Nasal Microbiome in Granulomatosis with Polyangiitis Compared to Chronic Rhinosinusitis
Abstract
:1. Introduction
2. Materials and Methods
2.1. Metagenomic Analysis
- Automatic sample demultiplexing;
- Generating FASTQ files containing raw reads.
- Checking the quality of the readings:
- Analysing the error profile of individual samples and the dynamic generation of parameters for quality control (FIGARO tool);
- Carrying out a quality control based on the coefficient of the maximum expected errors of the sample.
- Preprocessing data using the Cutadapt tool:
- Removing adapter sequences;
- Rejecting too short readings (a minimum length of 30 nt).
- Selecting unique ASV sequences (using the DADA2 package) by the following process:
- Filtering out the sequences containing errors created during the sequencing process (denoising);
- Combining paired readings—to increase the accuracy of sequencing, they are performed in paired-end mode so that at later stages of the analysis, the corresponding forward and reverse readings must be combined;
- Merging of identical, unique sequences while maintaining the number of springs and the quality profile (dereplicating);
- Removing constructs resulting from incorrect sequence assembly during PCR (filtering chimeras).
- Assigning a taxonomy to the ASV sequence based on the Silva reference base and using a hybrid approach:
- Comparing ASV sequences against the base in search of identical reference sequences (VSEARCH);
- Classifying the atypical sequences remaining after the previous step based on methods using machine learning (sklearn).
2.2. Statistical Analysis
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Kumpitsch, C.; Koskinen, K.; Schöpf, V.; Moissl-Eichinger, C. The Microbiome of the Upper Respiratory Tract in Health and Disease. BMC Biol. 2019, 17, 87. [Google Scholar] [CrossRef]
- Dimitri-Pinheiro, S.; Soares, R.; Barata, P. The Microbiome of the Nose-Friend or Foe? Allergy Rhinol. 2020, 11, 2152656720911605. [Google Scholar] [CrossRef] [PubMed]
- Psaltis, A.J.; Wormald, P.-J. Therapy of Sinonasal Microbiome in CRS: A Critical Approach. Curr. Allergy Asthma Rep. 2017, 17, 59. [Google Scholar] [CrossRef] [PubMed]
- De Boeck, I.; van den Broek, M.F.L.; Allonsius, C.N.; Spacova, I.; Wittouck, S.; Martens, K.; Wuyts, S.; Cauwenberghs, E.; Jokicevic, K.; Vandenheuvel, D.; et al. Lactobacilli Have a Niche in the Human Nose. Cell Rep. 2020, 31, 107674. [Google Scholar] [CrossRef]
- Yamanishi, S.; Pawankar, R. Current Advances on the Microbiome and Role of Probiotics in Upper Airways Disease. Curr. Opin. Allergy Clin. Immunol. 2020, 20, 30–35. [Google Scholar] [CrossRef] [PubMed]
- Xiao, H.; Hu, P.; Falk, R.J.; Jennette, J.C. Overview of the Pathogenesis of ANCA-Associated Vasculitis. Kidney Dis. 2016, 1, 205–215. [Google Scholar] [CrossRef]
- Jennette, J.C.; Falk, R.J.; Bacon, P.A.; Basu, N.; Gross, W.L.; Guillevin, L.; Hagen, E.C.; Hoffman, G.S.; Jayne, D.R.; Kallenberg, C.G.M.; et al. 2012 Revised International Chapel Hill Consensus Conference Nomenclature of Vasculitides. Arthritis Rheum. 2013, 65, 1–11. [Google Scholar] [CrossRef]
- Jennette, J.C.; Falk, R.J. Pathogenesis of Antineutrophil Cytoplasmic Autoantibody-Mediated Disease. Nat. Rev. Rheumatol. 2014, 10, 463–473. [Google Scholar] [CrossRef]
- Fokkens, W.J.; Lund, V.J.; Hopkins, C.; Hellings, P.W.; Kern, R.; Reitsma, S.; Toppila-Salmi, S.; Bernal-Sprekelsen, M.; Mullol, J.; Alobid, I.; et al. European Position Paper on Rhinosinusitis and Nasal Polyps 2020. Rhinology 2020, 58 (Suppl. S29), 1–464. [Google Scholar] [CrossRef]
- Wojciechowska, J.; Krajewski, W.; Krajewski, P.; Kręcicki, T. Granulomatosis With Polyangiitis in Otolaryngologist Practice: A Review of Current Knowledge. Clin. Exp. Otorhinolaryngol. 2016, 9, 8–13. [Google Scholar] [CrossRef]
- Brożek-Mądry, E.; Szopiński, K.; Życińska, K.; Krzeski, A. Granulomatosis with Polyangiitis and Nasal Involvement—What Radiological Markers Point the Disease. Pol. Arch. Intern. Med. 2021, 131, 649–657. [Google Scholar] [CrossRef] [PubMed]
- Stegeman, C.A.; Tervaert, J.W.; Sluiter, W.J.; Manson, W.L.; de Jong, P.E.; Kallenberg, C.G. Association of Chronic Nasal Carriage of Staphylococcus Aureus and Higher Relapse Rates in Wegener Granulomatosis. Ann. Intern. Med. 1994, 120, 12–17. [Google Scholar] [CrossRef] [PubMed]
- Laudien, M.; Gadola, S.D.; Podschun, R.; Hedderich, J.; Paulsen, J.; Reinhold-Keller, E.; Csernok, E.; Ambrosch, P.; Hellmich, B.; Moosig, F.; et al. Nasal Carriage of Staphylococcus Aureus and Endonasal Activity in Wegener s Granulomatosis as Compared to Rheumatoid Arthritis and Chronic Rhinosinusitis with Nasal Polyps. Clin. Exp. Rheumatol. 2010, 28 (Suppl. S57), S51–S55. [Google Scholar]
- Zycinska, K.; Wardyn, K.A.; Zielonka, T.M.; Demkow, U.; Traburzynski, M.S. Chronic Crusting, Nasal Carriage of Staphylococcus Aureus and Relapse Rate in Pulmonary Wegener’s Granulomatosis. J. Physiol. Pharmacol. 2008, 59 (Suppl. S6), 825–831. [Google Scholar] [PubMed]
- Salmela, A.; Rasmussen, N.; Tervaert, J.W.C.; Jayne, D.R.W.; Ekstrand, A.; European Vasculitis Study Group. Chronic Nasal Staphylococcus Aureus Carriage Identifies a Subset of Newly Diagnosed Granulomatosis with Polyangiitis Patients with High Relapse Rate. Rheumatology 2017, 56, 965–972. [Google Scholar] [CrossRef] [PubMed]
- Stegeman, C.A.; Tervaert, J.W.; de Jong, P.E.; Kallenberg, C.G. Trimethoprim-Sulfamethoxazole (Co-Trimoxazole) for the Prevention of Relapses of Wegener’s Granulomatosis. Dutch Co-Trimoxazole Wegener Study Group. N. Engl. J. Med. 1996, 335, 16–20. [Google Scholar] [CrossRef] [PubMed]
- Zycinska, K.; Wardyn, K.A.; Zielonka, T.M.; Krupa, R.; Lukas, W. Co-Trimoxazole and Prevention of Relapses of PR3-ANCA Positive Vasculitis with Pulmonary Involvement. Eur. J. Med. Res. 2009, 14 (Suppl. S4), 265–267. [Google Scholar] [CrossRef] [PubMed]
- Tan, B.K.; Crabol, Y.; Tasse, J.; Laurent, F.; Nekkab, N.; Vinter, C.; Puéchal, X.; Guillevin, L. No Evident Association of Nasal Carriage of Staphylococcus Aureus or Its Small-Colony Variants with Cotrimoxazole Use or ANCA-Associated Vasculitis Relapses. Rheumatology 2020, 59, 77–83. [Google Scholar] [CrossRef] [PubMed]
- Antonino, M.; Nicolò, M.; Jerome Renee, L.; Federico, M.; Chiara, V.; Stefano, S.; Maria, S.; Salvatore, C.; Antonio, B.; Calvo-Henriquez, C.; et al. Single-nucleotide Polymorphism in Chronic Rhinosinusitis: A Systematic Review. Clin. Otolaryngol. 2022, 47, 14–23. [Google Scholar] [CrossRef]
- Hopkins, C.; Gillett, S.; Slack, R.; Lund, V.J.; Browne, J.P. Psychometric Validity of the 22-Item Sinonasal Outcome Test. Clin. Otolaryngol. 2009, 34, 447–454. [Google Scholar] [CrossRef]
- Lund, V.J.; Kennedy, D.W. Staging for Rhinosinusitis. Otolaryngol. Head Neck Surg. 1997, 117, S35–S40. [Google Scholar] [CrossRef]
- Rhee, R.L.; Lu, J.; Bittinger, K.; Lee, J.-J.; Mattei, L.M.; Sreih, A.G.; Chou, S.; Miner, J.J.; Cohen, N.A.; Kelly, B.J.; et al. Dynamic Changes in the Nasal Microbiome Associated With Disease Activity in Patients With Granulomatosis With Polyangiitis. Arthritis Rheumatol. 2021, 73, 1703–1712. [Google Scholar] [CrossRef] [PubMed]
- Rhee, R.L.; Sreih, A.G.; Najem, C.E.; Grayson, P.C.; Zhao, C.; Bittinger, K.; Collman, R.G.; Merkel, P.A. Characterisation of the Nasal Microbiota in Granulomatosis with Polyangiitis. Ann. Rheum. Dis. 2018, 77, 1448–1453. [Google Scholar] [CrossRef] [PubMed]
- Wagner, J.; Harrison, E.M.; Martinez Del Pero, M.; Blane, B.; Mayer, G.; Leierer, J.; Gopaluni, S.; Holmes, M.A.; Parkhill, J.; Peacock, S.J.; et al. The Composition and Functional Protein Subsystems of the Human Nasal Microbiome in Granulomatosis with Polyangiitis: A Pilot Study. Microbiome 2019, 7, 137. [Google Scholar] [CrossRef]
- Lamprecht, P.; Fischer, N.; Huang, J.; Burkhardt, L.; Lütgehetmann, M.; Arndt, F.; Rolfs, I.; Kerstein, A.; Iking-Konert, C.; Laudien, M. Changes in the Composition of the Upper Respiratory Tract Microbial Community in Granulomatosis with Polyangiitis. J. Autoimmun. 2019, 97, 29–39. [Google Scholar] [CrossRef]
- Jennette, J.C. Overview of the 2012 Revised International Chapel Hill Consensus Conference Nomenclature of Vasculitides. Clin. Exp. Nephrol. 2013, 17, 603–606. [Google Scholar] [CrossRef] [PubMed]
- Claesen, J.; Spagnolo, J.B.; Ramos, S.F.; Kurita, K.L.; Byrd, A.L.; Aksenov, A.A.; Melnik, A.V.; Wong, W.R.; Wang, S.; Hernandez, R.D.; et al. A Cutibacterium Acnes Antibiotic Modulates Human Skin Microbiota Composition in Hair Follicles. Sci. Transl. Med. 2020, 12, eaay5445. [Google Scholar] [CrossRef]
- Nakamura, K.; O’Neill, A.M.; Williams, M.R.; Cau, L.; Nakatsuji, T.; Horswill, A.R.; Gallo, R.L. Short Chain Fatty Acids Produced by Cutibacterium Acnes Inhibit Biofilm Formation by Staphylococcus Epidermidis. Sci. Rep. 2020, 10, 21237. [Google Scholar] [CrossRef]
- Piewngam, P.; Otto, M. Staphylococcus Aureus Colonisation and Strategies for Decolonisation. Lancet Microbe 2024, 5, e606–e618. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Lei, H.; Li, Z.; Li, H.; Wang, Y.; Lai, Y. A Novel Lipopeptide from Skin Commensal Activates TLR2/CD36-P38 MAPK Signaling to Increase Antibacterial Defense against Bacterial Infection. PLoS ONE 2013, 8, e58288. [Google Scholar] [CrossRef]
- Zipperer, A.; Konnerth, M.C.; Laux, C.; Berscheid, A.; Janek, D.; Weidenmaier, C.; Burian, M.; Schilling, N.A.; Slavetinsky, C.; Marschal, M.; et al. Human Commensals Producing a Novel Antibiotic Impair Pathogen Colonization. Nature 2016, 535, 511–516. [Google Scholar] [CrossRef] [PubMed]
- Moodley, A.; Damborg, P.; Nielsen, S.S. Antimicrobial Resistance in Methicillin Susceptible and Methicillin Resistant Staphylococcus Pseudintermedius of Canine Origin: Literature Review from 1980 to 2013. Vet. Microbiol. 2014, 171, 337–341. [Google Scholar] [CrossRef] [PubMed]
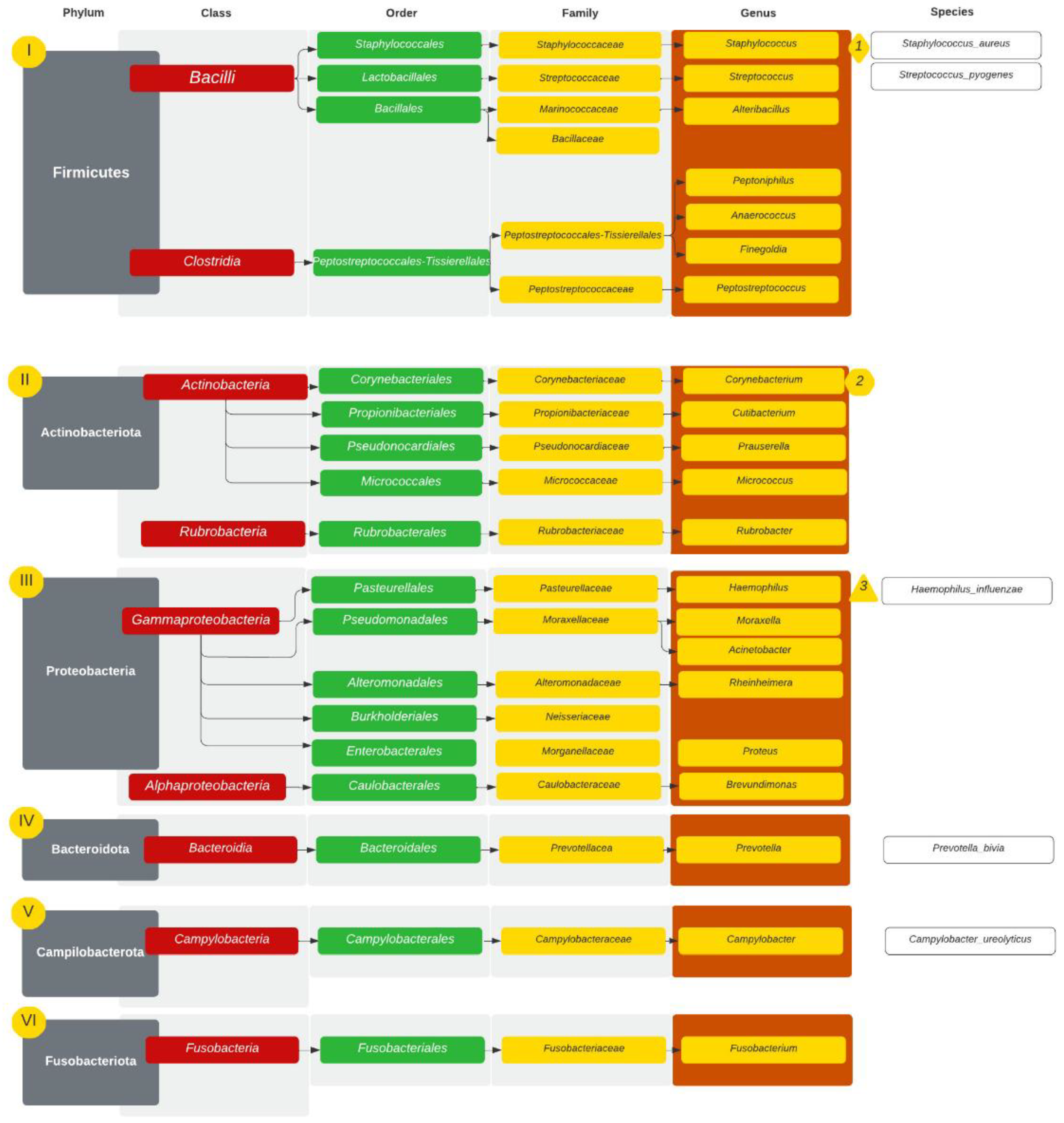
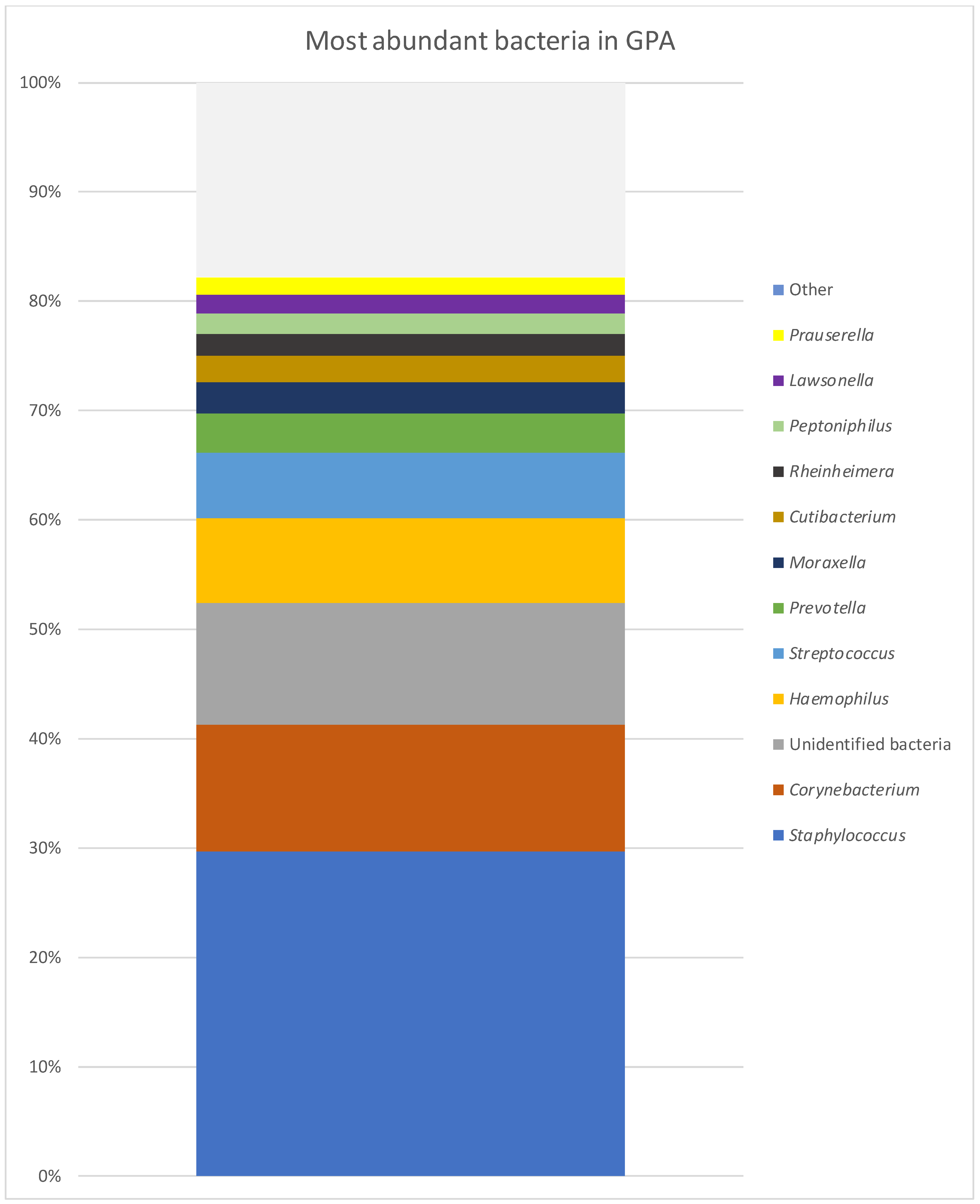

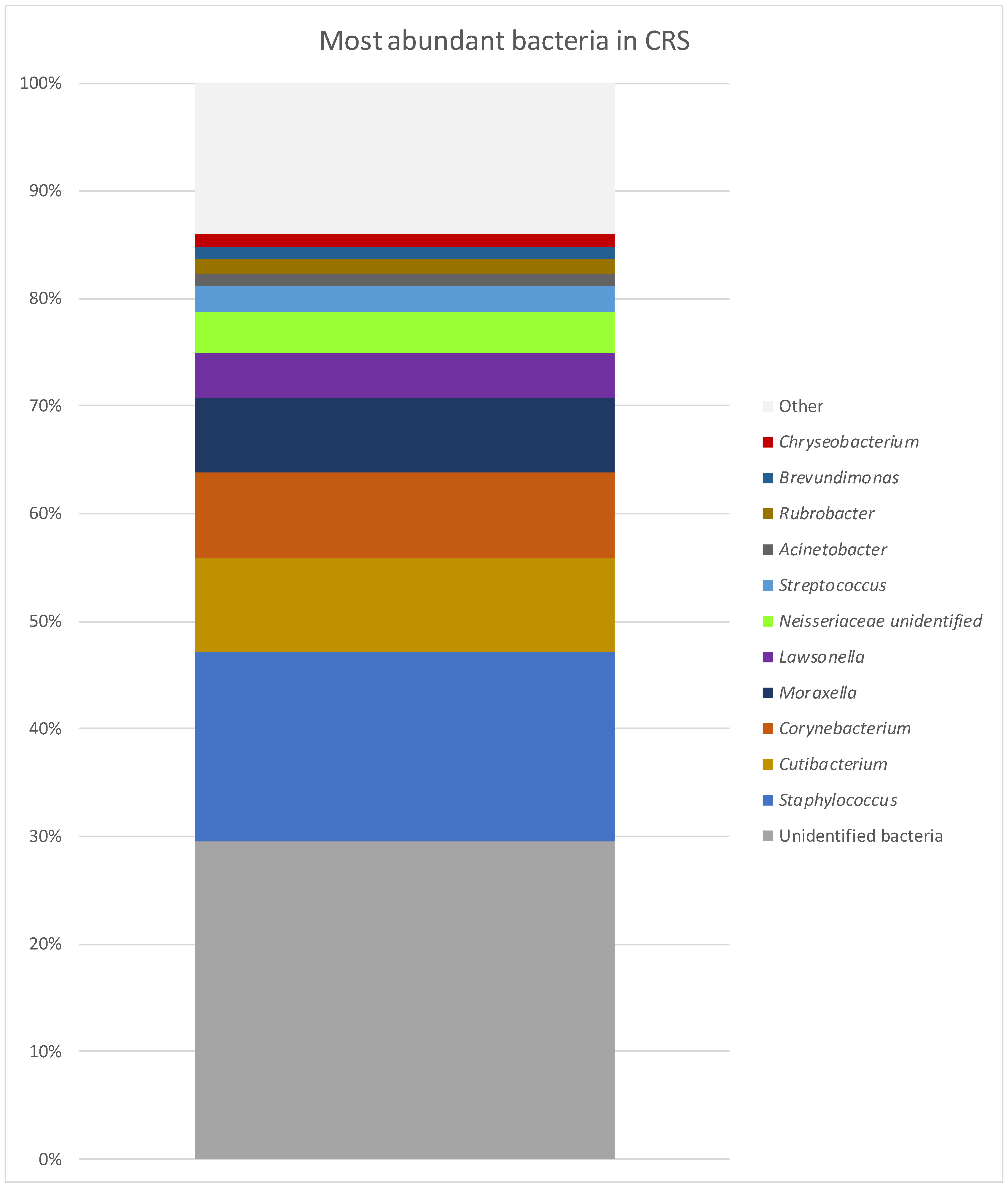
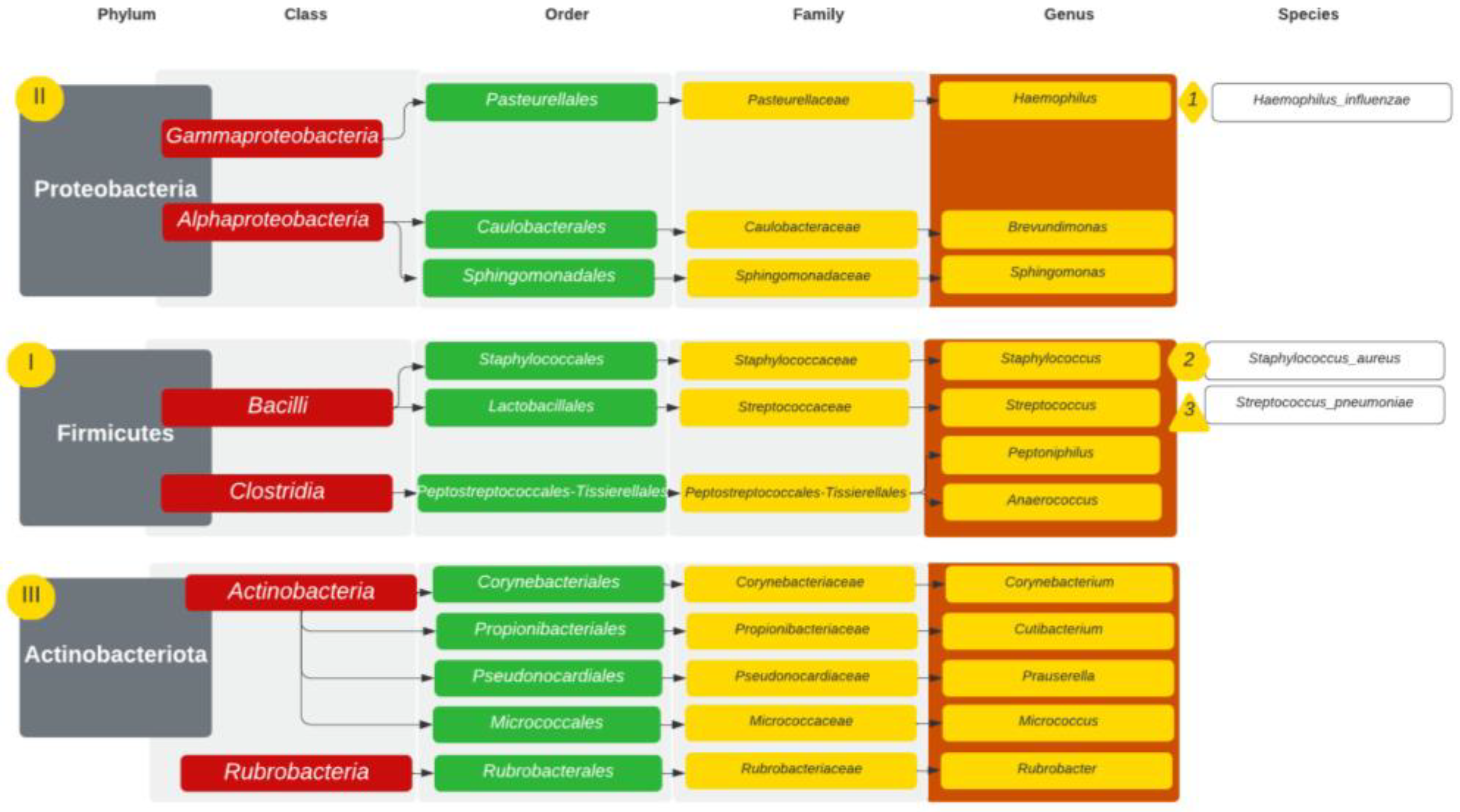
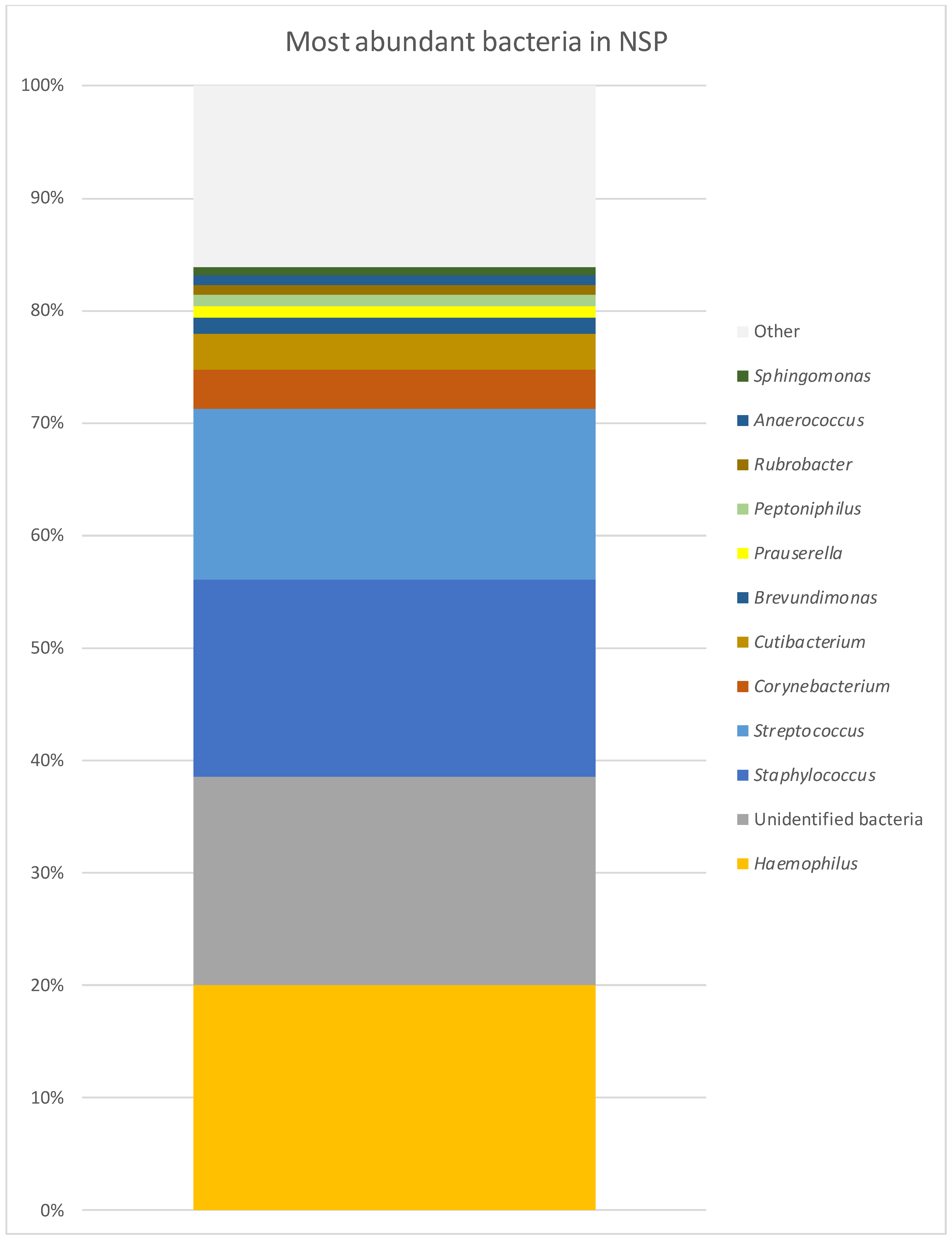
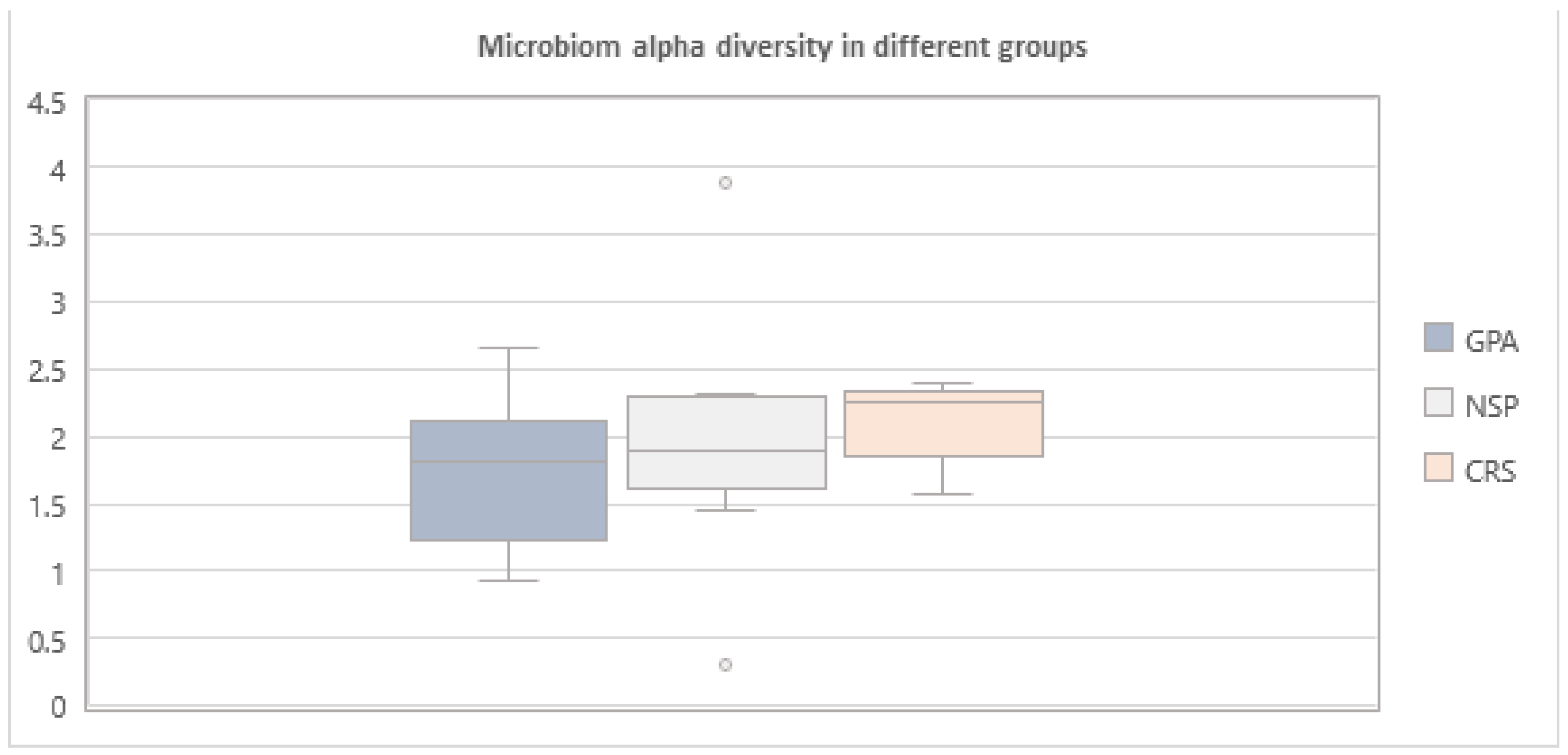
| GPA | CRS | NSP | |
| Staphylococcus | 29.65% | 17.47% | 17.42% |
| Corynebacterium | 11.65% | 7.93% | 3.58% |
| Unidentified bacteria | 11.16% | 29.60% | 18.47% |
| Haemophilus | 7.70% | 0.50% | 20.03% |
| Streptococcus | 5.97% | 2.93% | 15.16% |
| Prevotella | 3.68% | 0.12% | 0.21% |
| Moraxella | 2.77% | 7.05% | 0.32% |
| Cutibacterium | 2.49% | 8.80% | 3.20% |
| Rheinheimera | 1.95% | 0.03% | 0.01% |
| Peptoniphilus | 1.84% | 0.37% | 1.01% |
| Lawsonella | 1.71% | 4.10% | 0.48% |
| Prauserella | 1.70% | 0.50% | 1.04% |
| Alteribacillus | 1.56% | 0.46% | 0.23% |
| Rubrobacter | 1.39% | 1.26% | 0.94% |
| Anaerococcus | 1.06% | 0.83% | 0.75% |
| Neisseriaceae unidentified | 0.69% | 3.80% | 0.01% |
| Acinetobacter | 0.98% | 1.27% | 0.24% |
| Brevundimonas | 0.59% | 1.19% | 1.35% |
| Chryseobacterium | 0.00% | 1.18% | 0.22% |
| (A) | |||||
| Family | U Mann–Whitney Test | ||||
| Median GPA | Median NPS | p-Value | |||
| Actinomycetaceae | 0.00% | 0.05% | 0.016 | ||
| Flavobacteriaceae | 0.00% | 0.04% | 0.038 | ||
| Weeksellaceae | 0.00% | 0.13% | 0.016 | ||
| Streptococcaceae | 0.42% | 11.22% | 0.025 | ||
| Lachnospiraceae | 0.00% | 0.39% | 0.025 | ||
| Beijerinckiaceae | 0.00% | 0.23% | 0.020 | ||
| Rhodobacteraceae | 0.01% | 0.32% | 0.031 | ||
| (B) | |||||
| Family | U Mann–Whitney Test | ||||
| Median GPA | Median CRS | p-Value | |||
| Propionibacteriaceae | 1.15% | 4.78% | 0.026 | ||
| Weeksellaceae | 0.00% | 0.18% | <0.001 | ||
| Caulobacteraceae | 0.25% | 0.89% | 0.041 | ||
| Beijerinckiaceae | 0.00% | 0.14% | 0.041 | ||
| Xanthobacteraceae | 0.00% | 0.12% | 0.041 | ||
| Sphingomonadaceae | 0.02% | 0.28% | 0.015 | ||
| Moraxellaceae | 0.02% | 2.57% | 0.033 | ||
| (C) | |||||
| Family | U Mann–Whitney Test | ||||
| Median NPS | Median CRS | p-Value | |||
| Lachnospiraceae | 0.39% | 0.00% | 0.035 | ||
| Neisseriaceae | 0.13% | 0.78% | 0.005 | ||
| (A) | |||
| Genus | U Mann–Whitney Test | ||
| Median GPA | Median NSP | p-Value | |
| Actinomyces | 0.00% | 0.03% | 0.008 |
| Streptococcus | 0.17% | 11.22% | 0.008 |
| Methylobacterium-Methylorubrum | 0.00% | 0.11% | 0.014 |
| Paracoccus | 0.00% | 0.32% | 0.046 |
| (B) | |||
| Genus | U Mann–Whitney test | ||
| Median GPA | Median CRS | p-Value | |
| Kocuria | 0.00% | 0.17% | 0.022 |
| Rothia | 0.00% | 0.36% | 0.017 |
| Cutibacterium | 1.15% | 4.72% | 0.036 |
| Streptococcus | 0.17% | 1.64% | 0.046 |
| Methylobacterium-Methylorubrum | 0.00% | 0.14% | 0.022 |
| Tepidimonas | 0.00% | 0.25% | 0.046 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Brożek-Mądry, E.; Burska, Z.; Życińska, K.; Sierdziński, J. Nasal Microbiome in Granulomatosis with Polyangiitis Compared to Chronic Rhinosinusitis. Diagnostics 2024, 14, 1673. https://doi.org/10.3390/diagnostics14151673
Brożek-Mądry E, Burska Z, Życińska K, Sierdziński J. Nasal Microbiome in Granulomatosis with Polyangiitis Compared to Chronic Rhinosinusitis. Diagnostics. 2024; 14(15):1673. https://doi.org/10.3390/diagnostics14151673
Chicago/Turabian StyleBrożek-Mądry, Eliza, Zofia Burska, Katarzyna Życińska, and Janusz Sierdziński. 2024. "Nasal Microbiome in Granulomatosis with Polyangiitis Compared to Chronic Rhinosinusitis" Diagnostics 14, no. 15: 1673. https://doi.org/10.3390/diagnostics14151673





