Similar Gut Bacterial Microbiota in Two Fruit-Feeding Moth Pests Collected from Different Host Species and Locations
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection and DNA Extraction
2.2. 16S rRNA Gene Amplification and Sequencing
2.3. Quality Control and OTU Identification
2.4. Diversity Analysis
3. Results
3.1. Community Composition of the Gut Microbiota in PFM and OFM
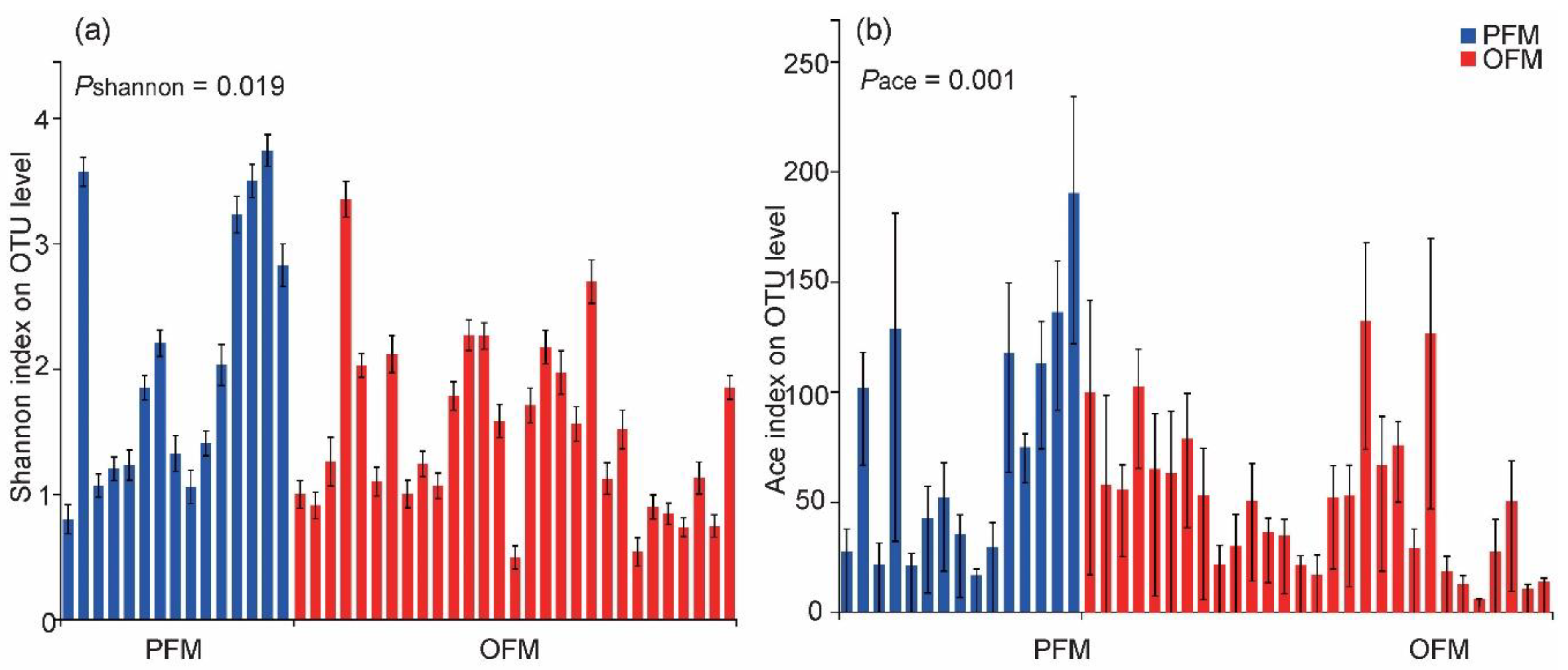
3.2. Comparison on Gut Microbiota between PFM and OFM
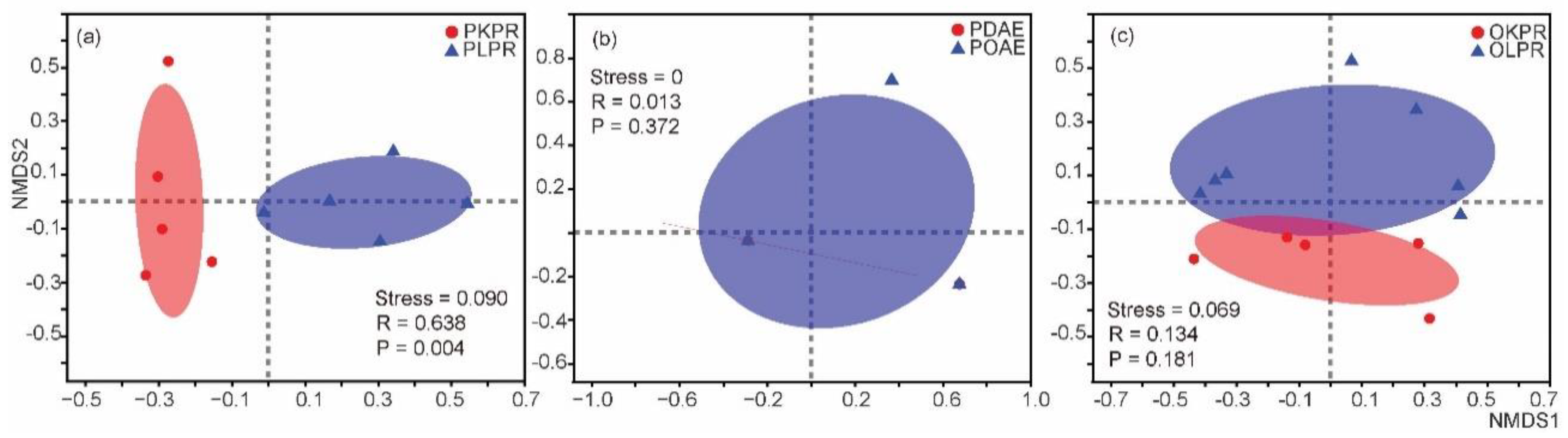
3.3. Influence of Orchard on Gut Microbiota within Species
4. Discussion
4.1. Comparison of Gut Microbiota from Two Fruit Borers
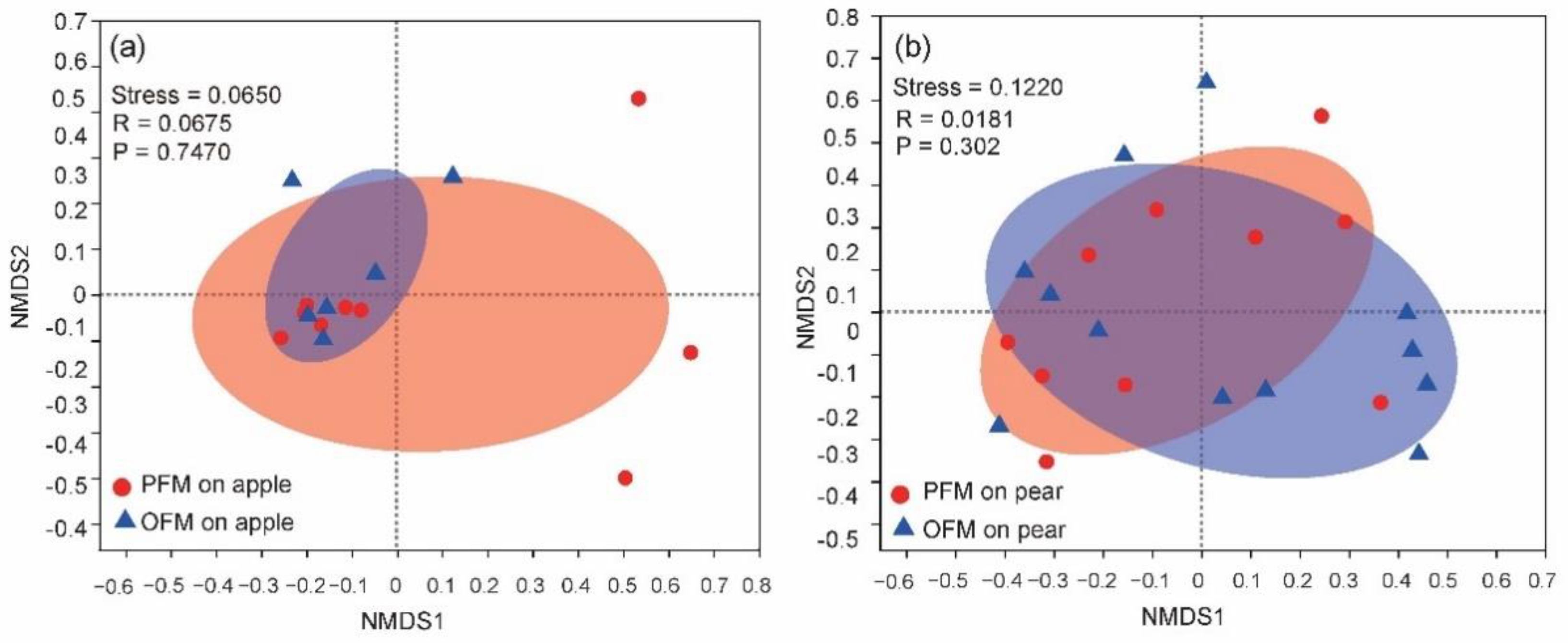
4.2. Influence of Orchard and Host Species on Gut Microbiota
4.3. Wolbachia in PFM and OFM
4.4. Implications for Pest Management
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Shapira, M. Gut microbiotas and host evolution: Scaling up symbiosis. Trends Ecol. Evol. 2016, 31, 539–549. [Google Scholar] [CrossRef]
- Dillon, R.; Dillon, V. The gut bacteria of insects: Nonpathogenic interactions. Annu. Rev. Entomol. 2004, 49, 71–92. [Google Scholar] [CrossRef]
- Engel, P.; Moran, N.A. The gut microbiota of insects–diversity in structure and function. FEMS Microbiol. Rev. 2013, 37, 699–735. [Google Scholar] [CrossRef] [PubMed]
- Douglas, A.E. Multiorganismal insects: Diversity and function of resident microorganisms. Annu. Rev. Entomol. 2015, 60, 17–34. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dicke, M.; Cusumano, A.; Poelman, E.H. Microbial symbionts of parasitoids. Annu. Rev. Entomol. 2020, 65, 171–190. [Google Scholar] [CrossRef] [PubMed]
- Berasategui, A.; Shukla, S.; Salem, H.; Kaltenpoth, M. Potential applications of insect symbionts in biotechnology. Appl. Microbiol. Biotechnol. 2016, 100, 1567–1577. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Walker, T.; Johnson, P.H.; Moreira, L.A.; Iturbe-Ormaetxe, I.; Frentiu, F.D.; McMeniman, C.J.; Leong, Y.S.; Dong, Y.; Axford, J.; Kriesner, P.; et al. The wMel Wolbachia strain blocks dengue and invades caged Aedes aegypti populations. Nature 2011, 476, 450–453. [Google Scholar] [CrossRef]
- Hansen, A.K.; Moran, N.A. The impact of microbial symbionts on host plant utilization by herbivorous insects. Mol. Ecol. 2014, 23, 1473–1496. [Google Scholar] [CrossRef]
- Adair, K.L.; Douglas, A.E. Making a microbiome: The many determinants of host-associated microbial community composition. Curr. Opin. Microbiol. 2017, 35, 23–29. [Google Scholar] [CrossRef]
- Szelagiewicz, M.; Marcolli, C.; Cianferani, S.; Hard, A.P.; Vit, A.; Burkhard, A.; von Raumer, M.; Hofmeier, U.C.; Zilian, A.; Francotte, E.; et al. In situ characterization of polymorphic forms the potential of Raman techniques. J. Therm. Anal. Calorim. 1999, 57, 23–43. [Google Scholar] [CrossRef]
- Priya, N.G.; Ojha, A.; Kajla, M.K.; Raj, A.; Rajagopal, R. Host plant induced variation in gut bacteria of Helicoverpa armigera. PLoS ONE 2012, 7, e30768. [Google Scholar]
- Renoz, F.; Pons, I.; Vanderpoorten, A.; Bataille, G.; Noel, C.; Foray, V.; Pierson, V.; Hance, T. Evidence for gut-associated Serratia symbiotica in wild aphids and ants provides new perspectives on the evolution of bacterial mutualism in insects. Microb. Ecol. 2019, 78, 159–169. [Google Scholar] [CrossRef]
- Hammer, T.J.; Janzen, D.H.; Hallwachs, W.; Jaffe, S.P.; Fierer, N. Caterpillars lack a resident gut microbiome. Proc. Natl. Acad. Sci. USA 2017, 114, 9641–9646. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mereghetti, V.; Chouaia, B.; Limonta, L.; Locatelli, D.P.; Montagna, M. Evidence for a conserved microbiota across the different developmental stages of Plodia interpunctella. Insect Sci. 2019, 26, 466–478. [Google Scholar] [CrossRef] [PubMed]
- Staudacher, H.; Kaltenpoth, M.; Breeuwer, J.A.; Menken, S.B.; Heckel, D.G.; Groot, A.T. Variability of bacterial communities in the moth Heliothis virescens indicates transient association with the host. PLoS ONE 2016, 11, e0154514. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, X.L.; Pan, Q.J.; Tian, H.G.; Douglas, A.E.; Liu, T.X. Bacteria abundance and diversity of different life stages of Plutella xylostella (Lepidoptera: Plutellidae), revealed by bacteria culture-dependent and PCR-DGGE methods. Insect Sci. 2015, 22, 375–385. [Google Scholar] [CrossRef] [PubMed]
- Montagna, M.; Mereghetti, V.; Gargari, G.; Guglielmetti, S.; Faoro, F.; Lozzia, G.; Locatelli, D.; Limonta, L. Evidence of a bacterial core in the stored products pest Plodia interpunctella: The influence of different diets. Environ. Microbiol. 2016, 18, 4961–4973. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mereghetti, V.; Chouaia, B.; Montagna, M. New insights into the microbiota of moth pests. Int. J. Mol. Sci. 2017, 18, 2450. [Google Scholar] [CrossRef] [Green Version]
- Jones, A.G.; Mason, C.J.; Felton, G.W.; Hoover, K. Host plant and population source drive diversity of microbial gut communities in two polyphagous insects. Sci. Rep. 2019, 9, 2792. [Google Scholar] [CrossRef] [Green Version]
- Paniagua Voirol, L.R.; Frago, E.; Kaltenpoth, M.; Hilker, M.; Fatouros, N.E. Bacterial symbionts in Lepidoptera: Their diversity, transmission, and impact on the host. Front. Microbiol. 2018, 9, 556. [Google Scholar] [CrossRef]
- Ma, G.; Tian, B.L.; Zhao, F.; Wei, G.S.; Hoffmann, A.A.; Ma, C.S. Soil moisture conditions determine phenology and success of larval escape in the peach fruit moth, Carposina sasakii (Lepidoptera, Carposinidae): Implications for predicting drought effects on a diapausing insect. Appl. Soil Ecol. 2017, 110, 65–72. [Google Scholar] [CrossRef]
- Quan, L.F.; Qiu, G.S.; Zhang, H.J.; Sun, L.N.; Li, Y.Y.; Yan, W.T. Sublethal concentration of beta-cypermethrin influences fecundity and mating behavior of Carposina sasakii (Lepidoptera: Carposinidae) adults. J. Econ. Entomol. 2016, 109, 2196–2204. [Google Scholar] [CrossRef] [PubMed]
- Rothschild, G.H.L.; Vickers, R.A. Biology, ecology and control of the oriental fruit moth. In World Crop Pests. Tortricid Pests: Their Biology, Natural Enemies and Control; van der Geest, L., Evenhuis, H., Eds.; Elsevier: Amsterdam, The Netherlands, 1991; Volume 5, pp. 389–412. [Google Scholar]
- Wang, Y.Z.; Li, B.Y.; Hoffmann, A.A.; Cao, L.J.; Gong, Y.J.; Song, W.; Zhu, J.Y.; Wei, S.J. Patterns of genetic variation among geographic and host-plant associated populations of the peach fruit moth Carposina sasakii (Lepidoptera: Carposinidae). BMC Evol. Biol. 2017, 17, 265. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kwon, D.; Kwon, H.; Kim, D.; Yang, C. Larval species composition and genetic structures of Carposina sasakii, Grapholita dimorpha, and Grapholita molesta from Korea. Bull. Entomol. Res. 2018, 108, 241–252. [Google Scholar] [CrossRef] [PubMed]
- Kwon, D.H.; Kim, D.H.; Kim, H.H.; Lee, S.H.; Yang, C.Y. Genetic diversity and structure in apple-infesting pests of Carposina sasakii, Grapholita dimorpha and Grapholita molesta in Korea. J. Asia-Pac. Entomol. 2017, 20, 13–16. [Google Scholar] [CrossRef]
- Sarker, S.; Lim, U.T. Development and fecundity performance of Grapholita molesta and Grapholita dimorpha (Lepidoptera: Tortricidae) on different immature fruits. PLoS ONE 2019, 14, e0217492. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Sun, L.; Tian, Z.; Haibin, H.; Zhang, H.; Guisheng, Q.; Yan, W.; Yue, Q. Gut bacterial community diversity in Carposina sasakii and Grapholitha molesta. Chin. J. Appl. Ecol. 2018, 29, 293–300. [Google Scholar]
- Liu, Y.; Shen, Z.; Yu, J.; Li, Z.; Liu, X.; Xu, H. Comparison of gut bacterial communities and their associations with host diets in four fruit borers. Pest Manag. Sci. 2020, 76, 1353–1362. [Google Scholar] [CrossRef]
- Kozich, J.J.; Westcott, S.L.; Baxter, N.T.; Highlander, S.K.; Schloss, P.D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl. Environ. Microbiol. 2013, 79, 5112–5120. [Google Scholar] [CrossRef] [Green Version]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc (accessed on 27 November 2020).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [Green Version]
- Magoc, T.; Salzberg, S.L. FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 2011, 27, 2957–2963. [Google Scholar] [CrossRef] [PubMed]
- Fondon, J.W., III; Martin, A.; Richards, S.; Gibbs, R.A.; Mittelman, D. Analysis of Microsatellite Variation in Drosophila melanogaster with Population-Scale Genome Sequencing. PLoS ONE 2012, 7, e33036. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. UPARSE: Highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 2013, 10, 996–998. [Google Scholar] [CrossRef]
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5567. [Google Scholar] [CrossRef] [Green Version]
- Yilmaz, P.; Parfrey, L.W.; Yarza, P.; Gerken, J.; Pruesse, E.; Quast, C.; Schweer, T.; Peplies, J.; Ludwig, W.; Glockner, F.O. The SILVA and “All-species Living Tree Project (LTP)” taxonomic frameworks. Nucleic Acids Res. 2014, 42, D643–D648. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef] [Green Version]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Pena, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [Green Version]
- Yun, J.H.; Roh, S.W.; Whon, T.W.; Jung, M.J.; Kim, M.S.; Park, D.S.; Yoon, C.; Nam, Y.D.; Kim, Y.J.; Choi, J.H.; et al. Insect gut bacterial diversity determined by environmental habitat, diet, developmental stage, and phylogeny of host. Appl. Environ. Microbiol. 2014, 80, 5254–5264. [Google Scholar] [CrossRef] [Green Version]
- Broderick, N.A.; Raffa, K.F.; Goodman, R.M.; Handelsman, J. Census of the bacterial community of the gypsy moth larval midgut by using culturing and culture-independent methods. Appl. Environ. Microbiol. 2004, 70, 293–300. [Google Scholar] [CrossRef] [Green Version]
- Hui, X.A.; Wei, G.F.; Jia, S.H.; Huang, J.H.; Miao, X.X.; Zhou, Z.; Zhao, L.P.; Huang, Y.P. Microbial communities in the larval midgut of laboratory and field populations of cotton bollworm (Helicoverpa armigera). Can. J. Microbiol. 2006, 52, 1085–1092. [Google Scholar]
- Chen, B.; Du, K.; Sun, C.; Vimalanathan, A.; Liang, X.; Li, Y.; Wang, B.; Lu, X.; Li, L.; Shao, Y. Gut bacterial and fungal communities of the domesticated silkworm (Bombyx mori) and wild mulberry-feeding relatives. ISME J. 2018, 12, 2252–2262. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xia, X.F.; Zheng, D.D.; Zhong, H.Z.; Qin, B.C.; Gurr, G.M.; Vasseur, L.; Lin, H.L.; Bai, J.L.; He, W.Y.; You, M.S. DNA sequencing reveals the midgut microbiota of diamondback moth, Plutella xylostella (L.) and a possible relationship with insecticide resistance. PLoS ONE 2013, 8, e68852. [Google Scholar] [CrossRef] [PubMed]
- Meriweather, M.; Matthews, S.; Rio, R.; Baucom, R.S. A 454 survey reveals the community composition and core microbiome of the common bed bug (Cimex lectularius) across an urban landscape. PLoS ONE 2013, 8, e61465. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chandler, J.A.; Lang, J.M.; Bhatnagar, S.; Eisen, J.A.; Kopp, A. Bacterial communities of diverse Drosophila species: Ecological context of a host-microbe model system. PLoS Genet. 2011, 7, e1002272. [Google Scholar] [CrossRef]
- Werren, J.H. Biology of Wolbachia. Annu. Rev. Entomol. 1997, 42, 587–609. [Google Scholar] [CrossRef] [Green Version]
- Ahmed, M.Z.; Araujo-Jnr, E.V.; Welch, J.J.; Kawahara, A.Y. Wolbachia in butterflies and moths: Geographic structure in infection frequency. Front. Zool. 2015, 12, 16. [Google Scholar] [CrossRef] [Green Version]
- Kageyama, D.; Nishimura, G.; Hoshizaki, S.; Ishikawa, Y. Feminizing Wolbachia in an insect, Ostrinia furnacalis (Lepidoptera: Crambidae). Heredity 2002, 88, 444. [Google Scholar] [CrossRef] [Green Version]
- Berec, L.; Maxin, D.; Bernhauerová, V.; Rohr, J. Male-killing bacteria as agents of insect pest control. J. Appl. Ecol. 2016, 53, 1270–1279. [Google Scholar] [CrossRef]
- Hurst, G.D.; Johnson, A.P.; Schulenburg, J.H.; Fuyama, Y. Male-killing Wolbachia in Drosophila: A temperature-sensitive trait with a threshold bacterial density. Genetics 2000, 156, 699–709. [Google Scholar]
- Kaiser, W.; Huguet, E.; Casas, J.; Commin, C.; Giron, D. Plant green-island phenotype induced by leaf-miners is mediated by bacterial symbionts. Proc. R. Soc. B Biol. Sci. 2010, 277, 2311–2319. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dittmer, J.; Bouchon, D. Feminizing Wolbachia influence microbiota composition in the terrestrial isopod Armadillidium vulgare. Sci. Rep. 2018, 8, 6998. [Google Scholar] [CrossRef] [PubMed]
- Simhadri, R.K.; Fast, E.M.; Guo, R.; Schultz, M.J.; Vaisman, N.; Ortiz, L.; Bybee, J.; Slatko, B.E.; Frydman, H.M. The gut commensal microbiome of Drosophila melanogaster is modified by the endosymbiont Wolbachia. mSphere 2017, 2. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shi, M.; White, V.L.; Schlub, T.; Eden, J.S.; Hoffmann, A.A.; Holmes, E.C. No detectable effect of Wolbachia wMel on the prevalence and abundance of the RNA virome of Drosophila melanogaster. Proc. R. Soc. B-Biol. Sci. 2018, 285, 1883. [Google Scholar]
- Bosch, T.J.; Welte, C.U. Detoxifying symbionts in agriculturally important pest insects. Microb. Biotechnol. 2017, 10, 531–540. [Google Scholar] [CrossRef] [PubMed]
- Itoh, H.; Tago, K.; Hayatsu, M.; Kikuchi, Y. Detoxifying symbiosis: Microbe-mediated detoxification of phytotoxins and pesticides in insects. Nat. Prod. Rep. 2018, 35, 434–454. [Google Scholar] [CrossRef]
- Banerjee, S.; Kamila, B.; Barman, S.; Joshi, S.R.; Mandal, T.; Haider, G. Interlining Cr(VI) remediation mechanism by a novel bacterium Pseudomonas brenneri isolated from coalmine wastewater. J. Environ. Manag. 2019, 233, 271–282. [Google Scholar] [CrossRef]
- He, M.Y.; Jiang, J.J.; Cheng, D.F. The plant pathogen Gluconobacter cerinus strain CDF1 is beneficial to the fruit fly Bactrocera dorsalis. Amb Express 2017, 7, 207. [Google Scholar] [CrossRef]
- Walterson, A.M.; Stavrinides, J. Pantoea: Insights into a highly versatile and diverse genus within the Enterobacteriaceae. FEMS Microbiol. Rev. 2015, 39, 968–984. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Q.; Acuna, J.J.; Inostroza, N.G.; Mora, M.L.; Radic, S.; Sadowsky, M.J.; Jorquera, M.A. Endophytic Bacterial Communities Associated with Roots and Leaves of Plants Growing in Chilean Extreme Environments. Sci. Rep. 2019, 9, 4950. [Google Scholar] [CrossRef] [Green Version]
- Bing, X.; Gerlach, J.; Loeb, G.; Buchon, N. Nutrient-dependent impact of microbes on Drosophila suzukii development. mBio 2018, 9. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schultz, J. Tannin-insect interactions. In Chemistry and Significance of Condensed Tannins; Hemingway, R.W., Karchesy, J.J., Branham, S.J., Eds.; Springer: Boston, MA, USA, 1989. [Google Scholar]
- Chung, S.H.; Parker, B.J.; Blow, F.; Brisson, J.A.; Douglas, A.E. Host and symbiont genetic determinants of nutritional phenotype in a natural population of the pea aphid. Mol. Ecol. 2020, 29, 848–858. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.H.; Shah, M.M.R.; Song, Y.; Liu, T.X. Host plant affects symbiont abundance in Bemisia tabaci (Hemiptera: Aleyrodidae). Insects 2020, 11, 501. [Google Scholar] [CrossRef] [PubMed]
- Ayayee, P.A.; Larsen, T.; Sabree, Z. Symbiotic essential amino acids provisioning in the American cockroach, Periplaneta americana (Linnaeus) under various dietary conditions. PeerJ 2016, 4, e2046. [Google Scholar] [CrossRef] [Green Version]
- Ayayee, P.A.; Larsen, T.; Rosa, C.; Felton, G.W.; Ferry, J.G.; Hoover, K. Essential amino acid supplementation by gut microbes of a wood-feeding cerambycid. Environ. Entomol. 2016, 45, 66–73. [Google Scholar] [CrossRef]
- Ceja-Navarro, J.A.; Vega, F.E.; Karaoz, U.; Hao, Z.; Jenkins, S.; Lim, H.C.; Kosina, P.; Infante, F.; Northen, T.R.; Brodie, E.L. Gut microbiota mediate caffeine detoxification in the primary insect pest of coffee. Nat. Commun. 2015, 6, 7618. [Google Scholar] [CrossRef] [Green Version]
- Vilanova, C.; Baixeras, J.; Latorre, A.; Porcar, M. The generalist inside the specialist: Gut bacterial communities of two insect species feeding on toxic plants are dominated by Enterococcus sp. Front. Microbiol. 2016, 7, 1005. [Google Scholar] [CrossRef]


| Code * | Species | Collecting Location | Host (code) | Coordinate | NO. |
|---|---|---|---|---|---|
| PKPR | PFM | Kaosanji of Pinggu district (K) | Pear (PR) | 40°12′ N, 117°19′ E | 5 |
| OKPR | OFM | 5 | |||
| PLPR | PFM | Lvfulong of Yanqing district (L) | Pear (PR) | 40°32′ N, 116°4′ E | 5 |
| OLPR | OFM | 7 | |||
| PGAE | PFM | Liugou of Yanqing district (G) | Apple (AE) | 40°27′ N, 116°6′ E | 4 |
| OGAE | OFM | 6 | |||
| PDAE | PFM | Dafengying of Yanqing district (D) | Apple (AE) | 40°26′ N, 115°54′ E | 5 |
| OYPH | OFM | Linguosuo of Haidian district (Y) | Peach shoot (PH) | 39°58′ N, 116°13′ E | 5 |
| OSPH | OFM | Shuangxin of Haidian district (S) | Peach shoot (PH) | 39°57′ N, 116°12′ E | 2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gong, Q.; Cao, L.-J.; Sun, L.-N.; Chen, J.-C.; Gong, Y.-J.; Pu, D.-Q.; Huang, Q.; Hoffmann, A.A.; Wei, S.-J. Similar Gut Bacterial Microbiota in Two Fruit-Feeding Moth Pests Collected from Different Host Species and Locations. Insects 2020, 11, 840. https://doi.org/10.3390/insects11120840
Gong Q, Cao L-J, Sun L-N, Chen J-C, Gong Y-J, Pu D-Q, Huang Q, Hoffmann AA, Wei S-J. Similar Gut Bacterial Microbiota in Two Fruit-Feeding Moth Pests Collected from Different Host Species and Locations. Insects. 2020; 11(12):840. https://doi.org/10.3390/insects11120840
Chicago/Turabian StyleGong, Qiang, Li-Jun Cao, Li-Na Sun, Jin-Cui Chen, Ya-Jun Gong, De-Qiang Pu, Qiong Huang, Ary Anthony Hoffmann, and Shu-Jun Wei. 2020. "Similar Gut Bacterial Microbiota in Two Fruit-Feeding Moth Pests Collected from Different Host Species and Locations" Insects 11, no. 12: 840. https://doi.org/10.3390/insects11120840








