Prognostic Model of ICU Admission Risk in Patients with COVID-19 Infection Using Machine Learning
Abstract
:1. Introduction
2. Related Work
3. Materials and Methods
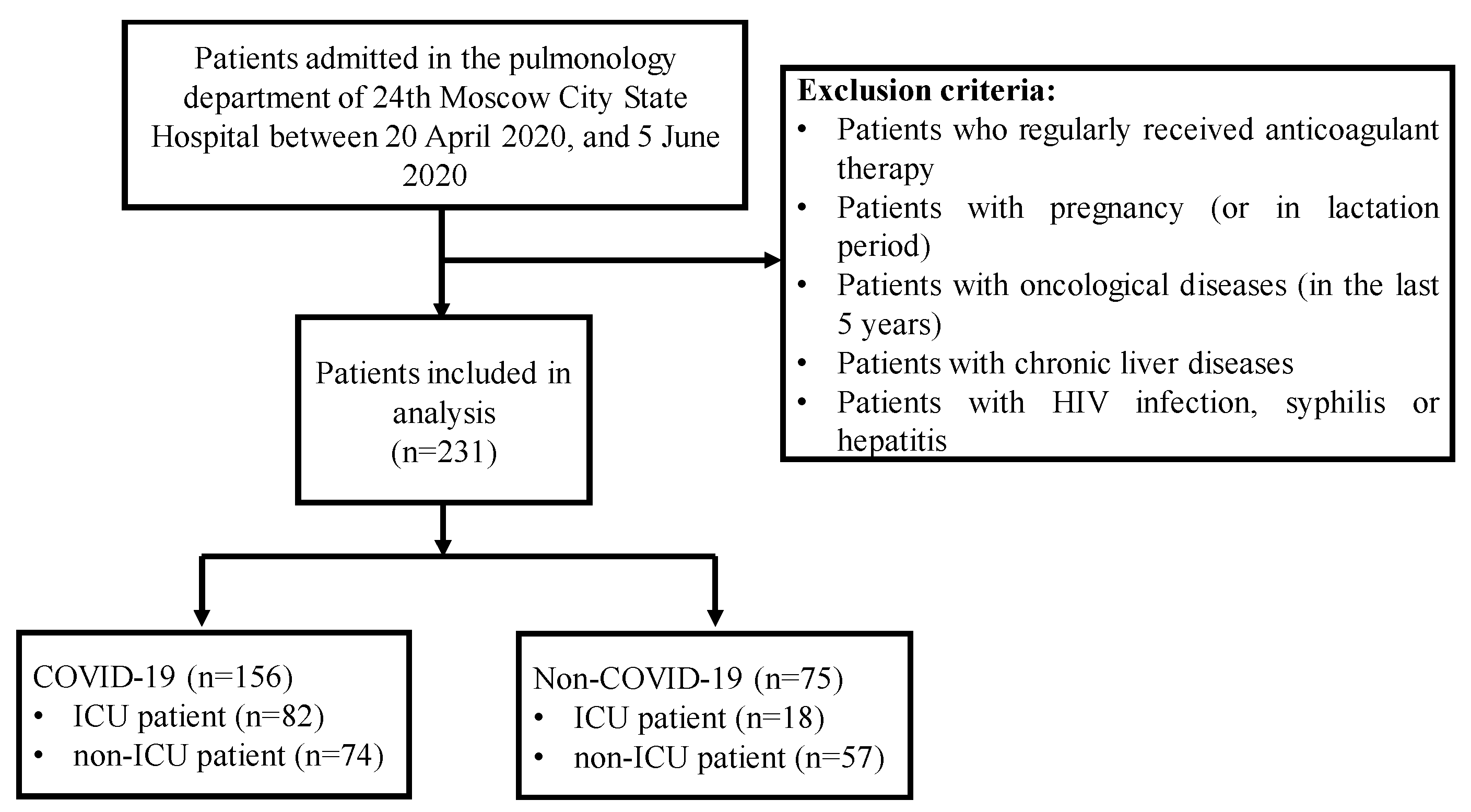
3.1. Study Population
3.2. Statistical Analysis
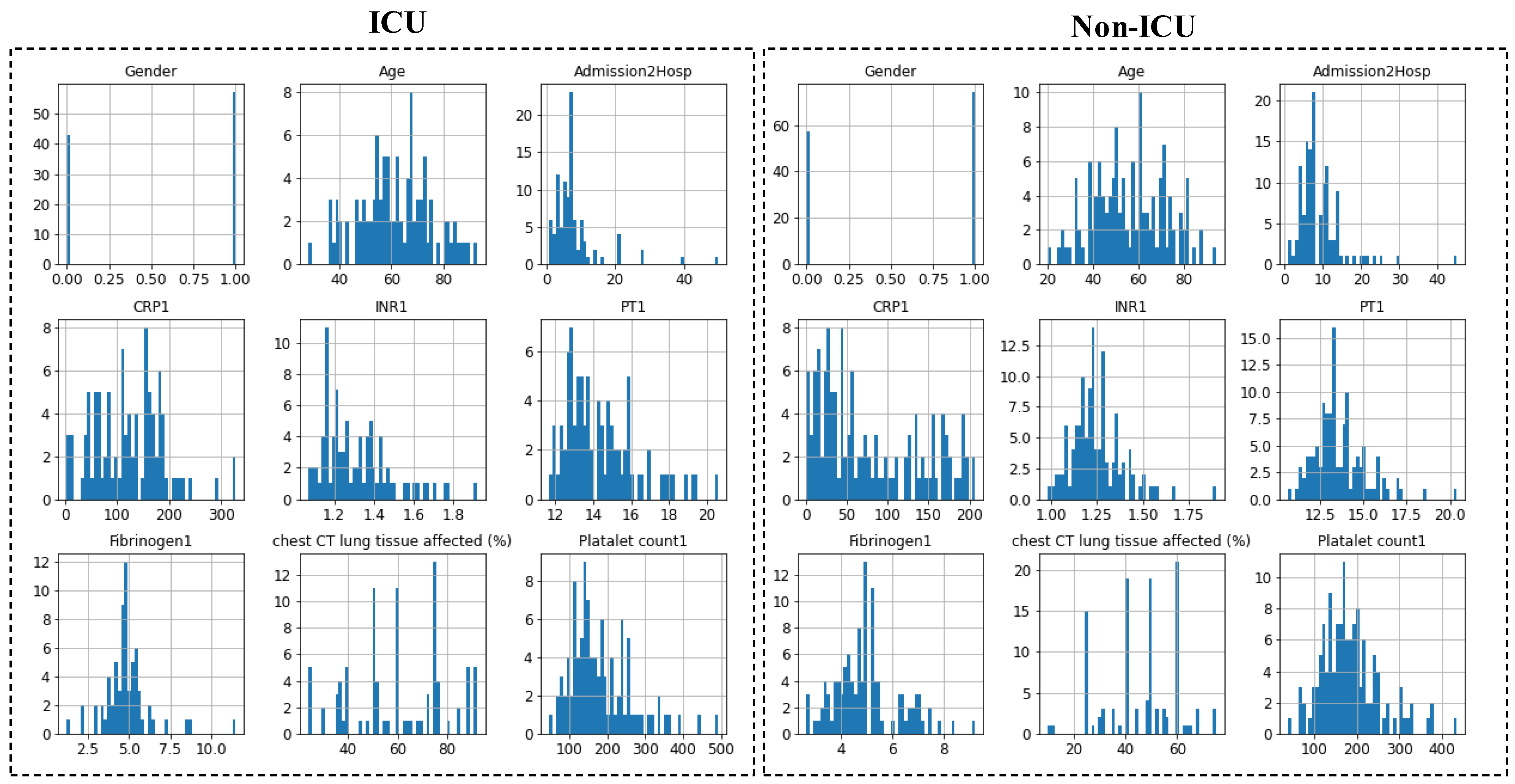
3.3. Data Preprocessing
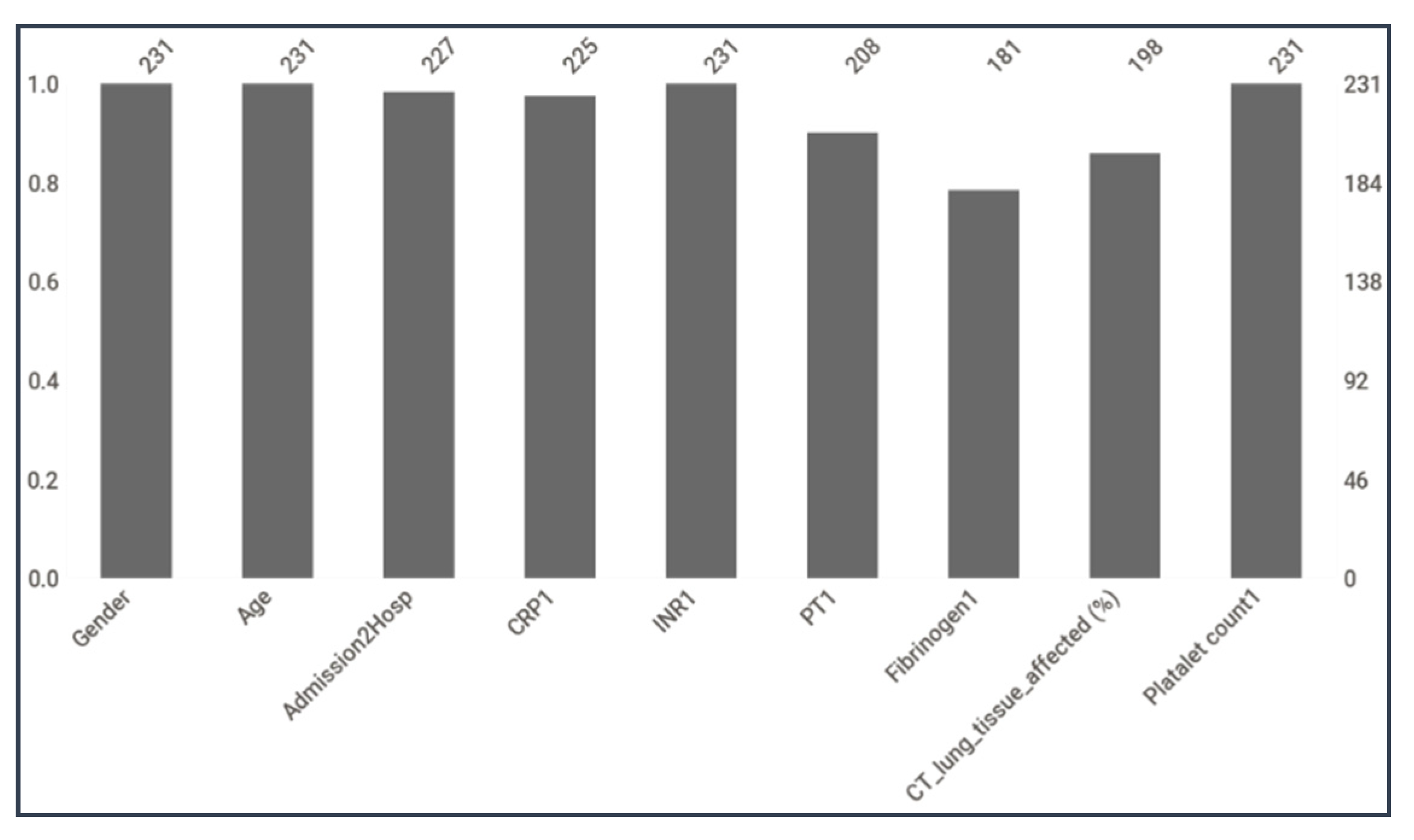
3.3.1. Missing Data Imputation
3.3.2. Balancing the Dataset
3.4. Feature Reduction
3.5. Feature Selection
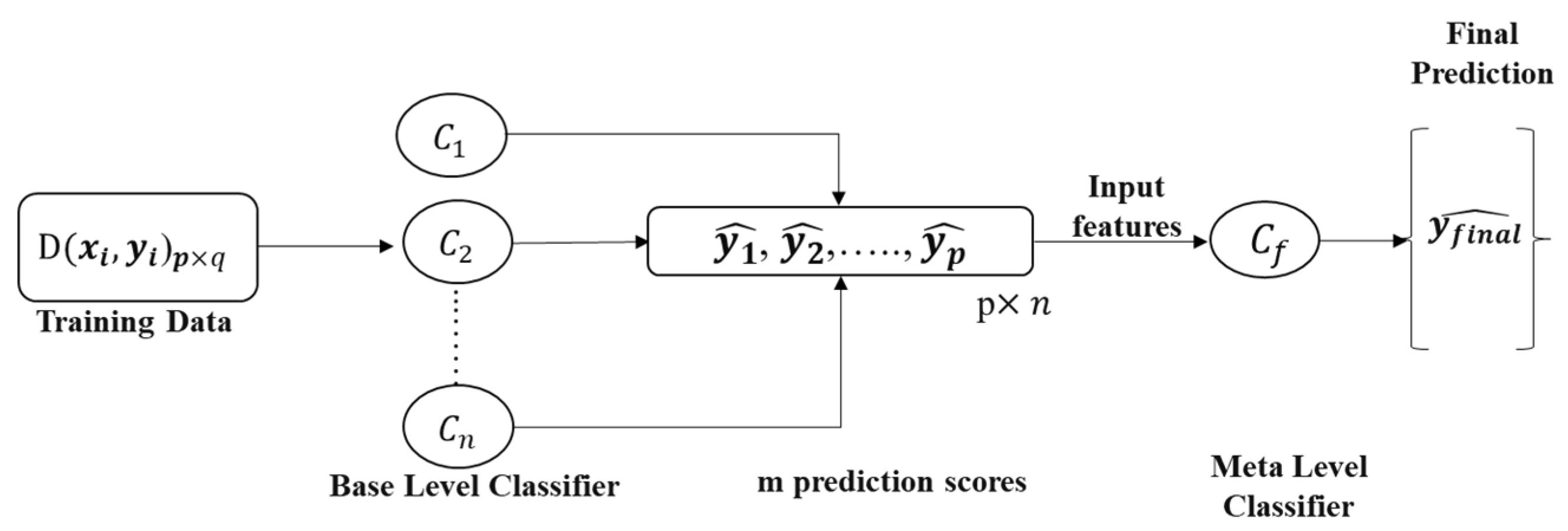
3.6. Stacking-Based Machine Learning Model
3.7. Development and Validation of Classification Model
4. Results
4.1. Characteristics and Outcomes
4.2. Best Feature Combination for Early Prediction of ICU
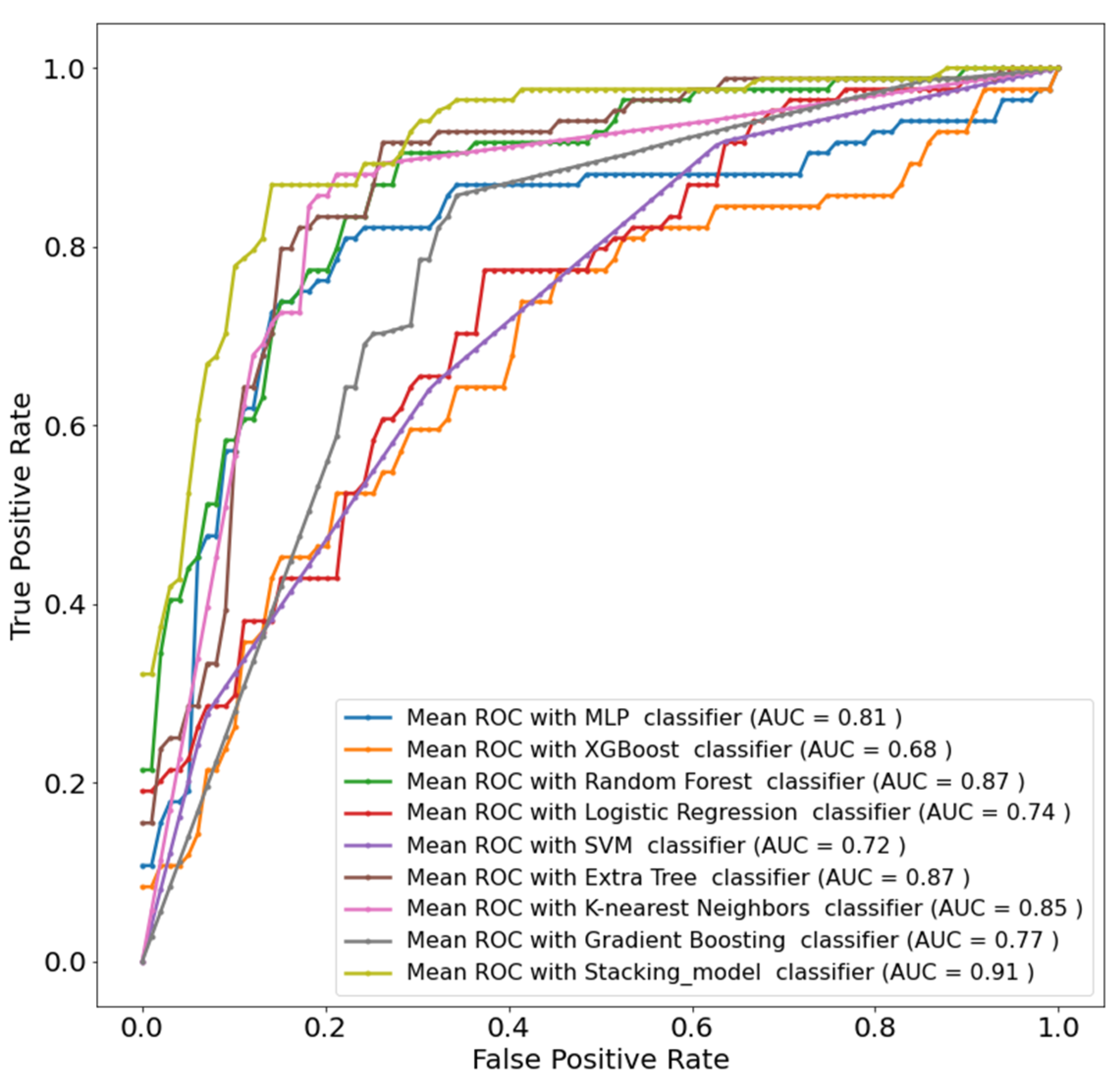
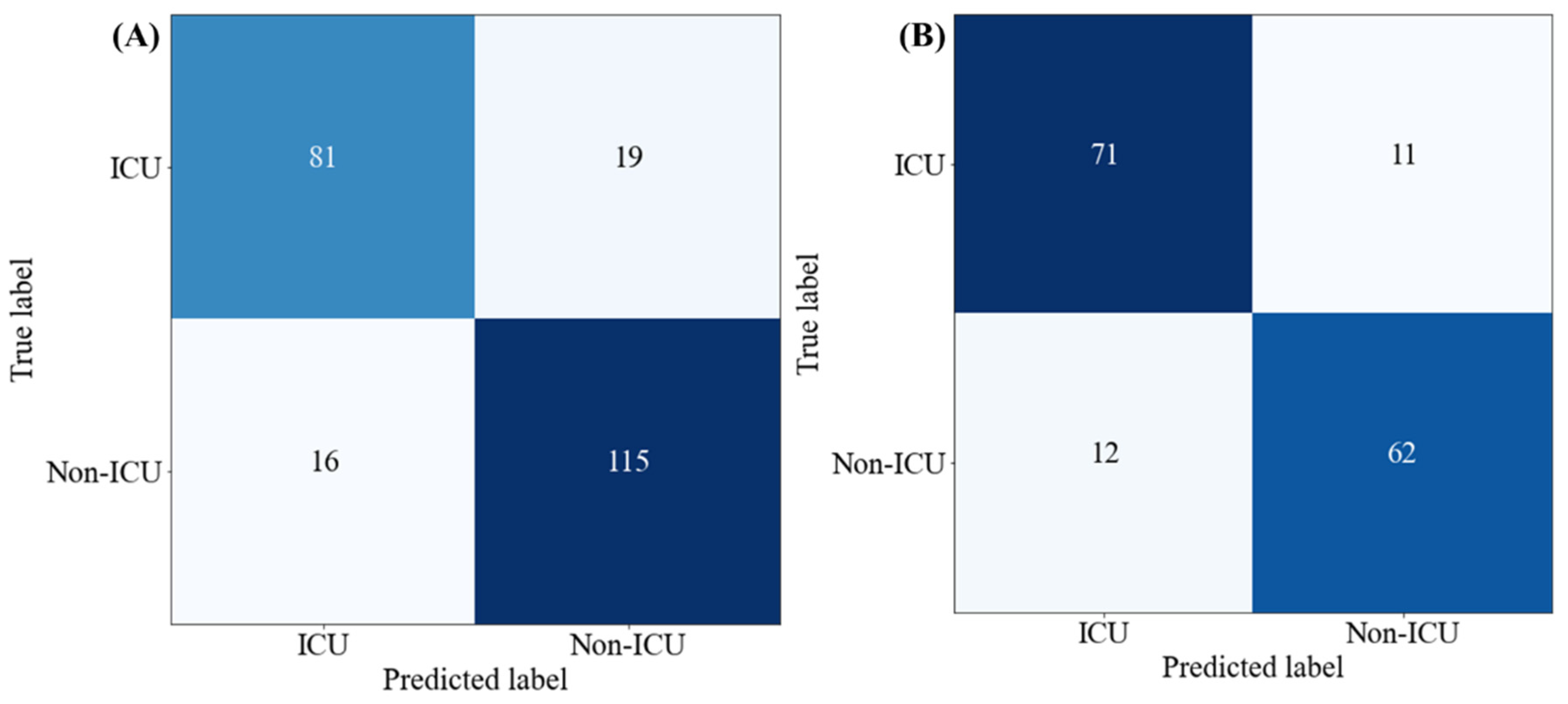
4.3. Development and Validation of the Stacking Model
4.4. Individual Feature as ICU Admission Predictor
5. Discussion
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Krit, B.; Kuvshinov, V.; Kukushkin, D.Y.; Morozova, N.; Omel’chuk, Y.A.; Revenok, T.; Sleptsov, V. The application of nanoclaster coatngs for modification of image receiving surface of thermophotoelectric energy converters. Surf. Eng. Appl. Electrochem. 2020, 56, 100–104. [Google Scholar] [CrossRef]
- COVID-19 Coronavirus Pandemic. Available online: https://www.worldometers.info/coronavirus/ (accessed on 1 July 2022).
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R. A novel coronavirus from patients with pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Liu, Y.; Yan, L.-M.; Wan, L.; Xiang, T.-X.; Le, A.; Liu, J.-M.; Peiris, M.; Poon, L.L.; Zhang, W. Viral dynamics in mild and severe cases of COVID-19. Lancet Infect. Dis. 2020, 20, 656–657. [Google Scholar] [CrossRef]
- Herring, A.A.; Ginde, A.A.; Fahimi, J.; Alter, H.J.; Maselli, J.H.; Espinola, J.A.; Sullivan, A.F.; Camargo, C.A., Jr. Increasing critical care admissions from US emergency departments, 2001–2009. Crit. Care Med. 2013, 41, 1197. [Google Scholar] [CrossRef] [PubMed]
- Halpern, N.A.; Pastores, S.M. Critical care medicine in the United States 2000–2005: An analysis of bed numbers, occupancy rates, payer mix, and costs. Crit. Care Med. 2010, 38, 65–71. [Google Scholar] [CrossRef]
- American College of Emergency Physicians. Boarding of Admitted and Intensive Care Patients in the Emergency Department. Available online: https://pubmed.ncbi.nlm.nih.gov/18655931/ (accessed on 1 February 2022).
- Goldstein, R.S. Management of the critically ill patient in the emergency department: Focus on safety issues. Crit. Care Clin. 2005, 21, 81–89. [Google Scholar] [CrossRef]
- Aslaner, M.A.; Akkaş, M.; Eroğlu, S.; Aksu, N.M.; Özmen, M.M. Admissions of critically ill patients to the ED intensive care unit. Am. J. Emerg. Med. 2015, 33, 501–505. [Google Scholar] [CrossRef]
- Frater, J.L.; Zini, G.; d’Onofrio, G.; Rogers, H.J. COVID-19 and the clinical hematology laboratory. Int. J. Lab. Hematol. 2020, 42, 11–18. [Google Scholar] [CrossRef]
- Agapakis, D.I.; Tsantilas, D.; Psarris, P.; Massa, E.V.; Kotsaftis, P.; Tziomalos, K.; Hatzitolios, A.I. Coagulation and inflammation biomarkers may help predict the severity of community-acquired pneumonia. Respirology 2010, 15, 796–803. [Google Scholar] [CrossRef]
- Giannis, D.; Ziogas, I.A.; Gianni, P. Coagulation disorders in coronavirus infected patients: COVID-19, SARS-CoV-1, MERS-CoV and lessons from the past. J. Clin. Virol. 2020, 127, 104362. [Google Scholar] [PubMed]
- Wang, D.; Yin, Y.; Hu, C.; Liu, X.; Zhang, X.; Zhou, S.; Jian, M.; Xu, H.; Prowle, J.; Hu, B. Clinical course and outcome of 107 patients infected with the novel coronavirus, SARS-CoV-2, discharged from two hospitals in Wuhan, China. Crit. Care 2020, 24, 1–9. [Google Scholar]
- Tan, C.; Huang, Y.; Shi, F.; Tan, K.; Ma, Q.; Chen, Y.; Jiang, X.; Li, X. C-reactive protein correlates with computed tomographic findings and predicts severe COVID-19 early. J. Med. Virol. 2020, 92, 856–862. [Google Scholar] [CrossRef] [PubMed]
- Luo, X.; Zhou, W.; Yan, X.; Guo, T.; Wang, B.; Xia, H.; Ye, L.; Xiong, J.; Jiang, Z.; Liu, Y. Prognostic value of C-reactive protein in patients with coronavirus 2019. Clin. Infect. Dis. 2020, 71, 2174–2179. [Google Scholar] [CrossRef] [PubMed]
- Zhai, Q.; Lin, Z.; Ge, H.; Liang, Y.; Li, N.; Ma, Q.; Ye, C. Using machine learning tools to predict outcomes for emergency department intensive care unit patients. Sci. Rep. 2020, 10, 20919. [Google Scholar]
- Hong, K.J.; Shin, S.D.; Ro, Y.S.; Song, K.J.; Singer, A.J. Development and validation of the excess mortality ratio–based Emergency Severity Index. Am. J. Emerg. Med. 2012, 30, 1491–1500. [Google Scholar] [PubMed]
- Purohit, K.; Kesarwani, A.; Kisku, D.R.; Dalui, M. COVID-19 detection on chest X-ray and ct scan images using multi-image augmented deep learning model. In Proceedings of the Seventh International Conference on Mathematics and Computing, Chongqing, China, 18–20 March 2022; pp. 395–413. [Google Scholar]
- Rahman, T.; Chowdhury, M.E.; Khandakar, A.; Mahbub, Z.B.; Hossain, M.S.A.; Alhatou, A.; Abdalla, E.; Muthiyal, S.; Islam, K.F.; Kashem, S.B.A. BIO-CXRNET: A Robust Multimodal Stacking Machine Learning Technique for Mortality Risk Prediction of COVID-19 Patients using Chest X-Ray Images and Clinical Data. arXiv 2022, arXiv:2206.07595. [Google Scholar]
- Tahir, A.M.; Qiblawey, Y.; Khandakar, A.; Rahman, T.; Khurshid, U.; Musharavati, F.; Islam, M.; Kiranyaz, S.; Al-Maadeed, S.; Chowdhury, M.E. Deep learning for reliable classification of COVID-19, MERS, and SARS from chest X-ray images. Cogn. Comput. 2022, 1–21. [Google Scholar] [CrossRef]
- Borghesi, A.; Golemi, S.; Scrimieri, A.; Nicosia, C.M.C.; Zigliani, A.; Farina, D.; Maroldi, R. Chest X-ray versus chest computed tomography for outcome prediction in hospitalized patients with COVID-19. La Radiol. Med. 2022, 127, 305–308. [Google Scholar] [CrossRef]
- Rahman, T.; Khandakar, A.; Qiblawey, Y.; Tahir, A.; Kiranyaz, S.; Kashem, S.B.A.; Islam, M.T.; Al Maadeed, S.; Zughaier, S.M.; Khan, M.S. Exploring the effect of image enhancement techniques on COVID-19 detection using chest X-ray images. Comput. Biol. Med. 2021, 132, 104319. [Google Scholar] [CrossRef]
- Qiblawey, Y.; Tahir, A.; Chowdhury, M.; Khandakar, A.; Kiranyaz, S.; Rahman, T.; Ibtehaz, N.; Mahmud, S.; Al Maadeed, S.; Musharavati, F. Detection and severity classification of COVID-19 in CT images using deep learning. Diagnostics 2021, 11, 893. [Google Scholar] [CrossRef] [PubMed]
- Poly, T.N.; Islam, M.M.; Li, Y.-C.J.; Alsinglawi, B.; Hsu, M.-H.; Jian, W.S.; Yang, H.-C. Application of artificial intelligence for screening covid-19 patients using digital images: Meta-analysis. JMIR Med. Inform. 2021, 9, e21394. [Google Scholar] [CrossRef] [PubMed]
- Feng, C.; Wang, L.; Chen, X.; Zhai, Y.; Zhu, F.; Chen, H.; Wang, Y.; Su, X.; Huang, S.; Tian, L. A novel artificial intelligence-assisted triage tool to aid in the diagnosis of suspected COVID-19 pneumonia cases in fever clinics. Ann. Transl. Med. 2021, 9, 201. [Google Scholar] [CrossRef] [PubMed]
- Calvet, J.; Berenguer-Llergo, A.; Gay, M.; Massanella, M.; Domingo, P.; Llop, M.; Sánchez-Jiménez, E.; Arévalo, M.; Carrillo, J.; Albiñana, N. Biomarker candidates for progression and clinical management of COVID-19 associated pneumonia at time of admission. Sci. Rep. 2022, 12, 640. [Google Scholar] [CrossRef]
- Halasz, G.; Sperti, M.; Villani, M.; Michelucci, U.; Agostoni, P.; Biagi, A.; Rossi, L.; Botti, A.; Mari, C.; Maccarini, M. A machine learning approach for mortality prediction in COVID-19 pneumonia: Development and evaluation of the Piacenza score. J. Med. Internet Res. 2021, 23, e29058. [Google Scholar]
- Alsinglawi, B.; Alnajjar, F.; Mubin, O.; Novoa, M.; Karajeh, O.; Darwish, O. Benchmarking predictive models in electronic health records: Sepsis length of stay prediction. In Proceedings of the International Conference on Advanced Information Networking and Applications, Caserta, Italy, 15–17 April 2020; pp. 258–267. [Google Scholar]
- Zou, Y.; Guo, H.; Zhang, Y.; Zhang, Z.; Liu, Y.; Wang, J.; Lu, H.; Qian, Z. Analysis of coagulation parameters in patients with COVID-19 in Shanghai, China. Biosci. Trends 2020, 14, 285–289. [Google Scholar] [CrossRef]
- Weng, Z.; Chen, Q.; Li, S.; Li, H.; Zhang, Q.; Lu, S.; Wu, L.; Xiong, L.; Mi, B.; Liu, D. ANDC: An early warning score to predict mortality risk for patients with coronavirus disease 2019. J. Transl. Med. 2020, 18, 328. [Google Scholar] [CrossRef]
- Gong, J.; Ou, J.; Qiu, X.; Jie, Y.; Chen, Y.; Yuan, L.; Cao, J.; Tan, M.; Xu, W.; Zheng, F. A tool for early prediction of severe coronavirus disease 2019 (COVID-19): A multicenter study using the risk nomogram in Wuhan and Guangdong, China. Clin. Infect. Dis. 2020, 71, 833–840. [Google Scholar] [CrossRef]
- Alabbad, D.A.; Almuhaideb, A.M.; Alsunaidi, S.J.; Alqudaihi, K.S.; Alamoudi, F.A.; Alhobaishi, M.K.; Alaqeel, N.A.; Alshahrani, M.S. Machine learning model for predicting the length of stay in the intensive care unit for COVID-19 patients in the eastern province of Saudi Arabia. Inform. Med. Unlocked 2022, 30, 100937. [Google Scholar] [CrossRef] [PubMed]
- Al Youha, S.; Doi, S.A.; Jamal, M.H.; Almazeedi, S.; Al Haddad, M.; AlSeaidan, M.; Al-Muhaini, A.; Al-Ghimlas, F.; Al-Sabah, S. Validation of the Kuwait Progression Indicator Score for predicting progression of severity in COVID19. MedRxiv 2020. [Google Scholar] [CrossRef]
- Ai, T.; Yang, Z.; Hou, H.; Zhan, C.; Chen, C.; Lv, W.; Tao, Q.; Sun, Z.; Xia, L. Correlation of chest CT and RT-PCR testing in coronavirus disease 2019 (COVID-19) in China: A report of 1014 cases. Radiology 2020, 296, E32–E40. [Google Scholar] [CrossRef]
- Chowdhury, M.E.; Rahman, T.; Khandakar, A.; Al-Madeed, S.; Zughaier, S.M.; Hassen, H.; Islam, M.T. An early warning tool for predicting mortality risk of COVID-19 patients using machine learning. Cogn. Comput. 2021, 1–16. [Google Scholar] [CrossRef]
- Rahman, T.; Al-Ishaq, F.A.; Al-Mohannadi, F.S.; Mubarak, R.S.; Al-Hitmi, M.H.; Islam, K.R.; Khandakar, A.; Hssain, A.A.; Al-Madeed, S.; Zughaier, S.M. Mortality prediction utilizing blood biomarkers to predict the severity of COVID-19 using machine learning technique. Diagnostics 2021, 11, 1582. [Google Scholar] [PubMed]
- Lorenzoni, G.; Sella, N.; Boscolo, A.; Azzolina, D.; Bartolotta, P.; Pasin, L.; Pettenuzzo, T.; De Cassai, A.; Baratto, F.; Toffoletto, F. COVID-19 ICU mortality prediction: A machine learning approach using SuperLearner algorithm. J. Anesth. Analg. Crit. Care 2021, 1, 3. [Google Scholar]
- Magunia, H.; Lederer, S.; Verbuecheln, R.; Gilot, B.J.; Koeppen, M.; Haeberle, H.A.; Mirakaj, V.; Hofmann, P.; Marx, G.; Bickenbach, J. Machine learning identifies ICU outcome predictors in a multicenter COVID-19 cohort. Crit. Care 2021, 25, 295. [Google Scholar]
- Liang, W.; Yao, J.; Chen, A.; Lv, Q.; Zanin, M.; Liu, J.; Wong, S.; Li, Y.; Lu, J.; Liang, H. Early triage of critically ill COVID-19 patients using deep learning. Nat. Commun. 2020, 11, 3543. [Google Scholar]
- Wang, C.; Deng, R.; Gou, L.; Fu, Z.; Zhang, X.; Shao, F.; Wang, G.; Fu, W.; Xiao, J.; Ding, X. Preliminary study to identify severe from moderate cases of COVID-19 using combined hematology parameters. Ann. Transl. Med. 2020, 8, 593. [Google Scholar] [PubMed]
- Cai, Y.-Q.; Zeng, H.-Q.; Zhang, X.-B.; Wei, X.-J.; Hu, L.; Zhang, Z.-Y.; Ming, Q.; Peng, Q.-P.; Chen, L.-D. Prognostic value of neutrophil-to-lymphocyte ratio, lactate dehydrogenase, D-Dimer and CT score in patients with COVID-19. Aging 2020, 13, 20896–20905. [Google Scholar] [CrossRef] [PubMed]
- McRae, M.P.; Simmons, G.W.; Christodoulides, N.J.; Lu, Z.; Kang, S.K.; Fenyo, D.; Alcorn, T.; Dapkins, I.P.; Sharif, I.; Vurmaz, D. Clinical decision support tool and rapid point-of-care platform for determining disease severity in patients with COVID-19. Lab Chip 2020, 20, 2075–2085. [Google Scholar]
- Wang, G.; Wu, C.; Zhang, Q.; Wu, F.; Yu, B.; Lv, J.; Li, Y.; Li, T.; Zhang, S.; Wu, C. C-reactive protein level may predict the risk of COVID-19 aggravation. In Open Forum Infectious Diseases; Oxford University Press: Oxford, MI, USA, 2020; p. ofaa153. [Google Scholar]
- Hegde, H.; Shimpi, N.; Panny, A.; Glurich, I.; Christie, P.; Acharya, A. MICE vs. PPCA: Missing data imputation in healthcare. Inform. Med. Unlocked 2019, 17, 100275. [Google Scholar]
- Baranovskii, D.S.; Klabukov, I.D.; Krasilnikova, O.A.; Nikogosov, D.A.; Polekhina, N.V.; Baranovskaia, D.R.; Laberko, L.A. Prolonged prothrombin time as an early prognostic indicator of severe acute respiratory distress syndrome in patients with COVID-19 related pneumonia. Curr. Med. Res. Opin. 2021, 37, 21–25. [Google Scholar] [CrossRef]
- Rahman, T.; Khandakar, A.; Hoque, M.E.; Ibtehaz, N.; Kashem, S.B.; Masud, R.; Shampa, L.; Hasan, M.M.; Islam, M.T.; Al-Maadeed, S. Development and Validation of an Early Scoring System for Prediction of Disease Severity in COVID-19 Using Complete Blood Count Parameters. IEEE Access 2021, 9, 120422–120441. [Google Scholar] [CrossRef]
- Stevens, J.R.; Suyundikov, A.; Slattery, M.L. Accounting for missing data in clinical research. JAMA 2016, 315, 517–518. [Google Scholar] [CrossRef]
- Speiser, J.L.; Miller, M.E.; Tooze, J.; Ip, E. A comparison of random forest variable selection methods for classification prediction modeling. Expert Syst. Appl. 2019, 134, 93–101. [Google Scholar] [CrossRef]
- Beretta, L.; Santaniello, A. Nearest neighbor imputation algorithms: A critical evaluation. BMC Med. Inform. Decis. Mak. 2016, 16, 74. [Google Scholar] [CrossRef]
- Chawla, N.V.; Bowyer, K.W.; Hall, L.O.; Kegelmeyer, W.P. SMOTE: Synthetic minority over-sampling technique. J. Artif. Intell. Res. 2002, 16, 321–357. [Google Scholar] [CrossRef]
- Khandakar, A.; Chowdhury, M.E.; Reaz, M.B.I.; Ali, S.H.M.; Hasan, M.A.; Kiranyaz, S.; Rahman, T.; Alfkey, R.; Bakar, A.A.A.; Malik, R.A. A machine learning model for early detection of diabetic foot using thermogram images. Comput. Biol. Med. 2021, 137, 104838. [Google Scholar]
- Khandakar, A.; Chowdhury, M.E.; Reaz, M.B.I.; Ali, S.H.M.; Kiranyaz, S.; Rahman, T.; Chowdhury, M.H.; Ayari, M.A.; Alfkey, R.; Bakar, A.A.A. A Novel Machine Learning Approach for Severity Classification of Diabetic Foot Complications Using Thermogram Images. Sensors 2022, 22, 4249. [Google Scholar] [PubMed]
- Tallarida, R.J.; Murray, R.B. Chi-square test. In Manual of Pharmacologic Calculations; Springer: Berlin/Heidelberg, Germany, 1987; pp. 140–142. [Google Scholar]
- Saidi, R.; Bouaguel, W.; Essoussi, N. Hybrid feature selection method based on the genetic algorithm and pearson correlation coefficient. In Machine Learning Paradigms: Theory and Application; Springer: Berlin/Heidelberg, Germany, 2019; pp. 3–24. [Google Scholar]
- Lin, X.; Yang, F.; Zhou, L.; Yin, P.; Kong, H.; Xing, W.; Lu, X.; Jia, L.; Wang, Q.; Xu, G. A support vector machine-recursive feature elimination feature selection method based on artificial contrast variables and mutual information. J. Chromatogr. B 2012, 910, 149–155. [Google Scholar] [CrossRef] [PubMed]
- Hosseini, S.; Khandakar, A.; Chowdhury, M.E.; Ayari, M.A.; Rahman, T.; Chowdhury, M.H.; Vaferi, B. Novel and robust machine learning approach for estimating the fouling factor in heat exchangers. Energy Rep. 2022, 8, 8767–8776. [Google Scholar]
- Romero Starke, K.; Petereit-Haack, G.; Schubert, M.; Kämpf, D.; Schliebner, A.; Hegewald, J.; Seidler, A. The age-related risk of severe outcomes due to COVID-19 infection: A rapid review, meta-analysis, and meta-regression. Int. J. Environ. Res. Public Health 2020, 17, 5974. [Google Scholar] [CrossRef] [PubMed]
- Booth, C.M.; Matukas, L.M.; Tomlinson, G.A.; Rachlis, A.R.; Rose, D.B.; Dwosh, H.A.; Walmsley, S.L.; Mazzulli, T.; Avendano, M.; Derkach, P. Clinical features and short-term outcomes of 144 patients with SARS in the greater Toronto area. JAMA 2003, 289, 2801–2809. [Google Scholar] [CrossRef] [PubMed]
- Scudiero, F.; Silverio, A.; Di Maio, M.; Russo, V.; Citro, R.; Personeni, D.; Cafro, A.; D’Andrea, A.; Attena, E.; Pezzullo, S. Pulmonary embolism in COVID-19 patients: Prevalence, predictors and clinical outcome. Thromb. Res. 2021, 198, 34–39. [Google Scholar] [CrossRef]
- Iba, T.; Levi, M.; Levy, J.H. Sepsis-induced coagulopathy and disseminated intravascular coagulation. In Seminars in Thrombosis and Hemostasis; Thieme Medical: Stuttgart, Germany, 2020; pp. 089–095. [Google Scholar]
- Thachil, J.; Tang, N.; Gando, S.; Falanga, A.; Cattaneo, M.; Levi, M.; Clark, C.; Iba, T. ISTH interim guidance on recognition and management of coagulopathy in COVID-19. J. Thromb. Haemost. 2020, 18, 1023–1026. [Google Scholar] [CrossRef]
- Marnell, L.; Mold, C.; Clos, T.W.D. C-reactive protein: Ligands, receptors and role in inflammation. Clin. Immunol. 2005, 117, 104–111. [Google Scholar] [CrossRef] [PubMed]
- Young, B.; Gleeson, M.; Cripps, A.W. C-reactive protein: A critical review. Pathology 1991, 23, 118–124. [Google Scholar] [CrossRef]
- Wang, Y.; Dong, C.; Hu, Y.; Li, C.; Ren, Q.; Zhang, X.; Shi, H.; Zhou, M. Temporal changes of CT findings in 90 patients with COVID-19 pneumonia: A longitudinal study. Radiology 2020, 296, E55–E64. [Google Scholar] [CrossRef]
- Francone, M.; Iafrate, F.; Masci, G.M.; Coco, S.; Cilia, F.; Manganaro, L.; Panebianco, V.; Andreoli, C.; Colaiacomo, M.C.; Zingaropoli, M.A. Chest CT score in COVID-19 patients: Correlation with disease severity and short-term prognosis. Eur. Radiol. 2020, 30, 6808–6817. [Google Scholar] [CrossRef]
- Simpson, S.; Kay, F.U.; Abbara, S.; Bhalla, S.; Chung, J.H.; Chung, M.; Henry, T.S.; Kanne, J.P.; Kligerman, S.; Ko, J.P. Radiological society of north America expert consensus document on reporting chest CT findings related to COVID-19: Endorsed by the society of thoracic Radiology, the American college of Radiology, and RSNA. Radiol. Cardiothorac. Imaging 2020, 2, e200152. [Google Scholar] [CrossRef]
- Sayeed, S.; Belqees, Y.F.; Aslam, S.; Masood, L.; Saeed, R. CT Chest Severity Score for COVID 19 Pneumonia: A Quantitative Imaging Tool for Severity Assessment of Disease. J. Coll. Physicians Surg.—Pak. JCPSP 2021, 30, 388–392. [Google Scholar]
- Mruk, B.; Plucińska, D.; Walecki, J.; Półtorak-Szymczak, G.; Sklinda, K. Chest Computed Tomography (CT) Severity Scales in COVID-19 Disease: A Validation Study. Med. Sci. Monit. Int. Med. J. Exp. Clin. Res. 2021, 27, e931283-1. [Google Scholar] [CrossRef] [PubMed]




| Item | ICU | Non-ICU | Total | Method | χ2 = 17.5 | p Value |
|---|---|---|---|---|---|---|
| Gender | Chi-square test | χ2 = 17.5 | <0.05 * | |||
| Male (%) | 57 (57%) | 74 (56.5%) | 131 (57%) | |||
| Female (%) | 43 (43%) | 57 (43.5%) | 100 (43%) | |||
| Age (years) | Rank-sum test | Z = −6.2 | <0.05 * | |||
| N(missing) | 100 (0) | 131 (0) | 231 (0) | |||
| Mean ± SD | 61.6 ± 13.8 | 55.9 ± 15.7 | 58.4 ± 15.1 | |||
| Q1, Q3 | 53, 70.2 | 44, 68.5 | 48.0, 69.0 | |||
| Min, Max | 28, 93 | 20, 94 | 20, 94 | |||
| The time between the disease and admission to hospital (Admission 2Hospital) (Days) | Rank-sum test | Z = −6.2 | <0.05 * | |||
| N(missing) | 99 (1) | 128 (3) | 227 (4) | |||
| Mean ± SD | 7.9 ± 7.45 | 9.1 ± 5.68 | 8.6 ± 6.53 | |||
| Q1, Q3 | 4, 8 | 6, 11 | 5.0, 10.0 | |||
| Min, Max | 1, 50 | 1, 45 | 1, 50 | |||
| C-reactive protein 1(CRP1) (mg/L) | Rank-sum test | Z = −4.34 | <0.05 * | |||
| N(missing) | 97 (3) | 128 (3) | 225 (6) | |||
| Mean ± SD | 123 ± 67.1 | 78 ± 61.7 | 97 ± 67.9 | |||
| Q1, Q3 | 64, 166 | 26, 134 | 40, 157 | |||
| Min, Max | 4, 328 | 1, 207 | 1, 328 | |||
| International normalized ratio (INR) | Rank-sum test | Z = 6.53 | 0.78 | |||
| N(missing) | 86 (14) | 122 (9) | 208 (23) | |||
| Mean ± SD | 1.32 ±0.18 | 1.25 ± 0.14 | 1.28 ± 0.16 | |||
| Q1, Q3 | 1.17, 1.4 | 1.16, 1.3 | 1.17, 1.36 | |||
| Min, max | 1.07,1.92 | 0.98, 1.9 | 0.98, 1.92 | |||
| Prothrombin time 1 (PT1) (s) | Rank-sum test | Z = 3.27 | <0.05 * | |||
| N(missing) | 86 (14) | 122 (9) | 208 (23) | |||
| Mean ± SD | 14.39 ± 1.91 | 13.63 ± 1.52 | 13.95 ± 1.73 | |||
| Q1, Q3 | 12.9, 15.3 | 12.7, 14.2 | 12.8, 14.9 | |||
| Min, max | 11.7, 20.6 | 10.7, 20.4 | 10.7, 20.6 | |||
| Fibrinogen 1 (mg/L) | Rank-sum test | Z = −5.89 | <0.05 * | |||
| N(missing) | 68 (32) | 113 (18) | 181 (50) | |||
| Mean ± SD | 4.9 ± 1.44 | 4.99 ± 1.25 | 4.9 ± 1.32 | |||
| Q1, Q3 | 4.2, 5.3 | 4.18, 5.45 | 4.2, 5.4 | |||
| Min, max | 1.2, 11.5 | 2.68, 9.21 | 1.2, 11.5 | |||
| Chest CT lung tissue affected (%) | Rank-sum test | Z = −1.11 | <0.05 * | |||
| N(missing) | 88 (12) | 110 (21) | 198 (33) | |||
| Mean ± SD | 59.9 ± 19 | 46.1 ± 14.1 | 52.2 ± 17.8 | |||
| Q1, Q3 | 49.5, 75 | 40, 60 | 40, 60 | |||
| Min, max | 24, 92 | 10, 75 | 10, 92 | |||
| Platelet count 1 (/L) | Rank-sum test | Z = 4.74 | 0.44 | |||
| N(missing) | 100 (0) | 131 (0) | 231 (0) | |||
| Mean ± SD | 182 ± 83.2 | 183 ± 68.8 | 183 ± 75.2 | |||
| Q1, Q3 | 126, 233 | 138, 216 | 129, 219 | |||
| Min, max | 47, 493 | 38, 436 | 38, 493 | |||
| Outcome | 100 (43%) | 131 (57%) | 231 (100%) |
| Features | Pearson Correlation Coefficient | Chi-Square Test | Recursive Feature Elimination | Total |
|---|---|---|---|---|
| CRP |  |  |  | 3 |
| Chest CT lung tissue affected (%) |  |  |  | 3 |
| Age |  |  |  | 3 |
| Admission2Hospital |  |  |  | 3 |
| Fibrinogen |  |  |  | 3 |
| Platelet Count |  |  | 2 | |
| Gender |  |  | 2 | |
| PT |  |  | 2 | |
| INR |  |  | 2 |
| Classifier | Overall | Weighted with 95% CI | |||
|---|---|---|---|---|---|
| Accuracy | Precision | Sensitivity | Specificity | F1-Score | |
| Support Vector Machine (SVM) | 61.21 ± 1.99 | 63.17 ± 1.97 | 61.21 ± 1.99 | 63.09 ± 1.97 | 61.29 ± 1.99 |
| XGBoost (XGB) | 65.52 ± 1.94 | 65.92 ± 1.93 | 65.52 ± 1.94 | 65.15 ± 1.94 | 65.64 ± 1.94 |
| MLP | 71.12 ± 1.85 | 70.98 ± 1.85 | 71.12 ± 1.85 | 69.4 ± 1.88 | 71.02 ± 1.85 |
| Logistic Regression (LR) | 71.12 ± 1.85 | 70.92 ± 1.85 | 71.12 ± 1.85 | 68.67 ± 1.89 | 70.86 ± 1.85 |
| K-Nearest Neighbors (KNN) | 71.55 ± 1.84 | 71.55 ± 1.84 | 71.55 ± 1.84 | 70.45 ± 1.86 | 71.55 ± 1.84 |
| Extra Trees (ET) | 79.74 ± 1.64 | 79.68 ± 1.64 | 79.74 ± 1.64 | 78.11 ± 1.69 | 79.64 ± 1.64 |
| Gradient Boosting (GB) | 81.03 ± 1.6 | 80.98 ± 1.6 | 81.04 ± 1.6 | 79.81 ± 1.64 | 80.98 ± 1.6 |
| Random Forest (RF) | 82.33 ± 1.56 | 82.33 ± 1.56 | 82.33 ± 1.56 | 80.55 ± 1.61 | 82.2 ± 1.56 |
| Stacking model (RF+ GB+ ET) | 84.48 ± 1.48 | 84.45 ± 1.48 | 84.48 ± 1.48 | 83.64 ± 1.51 | 84.47 ± 1.48 |
| Classifier | Overall | Weighted with 95% CI | |||
|---|---|---|---|---|---|
| Accuracy | Precision | Sensitivity | Specificity | F1-Score | |
| XGBoost (XGB) | 67.52 ± 2.32 | 67.86 ± 2.32 | 67.52 ± 2.32 | 67.82 ± 2.32 | 67.56 ± 2.32 |
| Support Vector Machine (SVM) | 71.97 ± 2.23 | 72.4 ± 2.22 | 71.97 ± 2.23 | 72.42 ± 2.22 | 72 ± 2.23 |
| MLP | 77.71 ± 2.07 | 77.92 ± 2.06 | 77.71 ± 2.07 | 77.93 ± 2.06 | 77.73 ± 2.06 |
| Gradient Boosting (GB) | 78.98 ± 2.02 | 79.34 ± 2.01 | 78.98 ± 2.02 | 79.4 ± 2.01 | 79 ± 2.02 |
| Logistic Regression (LR) | 80.25 ± 1.98 | 80.25 ± 1.98 | 80.25 ± 1.98 | 79.97 ± 1.99 | 80.25 ± 1.98 |
| K-Nearest Neighbors (KNN) | 82.17 ± 1.9 | 82.26 ± 1.9 | 82.16 ± 1.9 | 81.45 ± 1.93 | 82.09 ± 1.9 |
| Extra Tree (ET) | 82.80 ± 1.87 | 82.90 ± 1.87 | 82.80 ± 1.87 | 82.90 ± 1.87 | 82.82 ± 1.87 |
| Random Forest (RF) | 83.44 ± 1.84 | 83.49 ± 1.84 | 83.44 ± 1.84 | 83.45 ± 1.84 | 83.45 ± 1.84 |
| Stacking model (RF + ET+ KNN) | 85.35 ± 1.75 | 85.34 ± 1.76 | 85.35 ± 1.75 | 85.11 ± 1.77 | 85.34 ± 1.76 |
| (A) | |||||
| 95% Confidence Interval Results | |||||
| Feature | Overall Accuracy | Weighted Precision | Weighted Recall | Weighted Specificity | Weighted F1-Score |
| CRP | 71.11 ± 1.85 | 70.56 ± 1.86 | 71.11 ± 1.85 | 71.11 ± 1.85 | 70.58 ± 1.86 |
| Chest CT lung tissue affected (%) | 74.43 ± 1.78 | 77.65 ± 1.7 | 74.43 ± 1.78 | 74.43 ± 1.78 | 71.75 ± 1.84 |
| Age | 61.01 ± 1.99 | 36.46 ± 1.96 | 61.01 ± 1.99 | 61.01 ± 1.99 | 45.34 ± 2.03 |
| Admission2Hospital | 62.41 ± 1.98 | 37.86 ± 1.98 | 62.41 ± 1.98 | 62.41 ± 1.98 | 46.74 ± 2.03 |
| Fibrinogen | 65.31 ± 1.98 | 65.62 ± 1.97 | 65.31 ± 1.98 | 65.31 ± 1.98 | 65.47 ± 2.04 |
| Platelet Count | 67.01 ± 1.92 | 42.46 ± 2.02 | 67.01 ± 1.92 | 67.01 ± 1.92 | 51.34 ± 2.04 |
| Gender | 64.21 ± 1.95 | 61.51 ± 1.98 | 64.21 ± 1.95 | 64.21 ± 1.95 | 62.65 ± 2.04 |
| PT | 57.91 ± 1.9 | 43.36 ± 2.02 | 57.91 ± 1.9 | 57.91 ± 1.9 | 52.24 ± 2.04 |
| INR | 59.81 ± 2 | 35.26 ± 1.95 | 59.81 ± 2 | 59.81 ± 2 | 44.14 ± 2.02 |
| (B) | |||||
| 95% Confidence Interval Results | |||||
| Feature | Overall Accuracy | Weighted Precision | Weighted Recall | Weighted Specificity | Weighted F1-Score |
| CRP | 64.29 ± 2.38 | 64.6 ± 2.37 | 64.29 ± 2.38 | 64.29 ± 2.38 | 64.29 ± 2.38 |
| Chest CT lung tissue affected (%) | 68.77 ± 2.3 | 71.79 ± 2.23 | 68.77 ± 2.3 | 68.77 ± 2.3 | 67.88 ± 2.32 |
| Age | 55.95 ± 2.46 | 56.51 ± 2.46 | 55.95 ± 2.46 | 55.95 ± 2.46 | 55.75 ± 2.46 |
| Admission2Hospital | 73.9 ± 2.18 | 73.99 ± 2.18 | 73.9 ± 2.18 | 73.9 ± 2.18 | 73.92 ± 2.18 |
| Fibrinogen | 57.24 ± 2.46 | 57.15 ± 2.46 | 57.24 ± 2.46 | 57.24 ± 2.46 | 57.17 ± 2.46 |
| Platelet Count | 47.62 ± 2.48 | 47.96 ± 2.48 | 47.62 ± 2.48 | 47.62 ± 2.48 | 47.2 ± 2.48 |
| Gender | 52.75 ± 2.48 | 53.19 ± 2.48 | 52.75 ± 2.48 | 52.75 ± 2.48 | 52.57 ± 2.48 |
| PT | 51.47 ± 2.48 | 50.64 ± 2.48 | 51.47 ± 2.48 | 51.47 ± 2.48 | 50.43 ± 2.48 |
| INR | 53.39 ± 2.48 | 53.68 ± 2.47 | 53.39 ± 2.48 | 53.39 ± 2.48 | 53.37 ± 2.48 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Islam, K.R.; Kumar, J.; Tan, T.L.; Reaz, M.B.I.; Rahman, T.; Khandakar, A.; Abbas, T.; Hossain, M.S.A.; Zughaier, S.M.; Chowdhury, M.E.H. Prognostic Model of ICU Admission Risk in Patients with COVID-19 Infection Using Machine Learning. Diagnostics 2022, 12, 2144. https://doi.org/10.3390/diagnostics12092144
Islam KR, Kumar J, Tan TL, Reaz MBI, Rahman T, Khandakar A, Abbas T, Hossain MSA, Zughaier SM, Chowdhury MEH. Prognostic Model of ICU Admission Risk in Patients with COVID-19 Infection Using Machine Learning. Diagnostics. 2022; 12(9):2144. https://doi.org/10.3390/diagnostics12092144
Chicago/Turabian StyleIslam, Khandaker Reajul, Jaya Kumar, Toh Leong Tan, Mamun Bin Ibne Reaz, Tawsifur Rahman, Amith Khandakar, Tariq Abbas, Md. Sakib Abrar Hossain, Susu M. Zughaier, and Muhammad E. H. Chowdhury. 2022. "Prognostic Model of ICU Admission Risk in Patients with COVID-19 Infection Using Machine Learning" Diagnostics 12, no. 9: 2144. https://doi.org/10.3390/diagnostics12092144
APA StyleIslam, K. R., Kumar, J., Tan, T. L., Reaz, M. B. I., Rahman, T., Khandakar, A., Abbas, T., Hossain, M. S. A., Zughaier, S. M., & Chowdhury, M. E. H. (2022). Prognostic Model of ICU Admission Risk in Patients with COVID-19 Infection Using Machine Learning. Diagnostics, 12(9), 2144. https://doi.org/10.3390/diagnostics12092144









