Comparative Analysis of Image Processing Techniques for Enhanced MRI Image Quality: 3D Reconstruction and Segmentation Using 3D U-Net Architecture
Abstract
:1. Introduction
2. Related Works
2.1. Image Pre-Processing
2.2. Segmentation Technique Using CNN in Deep Learning
2.3. Summary of Previous Studies
3. Results
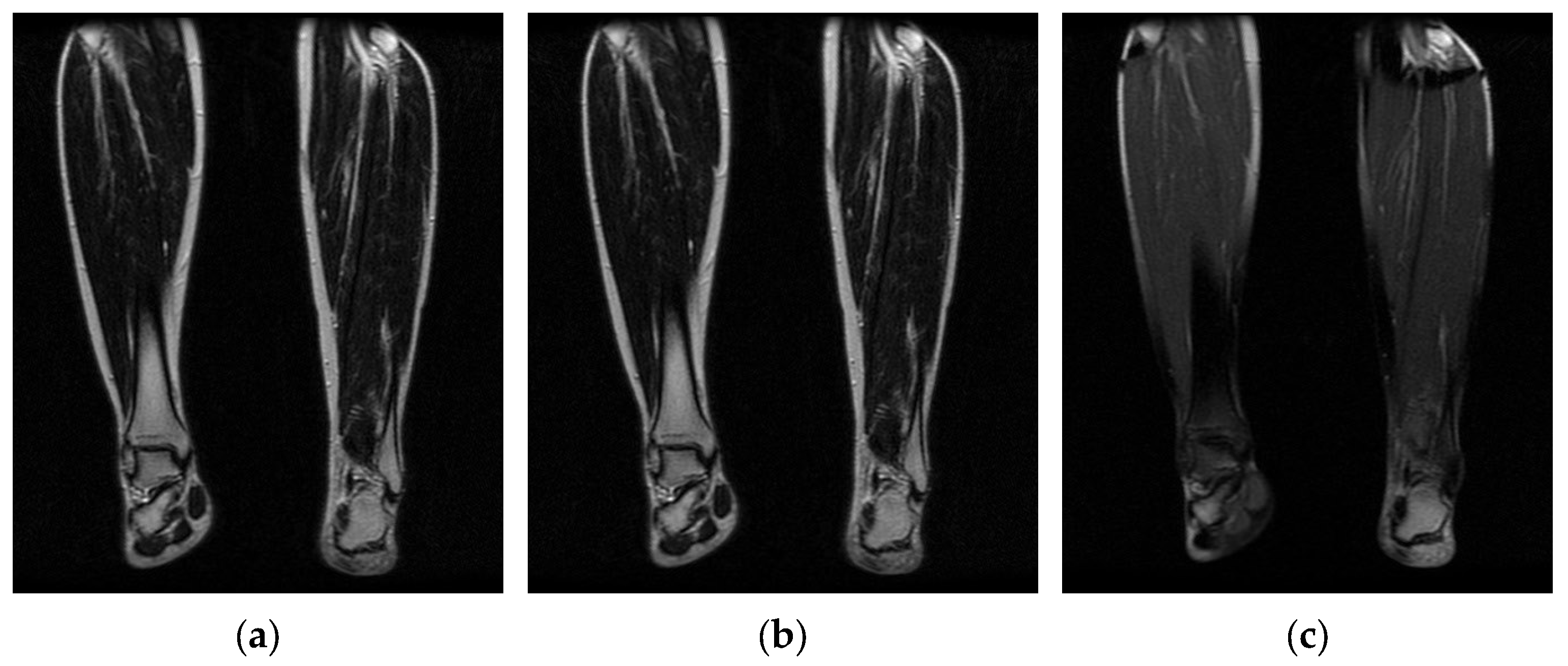
3.1. Image Acquisition
3.2. Image Enhancement
3.3. Image Denoising
3.4. Image Quality Assessment (IQA)
3.5. Reconstruct MRI Images into 3D Volumes
3.6. Segmentation Model for 3D Volumes
3.7. Image Segmentation Performance Validation
4. Discussion
4.1. Image Pre-Processing
4.2. Image Quality Assessment
4.3. Reconstruct MRI Images into 3D Model
4.4. Transformation of 3D Volumes
4.5. Quantitative and Qualitative Evaluation of 3D U-Net Model
4.6. Visualisation of the Predicted Output
4.7. Comparision DSC with Previous Research Works
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Mandava, R.; Alia, O.M.; Wei, B.C.; Ramachandram, D.; Aziz, M.E.; Shuaib, I.L. Osteosarcoma segmentation in MRI using dynamic Harmony Search based clustering. In Proceedings of the 2010 International Conference of Soft Computing and Pattern Recognition, Cergy-Pontoise, France, 7–10 December 2010; pp. 423–429. [Google Scholar] [CrossRef]
- Survival Rates for Osteosarcoma. Available online: https://www.cancer.org/cancer/osteosarcoma/detection-diagnosis-staging/survival-rates.html (accessed on 4 November 2021).
- Signs and Symptoms of Osteosarcoma. Available online: https://www.cancer.org/cancer/osteosarcoma/detection-diagnosis-staging/signs-and-symptoms.html (accessed on 4 November 2021).
- Nasor, M.; Obaid, W. Segmentation of osteosarcoma in MRI images by K-means clustering, Chan-Vese segmentation, and iterative Gaussian filtering. IET Image Process. 2021, 15, 1310–1318. [Google Scholar] [CrossRef]
- Rajeshwari, S.; Sharmila, T.S. Efficient quality analysis of MRI image using preprocessing techniques. In Proceedings of the 2013 IEEE Conference on Information & Communication Technologies, Thuckalay, India, 11–12 April 2013; p. 396. [Google Scholar] [CrossRef]
- Suhas, S.; Venugopal, C.R. MRI image preprocessing and noise removal technique using linear and nonlinear filters. In Proceedings of the 2017 International Conference on Electrical, Electronics, Communication, Computer, and Optimization Techniques (ICEECCOT), Mysuru, India, 15–16 December 2017; pp. 1–4. [Google Scholar] [CrossRef]
- Mohan, G.; Subashini, M.M. MRI based medical image analysis: Survey on brain tumor grade classification. Biomed. Signal Process. Control 2018, 39, 139–161. [Google Scholar] [CrossRef]
- Loizou, C.P.; Pantziaris, M.; Seimenis, I.; Pattichis, C.S. Brain MR image normalization in texture analysis of multiple sclerosis. In Proceedings of the 2009 9th International Conference on Information Technology and Applications in Biomedicine, Larnaka, Cyprus, 4–7 November 2009; pp. 1–5. [Google Scholar] [CrossRef]
- Singh, A.; Sengupta, S.; Lakshminarayanan, V. Explainable Deep Learning Models in Medical Image Analysis. J. Imaging 2020, 6, 52. [Google Scholar] [CrossRef] [PubMed]
- Debelee, T.G.; Kebede, S.R.; Schwenker, F.; Shewarega, Z.M. Deep Learning in Selected Cancers’ Image Analysis—A Survey. J. Imaging 2020, 6, 121. [Google Scholar] [CrossRef] [PubMed]
- Çiçek, Ö.; Abdulkadir, A.; Lienkamp, S.S.; Brox, T.; Ronneberger, O. 3D U-Net: Learning Dense Volumetric Segmentation from Sparse Annotation. In Medical Image Computing and Computer-Assisted Intervention—MICCAI 2016; Ourselin, S., Joskowicz, L., Sabuncu, M.R., Unal, G., Wells, W., Eds.; Springer International Publishing: Cham, Switzerland, 2016; Volume 9901, pp. 424–432. [Google Scholar] [CrossRef] [Green Version]
- Chen, H.; Dou, Q.; Yu, L.; Qin, J.; Heng, P.-A. VoxResNet: Deep voxelwise residual networks for brain segmentation from 3D MR images. NeuroImage 2018, 170, 446–455. [Google Scholar] [CrossRef] [PubMed]
- Holbrook, M.D.; Blocker, S.J.; Mowery, Y.M.; Badea, A.; Qi, Y.; Xu, E.S.; Kirsch, D.G.; Johnson, G.A.; Badea, C. MRI-Based Deep Learning Segmentation and Radiomics of Sarcoma in Mice. Tomography 2020, 6, 23–33. [Google Scholar] [CrossRef] [PubMed]
- Vaidyanathan, A.; van der Lubbe, M.F.J.A.; Leijenaar, R.T.H.; van Hoof, M.; Zerka, F.; Miraglio, B.; Primakov, S.; Postma, A.A.; Bruintjes, T.D.; Bilderbeek, M.A.L.; et al. Deep learning for the fully automated segmentation of the inner ear on MRI. Sci. Rep. 2021, 11, 2885. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Zhu, J.; Lv, B.; Yang, L.; Sun, W.; Dai, Z.; Gou, F.; Wu, J. Auxiliary Segmentation Method of Osteosarcoma MRI Image Based on Transformer and U-Net. Comput. Intell. Neurosci. 2022, 2022, 9990092. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Yang, S.; Gou, F.; Zhou, Z.; Xie, P.; Xu, N.; Dai, Z. Intelligent Segmentation Medical Assistance System for MRI Images of Osteosarcoma in Developing Countries. Comput. Math. Methods Med. 2022, 2022, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Labeeb, Y.; Morsy, M.; Abo-Elsoud, M. Preprocessing Technique for Enhancing the DICOM Kidney Images. Int. J. Eng. Res. Technol. 2018, 4, 836–841. [Google Scholar]
- Kaur, H.; Rani, J. MRI brain image enhancement using Histogram Equalization techniques. In Proceedings of the 2016 International Conference on Wireless Communications, Signal Processing and Networking (WiSPNET), Chennai, India, 23–25 March 2016; pp. 770–773. [Google Scholar] [CrossRef]
- Koonsanit, K.; Thongvigitmanee, S.; Pongnapang, N.; Thajchayapong, P. Image enhancement on digital X-ray images using N-CLAHE. In Proceedings of the 2017 10th Biomedical Engineering International Conference (BMEiCON), Hokkaido, Japan, 31 August 2017–2 September 2017; pp. 1–4. [Google Scholar] [CrossRef]
- Contrast Stretching Using Python and Pillow|Pythontic.com. Available online: https://pythontic.com/image-processing/pillow/contrast%20stretching (accessed on 29 December 2021).
- Nuruzzaman, F. Contrast Stretching in Image Processing Using Matlab. 10 February 2019. Available online: https://www.nzfaruqui.com/contrast-stretching-in-image-processing-using-matlab/ (accessed on 29 December 2021).
- Ali, H.M. MRI Medical Image Denoising by Fundamental Filters; IntechOpen: Rijeka, Croatia, 2018. [Google Scholar] [CrossRef] [Green Version]
- Pawar, M. MRI and CT Image Denoising using Gaussian Filter, Wavelet Transform and Curvelet Transform. Int. J. Eng. Sci. Comput. 2017, 7, 4. [Google Scholar]
- Gaussian Blur—An overview|ScienceDirect Topics. Available online: https://www.sciencedirect.com/topics/engineering/gaussian-blur (accessed on 27 December 2021).
- Young, I.T.; van Vliet, L.J. Recursive implementation of the Gaussian filter. Signal Process. 1995, 44, 139–151. [Google Scholar] [CrossRef] [Green Version]
- 26Evaluating Denoising Performances of Fundamental Filters for T2-Weighted MRI Images|Elsevier Enhanced Reader. Available online: https://reader.elsevier.com/reader/sd/pii/S1877050915023583?token=A8010A91375CC1FBFF21C5C99A2A2F13B2CC26F6A11AB2354554263835A78B6BE1AFA36E1F958D54E39C610CEC98FFB8&originRegion=eu-west-1&originCreation=20211227062644 (accessed on 27 December 2021).
- Kaur, R. Comparison of contrast enhancement techniques for medical image. In Proceedings of the 2016 Conference on Emerging Devices and Smart Systems (ICEDSS), Namakkal, India, 4–5 March 2016; pp. 155–159. [Google Scholar] [CrossRef]
- Mzoughi, H.; Njeh, I.; Slima, M.B.; Hamida, A.B.; Mhiri, C.; Mahfoudh, K.B. Denoising and contrast-enhancement approach of magnetic resonance imaging glioblastoma brain tumors. J. Med. Imaging 2019, 6, 044002. [Google Scholar] [CrossRef] [PubMed]
- Dnuggets, K. Medical Image Analysis with Deep Learning, Part 4. Available online: https://www.kdnuggets.com/medical-image-analysis-with-deep-learning-part-4.html/ (accessed on 3 January 2022).
- 30Neuroimaging in Python—Pipelines and Interfaces—Nipy Pipeline and Interfaces Package. Available online: https://nipype.readthedocs.io/en/latest/api/generated/nipype.interfaces.dcm2nii.html (accessed on 3 January 2022).
- Haque, I.R.I.; Neubert, J. Deep learning approaches to biomedical image segmentation. Inform. Med. Unlocked 2020, 18, 100297. [Google Scholar] [CrossRef]
- Soomro, M.H.; Coppotelli, M.; Conforto, S.; Schmid, M.; Giunta, G.; Del Secco, L.; Neri, E.; Caruso, D.; Rengo, M.; Laghi, A. Automated Segmentation of Colorectal Tumor in 3D MRI Using 3D Multiscale Densely Connected Convolutional Neural Network. J. Heal. Eng. 2019, 2019, 1–11. [Google Scholar] [CrossRef] [PubMed]
- 33MSD Manual Professional Edition. Magnetic Resonance Imaging—Special Subjects. Available online: https://www.msdmanuals.com/professional/special-subjects/principles-of-radiologic-imaging/magnetic-resonance-imaging (accessed on 16 June 2022).
- Nascimento, D.; Suchard, G.; Hatem, M.; de Abreu, A. The role of magnetic resonance imaging in the evaluation of bone tumours and tumour-like lesions. Insights Imaging 2014, 5, 419–440. [Google Scholar] [CrossRef] [PubMed]
- Lv, B.; Liu, F.; Gou, F.; Wu, J. Multi-Scale Tumor Localization Based on Priori Guidance-Based Segmentation Method for Osteosarcoma MRI Images. Mathematics 2022, 10, 2099. [Google Scholar] [CrossRef]
- Kayal, E.B.; Kandasamy, D.; Yadav, R.; Bakhshi, S.; Sharma, R.; Mehndiratta, A. Automatic segmentation and RECIST score evaluation in osteosarcoma using diffusion MRI: A computer aided system process. Eur. J. Radiol. 2020, 133, 109359. [Google Scholar] [CrossRef] [PubMed]


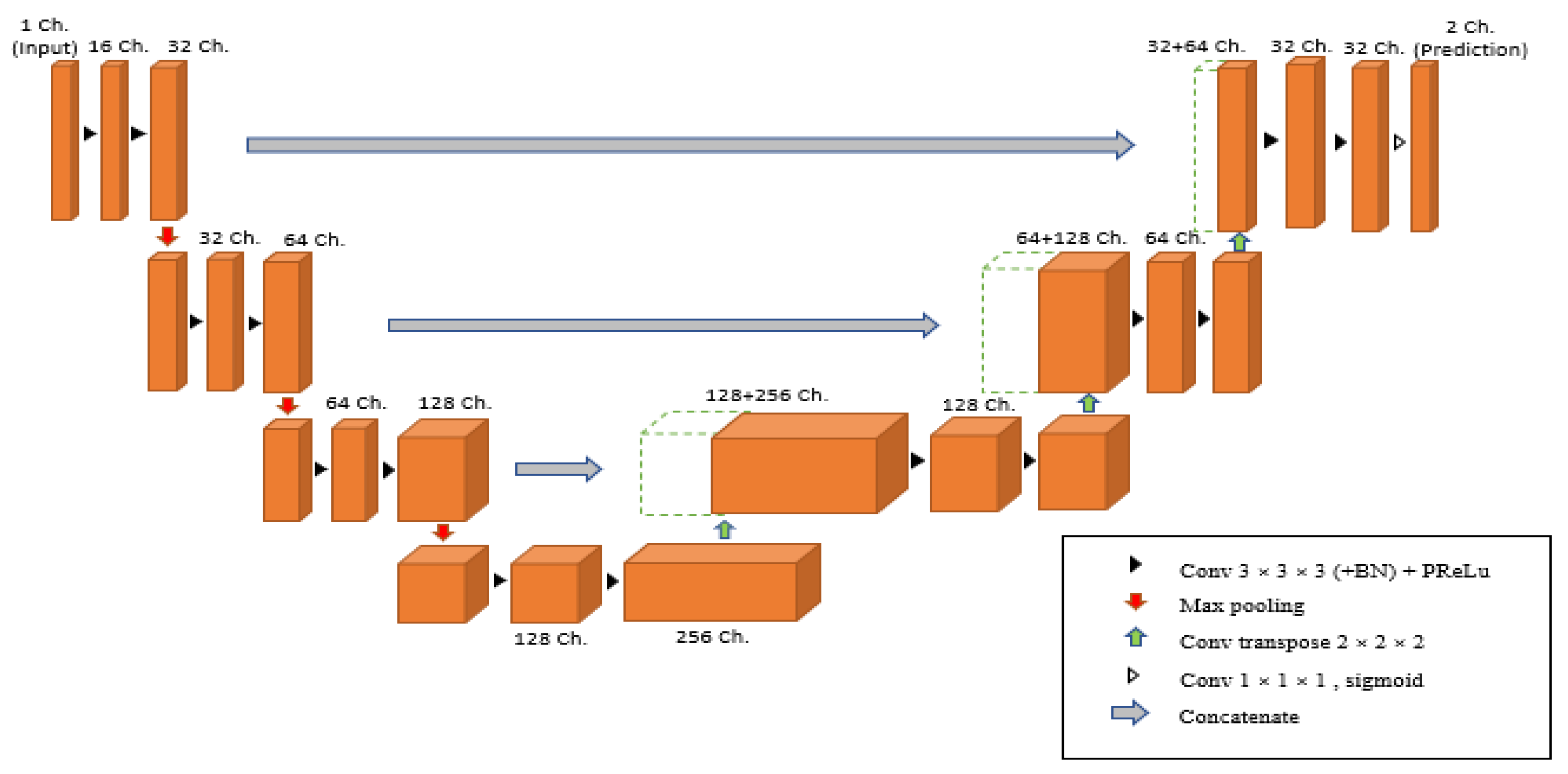


| Hyperparameters | Values |
|---|---|
| Data split ratio | 8:1:1 |
| Maximum epochs | 800 |
| Batch size | 2 |
| Optimizer | Adam |
| Loss function | Dice loss |
| Activation function | PReLU |
| Total number of parameters | 4,808,917 |
| T1W MRI Image after Contrast Enhancement | Combination of Pre-Processing Techniques | |||
|---|---|---|---|---|
| CLAHE + Gaussian Filter | CLAHE + Median Filter | Contrast Stretching + Gaussian Filter | Contrast Stretching + Median Filter | |
 |  |  |  |  |
 |  |  |  |  |
| T2W MRI Image after Contrast Enhancement | After Pre-Processing of T2W | |||
|---|---|---|---|---|
| CLAHE + Gaussian Filter | CLAHE + Median Filter | Contrast Stretching + Gaussian Filter | Contrast Stretching + Median Filter | |
 |  |  |  |  |
 |  |  |  |  |
| T1W + Gd MRI Image after Contrast Enhancement | After Pre-Processing of T1W + Gd | |||
|---|---|---|---|---|
| CLAHE + Gaussian Filter | CLAHE + Median Filter | Contrast Stretching + Gaussian Filter | Contrast Stretching + Median Filter | |
 |  |  |  |  |
 |  |  |  |  |
| MSE | PSNR | AMBE | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| T1W | T1W + Gd | T2W | T1W | T1W + Gd | T2W | T1W | T1W + Gd | T2W | ||
| CLAHE + Gaussian filter | MRI 1 | 99.5270 | 98.0830 | 110.4348 | 28.1514 | 28.2149 | 27.6997 | 0.06093 | 0.07008 | 0.08086 |
| MRI 2 | 77.6347 | 73.5121 | 106.9793 | 29.2302 | 29.4672 | 27.8378 | 0.07691 | 0.08538 | 0.10174 | |
| MRI 3 | 88.3859 | 86.8852 | 108.9995 | 28.6670 | 28.7413 | 27.7566 | 0.04468 | 0.06080 | 0.07764 | |
| MRI 4 | 93.6140 | 105.3020 | 108.0355 | 28.4174 | 27.9064 | 27.7951 | 0.04200 | 0.05408 | 0.05596 | |
| MRI 5 | 94.4030 | 100.5126 | 110.5212 | 28.3809 | 28.1086 | 27.6964 | 0.04347 | 0.05096 | 0.05848 | |
| AVG | 90.7129 | 92.8589 | 108.9940 | 28.5693 | 28.4876 | 27.7571 | 0.05359 | 0.06426 | 0.07493 | |
| CLAHE + Median filter | MRI 1 | 88.8244 | 88.3116 | 99.6605 | 28.6455 | 28.6706 | 28.1456 | 0.05805 | 0.06831 | 0.07490 |
| MRI 2 | 77.0120 | 72.0549 | 100.0506 | 29.2652 | 29.5542 | 28.1286 | 0.07475 | 0.08146 | 0.09748 | |
| MRI 3 | 81.6804 | 82.0138 | 97.0345 | 29.0096 | 28.9919 | 28.2615 | 0.04361 | 0.05940 | 0.06829 | |
| MRI 4 | 76.8616 | 91.1990 | 100.1020 | 29.2737 | 28.5309 | 28.1264 | 0.38308 | 0.05018 | 0.05279 | |
| MRI 5 | 83.4949 | 87.9688 | 101.8403 | 28.9142 | 28.6875 | 28.0516 | 0.04084 | 0.04781 | 0.05486 | |
| AVG | 81.5747 | 84.3096 | 99.7376 | 29.0216 | 28.8870 | 28.1427 | 0.1201 | 0.0614 | 0.0697 | |
| Contrast Stretching + Gaussian filter | MRI 1 | 22.5087 | 18.8500 | 24.1599 | 34.6073 | 35.3777 | 34.1599 | 0.01251 | 0.01570 | 0.02479 |
| MRI 2 | 40.4818 | 38.7746 | 38.2741 | 32.0582 | 32.2453 | 32.3018 | 0.04688 | 0.04477 | 0.04038 | |
| MRI 3 | 22.0822 | 21.9366 | 20.2009 | 34.6904 | 34.7191 | 35.0771 | 0.01152 | 0.01697 | 0.01564 | |
| MRI 4 | 7.3680 | 55.7069 | 8.9038 | 39.4573 | 30.6717 | 38.6351 | 0.01216 | 0.03440 | 0.01196 | |
| MRI 5 | 7.4555 | 15.0778 | 9.9041 | 39.4060 | 36.3474 | 38.1726 | 0.01061 | 0.02000 | 0.01385 | |
| AVG | 19.9792 | 30.0692 | 20.2886 | 36.0438 | 33.8722 | 35.6693 | 0.01874 | 0.02647 | 0.02132 | |
| Contrast Stretching + Median filter | MRI 1 | 21.7134 | 18.5888 | 23.6220 | 34.7635 | 35.4383 | 34.3976 | 0.01197 | 0.01522 | 0.02406 |
| MRI 2 | 39.5238 | 38.6358 | 36.7550 | 32.1622 | 32.2609 | 32.4776 | 0.04632 | 0.04444 | 0.03961 | |
| MRI 3 | 20.9480 | 20.8487 | 19.3480 | 34.9194 | 34.9400 | 35.2644 | 0.01099 | 0.01660 | 0.01492 | |
| MRI 4 | 7.6674 | 52.1598 | 9.1981 | 39.2844 | 30.9575 | 38.4938 | 0.01136 | 0.03300 | 0.01047 | |
| MRI 5 | 7.5921 | 14.8690 | 10.3506 | 39.3272 | 36.4080 | 37.9812 | 0.00986 | 0.01906 | 0.01293 | |
| AVG | 19.4889 | 29.0204 | 19.8547 | 36.0913 | 34.0009 | 35.7229 | 0.01810 | 0.02566 | 0.02040 | |
| Type of MRI | Plane | DICOM Image | 3D Model in NIfTI |
|---|---|---|---|
| T1W | Axial |  |  |
| Coronal |  | ||
| Sagittal |  | ||
| T1W + Gd | Axial |  |  |
| Coronal |  | ||
| Sagittal |  | ||
| T2W | Axial |  |  |
| Coronal |  | ||
| Sagittal |  |
| Slice of Original Image | Transformation for Ground Truth | After Transformation | ||
|---|---|---|---|---|
| T1W | T1W + Gd | T2W | ||
 |  |  |  |  |
 |  |  |  |  |
 |  |  |  |  |
| Types of MRI Image | Epoch | Training Time (Second) | Mean DSC | Epoch Average Dice Loss |
|---|---|---|---|---|
| T1W | 786 | 20,194.563 | 0.8375 | 0.1709 |
| T2W | 792 | 20,429.427 | 0.8545 | 0.1563 |
| T1W + Gd | 700 | 20,020.069 | 0.8762 | 0.1534 |
| Sample No. (MRI Slice) | Input | Ground Truth | Output | ||
|---|---|---|---|---|---|
| T1W | T2W | T1W + Gd | |||
| 1 (80th slice) |  |  |  |  |  |
| 2 (80th slice) |  |  |  |  |  |
| 3 (80th slice) |  |  |  |  |  |
| MRI Slice | Input | Ground Truth (G) | Output (O) | Overlaid Slices | ||
|---|---|---|---|---|---|---|
| Ground Truth | Output | Ground Truth and Output, (G ∩ O) | ||||
| 60th |  |  |  |  |  |  |
| 70th |  |  |  |  |  |  |
| 80th |  |  |  |  |  |  |
| 90th |  |  |  |  |  |  |
| 100th |  |  |  |  |  |  |
| MRI Slice | Input | Ground Truth | Output | Overlaid Slices | ||
|---|---|---|---|---|---|---|
| Ground Truth | Output | Ground Truth and Output | ||||
| 60th |  |  |  |  |  |  |
| 70th |  |  |  |  |  |  |
| 80th |  |  |  |  |  |  |
| 90th |  |  |  |  |  |  |
| 100th |  |  |  |  |  |  |
| MRI Slice | Input | Ground Truth | Output | Overlaid Slices | ||
|---|---|---|---|---|---|---|
| Ground Truth | Output | Ground Truth and Output | ||||
| 60th |  |  |  |  |  |  |
| 70th |  |  |  |  |  |  |
| 80th |  |  |  |  |  |  |
| 90th |  |  |  |  |  |  |
| 100th |  |  |  |  |  |  |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lim, C.C.; Ling, A.H.W.; Chong, Y.F.; Mashor, M.Y.; Alshantti, K.; Aziz, M.E. Comparative Analysis of Image Processing Techniques for Enhanced MRI Image Quality: 3D Reconstruction and Segmentation Using 3D U-Net Architecture. Diagnostics 2023, 13, 2377. https://doi.org/10.3390/diagnostics13142377
Lim CC, Ling AHW, Chong YF, Mashor MY, Alshantti K, Aziz ME. Comparative Analysis of Image Processing Techniques for Enhanced MRI Image Quality: 3D Reconstruction and Segmentation Using 3D U-Net Architecture. Diagnostics. 2023; 13(14):2377. https://doi.org/10.3390/diagnostics13142377
Chicago/Turabian StyleLim, Chee Chin, Apple Ho Wei Ling, Yen Fook Chong, Mohd Yusoff Mashor, Khalilalrahman Alshantti, and Mohd Ezane Aziz. 2023. "Comparative Analysis of Image Processing Techniques for Enhanced MRI Image Quality: 3D Reconstruction and Segmentation Using 3D U-Net Architecture" Diagnostics 13, no. 14: 2377. https://doi.org/10.3390/diagnostics13142377






