Transcriptomic Profiling of Bean Aphid Megoura crassicauda upon Exposure to the Aphid-Obligate Entomopathogen Conidiobolus obscurus (Entomophthoromycotina) and Screening of CytCo-Binding Aphid Proteins through a Pull-Down Assay
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Fungus Origin and Preparation
2.2. Aphid Inoculation Procedure
2.3. Sampling of Aphids and RNA Extraction
2.4. Transcriptome Analysis
2.5. The Real-Time Quantitative PCR (qPCR) Assay
2.6. Pull-Down Assay
2.7. Identification of the Potential CytCo-Binding Proteins by LC-MS/MS
3. Results
3.1. Transcriptome Assembly and Annotation
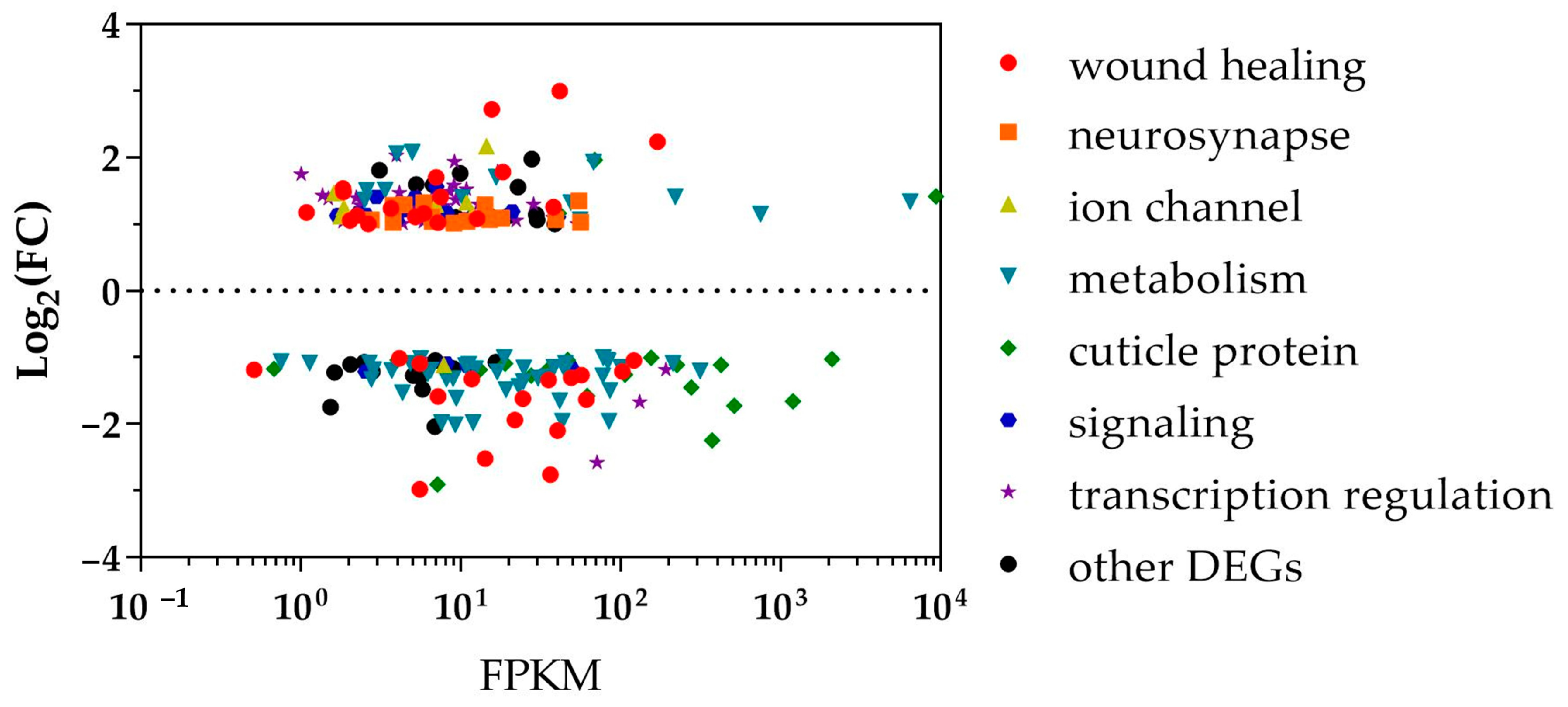
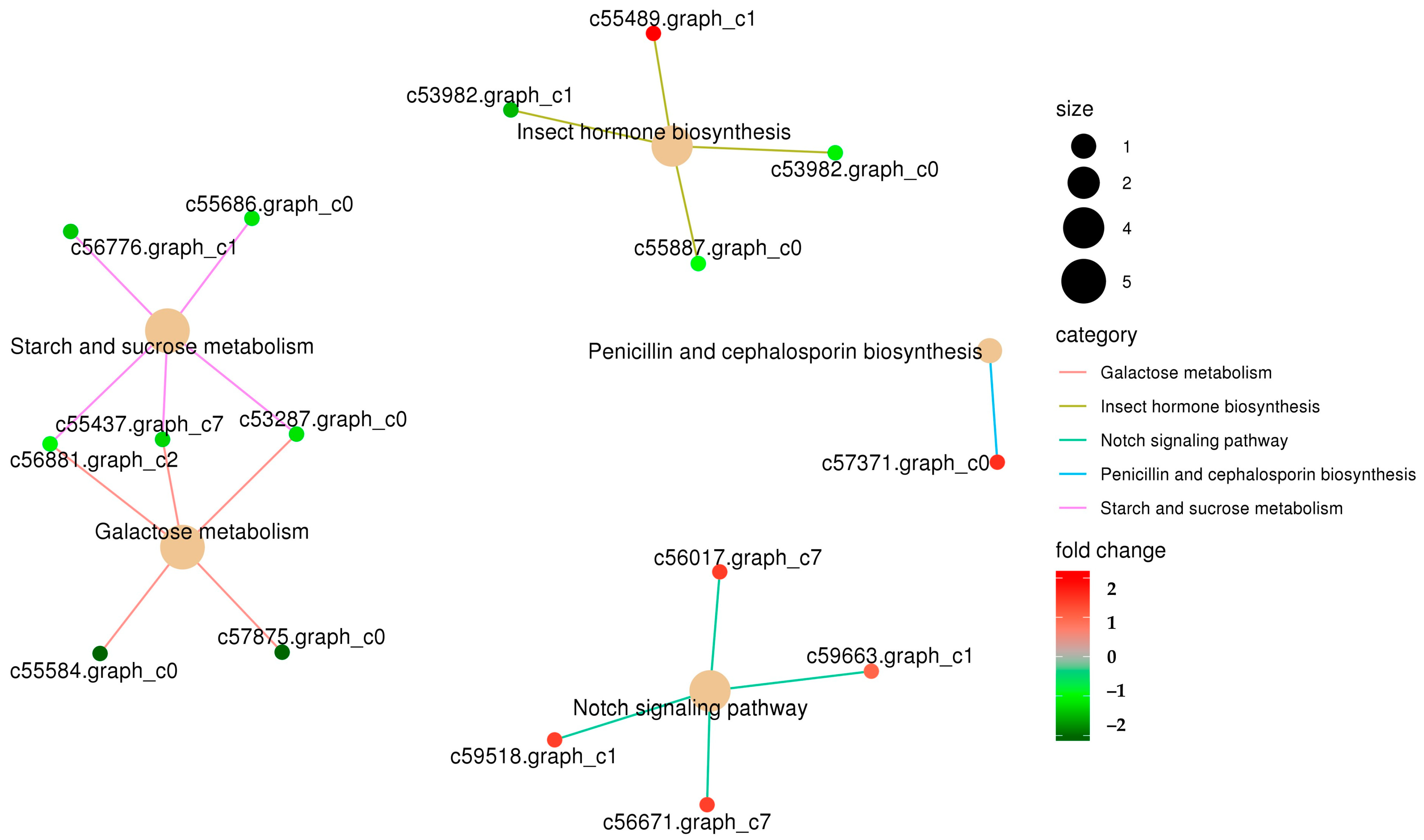
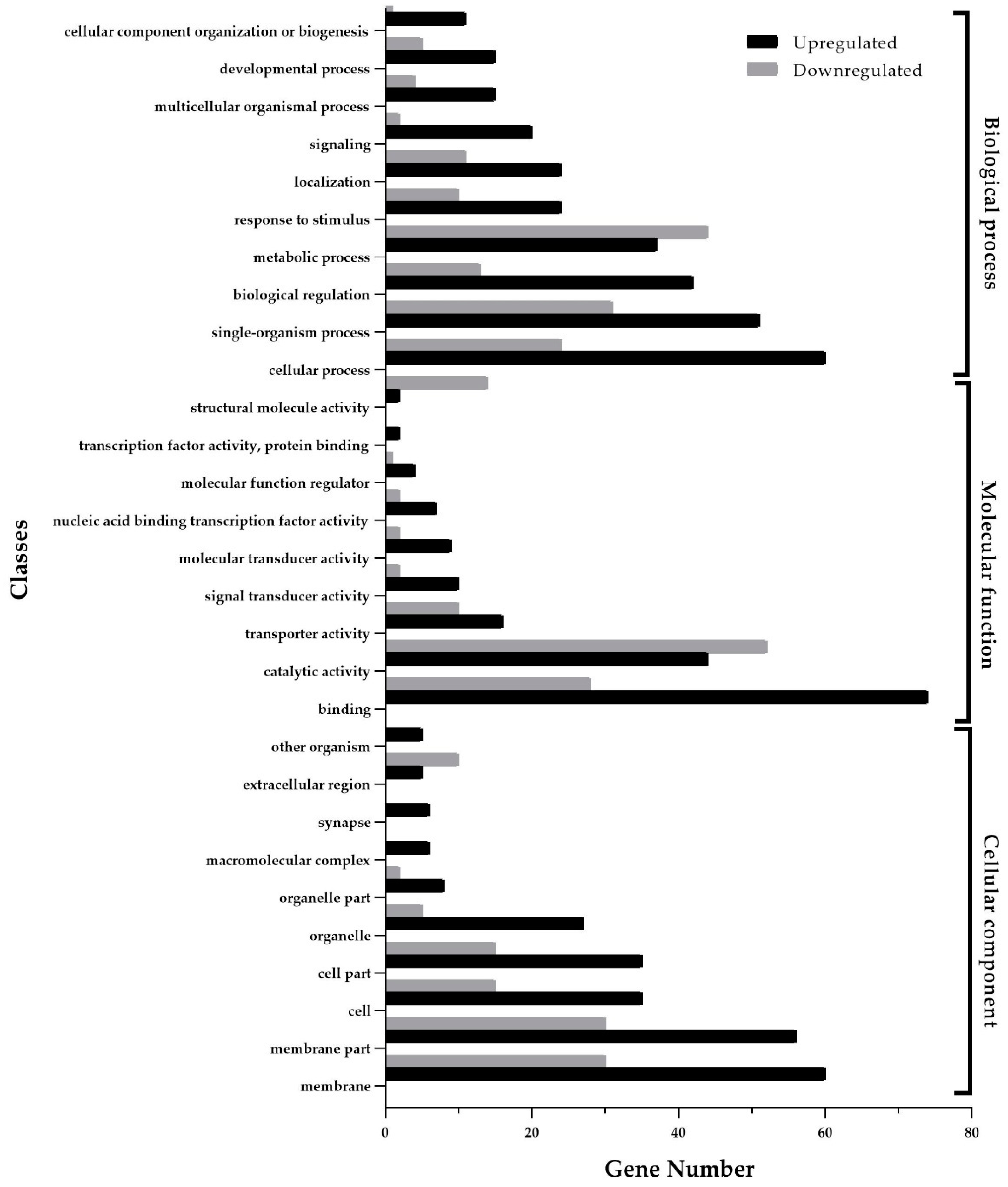
3.2. Functional Enrichment Analysis of Differentially Expressed Genes
3.3. Identification of CytCo-Binding Proteins
4. Discussion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Hajek, A.; Steinkraus, D.; Castrillo, L. Sleeping Beauties: Horizontal Transmission via Resting Spores of Species in the Entomophthoromycotina. Insects 2018, 9, 102. [Google Scholar] [CrossRef] [PubMed]
- Hong, S.; Shang, J.; Sun, Y.-W.; Tang, G.; Wang, C. Fungal Infection of Insects: Molecular Insights and Prospects. Trends Microbiol. 2024, 32, 302–316. [Google Scholar] [CrossRef] [PubMed]
- Gryganskyi, A.P.; Mullens, B.A.; Gajdeczka, M.T.; Rehner, S.A.; Vilgalys, R.; Hajek, A.E. Hijacked: Co-Option of Host Behavior by Entomophthoralean Fungi. PLoS Pathog. 2017, 13, e1006274. [Google Scholar] [CrossRef]
- Lovett, B.; St. Leger, R.J. The Insect Pathogens. Microbiol. Spectrum 2017, 5, 923–943. [Google Scholar] [CrossRef] [PubMed]
- Shen, D.; Nyawira, K.T.; Xia, A. New Discoveries and Applications of Mosquito Fungal Pathogens. Curr. Opin. Insect Sci. 2020, 40, 111–116. [Google Scholar] [CrossRef]
- Ortiz-Urquiza, A.; Luo, Z.; Keyhani, N.O. Improving Mycoinsecticides for Insect Biological Control. Appl. Microbiol. Biotechnol. 2014, 99, 1057–1068. [Google Scholar] [CrossRef] [PubMed]
- Mascarin, G.M.; Lopes, R.B.; Delalibera, Í.; Fernandes, É.K.; Luz, C.; Faria, M. Current Status and Perspectives of Fungal Entomopathogens Used for Microbial Control of Arthropod Pests in Brazil. J. Inverteb. Pathol. 2019, 165, 46–53. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhou, X.; Guo, K.; Zhang, X.; Lin, H.; Montalva, C. Transcriptomic Insight into Pathogenicity-Associated Factors of Conidiobolus Obscurus, an Obligate Aphid-Pathogenic Fungus Belonging to Entomopthoromycota. Pest Manag. Sci. 2018, 74, 1677–1686. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Wang, S. Insect Pathogenic Fungi: Genomics, Molecular Interactions, and Genetic Improvements. Annu. Rev. Entomol. 2017, 62, 73–90. [Google Scholar] [CrossRef]
- Gasmi, L.; Baek, S.; Kim, J.C.; Kim, S.; Lee, M.R.; Park, S.E.; Shin, T.Y.; Lee, S.J.; Parker, B.L.; Kim, J.S. Gene Diversity Explains Variation in Biological Features of Insect Killing Fungus, Beauveria Bassiana. Sci. Rep. 2021, 11, 91. [Google Scholar] [CrossRef]
- Lovett, B.; St. Leger, R.J. Fungal Biopesticides. Pest Manag. Sci. 2017, 74, 781–789. [Google Scholar] [CrossRef] [PubMed]
- Malagocka, J.; Eilenberg, J.; Jensen, A.B. Social Immunity Behaviour among Ants Infected by Specialist and Generalist Fungi. Curr. Opin. Insect Sci. 2019, 33, 99–104. [Google Scholar] [CrossRef]
- De Fine Licht, H.H.; Jensen, A.B.; Eilenberg, J. Comparative Transcriptomics Reveal Host-Specific Nucleotide Variation in Entomophthoralean Fungi. Mol. Ecol. 2016, 26, 2092–2110. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Yang, T.; Yu, W.; Wang, X.; Zhou, X.; Zhou, X. Genome-Wide Study of Conidiation-Related Genes in the Aphid-Obligate Fungal Pathogen Conidiobolus Obscurus (Entomophthoromycotina). J. Fungi 2022, 8, 389. [Google Scholar] [CrossRef] [PubMed]
- Stajich, J.E.; Lovett, B.; Lee, E.; Macias, A.M.; Hajek, A.E.; de Bivort, B.L.; Kasson, M.T.; De Fine Licht, H.H.; Elya, C. Signatures of Transposon-Mediated Genome Inflation, Host Specialization, and Photoentrainment in Entomophthora Muscae and Allied Entomophthoralean Fungi. eLife 2023, 12, RP92863. [Google Scholar] [CrossRef] [PubMed]
- Gryganskyi, A.P.; Hajek, A.E.; Voloshchuk, N.; Idnurm, A.; Eilenberg, J.; Manfrino, R.G.; Bushley, K.E.; Kava, L.; Kutovenko, V.B.; Anike, F.; et al. Potential for Use of Species in the Subfamily Erynioideae for Biological Control and Biotechnology. Microorganisms 2024, 12, 168. [Google Scholar] [CrossRef] [PubMed]
- Boomsma, J.J.; Jensen, A.B.; Meyling, N.V.; Eilenberg, J. Evolutionary Interaction Networks of Insect Pathogenic Fungi. Ann. Rev. Entomol. 2014, 59, 467–485. [Google Scholar] [CrossRef] [PubMed]
- Eilenberg, J.; Saussure, S.; Fekih, I.B.; Jensen, A.B.; Klingen, I. Factors Driving Susceptibility and Resistance in Aphids That Share Specialist Fungal Pathogens. Curr. Opin. Insect Sci. 2019, 33, 91–98. [Google Scholar] [CrossRef]
- Ma, M.; Luo, J.; Li, C.; Eleftherianos, I.; Zhang, W.; Xu, L. A Life-And-Death Struggle: Interaction of Insects with Entomopathogenic Fungi across Various Infection Stages. Front. Immunol. 2024, 14, 1329843. [Google Scholar] [CrossRef]
- Hillyer, J.F. Insect Immunology and Hematopoiesis. Dev. Comp. Immunol. 2016, 58, 102–118. [Google Scholar] [CrossRef]
- Stanley, D.; Haas, E.; Miller, J. Eicosanoids: Exploiting Insect Immunity to Improve Biological Control Programs. Insects 2012, 3, 492–510. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Wang, D.-W.; Zhang, X.; Wang, J.H. The Influence of the Aphid-Specific Pathogen Conidiobolus Obscurus (Entomophthoromycota: Entomophthorales) on the Mortality and Fecundity of Bamboo Aphids. J. For. Res. 2014, 19, 388–394. [Google Scholar] [CrossRef]
- Wang, Y.; Chen, S.; Wang, J.; Zhou, X. Characterization of a Cytolytic-like Gene from the Aphid-Obligate Fungal Pathogen Conidiobolus Obscurus. J. Invertebr. Pathol. 2020, 173, 107366. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Yang, T.; Su, X.; Zhang, X.; Zhou, X. Debilitation of Galleria Mellonella Hemocytes Using CytCo a Cytolytic-like Protein Derived from the Entomopathogen Conidiobolus Obscurus. Pestic. Biochem Phys. 2023, 193, 105418. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Guo, K.; Mao, S.-F.; He, W.; Zhu, Y.-Y. Modeling Analysis on Sporulation Capacity, Storage and Infectivity of the Aphid-Specific Pathogen Conidiobolus Obscurus (Entomophthoromycota: Entomophthorales). Mycoscience 2014, 55, 21–26. [Google Scholar] [CrossRef]
- Guo, K.; Lin, H.; Su, X.; Wang, J.; Montalva, C.; Shao, S.; Zhou, X. The Spatial–Temporal Dynamics of Bamboo Aphid Dispersal Flight along with Their Natural Enemies: Biocontrol Implication. Agroforest. Syst. 2017, 93, 631–639. [Google Scholar] [CrossRef]
- Guo, X.Q.; Chen, F.Z.; Gao, F.; Li, L.; Liu, K.; You, L.; Hua, C.; Yang, F.; Liu, W.; Peng, C.; et al. CNSA: A data repository for archiving omics data. Database 2020, 2020, baaa055. [Google Scholar] [CrossRef] [PubMed]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-Length Transcriptome Assembly from RNA-Seq Data without a Reference Genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Dewey, C.N. RSEM: Accurate Transcript Quantification from RNA-Seq Data with or without a Reference Genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential Gene and Transcript Expression Analysis of RNA-Seq Experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef] [PubMed]
- Shannon, P. Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Tan, K.; Vlisidou, I.; Wood, W. Ecdysone Mediates the Development of Immunity in the Drosophila Embryo. Curr. Biol. 2014, 24, 1145–1152. [Google Scholar] [CrossRef]
- Gerardo, N.M.; Altincicek, B.; Anselme, C.; Atamian, H.; Barribeau, S.M.; de Vos, M.; Duncan, E.J.; Evans, J.D.; Gabaldón, T.; Ghanim, M.; et al. Immunity and Other Defenses in Pea Aphids, Acyrthosiphon pisum. Genome Biol. 2010, 11, R21. [Google Scholar] [CrossRef]
- Buchon, N.; Poidevin, M.; Kwon, H.-M.; Guillou, A.; Sottas, V.; Lee, B.L.; Lemaitre, B. A Single Modular Serine Protease Integrates Signals from Pattern-Recognition Receptors Upstream of the Drosophila Toll Pathway. Proc. Natl. Acad. Sci. USA 2009, 106, 12442–12447. [Google Scholar] [CrossRef]
- Boguś, M.I.; Wieloch, W.; Ligęza-Żuber, M. Coronatin-2 from the Entomopathogenic Fungus Conidiobolus coronatus Kills Galleria mellonella Larvae and Incapacitates Hemocytes. Bull. Entomol. Res. 2016, 107, 66–76. [Google Scholar] [CrossRef] [PubMed]
- Wrońska, A.K.; Boguś, M.I. Harman and Norharman, Metabolites of the Entomopathogenic Fungus Conidiobolus coronatus (Entomophthorales), Affect the Serotonin Levels and Phagocytic Activity of Hemocytes, Insect Immunocompetent Cells, in Galleria mellonella (Lepidoptera). Cell Biosci. 2019, 9, 29. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Zhou, X.; Guo, K.; Chen, S.-N.; Su, X. Transcriptomic Insights into the Effects of CytCo, a Novel Nematotoxic Protein, on the Pine Wood Nematode Bursaphelenchus Xylophilus. BMC Genom. 2021, 22, 394. [Google Scholar] [CrossRef] [PubMed]
- Stammers, A.N.; Susser, S.E.; Hamm, N.C.; Hlynsky, M.W.; Kimber, D.E.; Kehler, D.S.; Duhamel, T.A. The Regulation of Sarco(Endo)Plasmic Reticulum Calcium-ATPases (SERCA). Can. J. Physiol. Pharmacol. 2015, 93, 843–854. [Google Scholar] [CrossRef]
- Sanyal, S.; Consoulas, C.; Kuromi, H.; Basole, A.; Mukai, L.; Kidokoro, Y.; Krishnan, K.S.; Ramaswami, M. Analysis of Conditional Paralytic Mutants in Drosophila Sarco-Endoplasmic Reticulum Calcium ATPase Reveals Novel Mechanisms for Regulating Membrane Excitability. Genetics 2005, 169, 737–750. [Google Scholar] [CrossRef]

| Unigene ID | Annotation | FPKM | Log2(FC) b | GO/KO ID | |
|---|---|---|---|---|---|
| Healthy | Inoculated | ||||
| Immunoglobulin | |||||
| c57868 | Immunoglobulin I-set domain | 1.81 | 6.99 | 1.70 | K23915 |
| c56461 | Immunoglobulin V-set domain | 0.39 | 1.84 | 1.48 | K06772 |
| c58593 | Immunoglobulin V-set domain | 1.33 | 3.66 | 1.23 | GO:0016021 |
| c56079 | Immunoglobulin I-set domain | 2.33 | 5.85 | 1.17 | GO:0016021 |
| c54209 | Immunoglobulin V-set domain | 0.84 | 2.25 | 1.13 | K22655 |
| c56508 | Immunoglobulin I-set domain | 1.06 | 2.64 | 1.01 | GO:0016021 |
| Peroxidase or ROS-related | |||||
| c55353 * | Chorion peroxidase | 7.95 | 4.12 | −1.01 | K19511 |
| c59130 | Animal haem peroxidase | 245.04 | 120.77 | −1.04 | K19511 |
| c59021 | Melanization protease | 125.49 | 56.73 | −1.27 | GO:0004252 |
| c55437 | Phenoloxidase-activating enzyme | 120.31 | 35.41 | −1.34 | GO:0004252 |
| c58805 | Glutamate transporter polyphemus | 25.40 | 7.20 | −1.58 | K14209 |
| Wound healing | |||||
| c54775 | Fibronectin type III | 5.48 | 18.32 | 1.78 | K17591 |
| c51444 | Mucin 2-like protein | 0.23 | 1.08 | 1.17 | / |
| c54290 | Fibronectin type III | 2.55 | 6.57 | 1.11 | GO:0005515 |
| c57232 | Ig-like and fibronectin type-III | 5.32 | 12.58 | 1.09 | K06774 |
| c52952 | Fibronectin type III | 1.68 | 0.64 | −1.18 | GO:0002376 |
| Toll pathway | |||||
| c57284 | NAD(+) hydrolase sarm1 | 2.31 | 7.01 | 1.56 | GO:0034128 |
| c53917 | Toll-like receptor | 0.88 | 2.09 | 1.05 | K10170 |
| c54414 | modular serine protease ModSP | 84.80 | 34.60 | −1.29 | K20674 |
| c57804 | modular serine protease ModSP | 34.66 | 7.57 | −1.97 | K20674 |
| c58774 | Serpin | 76.40 | 21.80 | −1.93 | K13963 |
| Ecdysone-mediated | |||||
| c55917 | Krueppel 1-like | 3.87 | 16.90 | 1.78 | K09228 |
| c56481 | Ecdysone-induced protein | 1.65 | 5.14 | 1.27 | K08701 |
| c55601 | PR domain zinc finger protein 1 isoform X1 | 9.68 | 22.20 | 1.06 | GO:0035075 |
| Unigene ID | Annotation | FPKM | Log2(FC) b | GO/KO ID | |
|---|---|---|---|---|---|
| Healthy | Inoculated | ||||
| Signaling | |||||
| c56897 | inositol 1,4,5-trisphosphate receptor | 0.76 | 2.95 | 1.41 | K04958 |
| c56671 | Protein serrate | 1.75 | 5.11 | 1.39 | GO:0007219 |
| c58773 | beta-1 adrenergic receptor | 1.70 | 5.26 | 1.39 | GO:0004930 |
| c58924 | receptor-type tyrosine-protein phosphatase | 1.89 | 4.44 | 1.21 | K06776 |
| c55654 | Rho GTPase-activating | 8.45 | 20.90 | 1.19 | K20655 |
| c54354 | ephrin-B1 | 2.89 | 8.17 | 1.17 | K05463 |
| c57822 | G-protein coupled receptor | 1.05 | 2.28 | 1.16 | GO:0004930 |
| c57839 | FMRFamide receptor | 0.64 | 1.74 | 1.14 | GO:0008528 |
| c54122 | WNT1-inducible-signaling pathway | 0.73 | 2.55 | 1.14 | K22471 |
| c56120 | G-protein coupled receptor | 0.57 | 1.68 | 1.13 | K08469 |
| c56130 | regulator of G-protein signaling | 17.00 | 40.40 | 1.11 | GO:0001965 |
| c57791 | dual 3′,5′-cyclic-AMP and -GMP phosphodiesterase | 4.82 | 11.60 | 1.08 | GO:0004114 |
| c56800 | JNK-interacting | 4.89 | 10.20 | 1.06 | K20317 |
| c54894 | arrestin homolog | 1.55 | 3.98 | 1.05 | K04439 |
| Neuromotor system | |||||
| c56627 | Neuropilin and tolloid-like | 5.08 | 14.20 | 1.29 | K19404 |
| c57981 | unc-80 homolog | 1.16 | 3.76 | 1.28 | K24015 |
| c54290 | Neural cell adhesion | 2.55 | 6.57 | 1.11 | K06491 |
| c57640 * | synaptotagmin 1 isoform X1 | 7.61 | 17.90 | 1.09 | GO:0007269 |
| c56243 | Acetylcholine receptor subunit beta-like | 18.70 | 39.30 | 1.07 | K05312 |
| c55778 | neuronal acetylcholine receptor subunit α-7 | 8.41 | 15.10 | 1.07 | GO:0022848 |
| c59372 | neurobeachin isoform X3 | 7.78 | 10.90 | 1.04 | K24183 |
| c58381 | locomotion-related protein Hikaru genki | 3.02 | 6.64 | 1.04 | K17495 |
| c59504 | regulating synaptic membrane exocytosis | 25.30 | 55.90 | 1.03 | K15297 |
| c55640 | serine/threonine-protein kinase Nek8 | 1.64 | 3.77 | 1.03 | K20877 |
| c57984 | myelin regulatory factor-like | 4.91 | 9.05 | 1.02 | / |
| Ion channel | |||||
| c58440 | Voltage-gated chloride channel | 2.11 | 14.4 | 2.17 | GO:0005247 |
| c58546 | voltage-dependent calcium channel | 0.35 | 1.61 | 1.47 | K04863 |
| c58915 | Cyclic nucleotide-gated cation channel α-3 | 4.15 | 10.80 | 1.33 | GO:0005249 |
| c55895 | Potassium channel domain | 2.50 | 6.95 | 1.30 | / |
| c57081 | Small conductance calcium-activated potassium channel | 2.32 | 5.82 | 1.30 | GO:0016286 |
| c56821 | potassium voltage-gated channel | 0.53 | 1.85 | 1.25 | GO:0008076 |
| c56460 | Potassium channel | 4.94 | 11.80 | 1.16 | GO:0005267 |
| c58522 | potassium voltage-gated channel | 0.66 | 1.77 | 1.12 | K04905 |
| c52823 | glutamate-gated chloride channel-like | 17.10 | 7.82 | −1.11 | GO:0005230 |
| Unigene ID | Annotation | Accession No. | Unused ProtScore | Coverage (%) | Peptides | TMRs |
|---|---|---|---|---|---|---|
| c55126 | calcium-transporting ATPase sarcoplasmic/endoplasmic reticulum type isoform X2 | gi|1028717492 | 37.86 | 24.9 | 20 | 8 |
| c55766 | AMP deaminase 2 isoform X2 | gi|1028703692 | 12.87 | 9.4 | 7 | 2 |
| c57640 | synaptotagmin 1 isoform X1 | gi|641664799 | 9.68 | 14.4 | 5 | 1 |
| c58629 | ATP-binding cassette sub-family F member 1 | gi|193676397 | 6.00 | 3 | 3 | 3 |
| c52852 | dihydroorotate dehydrogenase (quinone), mitochondrial | gi|641676995 | 5.23 | 8.3 | 3 | 4 |
| c55131 | probable hydroxyacid-oxoacid transhydrogenase, mitochondrial | gi|641678166 | 4.21 | 4.1 | 2 | 10 |
| c53223 | innexin inx7 | gi|328723643 | 4.08 | 6.7 | 2 | 5 |
| c59449 | transmembrane 9 superfamily member 2 | gi|1028716222 | 4.00 | 3.8 | 2 | 9 |
| c52282 | ubiquitin-like modifier-activating enzyme 5 | gi|193707023 | 4.00 | 3.4 | 3 | 2 |
| c57885 | Mo25 uncharacterized | gi|328720151 | 3.64 | 5.8 | 2 | 6 |
| c54081 | tumor suppressor candidate 3 | gi|641672251 | 1.49 | 5.8 | 2 | 6 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhu, J.; Fu, Y.; Zhang, L.; Yang, T.; Zhou, X. Transcriptomic Profiling of Bean Aphid Megoura crassicauda upon Exposure to the Aphid-Obligate Entomopathogen Conidiobolus obscurus (Entomophthoromycotina) and Screening of CytCo-Binding Aphid Proteins through a Pull-Down Assay. Insects 2024, 15, 388. https://doi.org/10.3390/insects15060388
Zhu J, Fu Y, Zhang L, Yang T, Zhou X. Transcriptomic Profiling of Bean Aphid Megoura crassicauda upon Exposure to the Aphid-Obligate Entomopathogen Conidiobolus obscurus (Entomophthoromycotina) and Screening of CytCo-Binding Aphid Proteins through a Pull-Down Assay. Insects. 2024; 15(6):388. https://doi.org/10.3390/insects15060388
Chicago/Turabian StyleZhu, Jiaqin, Yaqi Fu, Lvhao Zhang, Tian Yang, and Xiang Zhou. 2024. "Transcriptomic Profiling of Bean Aphid Megoura crassicauda upon Exposure to the Aphid-Obligate Entomopathogen Conidiobolus obscurus (Entomophthoromycotina) and Screening of CytCo-Binding Aphid Proteins through a Pull-Down Assay" Insects 15, no. 6: 388. https://doi.org/10.3390/insects15060388






