Identification of Novel Toxin Genes from the Stinging Nettle Caterpillar Parasa lepida (Cramer, 1799): Insights into the Evolution of Lepidoptera Toxins
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Caterpillars and RNA Isolation
2.2. RNA Sequencing and Transcriptome Analysis
2.3. Identification of Toxin Genes
2.4. Gene Orthology and Phylogenetic Analysis
3. Results
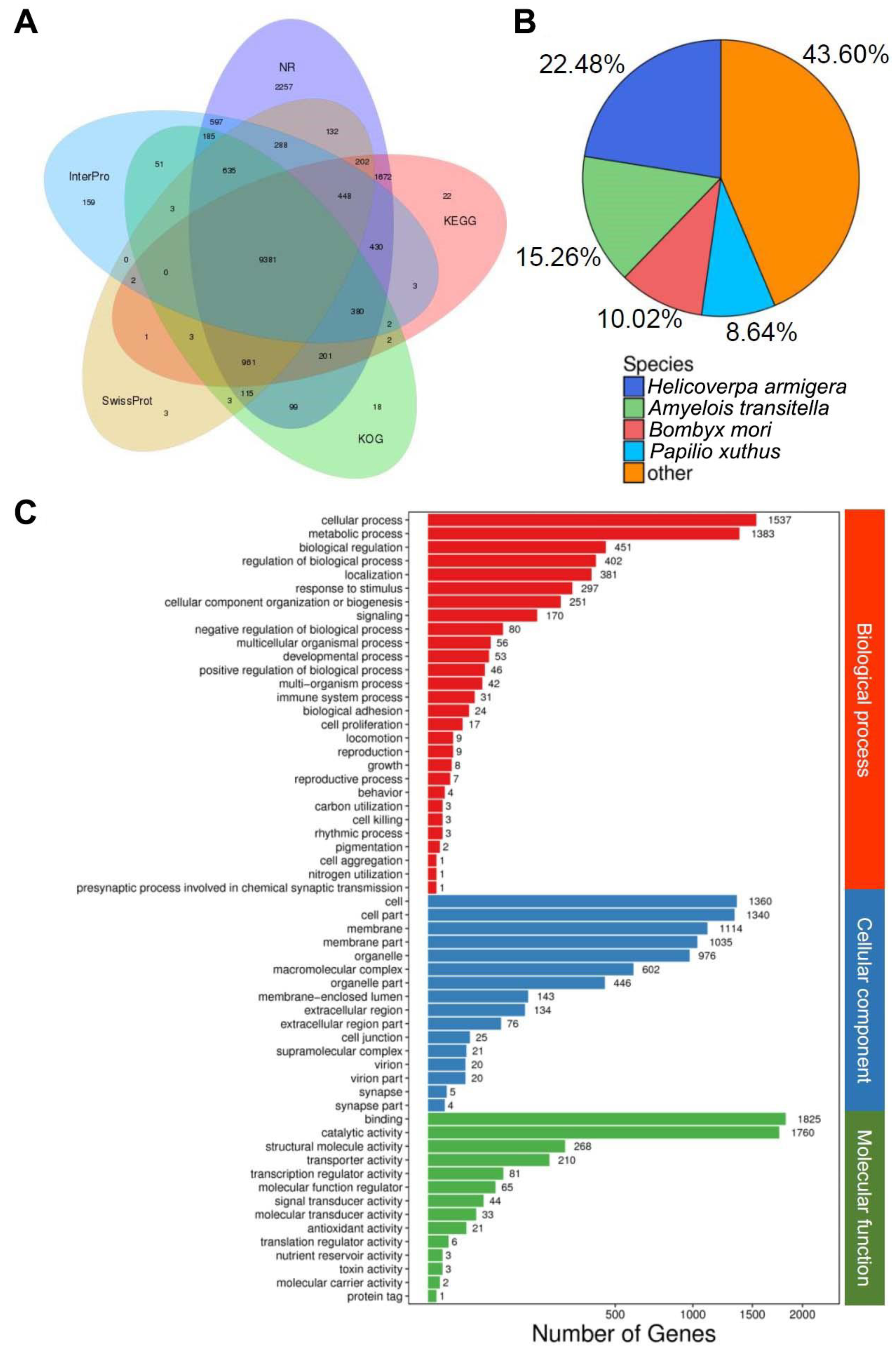
3.1. Transcriptome Assembly and Gene Annotation
3.2. Candidate Toxin Genes of Parasa lepida
3.3. Gene Family Expansion
3.4. The Expression Level of the Candidate Parasa lepida Toxin Genes
3.5. Putative Novel Toxin Proteins from P. lepida
4. Discussion
4.1. Predicted Functions of P. lepida Toxins
4.1.1. Knottin-Like Peptides
4.1.2. CAP Family
4.1.3. Venom Dipeptidyl Peptidase IV
4.1.4. Venom Acid Phosphatase
4.1.5. Carboxylesterase-6
4.1.6. Serine Proteases
4.1.7. Lipocalins
4.1.8. Serine Protease Inhibitor (Serpin)
4.1.9. Kazal-Type Protease Inhibitor
4.1.10. Kunitz-Type Protease Inhibitor
4.1.11. Trypsin Inhibitor
4.1.12. Serine Carboxypeptidase
4.1.13. C-Type Lectin
4.1.14. Phospholipase A2
4.1.15. Antimicrobial Peptides
4.2. P. lepida Toxins in Comparison with Other Venomous Lepidoptera
4.3. Evolution of P. lepida Toxin Genes
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Arbuckle, K. Evolutionary context of venom in animals. In Evolution of Venomous Animals and Their Toxins; Malhotra, A., Ed.; Toxinology; Springer: Dordrecht, The Netherlands, 2017; pp. 3–31. ISBN 978-94-007-6458-3. [Google Scholar]
- Schendel, V.; Rash, L.D.; Jenner, R.A.; Undheim, E.A.B. The diversity of venom: The importance of behavior and venom system morphology in understanding its ecology and evolution. Toxins 2019, 11, 666. [Google Scholar] [CrossRef] [Green Version]
- Fry, B.G.; Roelants, K.; Champagne, D.E.; Scheib, H.; Tyndall, J.D.A.; King, G.F.; Nevalainen, T.J.; Norman, J.A.; Lewis, R.J.; Norton, R.S.; et al. The toxicogenomic multiverse: Convergent recruitment of proteins into animal venoms. Annu. Rev. Genom. Human Genet. 2009, 10, 483–511. [Google Scholar] [CrossRef] [Green Version]
- Ericsson, C.D.; Hatz, C.; Junghanss, T.; Bodio, M. Medically important venomous animals: Biology, prevention, first aid, and clinical management. Clin. Infect. Dis. 2006, 43, 1309–1317. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ferraz, C.R.; Arrahman, A.; Xie, C.; Casewell, N.R.; Lewis, R.J.; Kool, J.; Cardoso, F.C. Multifunctional toxins in snake venoms and therapeutic implications: From pain to hemorrhage and necrosis. Front. Ecol. Evol. 2019, 7. [Google Scholar] [CrossRef] [Green Version]
- Primon-Barros, M.; José Macedo, A. Animal venom peptides: Potential for new antimicrobial agents. Curr. Top. Med. Chem. 2017, 17, 1119–1156. [Google Scholar] [CrossRef]
- Chatterjee, B. Animal venoms have potential to treat cancer. Curr. Top. Med. Chem. 2018, 18, 2555–2566. [Google Scholar] [CrossRef] [PubMed]
- Pennington, M.W.; Czerwinski, A.; Norton, R.S. Peptide therapeutics from venom: Current status and potential. Bioorganic Med. Chem. 2018, 26, 2738–2758. [Google Scholar] [CrossRef]
- King, G.F. Tying pest insects in knots: The deployment of spider-venom-derived knottins as bioinsecticides. Pest Manag. Sci. 2019, 75, 2437–2445. [Google Scholar] [CrossRef]
- Maatuf, Y.; Geron, M.; Priel, A. The role of toxins in the pursuit for novel analgesics. Toxins 2019, 11, 131. [Google Scholar] [CrossRef] [Green Version]
- Yang, X.; Wang, Y.; Wu, C.; Ling, E.-A. Animal venom peptides as a treasure trove for new therapeutics against neurodegenerative disorders. Curr. Med. Chem. 2019, 26, 4749–4774. [Google Scholar] [CrossRef] [PubMed]
- Gullan, P.J.; Cranston, P.S. The Insects: An Outline of Entomology; John Wiley & Sons: Hoboken, NJ, USA, 2014; ISBN 978-1-118-84615-5. [Google Scholar]
- Hossler, E.W. Caterpillars and moths. Dermatol. Ther. 2009, 22, 353–366. [Google Scholar] [CrossRef]
- Veiga, A.B.; Blochtein, B.; Guimarães, J.A. Structures Involved in production, secretion and injection of the venom produced by the caterpillar Lonomia obliqua (Lepidoptera, Saturniidae). Toxicon 2001, 39, 1343–1351. [Google Scholar] [CrossRef]
- Seldeslachts, A.; Peigneur, S.; Tytgat, J. Caterpillar venom: A health hazard of the 21st century. Biomedicines 2020, 8, 143. [Google Scholar] [CrossRef]
- Yao, Z.; Kamau, P.M.; Han, Y.; Hu, J.; Luo, A.; Luo, L.; Zheng, J.; Tian, Y.; Lai, R. The Latoia consocia caterpillar induces pain by targeting nociceptive Ion channel TRPV1. Toxins 2019, 11, 695. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kawamoto, F.; Kumada, N. Biology and venoms of Lepidoptera. In Handbook of Natural Toxins; Marcel Dekker: New York, NY, USA, 1984. [Google Scholar]
- Villas-Boas, I.M.; Bonfá, G.; Tambourgi, D.V. Venomous caterpillars: From inoculation apparatus to venom composition and envenomation. Toxicon 2018, 153, 39–52. [Google Scholar] [CrossRef]
- Villas-Boas, I.M.; Pidde, G.; Lichtenstein, F.; Ching, A.T.C.; de Junqueira-de-Azevedo, I.L.M.; DeOcesano-Pereira, C.; Madureira Trufen, C.E.; Chudzinski-Tavassi, A.M.; Morais, K.L.P.; Tambourgi, D.V. Human chondrocyte activation by toxins from Premolis semirufa, an amazon rainforest moth caterpillar: Identifying an osteoarthritis signature. Front. Immunol. 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Estrella, M.; Elston, D.M. What’s eating you? Megalopyge opercularis. Cutis 2020, 106, 23–24; 27. [Google Scholar] [CrossRef]
- Veiga, A.B.G.; Ribeiro, J.M.C.; Guimarães, J.A.; Francischetti, I.M.B. A catalog for the transcripts from the venomous structures of the caterpillar Lonomia obliqua: Identification of the proteins potentially involved in the coagulation disorder and hemorrhagic syndrome. Gene 2005, 355, 11–27. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Carrijo-Carvalho, L.C.; Chudzinski-Tavassi, A.M. The Venom of the Lonomia caterpillar: An overview. Toxicon 2007, 49, 741–757. [Google Scholar] [CrossRef]
- Quintana, M.A.; Sciani, J.M.; Auada, A.V.V.; Martínez, M.M.; Sánchez, M.N.; Santoro, M.L.; Fan, H.W.; Peichoto, M.E. Stinging caterpillars from the genera podalia, leucanella and lonomia in Misiones, Argentina: A preliminary comparative approach to understand their toxicity. Comp. Biochem. Physiol. C Toxicol. Pharmacol. 2017, 202, 55–62. [Google Scholar] [CrossRef] [PubMed]
- de Jong, M.C.J.M.; Kawamoto, F.; Bleumink, E.; Kloosterhuis, A.J.; Meijer, G.T. A comparative study of the spicule venom of euproctis caterpillars. Toxicon 1982, 20, 477–485. [Google Scholar] [CrossRef]
- Moneo, I.; Vega, J.M.; Caballero, M.L.; Vega, J.; Alday, E. Isolation and characterization of Tha p 1, a major allergen from the pine processionary caterpillar Thaumetopoea pityocampa. Allergy 2003, 58, 34–37. [Google Scholar] [CrossRef]
- Villas-Boas, I.M.; Gonçalves-de-Andrade, R.M.; Pidde-Queiroz, G.; Assaf, S.L.M.R.; Portaro, F.C.V.; Sant’Anna, O.A.; van den Berg, C.W.; Tambourgi, D.V. Premolis semirufa (Walker, 1856) envenomation, disease affecting rubber tappers of the amazon: Searching for caterpillar-bristles toxic components. PLoS Negl. Trop. Dis. 2012, 6, e1531. [Google Scholar] [CrossRef] [Green Version]
- Pidde, G.; Nishiyama, M.Y.; de Oliveira, U.C.; Villas-Boas, I.M.; Paes-Leme, A.F.; Junqueira-de-Azevedo, I.L.; Marques-Porto, R.; Squaiella-Baptistão, C.C.; Tambourgi, D.V. Integrative multiomics analysis of premolis semirufa caterpillar venom in the search for molecules leading to a joint disease. Sci. Rep. 2021, 11, 1995. [Google Scholar] [CrossRef]
- Rodriguez-Mahillo, A.I.; Gonzalez-Muñoz, M.; Vega, J.M.; López, J.A.; Yart, A.; Kerdelhué, C.; Camafeita, E.; Ortiz, J.C.G.; Vogel, H.; Toffolo, E.P.; et al. Setae from the pine processionary moth (Thaumetopoea pityocampa) contain several relevant allergens. Contact Dermat. 2012, 67, 367–374. [Google Scholar] [CrossRef]
- Kapoor, K.; Deobhakta, S.R.; Dhamdhere, S. Bionomics of the Slug Caterpillar, Latoia lepida (Cramer) (Lepidoptera: Limacodidae) on Mango. J. Entomol. Res. 1985, 9, 235–236. [Google Scholar]
- Arumugam, G.; Karuppiah, H.; Sreeramulu, B.; Paulchamy, R.; Sundaram, J. Occurrence of natural lectin with bacterial agglutination property in the serum of lepidopteran pest, Parasa lepida. Entomol. Sci. 2019, 22, 239–249. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [Green Version]
- Haas, B.J.; Papanicolaou, A.; Yassour, M.; Grabherr, M.; Blood, P.D.; Bowden, J.; Couger, M.B.; Eccles, D.; Li, B.; Lieber, M.; et al. De novo transcript sequence reconstruction from rna-seq using the trinity platform for reference generation and analysis. Nat. Protoc. 2013, 8, 1494–1512. [Google Scholar] [CrossRef]
- Pertea, G.; Huang, X.; Liang, F.; Antonescu, V.; Sultana, R.; Karamycheva, S.; Lee, Y.; White, J.; Cheung, F.; Parvizi, B.; et al. TIGR Gene Indices Clustering Tools (TGICL): A software system for fast clustering of large EST datasets. Bioinformatics 2003, 19, 651–652. [Google Scholar] [CrossRef] [Green Version]
- Conesa, A.; Götz, S.; García-Gómez, J.M.; Terol, J.; Talón, M.; Robles, M. Blast2GO: A universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 2005, 21, 3674–3676. [Google Scholar] [CrossRef] [Green Version]
- Langmead, B.; Salzberg, S.L. Fast Gapped-Read Alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [Green Version]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef] [Green Version]
- Verdes, A.; Simpson, D.; Holford, M. Are fireworms venomous? Evidence for the convergent evolution of toxin homologs in three species of fireworms (Annelida, Amphinomidae). Genome. Biol. Evol. 2017, 10, 249–268. [Google Scholar] [CrossRef] [Green Version]
- von Reumont, B.M.; Undheim, E.A.B.; Jauss, R.-T.; Jenner, R.A. Venomics of remipede crustaceans reveals novel peptide diversity and illuminates the venom’s biological role. Toxins 2017, 9, 234. [Google Scholar] [CrossRef] [Green Version]
- Chudzinski-Tavassi, A.M.; Alvarez Flores, M.P. Exploring new molecules and activities from Lonomia obliqua caterpillars. Pathophysiol. Haemost. Thromb. 2005, 34, 228–233. [Google Scholar] [CrossRef]
- Quevillon, E.; Silventoinen, V.; Pillai, S.; Harte, N.; Mulder, N.; Apweiler, R.; Lopez, R. InterProScan: Protein domains identifier. Nucleic Acids Res. 2005, 33, W116–W120. [Google Scholar] [CrossRef] [Green Version]
- Xu, L.; Dong, Z.; Fang, L.; Luo, Y.; Wei, Z.; Guo, H.; Zhang, G.; Gu, Y.Q.; Coleman-Derr, D.; Xia, Q.; et al. OrthoVenn2: A web server for whole-genome comparison and annotation of orthologous clusters across multiple species. Nucleic Acids Res. 2019, 47, W52–W58. [Google Scholar] [CrossRef] [Green Version]
- Guindon, S.; Dufayard, J.-F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New algorithms and methods to estimate maximum-likelihood phylogenies: Assessing the performance of PhyML 3.0. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef] [Green Version]
- Ramírez, D.; Gonzalez, W.; Fissore, R.A.; Carvacho, I. Conotoxins as tools to understand the physiological function of voltage-gated calcium (CaV) channels. Mar. Drugs 2017, 15, 313. [Google Scholar] [CrossRef] [Green Version]
- Takeuchi, K.; Park, E.; Lee, C.W.; Kim, J.I.; Takahashi, H.; Swartz, K.; Shimada, I. Solution structure of ω-grammotoxin SIA, A gating modifier of P/Q and N-Type Ca2+ Channel. J. Mol. Biol. 2002, 321, 517–526. [Google Scholar] [CrossRef]
- Xiao, Y.-C.; Liang, S.-P. Purification and characterization of hainantoxin-V, a tetrodotoxin-sensitive sodium channel inhibitor from the venom of the spider Selenocosmia hainana. Toxicon 2003, 41, 643–650. [Google Scholar] [CrossRef]
- Xiao, L.; Gurrola, G.B.; Zhang, J.; Valdivia, C.R.; SanMartin, M.; Zamudio, F.Z.; Zhang, L.; Possani, L.D.; Valdivia, H.H. Structure–function relationships of peptides forming the calcin family of ryanodine receptor ligands. J. Gen. Physiol. 2016, 147, 375–394. [Google Scholar] [CrossRef] [Green Version]
- Fajloun, Z.; Kharrat, R.; Chen, L.; Lecomte, C.; Di Luccio, E.; Bichet, D.; El Ayeb, M.; Rochat, H.; Allen, P.D.; Pessah, I.N.; et al. Chemical synthesis and characterization of maurocalcine, a scorpion toxin that activates Ca(2+) release channel/ryanodine receptors. FEBS Lett. 2000, 469, 179–185. [Google Scholar] [CrossRef] [Green Version]
- Osteen, J.D.; Herzig, V.; Gilchrist, J.; Emrick, J.J.; Zhang, C.; Wang, X.; Castro, J.; Garcia-Caraballo, S.; Grundy, L.; Rychkov, G.Y.; et al. Selective spider toxins reveal a role for Nav1.1 channel in mechanical pain. Nature 2016, 534, 494–499. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gibbs, G.M.; Roelants, K.; O’Bryan, M.K. The CAP superfamily: Cysteine-rich secretory proteins, antigen 5, and pathogenesis-related 1 proteins--roles in reproduction, cancer, and immune defense. Endocr. Rev. 2008, 29, 865–897. [Google Scholar] [CrossRef] [PubMed]
- Pantera, B.; Hoffman, D.R.; Carresi, L.; Cappugi, G.; Turillazzi, S.; Manao, G.; Severino, M.; Spadolini, I.; Orsomando, G.; Moneti, G.; et al. Characterization of the major allergens purified from the venom of the paper wasp Polistes gallicus. Biochim. Biophys. Acta (BBA) Gen. Subj. 2003, 1623, 72–81. [Google Scholar] [CrossRef] [PubMed]
- Müller, U.R.; Johansen, N.; Petersen, A.B.; Fromberg-Nielsen, J.; Haeberli, G. Hymenoptera venom allergy: Analysis of double positivity to honey bee and vespula venom by estimation of IgE antibodies to species-specific major allergens Api M1 and Ves V5. Allergy 2009, 64, 543–548. [Google Scholar] [CrossRef] [PubMed]
- Torres, A.F.C.; Huang, C.; Chong, C.-M.; Leung, S.W.; Prieto-da-Silva, Á.R.B.; Havt, A.; Quinet, Y.P.; Martins, A.M.C.; Lee, S.M.Y.; Rádis-Baptista, G. Transcriptome analysis in venom gland of the predatory giant ant Dinoponera quadriceps: Insights into the polypeptide toxin arsenal of hymenopterans. PLoS ONE 2014, 9, e87556. [Google Scholar] [CrossRef] [Green Version]
- Yamazaki, Y.; Morita, T. Structure and function of snake venom cysteine-rich secretory proteins. Toxicon 2004, 44, 227–231. [Google Scholar] [CrossRef]
- Milne, T.J.; Abbenante, G.; Tyndall, J.D.A.; Halliday, J.; Lewis, R.J. Isolation and characterization of a cone snail protease with homology to CRISP proteins of the pathogenesis-related protein superfamily. J. Biol. Chem. 2003, 278, 31105–31110. [Google Scholar] [CrossRef] [Green Version]
- Blank, S.; Seismann, H.; Bockisch, B.; Braren, I.; Cifuentes, L.; McIntyre, M.; Rühl, D.; Ring, J.; Bredehorst, R.; Ollert, M.W.; et al. Identification, recombinant expression, and characterization of the 100 KDa high molecular weight hymenoptera venom allergens Api m 5 and Ves v 3. J. Immunol. 2010, 184, 5403–5413. [Google Scholar] [CrossRef] [Green Version]
- Dani, M.P.; Richards, E.H.; Isaac, R.E.; Edwards, J.P. Antibacterial and proteolytic activity in venom from the endoparasitic wasp Pimpla hypochondriaca (Hymenoptera: Ichneumonidae). J. Insect Physiol. 2003, 49, 945–954. [Google Scholar] [CrossRef]
- Robinson, S.D.; Mueller, A.; Clayton, D.; Starobova, H.; Hamilton, B.R.; Payne, R.J.; Vetter, I.; King, G.F.; Undheim, E.A.B. A comprehensive portrait of the venom of the giant red bull ant, Myrmecia gulosa, reveals a hyperdiverse hymenopteran toxin gene family. Sci. Adv. 2018, 4, eaau4640. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schiener, M.; Hilger, C.; Eberlein, B.; Pascal, M.; Kuehn, A.; Revets, D.; Planchon, S.; Pietsch, G.; Serrano, P.; Moreno-Aguilar, C.; et al. The high molecular weight dipeptidyl peptidase IV Pol d 3 is a major allergen of Polistes dominula Venom. Sci. Rep. 2018, 8, 1318. [Google Scholar] [CrossRef] [PubMed]
- Matysiak, J.; Hajduk, J.; Pietrzak, Ł.; Schmelzer, C.E.H.; Kokot, Z.J. Shotgun proteome analysis of honeybee venom using targeted enrichment strategies. Toxicon 2014, 90, 255–264. [Google Scholar] [CrossRef]
- Pushparani, D.S. High acid phosphatase level in the gingival tissues of periodontitis subjects. J. Basic Clin. Pharm. 2015, 6, 59–63. [Google Scholar] [CrossRef] [Green Version]
- Zhu, J.; Ye, G.; Hu, C. Molecular cloning and characterization of acid phosphatase in venom of the endoparasitoid wasp Pteromalus puparum (Hymenoptera: Pteromalidae). Toxicon 2008, 51, 1391–1399. [Google Scholar] [CrossRef]
- Bonasio, R.; Zhang, G.; Ye, C.; Mutti, N.S.; Fang, X.; Qin, N.; Donahue, G.; Yang, P.; Li, Q.; Li, C.; et al. Genomic comparison of the ants Camponotus floridanus and Harpegnathos saltator. Science 2010, 329, 1068–1071. [Google Scholar] [CrossRef] [Green Version]
- Dhananjaya, B.L.; D’Souza, C.J.M. The pharmacological role of phosphatases (Acid and Alkaline Phosphomonoesterases) in snake venoms related to release of purines-a Multitoxin. Basic Clin. Pharmacol. Toxicol. 2011, 108, 79–83. [Google Scholar] [CrossRef]
- Heavner, M.E.; Gueguen, G.; Rajwani, R.; Pagan, P.E.; Small, C.; Govind, S. Partial venom gland transcriptome of a drosophila parasitoid wasp, Leptopilina heterotoma, reveals novel and shared bioactive profiles with stinging hymenoptera. Gene 2013, 526, 195–204. [Google Scholar] [CrossRef] [Green Version]
- Spillner, E.; Blank, S.; Jakob, T. Hymenoptera allergens: From venom to “venome”. Front. Immunol. 2014, 5. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Perkin, L.C.; Friesen, K.S.; Flinn, P.W.; Oppert, B. Venom gland components of the ectoparasitoid wasp, Anisopteromalus calandrae. J. Venom. Res. 2015, 6, 19–37. [Google Scholar]
- Chai, L.; Yang, X.; Liu, M.; Liu, C.; Han, L.; Guo, H.; Li, C.; Sun, Y.; Li, X.; Xiao, M.; et al. Biopanning of allergens from wasp sting patients. Biosci. Rep. 2018, 38, BSR20181113. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chertemps, T.; François, A.; Durand, N.; Rosell, G.; Dekker, T.; Lucas, P.; Maïbèche-Coisne, M. A carboxylesterase, esterase-6, modulates sensory physiological and behavioral response dynamics to pheromone in Drosophila. BMC Biol. 2012, 10, 56. [Google Scholar] [CrossRef] [Green Version]
- Whittington, C.M.; Papenfuss, A.T.; Locke, D.P.; Mardis, E.R.; Wilson, R.K.; Abubucker, S.; Mitreva, M.; Wong, E.S.; Hsu, A.L.; Kuchel, P.W.; et al. Novel venom gene discovery in the platypus. Genome Biol. 2010, 11, R95. [Google Scholar] [CrossRef] [Green Version]
- Vaiyapuri, S.; Wagstaff, S.C.; Harrison, R.A.; Gibbins, J.M.; Hutchinson, E.G. Evolutionary analysis of novel serine proteases in the venom gland transcriptome of Bitis gabonica rhinoceros. PLoS ONE 2011, 6, e21532. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ruder, T.; Sunagar, K.; Undheim, E.A.B.; Ali, S.A.; Wai, T.-C.; Low, D.H.W.; Jackson, T.N.W.; King, G.F.; Antunes, A.; Fry, B.G. Molecular phylogeny and evolution of the proteins encoded by coleoid (Cuttlefish, Octopus, and Squid) posterior venom glands. J. Mol. Evol. 2013, 76, 192–204. [Google Scholar] [CrossRef]
- Choo, Y.M.; Lee, K.S.; Yoon, H.J.; Kim, B.Y.; Sohn, M.R.; Roh, J.Y.; Je, Y.H.; Kim, N.J.; Kim, I.; Woo, S.D.; et al. Dual function of a bee venom serine protease: Prophenoloxidase-activating factor in arthropods and fibrin(Ogen)olytic enzyme in mammals. PLoS ONE 2010, 5, e10393. [Google Scholar] [CrossRef]
- Ribeiro, J.M.C.; Genta, F.A.; Sorgine, M.H.F.; Logullo, R.; Mesquita, R.D.; Paiva-Silva, G.O.; Majerowicz, D.; Medeiros, M.; Koerich, L.; Terra, W.R.; et al. An insight into the transcriptome of the digestive tract of the bloodsucking bug, Rhodnius prolixus. PLoS Negl. Trop. Dis. 2014, 8, e2594. [Google Scholar] [CrossRef] [Green Version]
- von Reumont, B.M.; Blanke, A.; Richter, S.; Alvarez, F.; Bleidorn, C.; Jenner, R.A. The first venomous crustacean revealed by transcriptomics and functional morphology: Remipede venom glands express a unique toxin cocktail dominated by enzymes and a neurotoxin. Mol. Biol. Evol. 2014, 31, 48–58. [Google Scholar] [CrossRef] [Green Version]
- Nakhleh, J.; El Moussawi, L.; Osta, M.A. Chapter Three-The melanization response in insect immunity. In Advances in Insect Physiology; Ligoxygakis, P., Ed.; Insect Immunity; Academic Press: Cambridge, MA, USA, 2017; Volume 52, pp. 83–109. [Google Scholar]
- Flower, D.R. The lipocalin protein family: Structure and function. Biochem. J. 1996, 318, 1–14. [Google Scholar] [CrossRef]
- Paddock, C.D.; McKerrow, J.H.; Hansell, E.; Foreman, K.W.; Hsieh, I.; Marshall, N. Identification, cloning, and recombinant expression of procalin, a major triatomine allergen. J. Immunol. 2001, 167, 2694–2699. [Google Scholar] [CrossRef] [Green Version]
- Low, D.H.W.; Sunagar, K.; Undheim, E.A.B.; Ali, S.A.; Alagon, A.C.; Ruder, T.; Jackson, T.N.W.; Pineda Gonzalez, S.; King, G.F.; Jones, A.; et al. Dracula’s children: Molecular evolution of vampire bat venom. J. Proteom. 2013, 89, 95–111. [Google Scholar] [CrossRef]
- Junqueira-de-Azevedo, I.L.M.; Campos, P.F.; Ching, A.T.C.; Mackessy, S.P. Colubrid venom composition: An-omics perspective. Toxins 2016, 8, 230. [Google Scholar] [CrossRef] [PubMed]
- Reis, C.V.; Portaro, F.C.; Andrade, S.A.; Fritzen, M.; Fernandes, B.L.; Sampaio, C.A.; Camargo, A.C.; Chudzinski-Tavassi, A.M. A prothrombin activator serine protease from the Lonomia obliqua caterpillar venom (Lopap) biochemical characterization. Thromb. Res. 2001, 102, 427–436. [Google Scholar] [CrossRef]
- Law, R.H.; Zhang, Q.; McGowan, S.; Buckle, A.M.; Silverman, G.A.; Wong, W.; Rosado, C.J.; Langendorf, C.G.; Pike, R.N.; Bird, P.I.; et al. An overview of the serpin superfamily. Genome Biol. 2006, 7, 216. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wong, E.S.W.; Belov, K. Venom evolution through gene duplications. Gene 2012, 496, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Wang, L.; Yang, L.; Qian, C.; Wei, G.; Dai, L.; Li, J.; Zhu, B.; Liu, C. Serpin-15 from Bombyx mori Inhibits Prophenoloxidase activation and expression of antimicrobial peptides. Dev. Comp. Immunol. 2015, 51, 22–28. [Google Scholar] [CrossRef]
- Sigle, L.T.; Ramalho-Ortigão, M. Kazal-type serine proteinase inhibitors in the midgut of Phlebotomus papatasi. Mem. Inst. Oswaldo Cruz. 2013, 108, 671–678. [Google Scholar] [CrossRef] [Green Version]
- Francischetti, I.M.B.; Assumpção, T.C.F.; Ma, D.; Li, Y.; Vicente, E.C.; Uieda, W.; Ribeiro, J.M.C. The “Vampirome”: Transcriptome and proteome analysis of the principal and accessory submaxillary glands of the vampire bat Desmodus rotundus, a Vector of human rabies. J. Proteom. 2013, 82, 288–319. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wan, H.; Lee, K.S.; Kim, B.Y.; Zou, F.M.; Yoon, H.J.; Je, Y.H.; Li, J.; Jin, B.R. A spider-derived kunitz-type serine protease inhibitor that acts as a plasmin inhibitor and an elastase inhibitor. PLoS ONE 2013, 8, e53343. [Google Scholar] [CrossRef] [PubMed]
- Zhao, R.; Dai, H.; Qiu, S.; Li, T.; He, Y.; Ma, Y.; Chen, Z.; Wu, Y.; Li, W.; Cao, Z. SdPI, The first functionally characterized kunitz-type trypsin inhibitor from scorpion venom. PLoS ONE 2011, 6, e27548. [Google Scholar] [CrossRef] [PubMed]
- Kozlov, S.A.; Andreev, Y.A.; Murashev, A.N.; Skobtsov, D.I.; D’yachenko, I.A.; Grishin, E.V. New polypeptide components from the Heteractis crispa sea anemone with analgesic activity. Russ. J. Bioorg. Chem. 2009, 35, 711. [Google Scholar] [CrossRef]
- Mourão, C.B.F.; Schwartz, E.F. Protease inhibitors from marine venomous animals and their counterparts in terrestrial venomous animals. Mar. Drugs 2013, 11, 2069–2112. [Google Scholar] [CrossRef] [Green Version]
- Yuan, C.-H.; He, Q.-Y.; Peng, K.; Diao, J.-B.; Jiang, L.-P.; Tang, X.; Liang, S.-P. Discovery of a distinct superfamily of kunitz-type toxin (KTT) from tarantulas. PLoS ONE 2008, 3, e3414. [Google Scholar] [CrossRef]
- Wang, H.; Wang, L.; Zhou, M.; Yang, M.; Ma, C.; Chen, T.; Zhang, Y.; Zeller, M.; Hornshaw, M.; Shaw, C. Functional peptidomics of amphibian skin secretion: A novel kunitz-type chymotrypsin inhibitor from the african hyperoliid frog, Kassina senegalensis. Biochimie 2012, 94, 891–899. [Google Scholar] [CrossRef]
- Zeng, X.-C.; Wang, S.-X.; Li, W.-X. Identification of BmKAPi, a novel type of scorpion venom peptide with peculiar disulfide bridge pattern from Buthus martensii Karsch. Toxicon 2002, 40, 1719–1722. [Google Scholar] [CrossRef]
- de Oliveira, U.C.; Candido, D.M.; Coronado Dorce, V.A.; de Junqueira-de-Azevedo, I.L.M. The transcriptome recipe for the venom cocktail of Tityus bahiensis scorpion. Toxicon 2015, 95, 52–61. [Google Scholar] [CrossRef]
- Li, R.; Zhang, L.; Fang, Y.; Han, B.; Lu, X.; Zhou, T.; Feng, M.; Li, J. Proteome and Phosphoproteome analysis of honeybee (Apis mellifera) venom collected from electrical stimulation and manual extraction of the venom gland. BMC Genom. 2013, 14, 766. [Google Scholar] [CrossRef] [Green Version]
- Zelensky, A.N.; Gready, J.E. The C-type lectin-like domain superfamily. FEBS J. 2005, 272, 6179–6217. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Feng, J.-L.; Jin, M.; Ren, Q.; Wang, W. C-type lectin (MrCTL) from the giant freshwater prawn Macrobrachium rosenbergii participates in innate immunity. Fish Shellfish Immunol. 2016, 58, 136–144. [Google Scholar] [CrossRef] [PubMed]
- Morita, T. Structures and functions of snake venom CLPs (C-Type Lectin-like Proteins) with anticoagulant-, procoagulant-, and platelet-modulating activities. Toxicon 2005, 45, 1099–1114. [Google Scholar] [CrossRef] [PubMed]
- Magalhães, G.S.; Junqueira-de-Azevedo, I.L.M.; Lopes-Ferreira, M.; Lorenzini, D.M.; Ho, P.L.; Moura-da-Silva, A.M. Transcriptome analysis of expressed sequence tags from the venom glands of the fish Thalassophryne nattereri. Biochimie 2006, 88, 693–699. [Google Scholar] [CrossRef] [PubMed]
- Kvist, S.; Oceguera-Figueroa, A.; Tessler, M.; Jiménez-Armenta, J.; Freeman, R.M., Jr.; Giribet, G.; Siddall, M.E. When predator becomes prey: Investigating the salivary transcriptome of the shark-feeding leech Pontobdella macrothela (Hirudinea: Piscicolidae). Zool. J. Linn. Soc. 2017, 179, 725–737. [Google Scholar] [CrossRef]
- Kini, R.M. Excitement ahead: Structure, function and mechanism of snake venom phospholipase A2 enzymes. Toxicon 2003, 42, 827–840. [Google Scholar] [CrossRef]
- Nevalainen, T.J.; Graham, G.G.; Scott, K.F. Antibacterial actions of secreted phospholipases A2. review. Biochim. Biophys. Acta 2008, 1781, 1–9. [Google Scholar] [CrossRef]
- Perez-Riverol, A.; Lasa, A.M.; dos Santos-Pinto, J.R.A.; Palma, M.S. Insect venom phospholipases A1 and A2: Roles in the envenoming process and allergy. Insect Biochem. Mol. Biol. 2019, 105, 10–24. [Google Scholar] [CrossRef]
- Li, M.; Al Souhail, Q.; Veerapandian, R.; Vediyappan, G.; Kanost, M. Investigation of an antifungal peptide, diapausin, from Manduca sexta. FASEB J. 2019, 33, 800.2. [Google Scholar] [CrossRef]
- Tanaka, H.; Sato, K.; Saito, Y.; Yamashita, T.; Agoh, M.; Okunishi, J.; Tachikawa, E.; Suzuki, K. Insect diapause-specific peptide from the leaf beetle has consensus with a putative iridovirus peptide. Peptides 2003, 24, 1327–1333. [Google Scholar] [CrossRef]
- Kouno, T.; Mizuguchi, M.; Tanaka, H.; Yang, P.; Mori, Y.; Shinoda, H.; Unoki, K.; Aizawa, T.; Demura, M.; Suzuki, K.; et al. The structure of a novel insect peptide explains its Ca2+ channel blocking and antifungal activities. Biochemistry 2007, 46, 13733–13741. [Google Scholar] [CrossRef] [PubMed]
- Faddeeva-Vakhrusheva, A.; Derks, M.F.L.; Anvar, S.Y.; Agamennone, V.; Suring, W.; Smit, S.; van Straalen, N.M.; Roelofs, D. Gene family evolution reflects adaptation to soil environmental stressors in the genome of the collembolan Orchesella cincta. Genome. Biol. Evol. 2016, 8, 2106–2117. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Al Souhail, Q.; Hiromasa, Y.; Rahnamaeian, M.; Giraldo, M.C.; Takahashi, D.; Valent, B.; Vilcinskas, A.; Kanost, M.R. Characterization and regulation of expression of an antifungal peptide from hemolymph of an insect, Manduca sexta. Dev. Comp. Immunol. 2016, 61, 258–268. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hoffmann, J.A. Innate immunity of insects. Curr. Opin. Immunol. 1995, 7, 4–10. [Google Scholar] [CrossRef]
- Carlsson, A.; Engström, P.; Palva, E.T.; Bennich, H. Attacin, an Antibacterial Protein from Hyalophora cecropia, Inhibits Synthesis of Outer Membrane Proteins in Escherichia coli by Interfering with Omp Gene Transcription. Infect. Immun. 1991, 59, 3040–3045. [Google Scholar] [CrossRef] [Green Version]
- Wu, Q.; Patočka, J.; Kuča, K. Insect antimicrobial peptides, a mini review. Toxins 2018, 10, 461. [Google Scholar] [CrossRef]
- Alvarez Flores, M.P.; Fritzen, M.; Reis, C.V.; Chudzinski-Tavassi, A.M. Losac, a factor X activator from Lonomia obliqua bristle extract: Its role in the pathophysiological mechanisms and cell survival. Biochem. Biophys. Res. Commun. 2006, 343, 1216–1223. [Google Scholar] [CrossRef]
- Alvarez Flores, M.P.; Zannin, M.; Chudzinski-Tavassi, A.M. New insight into the mechanism of Lonomia obliqua envenoming: Toxin involvement and molecular approach. Pathophysiol. Haemost. Thromb. 2010, 37, 1–16. [Google Scholar] [CrossRef]
- Suranse, V.; Srikanthan, A.; Sunagar, K. Animal venoms: Origin, diversity and evolution. In ELS; American Cancer Society: New York, NY, USA, 2018; pp. 1–20. ISBN 978-0-470-01590-2. [Google Scholar]
- Chang, D.; Duda, T.F. Extensive and continuous duplication facilitates rapid evolution and diversification of gene families. Mol. Biol. Evol. 2012, 29, 2019–2029. [Google Scholar] [CrossRef] [Green Version]
- Undheim, E.A.B.; Jones, A.; Clauser, K.R.; Holland, J.W.; Pineda, S.S.; King, G.F.; Fry, B.G. Clawing through evolution: Toxin diversification and convergence in the ancient lineage chilopoda (Centipedes). Mol. Biol. Evol. 2014, 31, 2124–2148. [Google Scholar] [CrossRef] [Green Version]
- Hargreaves, A.D.; Swain, M.T.; Hegarty, M.J.; Logan, D.W.; Mulley, J.F. Restriction and recruitment—Gene duplication and the origin and evolution of snake venom toxins. Genome Biol. Evol. 2014, 6, 2088–2095. [Google Scholar] [CrossRef] [Green Version]
- Dowell, N.L.; Giorgianni, M.W.; Kassner, V.A.; Selegue, J.E.; Sanchez, E.E.; Carroll, S.B. The deep origin and recent loss of venom toxin genes in rattlesnakes. Curr. Biol. 2016, 26, 2434–2445. [Google Scholar] [CrossRef] [Green Version]
- Sunagar, K.; Moran, Y. The rise and fall of an evolutionary innovation: Contrasting strategies of venom evolution in ancient and young animals. PLoS Genet. 2015, 11, e1005596. [Google Scholar] [CrossRef] [Green Version]
- Malhotra, A. Mutation, duplication, and more in the evolution of venomous animals and their toxins. In Evolution of Venomous Animals and Their Toxins; Malhotra, A., Ed.; Toxinology; Springer: Dordrecht, The Netherlands, 2017; pp. 33–45. ISBN 978-94-007-6458-3. [Google Scholar]
- Sanggaard, K.W.; Bechsgaard, J.S.; Fang, X.; Duan, J.; Dyrlund, T.F.; Gupta, V.; Jiang, X.; Cheng, L.; Fan, D.; Feng, Y.; et al. Spider genomes provide insight into composition and evolution of venom and silk. Nat. Commun. 2014, 5, 3765. [Google Scholar] [CrossRef]
- Cao, Z.; Yu, Y.; Wu, Y.; Hao, P.; Di, Z.; He, Y.; Chen, Z.; Yang, W.; Shen, Z.; He, X.; et al. The genome of Mesobuthus martensii reveals a unique adaptation model of arthropods. Nat. Commun. 2013, 4. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Osaki, T.; Omotezako, M.; Nagayama, R.; Hirata, M.; Iwanaga, S.; Kasahara, J.; Hattori, J.; Ito, I.; Sugiyama, H.; Kawabata, S. Horseshoe crab hemocyte-derived antimicrobial polypeptides, tachystatins, with sequence similarity to spider neurotoxins. J. Biol. Chem. 1999, 274, 26172–26178. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shatters, R.G., Jr.; McKenzie, C.L.; Boykin, L.M.; Gazit, S.; Sinisterra, X.; Weathersbee, A.A., III; Brown, J.K.; Czosnek, H. A knottin-like putative antimicrobial gene family in the whitefly Bemisia tabaci biotype B: Cloning and transcript regulation. J. Insect Sci. 2008, 8, 44–45. [Google Scholar]
- Hariton Shalev, A.; Sobol, I.; Ghanim, M.; Liu, S.-S.; Czosnek, H. The whitefly Bemisia tabaci Knottin-1 Gene is implicated in regulating the quantity of tomato yellow leaf curl virus ingested and transmitted by the insect. Viruses 2016, 8, 205. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, C.-R.; Zhang, S.; Xia, J.; Li, F.-F.; Xia, W.-Q.; Liu, S.-S.; Wang, X.-W. The immune strategy and stress response of the mediterranean species of the Bemisia tabaci complex to an orally delivered bacterial pathogen. PLoS ONE 2014, 9, e94477. [Google Scholar] [CrossRef]






| Sequence Analysis | Counts | Length (bp) |
|---|---|---|
| Raw sequence reads acquired | 67,876,696 | 6,787,669,600 |
| Sequence reads after QC | 66,361,788 | 6,636,178,800 |
| Contigs after trinity assembly | 48,761 | 34,507,843 |
| Unique contigs (Unigene) | 34,968 | 27,807,171 |
| GC content of Unigene | 36.73% | |
| Average length | 795 | |
| N50 | 1315 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mitpuangchon, N.; Nualcharoen, K.; Boonrotpong, S.; Engsontia, P. Identification of Novel Toxin Genes from the Stinging Nettle Caterpillar Parasa lepida (Cramer, 1799): Insights into the Evolution of Lepidoptera Toxins. Insects 2021, 12, 396. https://doi.org/10.3390/insects12050396
Mitpuangchon N, Nualcharoen K, Boonrotpong S, Engsontia P. Identification of Novel Toxin Genes from the Stinging Nettle Caterpillar Parasa lepida (Cramer, 1799): Insights into the Evolution of Lepidoptera Toxins. Insects. 2021; 12(5):396. https://doi.org/10.3390/insects12050396
Chicago/Turabian StyleMitpuangchon, Natrada, Kwan Nualcharoen, Singtoe Boonrotpong, and Patamarerk Engsontia. 2021. "Identification of Novel Toxin Genes from the Stinging Nettle Caterpillar Parasa lepida (Cramer, 1799): Insights into the Evolution of Lepidoptera Toxins" Insects 12, no. 5: 396. https://doi.org/10.3390/insects12050396








