Cell Death Dis. 2019, 10, 534. [Google Scholar] [CrossRef]
- Fernandez-Cuesta, L.; Perdomo, S.; Avogbe, P.H.; Leblay, N.; Delhomme, T.M.; Gaborieau, V.; Abedi-Ardekani, B.; Chanudet, E.; Olivier, M.; Zaridze, D.; et al. Identification of Circulating Tumor DNA for the Early Detection of Small-cell Lung Cancer. EBioMedicine 2016, 10, 117–123. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lam, S.N.; Zhou, Y.C.; Chan, Y.M.; Foo, C.M.; Lee, P.Y.; Mok, W.Y.; Wong, W.S.; Fung, Y.Y.; Wong, K.Y.; Huang, J.Y.; et al. Comparison of Target Enrichment Platforms for Circulating Tumor DNA Detection. Sci. Rep. 2020, 10, 4124. [Google Scholar] [CrossRef] [PubMed] [Green Version]




| 0.1% Multiplex I cfDNA Reference Standard (Part No.: HD779) | ||||||
|---|---|---|---|---|---|---|
| Gene | Variant | Expected VAF (%) | AVENIO_1 Replicate | AVENIO_2 Replicate | ||
| Detected | VAF | Detected | VAF | |||
| EGFR | p.L858R | 0.10% | NO | YES | 0.09% | |
| EGFR | p.ΔE746-A750 | 0.10% | YES | 0.11% | NO | |
| EGFR | p.T790M | 0.10% | NO | YES | 0.12% | |
| EGFR | p.V769-D770insASV | 0.10% | NO | YES | 0.12% | |
| KRAS | p.G12D | 0.13% | YES | 0.14% | YES | 0.16% |
| NRAS | p.Q61K | 0.13% | YES | 0.26% | YES | 0.13% |
| NRAS | p.A59T | 0.13% | YES | 0.19% | NO | |
| PIK3CA | p.E545K | 0.13% | NO | YES | 0.13% | |
| Sample with an Expected VAF of 0.5%, Diluted from 1% Multiplex I cfDNA Reference Standard (Part No.: HD778) | ||||||
| Gene | Variant | Expected VAF (%) | AVENIO Results | |||
| Detected | VAF | |||||
| EGFR | p.L858R | 0.50% | YES | 0.49% | ||
| EGFR | p.ΔE746-A750 | 0.50% | YES | 0.42% | ||
| EGFR | p.T790M | 0.50% | YES | 0.30% | ||
| EGFR | p.V769-D770insASV | 0.50% | YES | 0.29% | ||
| KRAS | p.G12D | 0.65% | YES | 0.40% | ||
| NRAS | p.Q61K | 0.65% | YES | 0.71% | ||
| NRAS | p.A59T | 0.65% | YES | 0.72% | ||
| PIK3CA | p.E545K | 0.65% | YES | 0.43% | ||
| HORIZON Structural Multiplex cfDNA Reference Standard | ||||||
| Gene | Variant | Expected VAF (%) | AVENIO Results | |||
| Detected | VAF | |||||
| GNA11 | p.Q209L | 5.6 | YES | 4.17% | ||
| AKT1 | p.E17K | 5.0 | YES | 3.49% | ||
| PIK3CA | p.E545K | 5.6 | YES | 4.63% | ||
| EGFR | p.V769_D770insASV | 5.6 | YES | 4.50% | ||
| EGFR | p.∆E746-A750 | 5.3 | YES | 4.70% | ||
| ROS1 | SLC34A2/ROS1 fusion | 5.6 | YES | |||
| RET | CCDC6/RET fusion | 5.0 | YES | |||
| MET | amplification | 4.5 copies | YES | |||
| KRAS | p.G13D | 5.6 | YES | 4.41% | ||
| MET | p.V237fs | 2.5 | DEL NOT COVERED BY THE PANEL | |||
| FLT3 | p.S985fs | 5.6 | DEL NOT COVERED BY THE PANEL | |||
| BRCA2 | p.A1689fs | 5.6 | DEL NOT COVERED BY THE PANEL | |||
| FBXW7 | p.G667fs | 5.6 | DEL NOT COVERED BY THE PANEL | |||
| EGFR | p.G719S | 5.3 | YES | 5.07% | ||
| BRAF | p.V600E | 18.2 | YES | 15.58% | ||
| PIK3CA | p.H1047R | 16.7 | YES | 15.73% | ||
| MYC-N | amplification | 9.5 copies | GENE NOT INCLUDED IN THE PANEL | |||
| NOTCH1 | p.P668S | 5.0 | GENE NOT INCLUDED IN THE PANEL | |||
| Sample | Ng Inpu | Gene | SNV | VAF (%) | Mean | St dev | ||
|---|---|---|---|---|---|---|---|---|
| RUN 1 | RUN 2 | RUN 1 | RUN 2 | |||||
| 1 | 50 | 50 | KRAS | p.Gly12Ala | 52.88 | 54.63 | 53.76 | 1.24 |
| TP53 | p.Arg306 * | 41.22 | 41.18 | 41.20 | 0.03 | |||
| FLT1 | p.Glu201Asp | 0.86 | 0.90 | 0.88 | 0.03 | |||
| MSH2 | p.Arg621Leu | 16.17 | 16.95 | 16.56 | 0.55 | |||
| MTOR | p.Ala1792Pro | 0.70 | 0.74 | 0.72 | 0.03 | |||
| PTCH1 | p.Val1386Ile | 0.50 | 0.48 | 0.49 | 0.01 | |||
| 2 | 50 | 50 | KRAS | p.Gly12Ala | 8.91 | 8.81 | 8.86 | 0.07 |
| TP53 | p.Arg306 * | 5.79 | 6.31 | 6.05 | 0.37 | |||
| MSH2 | p.Arg621Leu | 2.89 | 3.11 | 3.00 | 0.16 | |||
| 3 | 50 | 27 | TP53 | p.Gln331 * | 1.92 | 1.72 | 1.82 | 0.14 |
| TP53 | p.Arg248Gln | 0.95 | 0.95 | 0.95 | 0.00 | |||
| APC | p.Gln1611Glu | 1.70 | 1.89 | 1.80 | 0.13 | |||
| 4 | 17,5 | 23 | BRAF | p.Gly596Arg | 5.53 | 5.54 | 5.54 | 0.01 |
| TP53 | p.Gly245Val | 3.71 | 3.26 | 3.49 | 0.32 | |||
| DDR2 | p.Glu583Lys | 2.26 | 2.34 | 2.30 | 0.06 | |||
| KEAP1 | p.Arg536Pro | 4.71 | 4.43 | 4.57 | 0.20 | |||
| TSC2 | p.Ser740Cys | 3.94 | 4.68 | 4.31 | 0.52 | |||
| 5 | 50 | 20 | RB1 | p.Arg262Gln | 15.61 | 16.80 | 16.21 | 0.84 |
| TP53 | p.His179Arg | 33.71 | 36.57 | 35.14 | 2.02 | |||
| KDR | p.Ser968Ile | 2.72 | 3.00 | 2.86 | 0.20 | |||
| MSH6 | p.Leu1264Val | 15.64 | 16.15 | 15.90 | 0.36 | |||
| TERT | c.-146C > T | 6.50 | 7.07 | 6.79 | 0.40 | |||
| 6 | 50 | 50 | TP53 | c.993 + 1G > A | 5.09 | 4.98 | 5.04 | 0.08 |
| Sample | Gene | SNV | AVENIO cfDNA Expanded Assay | AVENIO Tumor Tissue Expanded Assay |
|---|---|---|---|---|
| 1 | PTCH1 | p.Ala793Ser | detected | detected |
| SMO | p.Asp25Gly | detected | detected | |
| MLH1 | c.1558 + 3A> | detected | detected | |
| 2 | TP53 | p.Ile255Phe | detected | detected |
| RB1 | p.Arg251 * | not detected | detected | |
| 3 | KRAS | p.Gly12Ala | detected | detected |
| TP53 | p.Arg306 * | detected | detected | |
| MSH2 | p.Arg621Leu | detected | detected | |
| MTOR | p.Ala1792Pro | detected | not detected | |
| MSH6 | p.Trp55Arg | not detected | detected |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zulato, E.; Tosello, V.; Nardo, G.; Bonanno, L.; Del Bianco, P.; Indraccolo, S. Implementation of Next Generation Sequencing-Based Liquid Biopsy for Clinical Molecular Diagnostics in Non-Small Cell Lung Cancer (NSCLC) Patients. Diagnostics 2021, 11, 1468. https://doi.org/10.3390/diagnostics11081468
Zulato E, Tosello V, Nardo G, Bonanno L, Del Bianco P, Indraccolo S. Implementation of Next Generation Sequencing-Based Liquid Biopsy for Clinical Molecular Diagnostics in Non-Small Cell Lung Cancer (NSCLC) Patients. Diagnostics. 2021; 11(8):1468. https://doi.org/10.3390/diagnostics11081468
Chicago/Turabian StyleZulato, Elisabetta, Valeria Tosello, Giorgia Nardo, Laura Bonanno, Paola Del Bianco, and Stefano Indraccolo. 2021. "Implementation of Next Generation Sequencing-Based Liquid Biopsy for Clinical Molecular Diagnostics in Non-Small Cell Lung Cancer (NSCLC) Patients" Diagnostics 11, no. 8: 1468. https://doi.org/10.3390/diagnostics11081468
APA StyleZulato, E., Tosello, V., Nardo, G., Bonanno, L., Del Bianco, P., & Indraccolo, S. (2021). Implementation of Next Generation Sequencing-Based Liquid Biopsy for Clinical Molecular Diagnostics in Non-Small Cell Lung Cancer (NSCLC) Patients. Diagnostics, 11(8), 1468. https://doi.org/10.3390/diagnostics11081468







