1. Introduction
Leishmaniasis is a vector-borne disease caused by an intracellular parasite of the genus
Leishmania, which is transmitted by bloodsucking dipterans. The disease is among the top three vector-borne diseases, along with malaria and filariasis [
1].
Cutaneous leishmaniasis (CL) is the most widespread and common form of the disease and causes three forms of skin lesions (ulcers, ulcero-crusted lesions, and nodular lesions) on exposed parts of the body, leaving lifelong scars and serious disability or stigma, especially among young women. The number of CL cases in the world is as high as one million, according to the World Health Organization [
2].
In Morocco, three species of
Leishmania are known to be causative agents of CL: the two most dominant species are
L.
major and
L.
tropica, the third species is
L.
infantum, which is less common than the other two
Leishmania species. The distribution of CL due to
L.
infantum in Morocco is not well defined; it is frequently found in
L.
tropica foci and even in
L.
major foci in pre-Saharan areas [
3].
Amongst the three clinically important
Leishmania species,
L. tropica has the largest geographic distribution and is considered to be a public health threat by the Moroccan Ministry of Health [
4].
L. tropica was reported for the first time in the rural locality of Tanant (Azilal province, High Atlas) in 1987 [
5]. Subsequently,
L. tropica cases have increasingly been reported from different regions of the country; in 1991, Pratlong et al. [
6] unveiled a vast
L. tropica focus in central and southern areas such as Guelmim, Agadir, and Essaouira. Since that time and until today, the
L. tropica-induced CL has continued to spread throughout the country, affecting Taza, Zouagha Moulay Yaacoub, Chichaoua, Settat, and Lbrouj [
7,
8,
9,
10]. There has also been an overlap and an integration of
L. tropica in certain
L. infantum foci, such as Ouazzane, and
L. major foci, such as Toundout, Agdz, and Ouarzazate [
11,
12]. In Morocco, anthroponotic transmission is, so far, the only recognized mode, in spite of recordings of
L. tropica infection in animal hosts [
13]; therefore, the transmission cycle of
L. tropica involves a vector
Phlebotomus sergenti, which is essential for the emergence of the disease, and a vertebrate host, which is the human.
P. sergenti is characterized by a wide geographical distribution in the Mediterranean basin; its density is significant in arid and Saharan zones, and it can be found in mid-to-high altitudes [
13].
The pathogens, hosts, and vectors involved in the transmission cycle of vector-borne diseases are environmentally sensitive [
14]. Leishmaniasis has been reported to be impacted by climate change and human modification of ecosystems [
3].
Valero and Uriarte [
15] divided risk factors into three categories: socioeconomic, environmental, and climatic variables; the latter showed significant associations with the incidence of both visceral leishmaniasis (VL) and CL. Socioeconomic factors were also associated with disease incidence in the vulnerable human populations of arid and tropical developing regions [
15]. The conversion of natural forest to other land uses in the last few decades has led to habitat fragmentation and altered landscape composition [
16]. The spread of the vector and disease at macroscales is associated with migration and the expansion of human populations into natural areas, the creation of roadways, energy networks, new farmlands, and poorly planned urban development [
17,
18,
19,
20]; climatic conditions are generally important risk factors for vector-borne diseases [
21]. However, overall, it is not well understood how the interactions among all of the climatic, environmental, and socioeconomic factors influence the risk of leishmaniasis.
In Morocco, a few studies have been carried out to elucidate the impact of biotic and abiotic factors on the occurrence and repartition of CL [
22,
23]. However, it is difficult to make a definitive statement about the link between these risk factors and the epidemiology of leishmaniasis, especially CL, due to
L. tropica, which continues to evolve in terms of both the number of human cases and the repartition area.
To contribute to a better understanding of the epidemiology of CL induced by L. tropica and the impact of climatic and environment factors on this form of leishmaniasis, we carried out a survey in three CL foci: Foum Jemaa, Imintanout, and Ouazzane. In addition to genotyping, we studied the population structure of L. tropica collected in the three regions, using the evolutionary marker internal transcribed spacer 1 (ITS1). We also investigated whether the mean temperature, annual rainfall, relative humidity, average wind speed, and vegetation index influenced the increase in leishmaniasis incidence in the three CL foci.
2. Materials and Methods
2.1. Study Area
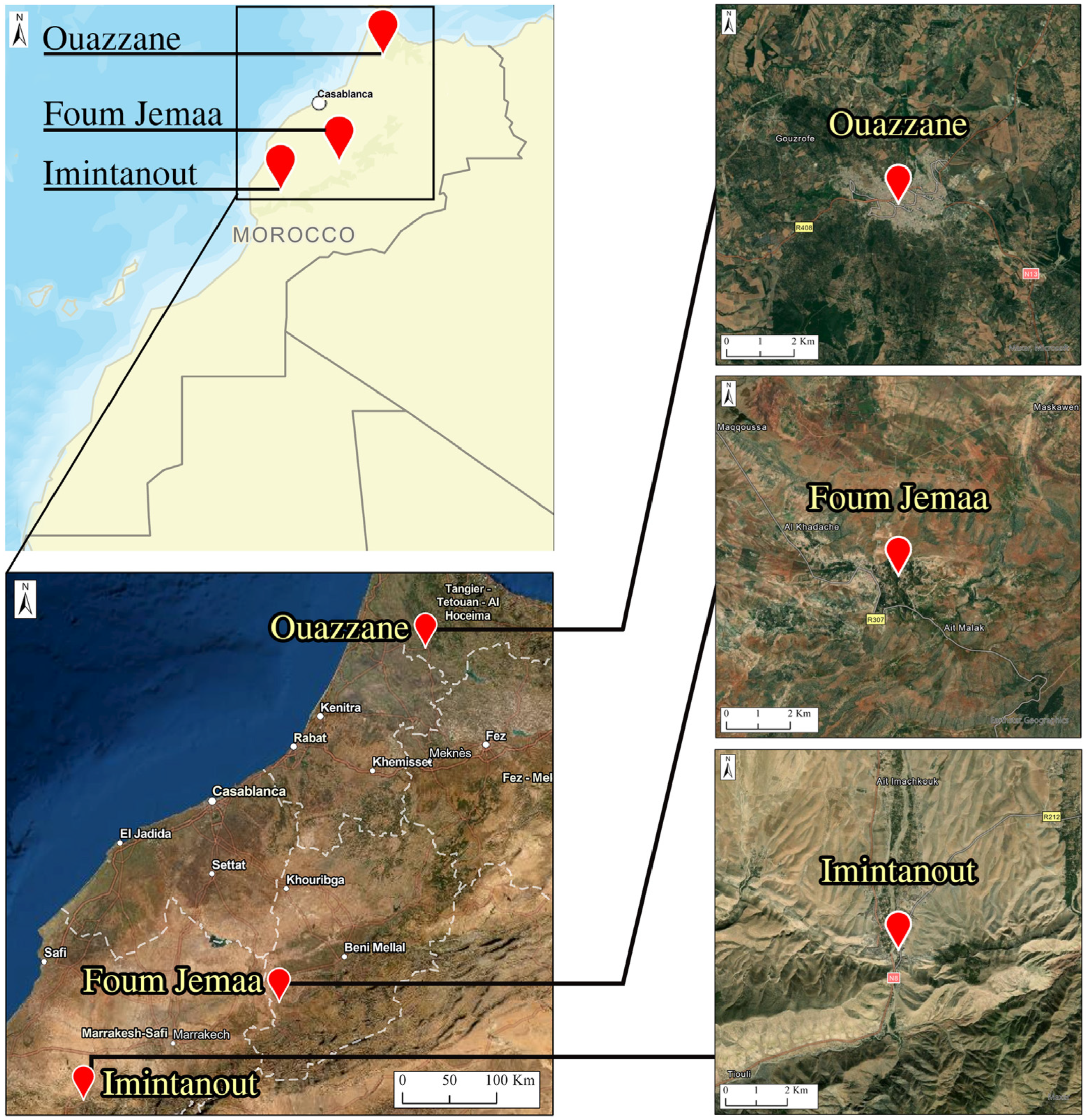
This study enrolled patients originating from three regions known as CL endemic foci: the Foum Jemaa locality in the center of Morocco (31°57′52.03″ N, 6°59′26.22″ W) (744 m asl), High Atlas in Azilal Province; Imintanout village (31°10′13.90″ N, 8°50′45.86″ W) (912 m asl), located in the southwest of the country, in the province of Chichaoua; and the periphery of the city of Ouazzane (34°47′40.89″ N, 5°34′4.49″ W) (614 m asl), in northern Morocco (
Figure 1).
2.2. Ethical Considerations
Informed consent was obtained from all adults who participated in the study. Consent for the inclusion of young children was obtained from parents or guardians. The study was reviewed and approved by the Ethical Review Committee for Biomedical Research (CERB) of the Faculty of Medicine and Pharmacy, Rabat, Morocco (IORG 0006594 FWA 00024287).
2.3. Patients and Sampling
During an active screening campaign for CL cases performed for 1 week each year between 2018 and 2020 in the three CL foci, samples were collected from 80 patients. All recruited patients presented skin lesions clinically suggestive of CL, and were never treated by Glucantime injection. Pregnant women and patients presenting chronic illness (e.g., blood pressure issues, diabetes, etc.) were not skin sampled.
For each consenting patient, before sampling, we acquired a completed structured questionnaire that included the relevant personal, epidemiological, and clinical data.
After cleaning the lesions, a gentle scraping at the edge of the lesion was performed, and the collected skin scraping was spread on slide smears for amastigote detection. A cotton swab was also used for molecular studies, as described by Daoui, O et al. [
11].
We cultured Leishmania on an RPMI 1640 medium (Biowest, Nuaille, France) supplemented with 2 mM l-glutamine (Eurobio, Les Ulis, France), 10% fetal bovine serum (Biowest, Nuaille, France), and 1% penicillin/streptomycin (100 U/mL penicillin and 100 μg/mL streptomycin; Biowest, Nuaille, France), followed by incubation at 25 °C.
2.4. Population at Risk
The population census estimation in each province was obtained from the Epidemiology and Disease Control Department, Moroccan Ministry of Health, for the period 2000–2019, except for Ouazzane, where the period was shorter (2009 to 2019) due to a lack of information, justified by the fact that CL cases in Ouazzane were very low and were, therefore, added to the closest province to Ouazzane. The annual incidence rate of CL in each province was defined as follows: incidence rate = (total number of CL cases per year/total population at risk) × 100,000 (
Table S1).
2.5. Meteorological Parameters
The meteorological data were obtained from the National Meteorology Department of Morocco and from TuTiempo.net (powered by Tutiempo Network, S.L., Madrid, Spain), which is a low-cost, citizen-based PM sensor network system that has been deployed globally. TuTiempo.net offers measurements of climate data and meteorological parameters (temperature, rainfall, humidity, and wind speed) using satellite images.
2.6. Normalized Difference Vegetation Index
This work was based on a time series of Landsat 5 and 8 satellite images, which were downloaded from USGS, Earth explorer (NASA),
https://earthexplorer.usgs.gov (accessed on 10 May 2021). We used a set of images covering two decades. This allowed us to obtain at least one image every 5 or 6 years from 2000 to 2019, for the whole study area of the three foci; after an atmospheric correction of the images, those from Landsat 5 were used to calculate the NDVI for the years 2000, 2005, and 2010, whereas those from Landsat 8 were used to calculate the NDVI for the years 2015 and 2019 with ArcMap 10.8 software (Casablanca, Morocco). The methodology is summarized in
Figure 2.
The NDVI calculation formula is as follows: (NIR − R)/(NIR + R), where NIR is the near-infrared band, and R is the red band.
2.7. Statistical Analyses
The clinical data and the CL data reported between 2000 and 2019 were analyzed using the statistical software R, version 4.1.2 (
http://www.R-project.org, accessed on 15 June 2021). The linear regression model was applied to assess the impact of some of the investigated factors. The relationship between the incidence of CL and environmental factors (temperature, rainfall, humidity, wind speed, and vegetation index) was tested using Pearson’s rank correlation, as previously described by [
24]. Differences were considered significant when
p < 0.05. Standardized principal component analysis (PCA) was carried out to generate an integrative description of the data (incidence of the disease and the environmental data) over the years. The incidence of the disease in each year was analyzed using the environmental data of the year before (T − 1), because, according to a normal
Leishmania cycle of infection, sandflies only start their activities in the summer; thus, counting the incubation time in human hosts, cases tend to appear at the end of the year and are, therefore, counted the year after. Additionally, the clinical data were analyzed using Fisher’s exact test; a comparison between the number of males and females carrying CL lesions, as well as the distribution of lesions (face and upper limbs), was performed. One sample
t-test was used to calculate the statistically significant differences between the distribution of CL, age groups, and type of lesions. One sample
t-test was also used to calculate the statistically significant differences between the three diagnostic methods performed in this study. Differences between groups were considered to be statistically significant at
p < 0.05.
2.8. Parasitological and Molecular Study
The smear slides were analyzed under a microscope with a 100× immersion objective to determine the parasite load. All slides were screened more than once before giving a final result. DNA extraction was performed using swabs after phenol–chloroform extraction followed by ethanol precipitation, as described by [
25]. The DNA sample quantification was determined using NanoDrop (Thermo Fisher Scientific, Waltham, MA, USA), adjusting the final concentration of each sample to 50 ng/μL.
For
Leishmania detection and identification, we used the Nested KDNA-PCR, with two pairs of primers: the forward primer CSB2XF with the reverse primer CSB1XR for the first stage of the reaction, and the forward primer 13Z with the reverse primer LiR for the second stage of the reaction, as described by Noyes et al. [
26]. ITS1-PCR was used to study the population structure of
L. tropica collected in the three areas. The two protocols of PCR were detailed by [
11].
Electrophoresis of PCR products was performed on 1.5% and 1% agarose gels for ITS1-PCR and Nested KDNA-PCR, respectively, to which 2 μL of ethidium bromide solution (10 mg/mL) was added (Promega, Madison, WI, USA). The electrophoretic migration was carried out in 0.5× TBE buffer, and the gel was visualized under UV light.
The amplified KDNA fragments on the agarose gel were compared with standard and marker bands (100 bp DNA ladder marker, Bioline) to identify the Leishmania species.
Sequencing: the final ITS1-PCR products of about 350 bp were purified using the Exs Pure Enzymatic PCR cleanup kit (NIMAGEN, Nijmegen, The Netherlands), and then sequenced using the BrillantDye Terminator Cycle Sequencing Kit v1.1 (Nimagen, The Netherlands) and ABI PRISM 3130xL DNA automated sequencer (Applied Biosystems, Waltham, MA, USA).
Sequencing data were analyzed using Chromas v.2.6.2 software (Technelysium), aligned, and compared to entries retrieved from Genbank, using the multiple alignment program MEGA7.
2.9. Basic Statistics, Tests for Selection, and Phylogeny
Estimates of genetic diversity were assessed using DnaSP v. 4.0 and Arlequin v. 3.5 [
27]. The number of segregating sites, the number of haplotypes (H), haplotype diversity, the average number of nucleotide differences, and nucleotide diversity were calculated. To deduce genetic diversity, we used the nuclear gene ITS1. Pairwise FST values were calculated using DnaSP [
28]. Three geographic groups were defined for the comparative study: (i) Foum Jemaa samples, (ii) Imintanout samples, and (iii) Ouazzane samples. To assess whether the examined gene evolved randomly or not, Tajima’s D test [
29] and Fu’s F test [
30] were performed on DnaSP and Arlequin.
To reconstruct the phylogenetic tree, we used MrBayes v. 3.2.1 [
31]. In the first step, the optimal substitution model was estimated using MrModeltest v.2.4 [
32]. Runs were computed in MrBayes for 200,000 generations while sampling every 100 generations. Then, the tree was visualized and edited using FigTree v.1.4.2 [
33]. A haplotype network was constructed and visualized using PopArt [
34].
4. Discussion
Since the first case was reported in 1987 in Azilal Province (Tanant) [
5], CL caused by
L. tropica has spread all over the country, becoming a serious public health problem; it is also the most difficult
Leishmaniasis form to control due to its wide geographical distribution [
13]. The spread of
L. tropica is probably related to the high ecological plasticity of its vector,
Phlebotomus sergenti [
35].
Our data revealed that children under 10 years old displayed the highest rate of infection (57.5%), in concordance with previously published data on the incidence of CL in other foci throughout Morocco [
36] and in other endemic countries, where the majority of infected people were under 16 years old [
37,
38]. The prevalence of CL is reported to increase generally with age up to 15 years, after which it stabilizes, probably reflecting the progressive buildup of immune protective status [
39]. On the other hand, in line with widely known evidence reported for
L. tropica-induced CL, our results showed that the face was the most affected site, generally with single lesions [
40,
41], in contrast to
L. major, which causes multiple lesions generally located on the limbs [
42]. Analysis also showed that women were more affected than men, which can be explained by the fact that, during the hot summer nights characterizing these regions, men are known to stay and sleep outdoors (i.e., on balconies or terraces), unlike women, who are often indoors;
P. sergenti is endophilic and anthropophagic [
43]. In addition, the most common form of lesion in our study was the papulonodular form. El Hamouchi, A et al., highlighted that, in the case of CL caused by
L.
major, the most frequent clinical lesion form was the ulcero-crusted form [
36]. Clinical manifestations of
Leishmania infections depend on multifactorial parameters, such as human genetic susceptibility and the genetic background of the parasite. Factors related to the vector may also affect CL manifestations [
44].
The World Health Organization considers leishmaniasis to be a climate-sensitive disease, occupying a characteristic ‘climate space’ that is strongly affected by changes in rainfall, atmospheric temperature, and humidity [
45]. According to various studies, the incidence of leishmaniasis is influenced by a variety of environmental, landscape, and socioeconomic factors [
15,
46]. Due to the dynamic nature of leishmaniasis as a vector-borne disease, demographic factors and human activity and behaviors are factors that need to be closely monitored [
21,
47]. CL transmission is related to the various
Leishmania spp., particularly vector dynamics, which determines the presence of leishmaniasis as a function of climatic and environmental changes [
21].
Using a linear regression model, we analyzed the impact of diverse bioclimatic and environmental variables, including mean temperature, annual rainfall, relative humidity, average wind speed, and vegetation index, on the incidence of CL over 20 years. Our results differed according to the foci. Indeed, while temperature and humidity were strongly intercorrelated and significantly associated with the incidence of CL in Imintanout, southwestern Morocco, none of the risk factors studied were significantly correlated with the incidence of the disease in the other two foci: Foum Jemaa and Ouazzane in central and northern Morocco, respectively. Focusing on another aspect of
L. tropica-induced CL in the southwest of the country, no significant correlation was established between some environmental predictors such as temperature, rainfall, and altitude and the incidence of leishmaniasis [
22].
Temperature has been identified as an important risk factor associated with leishmaniasis in Mediterranean, tropical, and arid regions. Indeed, small changes in temperature have a profound impact on the developmental time and metabolism of sandflies, as well as on the
Leishmania development cycle within its vector [
48]. This could potentially affect the distribution of leishmaniasis and allow transmission of the
Leishmania parasite in areas that were previously free of leishmaniasis [
45].
In Ethiopia, a temperature range between 17.2 and 23.8 °C was associated with CL occurrence [
49]. In South America, a peak of incidence of CL was found in Chaparral, Colombia, with a mean temperature of 20.6 ± 1.4 °C [
50]; in Tunisia, a temperature of 9.4–22.1 °C contributed to 20.7% of the variation in an ecological niche model of the vector [
51].
Humidity plays an important role in the survival, development, and activity of sandflies. Indeed, the humidity level during the night influences the growth of flies and, consequently, the occurrence and distribution of leishmaniasis [
52]. In Tunisia, it was found that a relative humidity between 30% and 45% increased the ZCL incidence in an
L. major focus [
53]. In Iran, a humidity range between 27% and 30% was registered in the CL hotspot areas in Isfahan [
54].
Other environmental and climatic factors have been reported to impact the incidence and distribution of leishmaniasis in different regions of the world. Indeed, in the Andean region of Colombia and in Brazil, annual rainfall is an important risk factor for CL [
55,
56]. Furthermore, in Iran and Turkey, high NDVI values have been associated with the occurrence of CL [
57,
58].
In Sri Lanka, as in Iran, the wind speed was identified as a predictor, revealing a positive correlation with the incidence of leishmaniasis [
59,
60]. However, a negative association was demonstrated between maximum wind speed, rainfall, altitude, and vegetation cover and CL incidence in Iran [
61]. In contrast to these studies, the analysis of our data revealed that vegetation index, precipitation, and wind speed had no impact on the incidence of CL in the three study sites.
Around the world, many studies have tried to find an association between climatic and environmental factors and the occurrence of leishmaniasis; some have been able to find associations, while others have not been able to draw any significant conclusions. This discord can be explained by the fact that leishmaniasis is a multifactorial disease [
15], and climatic factors are not the only ones influencing the occurrence of the disease; other factors need to be monitored, such as demographic and social factors, as well as the density of the vector and the matter of hygiene, which is generally neglected in rural areas. The presence of waste or stables in the vicinity of houses, as well as the use of traditional building materials for house construction, are all factors that favor the development of the vector and reservoirs, resulting in an increase in leishmaniasis cases.
In addition to elucidating the climatic and environmental factors that may influence the distribution of CL, we tried to shed light on the population structure of the L. tropica parasite isolated in the three geographically distant CL foci.
Our results showed low to medium geographic differentiation among the L. tropica populations, using pairwise differentiation. The molecular diversity index values were roughly similar among the Foum Jemaa and Imintanout populations, in contrast to the Ouazzane strains. Indeed, in the latter foci, only one single haplotype was identified among the three strains collected in this region. This should be explored with a larger sample.
Haplotype diversity ranged between 0.784 and 0.789, with a total of 13 haplotypes. The analysis of ITS1-5.8S rRNA gene and ITS2 sequences from 31
P. sergenti revealed a great heterogeneity of
L. tropica isolated from
P. sergenti, segregated into 16 haplotypes with phylogenetic relatedness to Indian strains and one Moroccan strain isolated from a CL patient [
62].
By analyzing a combination of mitochondrial and nucleic genes, Fotouhi-Ardakani, et al. [
63], demonstrated that
L. tropica has high haplotype diversity. An important haplotype diversity was also highlighted by Arroub et al. [
42], who analyzed the internal transcribed spacer ITS1 and 5.8S rDNA gene sequences of
L. tropica isolated from Moroccan patients. Recently, El Kacem et al. [
64], confirmed the high intraspecific variability of
L. tropica in Morocco using the MLST approach; moreover, the MLST analysis allowed for a distinct separation of
L. tropica strains according to their geographical origin.
While haplotype diversity was high in our study, low nucleotide diversity values indicated only small differences between haplotypes. This was also clear when examining the minimum-spanning haplotype network, which mostly showed single-nucleotide differences between haplotypes.
Regarding the population differentiation, the Foum Jemaa and Imintanout populations and the Ouazzane and Imintanout populations are remotely related to each other, as indicated by the high and significant FST value.
To evaluate the population expansion of
L. tropica, multiple selective neutrality tests were performed. Statistical tests originally developed to test the selective neutrality of a mutation were implemented [
65]. We selected two tests that are frequently used to detect population growth and vary somewhat in their approach. Tajima’s D test [
29] is a common way to quantify the demographic signatures of population growth on the basis of the distribution of allele frequency of segregating nucleotide sites. A positive value points to a population with a deficiency of rare haplotypes that has experienced a recent bottleneck, whereas a negative value indicates a bias toward rare alleles, with the latter being a signature of recent population expansion. Fu’s F test is based on the distribution of alleles or haplotypes, with negative values indicating possible recent population growth, while positive values are evidence of allele deficiency, as would be expected from a recent population bottleneck [
30]. In our study, Tajima’s D test showed nonsignificant negative values for the
L. tropica population, except for the Ouazzane population, which showed a null test value. Furthermore, Fu’s F test gave nonsignificant positive values for all populations except for Ouazzane’s
L. tropica, represented by a single allele, which prevented results from being inferred.
The negative values obtained using Tajima’s D test signify an excess of low-frequency polymorphisms relative to expectation; on the other hand, the positive values obtained using Fu’s F test are evidence of allele deficiency, as would be expected from a recent population bottleneck. However, overall, in all populations, Tajima’s D and Fu’s F test statistics were not statistically significant, indicating consistency with a population at a drift–mutation equilibrium.
Further analysis, including additional DNA markers and a larger sample size, could provide a more complete perspective on L. tropica’s population structure in these three regions.