Effects of Isolated LAB on Chemical Composition, Fermentation Quality and Bacterial Community of Stipa grandis Silage
Abstract
1. Introduction
2. Materials and Methods
2.1. Samples and Bacterial Isolates
2.2. Physiological and Morphological Tests
2.3. Identification of LAB Strains by 16S rRNA Sequence Analysis
2.4. Preparation of the Experimental Silages
2.5. Chemical and Microbial Analyses
2.6. Bacterial Community Sequencing Analysis
2.7. Bioinformatics Analysis
2.8. Statistical Analysis
3. Results
3.1. Lactic Acid Bacteria Strain Characteristics
3.2. Silage Characteristics of Fresh Stipa Grandis
3.3. Effect of Lactic Acid Bacteria Inoculant
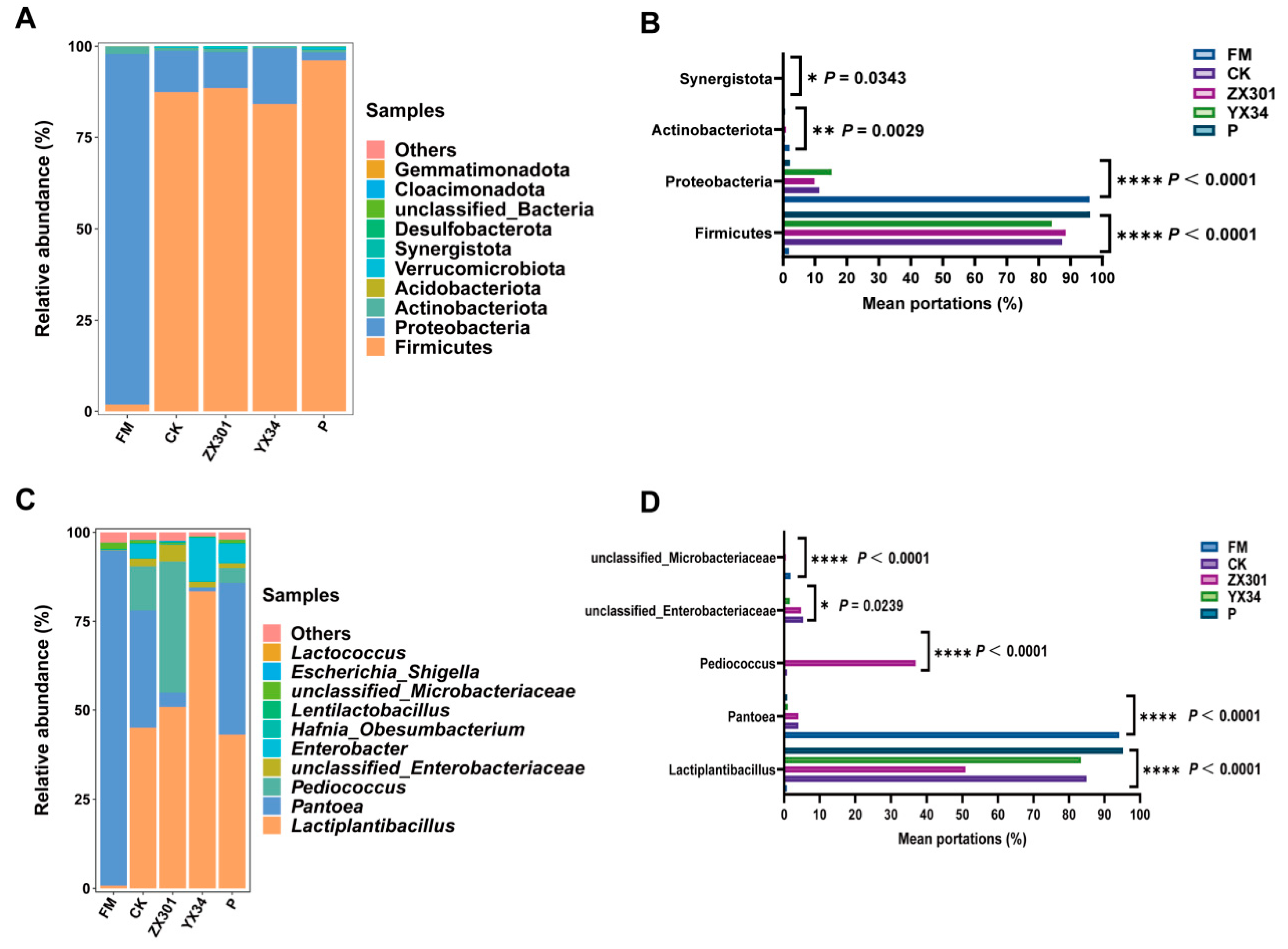
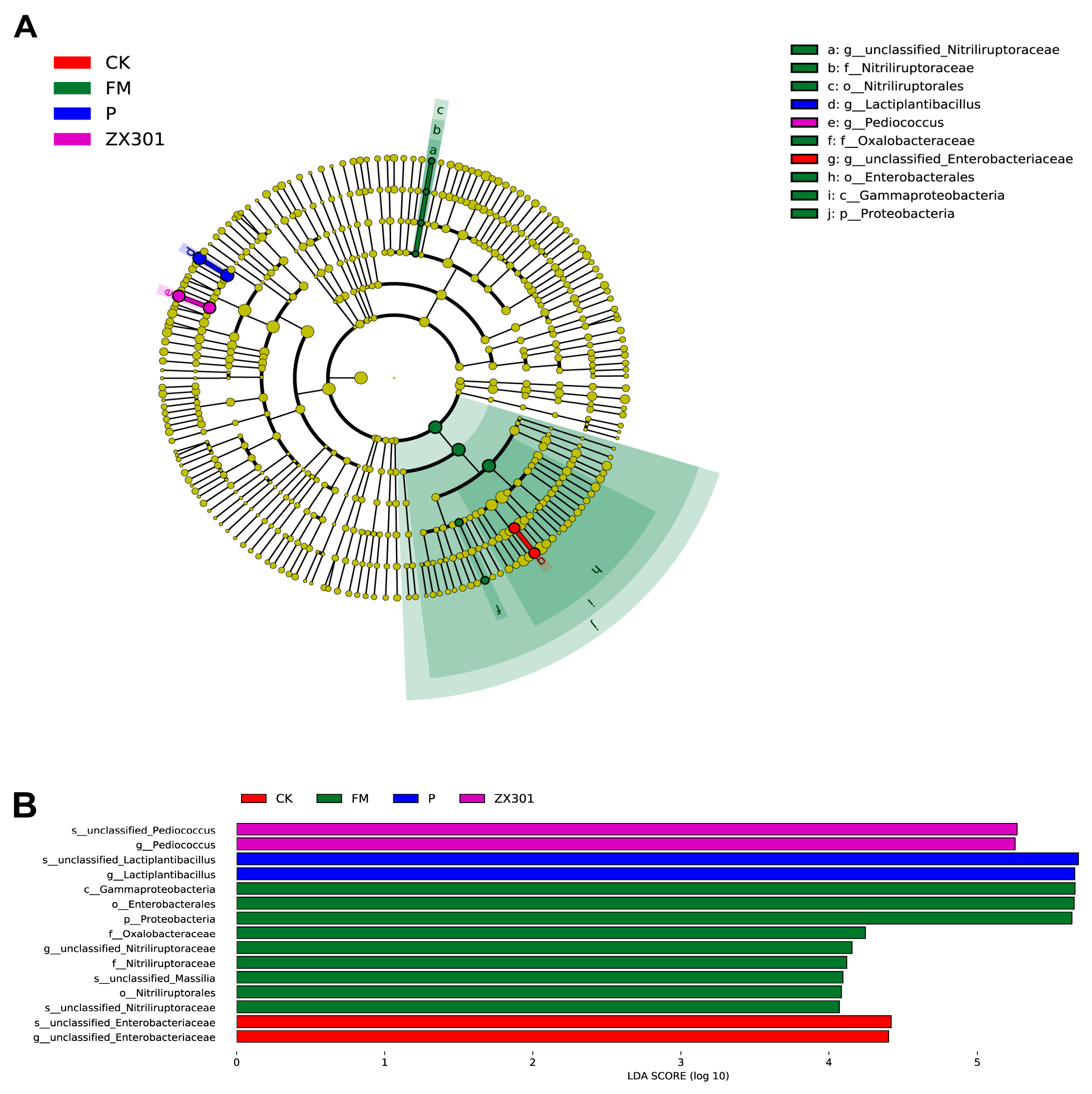
3.4. Microbial Diversity of Fresh Materials and Stipa Grandis after Ensiling
3.5. Relationships between Chemical Compositions, Fermentation Parameters, and Bacterial Community
3.6. Relationships between Chemical Compositions, Fermentation Parameters, and Bacterial Community
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Wang, Y.X.; Duan, L.M.; Liu, T.X.; Lou, Y.Y.; Li, D.F.; Tong, X.; Li, W.; Lei, H.M.; Singh, V.P. Evaluation of non-stationarity in summer precipitation and the response of vegetation over the typical steppe in Inner Mongolia. Clim. Dyn. 2022, 58, 2227–2247. [Google Scholar] [CrossRef]
- Wang, F.F.; Li, Z.M.; Fu, B.J.; Lü, Y.H.; Liu, G.P.; Wang, D.B.; Wu, X. Short-term grazing exclusion alters soil bacterial co-occurrence patterns rather than community diversity or composition in temperate grasslands. Front. Microbiol. 2022, 13, 824192. [Google Scholar] [CrossRef] [PubMed]
- You, S.H.; Du, S.; Ge, G.T.; Wan, T.; Jia, Y.S. Microbial community and fermentation characteristics of native grass prepared without or with isolated lactic acid bacteria on the Mongolian Plateau. Front. Microbiol. 2021, 12, 731770. [Google Scholar] [CrossRef] [PubMed]
- Li, D.X.; Ni, K.K.; Pang, H.L.; Wang, Y.P.; Cai, Y.M.; Jin, Q.S. Identification and antimicrobial activity detection of lactic acid bacteria isolated from corn stover silage. Asian-Australas. J. Anim. Sci. 2015, 28, 620–631. [Google Scholar] [CrossRef]
- Ren, H.Y.; Han, G.D.; Schönbach, P.; Gierus, M.; Taube, F. Forage nutritional characteristics and yield dynamics in a grazed semiarid steppe ecosystem of Inner Mongolia, China. Ecol. Indic. 2016, 60, 460–469. [Google Scholar] [CrossRef]
- Du, S.; You, S.H.; Bao, J.; Ge, G.T.; Jia, Y.S.; Cai, Y.M. Evaluation of the growth performance and meat quality of Mongolian lamb fed grass, hay or pellets of Inner Mongolian native grass. Small Rumin. Res. 2019, 181, 34–38. [Google Scholar] [CrossRef]
- Wang, S.R.; Li, J.F.; Zhao, J.; Dong, Z.H.; Dong, D.; Shao, T. Effect of epiphytic microbiota from napiergrass and Sudan grass on fermentation characteristics and bacterial community in oat silage. J. Appl. Microbiol. 2022, 132, 919–932. [Google Scholar] [CrossRef]
- Hou, M.L.; Ge, G.T.; Liu, T.Y.; Jia, Y.S.; Cai, Y.M. Silage preparation and fermentation quality of natural grasses treated with lactic acid bacteria and cellulase in meadow steppe and typical steppe. Asian-Australas. J. Anim. Sci. 2017, 30, 788–796. [Google Scholar] [CrossRef]
- Jaipolsaen, N.; Sangsritavong, S.; Uengwetwanit, T.; Angthong, P.; Plengvidhya, V.; Rungrassamee, W.; Yammuenart, S. Comparison of the effects of microbial inoculants on fermentation quality and microbiota in napier grass Pennisetum purpureum and corn Zea mights L. silage. Front. Microbiol. 2022, 12, 784535. [Google Scholar] [CrossRef]
- Xian, Z.Y.; Wu, J.Q.; Deng, M.; Wang, M.; Tian, H.C.; Liu, D.W.; Li, Y.K.; Liu, G.B.; Sun, B.L.; Guo, Y.Q. Effects of cellulase and Lactiplantibacillus plantarum on the fermentation parameters, nutrients, and bacterial community in cassia alata silage. Front. Microbiol. 2022, 13, 926065. [Google Scholar] [CrossRef]
- Zi, X.J.; Li, M.; Chen, Y.Y.; Lv, R.L.; Zhou, H.L.; Tang, J. Effects of citric acid and Lactobacillus plantarum on silage quality and bacterial diversity of king grass silage. Front. Microbiol. 2021, 12, 631096. [Google Scholar] [CrossRef] [PubMed]
- Tohno, M.; Kobayashi, H.; Nomura, M.; Kitahara, M.; Ohkuma, M.; Uegaki, R.; Cai, Y. Genotypic and phenotypic characterization of lactic acid bacteria isolated from Italian ryegrass silage. Anim. Sci. J. 2012, 83, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Fijałkowska, M.; Przemieniecki, S.W.; Purwin, C.; Lipiński, K.; Kurowski, T.P.; Karwowska, A. The effect of an additive containing three Lactobacillus species on the fermentation pattern and microbiological status of silage. J. Sci. Food Agric. 2020, 100, 1174–1184. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Q.M.; Li, M.Y.; Fan, X.Y.; Chen, Y.L.; Sun, H.; Xie, Y.X.; Zheng, Y.L.; Chen, C.; Li, P. Effects of epiphytic and exogenous lactic acid bacteria on fermentation quality and microbial community compositions of paper mulberry silage. Front. Microbiol. 2022, 13, 973500. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.L.; Du, Z.M.; Xiong, Y.; Wang, N.W.; Wang, X.K.; Zhou, X.L.; Yang, F.Y.; Ni, K.K. Genotypic and phenotypic characterization of lactic acid bacteria associated with silage fermentation of pineapple residue. Diversity 2022, 14, 631. [Google Scholar] [CrossRef]
- Yang, J.S.; Tang, K.; Tan, H.S.; Cai, Y.M. Characterization of lactic acid bacteria isolated from banana and its application in silage fermentation of defective banana. Microorganisms 2022, 10, 1185. [Google Scholar] [CrossRef]
- Sifeeldein, A.; Wang, S.; Li, J.; Dong, Z.; Chen, L.; Kaka, N.A.; Shao, T. Phylogenetic identification of lactic acid bacteria isolates and their effects on the fermentation quality of sweet sorghum Sorghum bicolor silage. J. Appl. Microbiol. 2019, 126, 718–729. [Google Scholar] [CrossRef]
- Zhao, S.S.; Wang, Y.P.; Yang, F.Y.; Wang, Y.; Zhang, H. Screening a Lactobacillus plantarum strain for good adaption in alfalfa ensiling and demonstrating its improvement of alfalfa silage quality. J. Appl. Microbiol. 2020, 129, 233–242. [Google Scholar] [CrossRef]
- Cai, Y.; Benno, Y.; Ogawa, M.; Ohmomo, S.; Kumai, S.; Nakase, T. Influence of Lactobacillus spp. from an inoculant and of Weissella and Leuconostoc spp. from forage crops on silage fermentation. Appl. Environ. Microbiol. 1998, 64, 2982–2987. [Google Scholar] [CrossRef]
- Kozaki, M.; Uchimura, T.; Okada, S. Experimentalmanual for Lactic Acid Bacteria; Asakurasyoten: Tokyo, Japan, 1992. [Google Scholar]
- Pang, H.L.; Tan, Z.F.; Qin, G.Y.; Wang, Y.P.; Li, Z.W.; Jin, Q.S.; Cai, Y.M. Phenotypic and phylogenetic analysis of lactic acid bacteria isolated from forage crops and grasses in the Tibetan Plateau. J. Microbiol. 2012, 50, 63–71. [Google Scholar] [CrossRef]
- Liu, Q.H.; Chen, M.X.; Zhang, J.G.; Shi, S.L.; Cai, Y.M. Characteristics of isolated lactic acid bacteria and their effectiveness to improve stylo Stylosanthes guianensis Sw. silage quality at various temperatures. Anim. Sci. J. 2012, 83, 128–135. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.R.; Yuan, X.J.; Dong, Z.H.; Li, J.F.; Shao, T. Isolating and evaluating lactic acid bacteria strains for effectiveness on silage quality at low temperatures on the Tibetan Plateau. Anim. Sci. J. 2017, 88, 1722–1729. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Yu, Z.; Wang, X.G. Isolating and evaluating lactic acid bacteria strains with or without sucrose for effectiveness of silage fermentation. Grassl. Sci. 2015, 61, 167–176. [Google Scholar] [CrossRef]
- Pitiwittayakul, N.; Bureenok, S.; Schonewille, J.T. Selective thermotolerant lactic acid bacteria isolated from fermented juice of epiphytic lactic acid bacteria and their effects on fermentation quality of stylo silages. Front. Microbiol. 2021, 12, 673946. [Google Scholar] [CrossRef]
- Ennahar, S.; Cai, Y.M.; Fujita, Y. Phylogenetic diversity of lactic acid bacteria associated with paddy rice silage as determined by 16S ribosomal DNA analysis. Appl. Environ. Microbiol. 2003, 69, 444–451. [Google Scholar] [CrossRef] [PubMed]
- AOAC. Oficial Methods of Analysis; Association of Official Analytical Chemists: Arlington, TX, USA, 1995. [Google Scholar]
- Thomas, T.A. An automated procedure for the determination of soluble carbohydrates in herbage. J. Sci. Food Agric. 1977, 28, 639–642. [Google Scholar] [CrossRef]
- Van Soest, P.J.; Robertson, J.B.; Lewis, B.A. Methods for dietary fiber, neutral detergent fiber, and nonstarch polysaccharides in relation to animal nutrition. J. Dairy Sci. 1991, 74, 3583–3597. [Google Scholar] [CrossRef] [PubMed]
- You, S.H.; Du, S.; Ge, G.T.; Wan, T.; Jia, Y.S. Selection of lactic acid bacteria from native grass silage and its effects as inoculant on silage fermentation. Agron. J. 2021, 113, 3169–3177. [Google Scholar] [CrossRef]
- Broderick, G.A.; Kang, J.H. Automated simultaneous determination of ammonia and total amino acids in ruminal fluid and in vitro media. J. Dairy Sci. 1980, 63, 64–75. [Google Scholar] [CrossRef]
- Guo, X.; Zheng, P.; Zou, X.; Chen, X.Y.; Zhang, Q. Influence of pyroligneous acid on fermentation parameters, CO2 production and bacterial communities of rice straw and stylo silage. Front. Microbiol. 2021, 12, 701434. [Google Scholar] [CrossRef]
- Huang, R.Z.; Zhang, F.F.; Wang, T.; Zhang, Y.L.; Li, X.; Chen, Y.C.; Ma, C.H. Effect of intrinsic tannins on the fermentation quality and associated with the bacterial and fungal community of sainfoin silage. Microorganisms 2022, 10, 844. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Dong, L.; Lian, W.H.; Lin, Z.L.; Lu, C.Y.; Xu, L.; Li, L.; Hozzein, W.N.; Li, W.J. Exploring untapped potential of Streptomyces spp. in gurbantunggut desert by use of highly selective culture strategy. Sci. Total Environ. 2021, 790, 148235. [Google Scholar] [CrossRef] [PubMed]
- Segata, N.; Izard, J.; Waldron, L.; Gevers, D.; Miropolsky, L.; Garrett, W.S.; Huttenhower, C. Metagenomic biomarker discovery and explanation. Genome Biol. 2011, 12, R60. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2014. [Google Scholar]
- Douglas, G.M.; Maffei, V.J.; Zaneveld, J.; Yurgel, S.N.; Brow, J.R.; Taylor, C.M.; Huttenhower, C.; Langille, M.G.I. PICRUSt2: An im proved and extensible approach for metagenome inference. BioRxiv 2020. [Google Scholar] [CrossRef]
- Burns, P.; Borgo, M.F.; Binetti, A.; Puntillo, M.; Bergamini, C.; Páez, R.; Mazzoni, R.; Reinheimer, J.; Vinderola, G. Isolation, characterization and performance of autochthonous spray dried lactic acid bacteria in maize micro and bucket-silos. Front. Microbiol. 2018, 9, 2861. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.R.; Dong, Z.H.; Li, J.F.; Chen, L.; Shao, T. Pediococcus acidilactici strains as silage inoculants for improving the fermentation quality, nutritive value and in vitro ruminal digestibility in different forages. J. Appl. Microbiol. 2019, 126, 424–434. [Google Scholar] [CrossRef]
- Yang, J.; Tan, H.S.; Cai, Y.M. Characteristics of lactic acid bacteria isolates and their effect on silage fermentation of fruit residues. J. Dairy Sci. 2016, 99, 5325–5334. [Google Scholar] [CrossRef]
- Agarussi, M.C.N.; Pereira, O.G.; da Silva, V.P.; Leandro, E.S.; Ribeiro, K.G.; Santos, S.A. Fermentative profile and lactic acid bacterial dynamics in non-wilted and wilted alfalfa silage in tropical conditions. Mol. Biol. Rep. 2019, 46, 451–460. [Google Scholar] [CrossRef]
- McDonald, P.; Henderson, A.R.; Heron, S.J.E. The Biochemistry of Silage; Chalcombe Publications: Marlow, UK, 1991. [Google Scholar]
- Ge, G.T.; Hou, M.L.; Liu, T.Y.; Jia, Y.S.; Cai, Y.M. Microbial population, chemical composition and silage fermentation of native grasses growing on the Inner Mongolian Plateau. Grassl. Sci. 2018, 64, 226–233. [Google Scholar] [CrossRef]
- Amer, S.; Hassanat, F.; Berthiaume, R.; Seguin, P.; Mustafa, A.F. Effects of water soluble carbohydrate content on ensiling characteristics, chemical composition and in vitro gas production of forage millet and forage sorghum silages. Anim. Feed Sci. Tech. 2012, 177, 23–29. [Google Scholar] [CrossRef]
- Chiy, P.C.; Phillips, C.J. Effects of sodium fertiliser on the distribution of trace elements, toxic metals and water-soluble carbohydrates in grass and clover fractions. J. Sci. Food Agric. 1999, 79, 2017–2024. [Google Scholar] [CrossRef]
- Bu, Z.K.; Ge, G.T.; Jia, Y.S.; Du, S. Effect of hay with or without concentrate or pellets on growth performance and meat quality of Ujimqin lambs on the Inner Mongolian Plateau. Anim. Sci. J. 2021, 92, e13553. [Google Scholar] [CrossRef] [PubMed]
- Agarussi, M.C.N.; Pereira, O.G.; de Paula, R.A.; de Silva, V.P.; Roseira, J.P.S.; ESilva, F.F. Novel lactic acid bacteria strains as inoculants on alfalfa silage fermentation. Sci. Rep. 2019, 9, 8007. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.C.; Li, Y.Y.; Lu, Q.; Sun, L.; Du, S.; Liu, T.; Hou, M.; Ge, G.T.; Wang, Z.J.; Jia, Y.S. Effects of lactic acid bacteria additives on the quality, volatile chemicals and microbial community of leymus chinensis silage during aerobic exposure. Front. Microbiol. 2022, 13, 938153. [Google Scholar] [CrossRef] [PubMed]
- You, L.J.; Bao, W.C.; Yao, C.Q.; Zhao, F.Y.; Hao, J.; Huang, W.Q.; Li, B.H.; Kwok, L.Y.; Liu, W.J. Changes in chemical composition, structural and functional microbiome during alfalfa Medicago sativa ensilage with Lactobacillus plantarum PS-8. Anim. Nutr. 2022, 9, 100–109. [Google Scholar] [CrossRef] [PubMed]
- Larsen, S.U.; Hjort-Gregersen, K.; Vazifehkhoran, A.H.; Triolo, J.M. Co-ensiling of straw with sugar beet leaves increases the methane yield from straw. Bioresour. Technol. 2017, 245, 106–115. [Google Scholar] [CrossRef]
- Jiang, F.G.; Cheng, H.J.; Liu, D.; Wei, C.; An, W.J.; Wang, Y.F.; Sun, H.T.; Song, E.L. Treatment of whole-plant corn silage with lactic acid bacteria and organic acid enhances quality by elevating acid content, reducing pH, and inhibiting undesirable microorganisms. Front. Microbiol. 2020, 11, 593088. [Google Scholar] [CrossRef]
- Muck, R.E. Silage microbiology and its control through additives. Rev. Bras. Zootecn. 2010, 39, 183–191. [Google Scholar] [CrossRef]
- Sousa, D.O.; Hansen, H.H.; Nussio, L.G.; Nadeau, E. Effects of wilting and ensiling with or without additive on protein quality and fermentation of a lucerne-white clover mixture. Anim. Feed Sci. 2019, 258, 114301. [Google Scholar] [CrossRef]
- Muck, R.E. Factors influencing silage quality and their implications for management. J. Dairy Sci. 1988, 71, 2992–3002. [Google Scholar] [CrossRef]
- Huang, R.Z.; Wang, X.Z.; Ma, C.H.; Zhang, F.F. Effects of intrinsic tannins on proteolysis dynamics, protease activity, and metabolome during sainfoin ensiling. Front. Microbiol. 2022, 13, 976118. [Google Scholar] [CrossRef] [PubMed]
- Hashemzadeh-Cigari, F.; Khorvash, M.; Ghorbani, G.R.; Ghasemi, E.; Taghizadeh, A.; Kargar, S.; Yang, W.Z. Interactive effects of molasses by homofermentative and heterofermentative inoculants on fermentation quality, nitrogen fractionation, nutritive value and aerobic stability of wilted alfalfa Medicago sativa L. silage. J. Anim. Physiol. Anim. Nutr. 2013, 98, 290–299. [Google Scholar] [CrossRef] [PubMed]
- Kung, L.M.; Shaver, R.D.; Grant, R.J.; Schmidt, R.J. Silage review: Interpretation of chemical, microbial, and organoleptic components of silages. J. Dairy Sci. 2018, 101, 4020–4033. [Google Scholar] [CrossRef] [PubMed]
- Dong, J.N.; Li, S.Z.; Chen, X.; Sun, Z.; Sun, Y.; Zhen, Y.G.; Qin, G.X.; Wang, T.; Demelash, N.; Zhang, X.F. Effects of Lactiplantibacillus plantarum inoculation on the quality and bacterial community of whole-crop corn silage at different harvest stages. Chem. Biol. Technol. Agric. 2022, 9, 57. [Google Scholar] [CrossRef]
- Lu, Y.X.; Li, P.; Bai, S.Q.; Chen, S.Y.; Zhao, M.; Gou, W.L.; You, M.H.; Cheng, Q.M. Effect of phenyllactic acid on silage fermentation and bacterial community of reed canary grass on the Qinghai Tibetan Plateau. BMC Microbiol. 2022, 22, 83. [Google Scholar] [CrossRef] [PubMed]
- Adesogan, A.T.; Salawu, M.B. The effect of different additives on the fermentation quality, aerobic stability and in vitro digestibility of pea/wheat bi-crop silages containing contrasting pea to wheat ratios. Grass Forage Sci. 2002, 57, 25–32. [Google Scholar] [CrossRef]
- Sucu, E.; Çifci, E.A. Effects of lines and inoculants on nutritive value and production costs of triticale silages. Rev. Bras. Zootec. 2016, 45, 355–364. [Google Scholar] [CrossRef]
- Yang, L.L.; Yuan, X.J.; Li, J.F.; Dong, Z.H.; Shao, T. Dynamics of microbial community and fermentation quality during ensiling of sterile and nonsterile alfalfa with or without Lactobacillus plantarum inoculant. Bioresour. Technol. 2019, 275, 280–287. [Google Scholar] [CrossRef]
- Cui, X.P.; Yang, Y.X.; Zhang, M.J.; Jiao, F.; Gan, T.T.; Lin, Z.W.; Huang, Y.Z.; Wang, H.X.; Liu, S.; Bao, L.J.; et al. Optimized ensiling conditions and microbial community in mulberry leaves silage with inoculants. Front. Microbiol. 2022, 13, 813363. [Google Scholar] [CrossRef]
- Du, Z.M.; Sun, L.; Chen, C.; Lin, J.; Yang, F.Y.; Cai, Y.M. Exploring microbial community structure and metabolic gene clusters during silage fermentation of paper mulberry, a high-protein woody plant. Anim. Feed Sci. Tech. 2020, 275, 114766. [Google Scholar] [CrossRef]
- Tian, H.C.; Zhu, Y.C.; Dai, M.X.; Li, T.; Guo, Y.Q.; Deng, M.; Sun, B.L. Additives altered bacterial communities and metabolic profiles in silage hybrid pennisetum. Front. Microbiol. 2022, 12, 770728. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Yin, X.J.; Wang, S.R.; Li, J.F.; Dong, Z.H.; Shao, T. Changes in the fermentation products, taxonomic and functional profiles of microbiota during high-moisture sweet sorghum silage fermentation. Front. Microbiol. 2022, 13, 967624. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.L.; Li, M.X.; Xu, J.Y.; Sun, H.; Cheng, Q.M.; Xie, Y.X.; Wang, C.M.; Chen, C.; Li, P. Effects of different cutting methods and additives on the fermentation quality and microbial community of Saccharum arundinaceum silage. Front. Microbiol. 2022, 13, 999881. [Google Scholar] [CrossRef] [PubMed]
- Ren, H.W.; Feng, Y.P.; Pei, J.W.; Li, J.P.; Wang, Z.Y.; Fu, S.F.; Zheng, Y.; Li, Z.Z.; Peng, Z.P. Effects of Lactobacillus plantarum additive and temperature on the ensiling quality and microbial community dynamics of cauliflower leaf silages. Bioresour. Technol. 2020, 307, 123238. [Google Scholar] [CrossRef]
- Madigan, M.T. Brock Biology of Microorganisms; Pearson/Prentice Hall: Hoboken, NJ, USA, 2006. [Google Scholar]
- Zhao, X.L.; Liu, J.H.; Liu, J.J.; Yang, F.Y.; Zhu, W.B.; Yuan, X.F.; Hu, Y.G.; Cui, Z.J.; Wang, X.F. Effect of ensiling and silage additives on biogas production and microbial community dynamics during anaerobic digestion of switchgrass. Bioresour. Technol. 2017, 241, 349–359. [Google Scholar] [CrossRef]
- Henderson, G.; Cox, F.; Ganesh, S.; Jonker, A.; Young, W.; Collaborators, G.R.C.; Janssen, P.H. Rumen microbial community composition varies with diet and host, but a core microbiome is found across a wide geographical range. Sci. Rep. 2015, 5, 14567. [Google Scholar] [CrossRef]
- You, J.T.; Zhang, H.; Zhu, H.F.; Xue, Y.L.; Cai, Y.M.; Zhang, G.J. Microbial community, fermentation quality, and in vitro degradability of ensiling caragana with lactic acid bacteria and rice bran. Front. Microbiol. 2022, 13, 804429. [Google Scholar] [CrossRef]
- Li, H.L.; Zeng, T.R.; Du, Z.C.; Dong, X.T.; Xin, Y.F.; Wu, Y.S.; Huang, L.K.; Liu, L.; Kang, B.; Jiang, D.M.; et al. Assessment on the fermentation quality and bacterial community of mixed silage of faba bean with forage wheat or oat. Front. Microbiol. 2022, 13, 875819. [Google Scholar] [CrossRef]
- Ogunade, I.M.; Jiang, Y.; Pech Cervantes, A.A.; Kim, D.H.; Oliveira, A.S.; Vyas, D.; Weinberg, Z.G.; Jeong, K.C.; Adesogan, A.T. Bacterial diversity and composition of alfalfa silage as analyzed by Illumina MiSeq sequencing: Effects of Escherichia coli O157:H7 and silage additives. J. Dairy Sci. 2018, 101, 2048–2059. [Google Scholar] [CrossRef]
- Wang, X.F.; Haruta, S.; Wang, P.; Ishii, M.; Igarashi, Y.; Cui, Z. Diversity of a stable enrichment culture which is useful for silage inoculant and its succession in alfalfa silage. FEMS Microbiol. Ecol. 2006, 57, 106–115. [Google Scholar] [CrossRef]
- Ostling, C.E.; Lindgren, S.E. Inhibition of enterobacteria and listeria growth by lactic, acetic and formic acids. J. Bacteriol. 1993, 75, 18–24. [Google Scholar] [CrossRef]
- Xu, J.Y.; Zhang, K.Y.; Lin, Y.F.; Li, M.X.; Wang, X.K.; Yu, Q.; Sun, H.; Cheng, Q.M.; Xie, Y.X.; Wang, C.M.; et al. Effect of cellulase and lactic acid bacteria on the fermentation quality, carbohydrate conversion, and microbial community of ensiling oat with different moisture contents. Front. Microbiol. 2022, 13, 1013258. [Google Scholar] [CrossRef]
- Fang, D.; Dong, Z.H.; Wang, D.L.; Lin, B.; Shi, P.B.; Yan, J.; Zhuang, D.Y.; Shao, T.; Wang, W.Y.; Gu, M.F. Evaluating the fermentation quality and bacterial community of high-moisture whole-plant quinoa silage ensiled with different additives. J. Appl. Microbiol. 2022, 132, 3578–3589. [Google Scholar] [CrossRef] [PubMed]
- Du, S.; You, S.H.; Jiang, X.W.; Li, Y.Y.; Jia, Y.S. Dynamics of the fermentation quality and microbiotsa in Ephedra sinica treated native grass silage. J. Appl. Microbiol. 2022. online ahead of print. [Google Scholar] [CrossRef]
- Tambong, J.T. Taxogenomics and systematics of the genus pantoea. Front. Microbiol. 2019, 10, 2463. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.W.; Tao, X.Y.; Liu, Q.S.; Zhang, Y.J.; Xu, J.B.; Zhang, W.Y.; Wang, J.; Zhang, D.D.; Li, B.; Wang, L.L.; et al. Succession changes of fermentation parameters, nutrient components and bacterial community of sorghum stalk silage. Front. Microbiol. 2022, 13, 982489. [Google Scholar] [CrossRef] [PubMed]
- Koivula, T.T.; Juvonen, R.; Haikara, A.; Suihko, M.L. Characterization of the brewery spoilage bacterium obesumbacterium proteus by automated ribotyping and development of PCR methods for its biotype 1. J. Appl. Microbiol. 2006, 100, 398–406. [Google Scholar] [CrossRef]
- Yuan, X.J.; Li, J.F.; Dong, Z.H.; Shao, T. The reconstitution mechanism of napier grass microiota during the ensiling of alfalfa and their contributions to fermentation quality of silage. Bioresour. Technol. 2020, 297, 122391. [Google Scholar] [CrossRef]
- Bai, J.; Ding, Z.T.; Ke, W.C.; Xu, D.M.; Wang, M.S.; Huang, W.K.; Zhang, Y.X.; Liu, F.; Guo, X.S. Different lactic acid bacteria and their combinations regulated the fermentation process of ensiled alfalfa: Ensiling characteristics, dynamics of bacterial community and their functional shifts. Microb. Biotechnol. 2021, 14, 1171–1182. [Google Scholar] [CrossRef]
- Xu, D.M.; Wang, N.; Rinne, M.; Ke, W.C.; Weinberg, Z.G.; Da, M.; Bai, J.; Zhang, Y.X.; Li, F.H.; Guo, X.S. The bacterial community and metabolome dynamics and their interactions modulate fermentation process of whole crop corn silage prepared with or without inoculants. Microb. Biotechnol. 2021, 14, 561–576. [Google Scholar] [CrossRef]
- Su, W.F.; Jiang, Z.P.; Hao, L.H.; Li, W.T.; Gong, T.; Zhang, Y.; Du, S.; Wang, C.; Lu, Z.P.; Jin, M.L.; et al. Variations of soybean meal and corn mixed substrates in physicochemical characteristics and microbiota during two-stage solid-state fermentation. Front. Microbiol. 2021, 12, 688839. [Google Scholar] [CrossRef] [PubMed]
- Flythe, M.D.; Russell, J.B. The effect of pH and a bacteriocin bovicin HC5 on clostridium sporogenes MD1, a bacterium that has the ability to degrade amino acids in ensiled plant materials. FEMS Microbiol. Ecol. 2004, 47, 215–222. [Google Scholar] [CrossRef] [PubMed]
- Bai, J.; Ding, Z.T.; Su, R.N.; Wang, M.S.; Cheng, M.Y.; Xie, D.M.; Guo, X.S. Storage temperature is more effective than lactic acid bacteria inoculations in manipulating fermentation and bacterial community diversity, co-occurrence and functionality of the whole-plant corn silage. Microbiol. Spectrum. 2022, 10, e0010122. [Google Scholar] [CrossRef]
- Kilstrup, M.; Hammer, K.; Jensen, P.R.; Martinussen, J. Nucleotide metabolism and its control in lactic acid bacteria. FEMS Microbiol. Rev. 2005, 29, 555–590. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.Y.; Du, S.; Sun, L.; Cheng, Q.M.; Hao, J.F.; Lu, Q.; Ge, G.T.; Wang, Z.J.; Jia, Y.S. Effects of lactic acid bacteria and molasses additives on dynamic fermentation quality and microbial community of native grass silage. Front. Microbiol. 2022, 13, 830121. [Google Scholar] [CrossRef] [PubMed]



| Characteristics | ZX301 | YX34 |
|---|---|---|
| Sources | Stipa grandis silage | Stipa grandis silage |
| Shape | Cocci | Rod |
| Gram stain | + | + |
| Gas for glucose | - | - |
| Catalase | - | - |
| Fermentation type | Homo | Homo |
| Growth at temperature | ||
| 5 | - | - |
| 10 | + | + |
| 15 | + | + |
| 30 | + | + |
| 45 | w | - |
| 50 | - | - |
| Growth at pH | ||
| 3 | - | - |
| 3.5 | + | + |
| 4 | + | + |
| 5 | + | + |
| 6 | + | + |
| 7 | + | + |
| 8 | + | + |
| Growth in NaCl (%) | ||
| 3 | + | + |
| 6.5 | + | + |
| Items | ZX301 | YX34 |
|---|---|---|
| L-arabinose | + | + |
| Ribose | + | + |
| D-xylose | + | - |
| D-Galactose | + | + |
| D-Glucose | + | + |
| D-Fructose | + | + |
| D-Mannose | + | + |
| L-Sorbose | - | w |
| Mannitol | + | - |
| Sorbitol | + | - |
| Methyl-α D-Mannopyranoside | + | - |
| Methyl-α D-Glucopyranoside | + | - |
| N-Acetyl Glucosamine | + | + |
| Laetrile | + | + |
| Arbutin | + | + |
| Aescin | + | + |
| Salicin | + | + |
| Cellobiose | + | + |
| Maltose | + | + |
| Lactose | + | + |
| Honey disaccharide | + | + |
| Sucrose | + | + |
| Trehalose | + | + |
| Trisaccharide | + | - |
| Raffinose | + | - |
| Starch | w | - |
| Xylitol | + | - |
| Gentiobiose | w | + |
| D-Arabinitol | w | - |
| Gluconate | w | - |
| Strain | Accession Number | 16S rRNA Gene Sequencing Data (Closest Relative) | Similarity (%) |
|---|---|---|---|
| ZX301 | KX886792.1 | Pediococcus pentosaceus DSM 20336 T | 100 |
| YX34 | NR_115605.1 | Lactobacillus plantarum JCM 1149 | 100 |
| Items | Stipa grandis |
|---|---|
| Dry matter (g/kg FW) | 50.71 ± 0.22 |
| Crude protein (g/kg DM) | 8.91 ± 0.10 |
| Neutral detergent fiber (g/kg DM) | 75.46 ± 0.72 |
| Acid detergent fiber (g/kg DM) | 39.79 ± 0.78 |
| Water-soluble carbohydrates (g/kg DM) | 2.52 ± 0.09 |
| LAB (log10 cfu/g FM) | 5.56 ± 0.10 |
| Aerobic bacteria (log10 cfu/g FM) | 8.03 ± 0.45 |
| Coliform bacteria (log10 cfu/g FM) | 6.25 ± 0.27 |
| Yeasts (log10 cfu/g FM) | 7.01 ± 1.33 |
| Mold (log10 cfu/g FM) | ND |
| Items | CK | ZX301 | YX34 | P | SEM | p-Value |
|---|---|---|---|---|---|---|
| DM (%) | 50.97 | 50.41 | 50.99 | 50.29 | 0.0024 | 0.7162 |
| WSC (% DM) | 1.27 b | 1.55 a | 1.76 a | 1.83 a | 0.0007 | 0.0001 |
| CP (% DM) | 7.99 | 7.92 | 7.93 | 7.96 | 0.0308 | 0.0785 |
| ADF (% DM) | 70.35 | 70.15 | 72.09 | 69.62 | 0.0042 | 0.2207 |
| NDF (% DM) | 38.67 | 38.97 | 39.17 | 38.03 | 0.0031 | 0.7144 |
| pH | 4.15 a | 4.00 b | 4.01 b | 4.01 b | 0.0191 | 0.0007 |
| Lactic acid (g/kg) | 7.42 b | 10.96 a | 11.14 a | 11.23 a | 0.0479 | 0.0001 |
| Acetic acid (g/kg) | 4.68 a | 3.79 b | 3.95 b | 3.07 c | 0.0172 | 0.0001 |
| Propionic acid (g/kg) | ND | ND | ND | ND | ||
| Butyric acid (g/kg) | ND | ND | ND | ND | ||
| NH3-N | 1.38 a | 0.87 b | 0.74 b | 0.88 b | 0.0768 | 0.0015 |
| LAB (log10 cfu/g FM) | 8.20 a | 7.28 ab | 6.36 b | 5.69 b | 0.3468 | 0.0396 |
| Aerobic bacteria (log10 cfu/g FM) | 4.13 | 4.21 | 4.06 | 3.59 | 0.1293 | 0.3999 |
| Coliform bacteria (log10 cfu/g FM) | ND | ND | ND | ND | ||
| Yeasts (log10 cfu/g FM) | ND | ND | ND | ND | ||
| Mold (log10 cfu/g FM) | ND | ND | ND | ND |
| Items | FM | CK | ZX301 | YX34 | P | SEM | p-Value |
|---|---|---|---|---|---|---|---|
| OTUs | 85 | 103 | 105 | 99 | 121 | 4.8563 | 0.2799 |
| ACE | 144.15 | 116.38 | 132.13 | 131.77 | 135.80 | 5.4811 | 0.7038 |
| Chao1 | 130.11 | 118.12 | 129.53 | 123.04 | 134.34 | 4.5630 | 0.8751 |
| Simpson | 0.1140 b | 0.2717 b | 0.5979 a | 0.2277 b | 0.0924 b | 0.0536 | 0.0034 |
| Shannon | 0.5305 b | 1.0725 b | 1.7554 a | 0.7079 b | 0.4884 b | 0.1374 | 0.0025 |
| Goods’ coverage | 99.92 b | 99.96 a | 99.91 b | 99.95 a | 99.95 a | 0.0001 | 0.0173 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, M.; Wang, Y.; Wang, Z.; Bao, J.; Zhao, M.; Ge, G.; Jia, Y.; Du, S. Effects of Isolated LAB on Chemical Composition, Fermentation Quality and Bacterial Community of Stipa grandis Silage. Microorganisms 2022, 10, 2463. https://doi.org/10.3390/microorganisms10122463
Liu M, Wang Y, Wang Z, Bao J, Zhao M, Ge G, Jia Y, Du S. Effects of Isolated LAB on Chemical Composition, Fermentation Quality and Bacterial Community of Stipa grandis Silage. Microorganisms. 2022; 10(12):2463. https://doi.org/10.3390/microorganisms10122463
Chicago/Turabian StyleLiu, Mingjian, Yu Wang, Zhijun Wang, Jian Bao, Muqier Zhao, Gentu Ge, Yushan Jia, and Shuai Du. 2022. "Effects of Isolated LAB on Chemical Composition, Fermentation Quality and Bacterial Community of Stipa grandis Silage" Microorganisms 10, no. 12: 2463. https://doi.org/10.3390/microorganisms10122463
APA StyleLiu, M., Wang, Y., Wang, Z., Bao, J., Zhao, M., Ge, G., Jia, Y., & Du, S. (2022). Effects of Isolated LAB on Chemical Composition, Fermentation Quality and Bacterial Community of Stipa grandis Silage. Microorganisms, 10(12), 2463. https://doi.org/10.3390/microorganisms10122463






