Metagenomics Reveal Microbial Effects of Lotus Root–Fish Co-Culture on Nitrogen Cycling in Aquaculture Pond Sediments
Abstract
:1. Introduction
2. Materials and Methods
2.1. Experiment Design
2.2. Sample Sampling and Pre-Treatment
2.3. Physicochemical Analysis
2.4. DNA Extraction and Metagenomic Sequencing
2.5. Statistical Analysis
3. Results
3.1. Sediment Physicochemical Properties
3.2. Microbial Community Structure
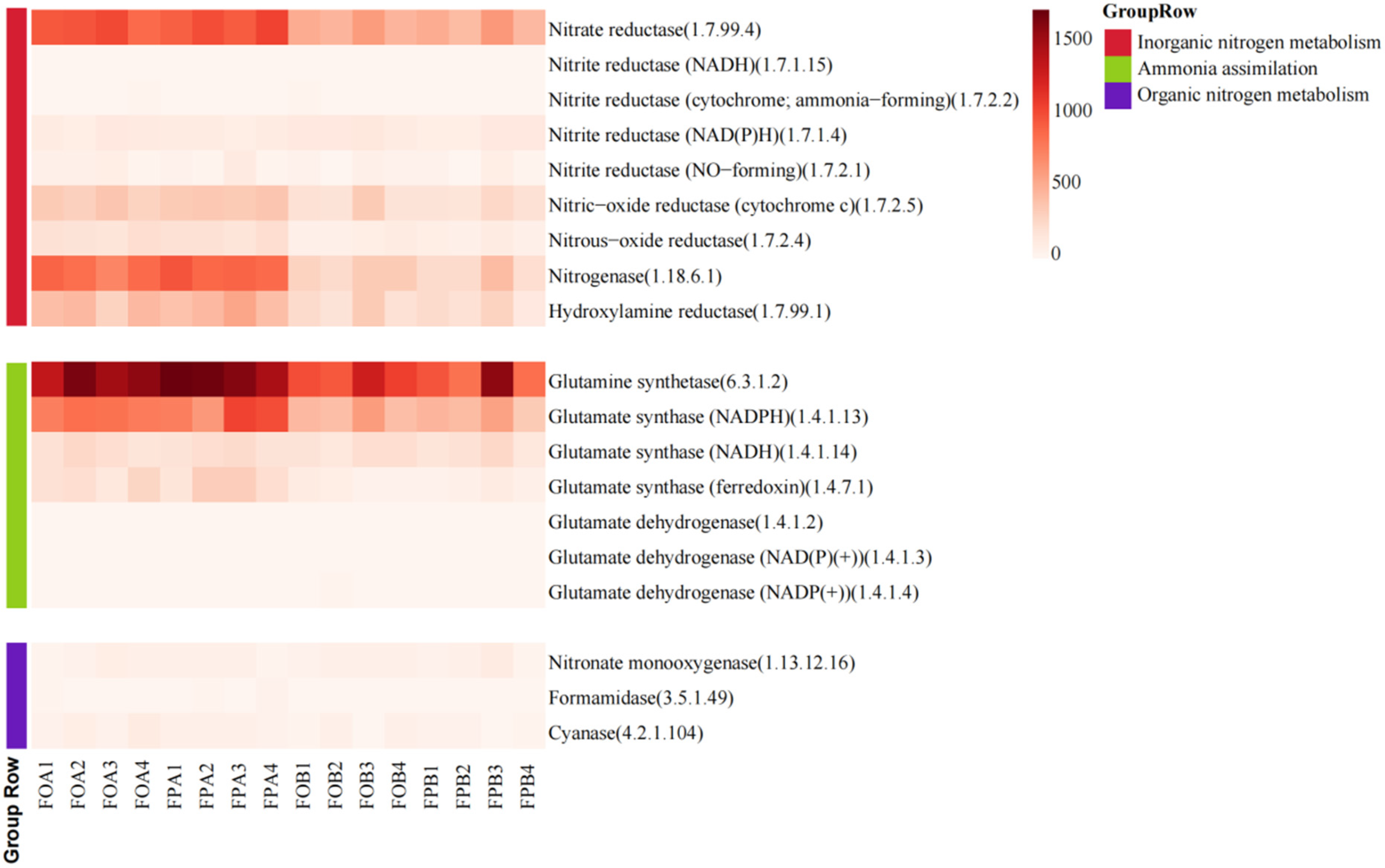
3.3. Nitrogen Cycling Enzyme Family
3.4. Nitrogen Cycle Pathways and Functional Gene Family
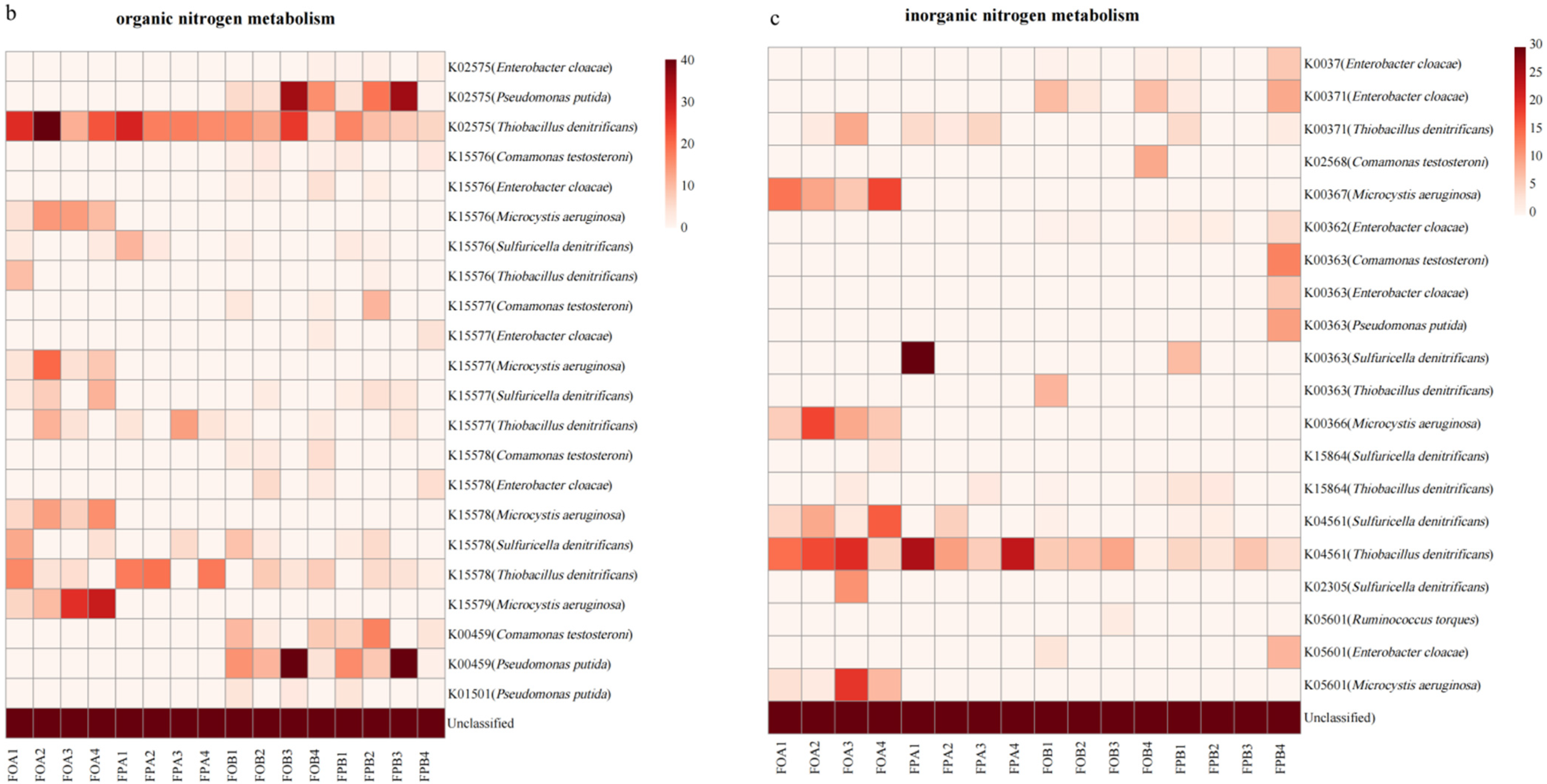
3.5. Gene Abundance of Nitrogen Cycle Microorganisms
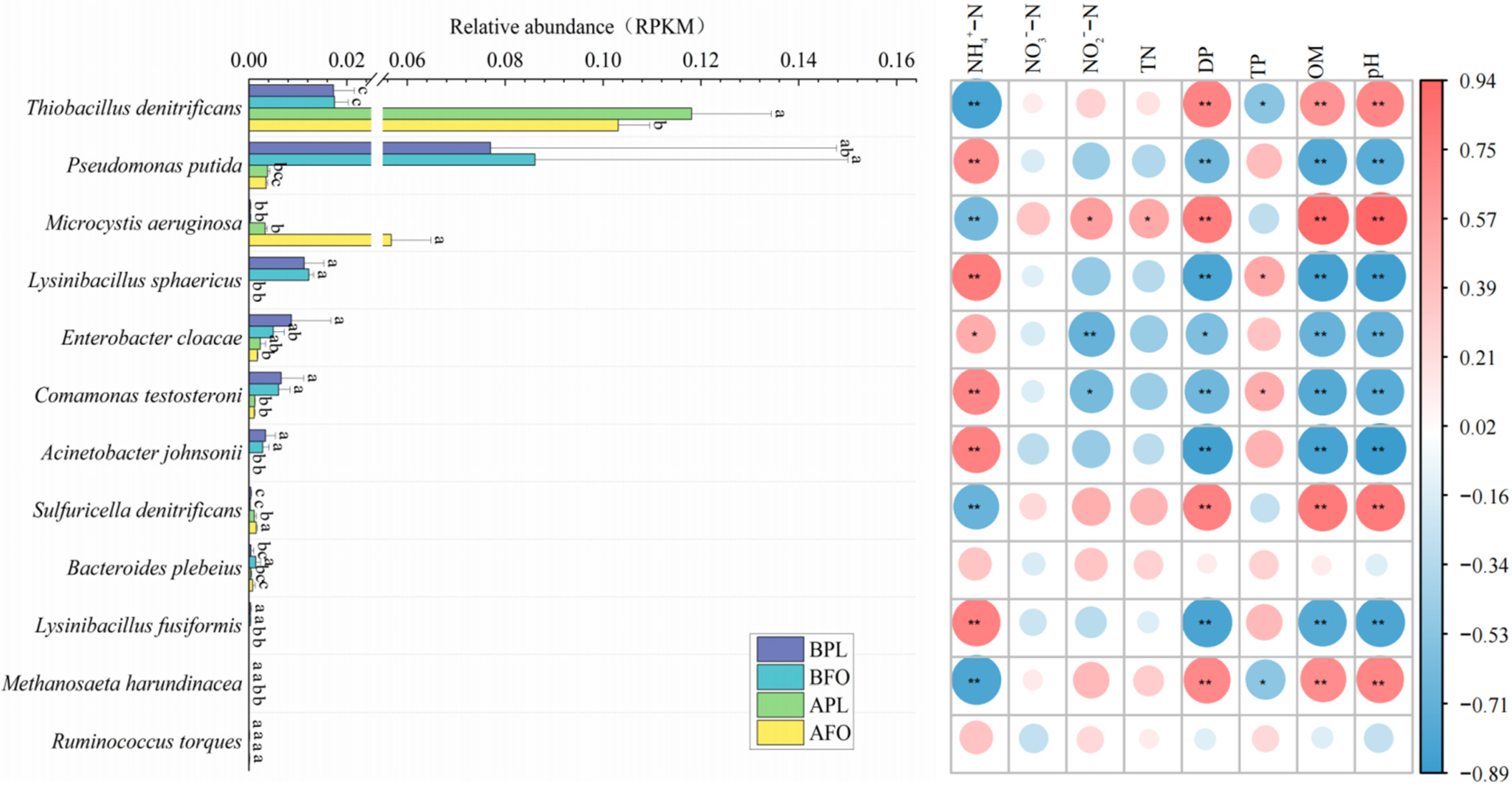
3.6. Differences in Microbial Abundance Involved in Nitrogen Cycle
4. Discussion
4.1. Differences of Nutrient Content and Microbial Abundance in Sediments before and after Culture in Different Culture Models
4.2. Differences in Nitrogen Cycle-Related Enzymes in Sediments before and after Culture in Different Culture Models
4.3. Differences in Nitrogen Cycle-Related Metabolic Pathways in the Sediments before and after Culture in Different Culture Models
4.4. Differences in Nitrogen Cycle-Related Microorganism in the Sediments before and after Culture in Different Culture Models
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Cripps, S.J.; Bergheim, A. Solids management and removal for intensive land-based aquaculture production systems. Aquac. Eng. 2000, 22, 33–56. [Google Scholar] [CrossRef]
- Gondwe, M.J.; Guildford, S.J.; Hecky, R.E. Tracing the flux of aquaculture-derived organic wastes in the southeast arm of Lake Malawi using carbon and nitrogen stable isotopes. Aquaculture 2012, 350, 8–18. [Google Scholar] [CrossRef]
- Lananan, F.; Hamid, S.H.A.; Din, W.N.S.; Khatoon, H.; Jusoh, A.; Endut, A. Symbiotic bioremediation of aquaculture wastewater in reducing ammonia and phosphorus utilizing Effective Microorganism (EM-1) and microalgae (Chlorella sp.). Int. Biodeterior. Biodegrad. 2014, 95, 127–134. [Google Scholar] [CrossRef]
- Conley, D.J.; Paerl, H.W.; Howarth, R.W.; Boesch, D.F.; Seitzinger, S.P.; Havens, K.E.; Lancelot, C.; Likens, G.E. Controlling eutrophication: Nitrogen and phosphorus. Am. Assoc. Adv. Sci. 2009, 323, 1014–1015. [Google Scholar] [CrossRef]
- Jin, J.; Wang, Y.; Wu, Z.; Hergazy, A.; Lan, J.; Zhao, L.; Liu, X.; Chen, N.; Lin, L. Transcriptomic analysis of liver from grass carp (Ctenopharyngodon idellus) exposed to high environmental ammonia reveals the activation of antioxidant and apoptosis pathways. Fish Shellfish Immunol. 2017, 63, 444–451. [Google Scholar] [CrossRef]
- Sun, R.; Wang, F.; Hu, C.; Liu, B. Metagenomics reveals taxon-specific responses of the nitrogen-cycling microbial community to long-term nitrogen fertilization. Soil Biol. Biochem. 2021, 156, 108214. [Google Scholar] [CrossRef]
- Chen, C.; Zhao, T.; Liu, R.; Luo, L. Performance of five plant species in removal of nitrogen and phosphorus from an experimental phytoremediation system in the Ningxia irrigation area. Environ. Monit. Assess. 2017, 189, 497. [Google Scholar] [CrossRef]
- Mohd Nizam, N.U.; Mohd Hanafiah, M.; Mohd Noor, I.; Abd Karim, H.I. Efficiency of five selected aquatic plants in phytoremediation of aquaculture wastewater. Appl. Sci. 2020, 10, 2712. [Google Scholar] [CrossRef]
- Liu, Y.; Lv, J.; Feng, J.; Liu, Q.; Nan, F.; Xie, S. Treatment of real aquaculture wastewater from a fishery utilizing phytoremediation with microalgae. J. Chem. Technol. Biotechnol. 2019, 94, 900–910. [Google Scholar] [CrossRef]
- Lu, S.; Liu, X.; Liu, C.; Wang, X.; Cheng, G. Review of ammonia-oxidizing bacteria and archaea in freshwater ponds. Rev. Environ. Sci. Bio/Technol. 2019, 18, 1–10. [Google Scholar] [CrossRef]
- Brown, M.N.; Briones, A.; Diana, J.; Raskin, L. Ammonia-oxidizing archaea and nitrite-oxidizing nitrospiras in the biofilter of a shrimp recirculating aquaculture system. FEMS Microbiol. Ecol. 2013, 83, 17–25. [Google Scholar] [CrossRef]
- Cao, L.; Wang, W.; Yang, Y.; Yang, C.; Yuan, Z.; Xiong, S.; Diana, J. Environmental impact of aquaculture and countermeasures to aquaculture pollution in China. Environ. Sci. Pollut. Res. Int. 2007, 14, 452–462. [Google Scholar] [CrossRef]
- Shen, L.-D.; Wu, H.-S.; Gao, Z.-Q.; Ruan, Y.-J.; Xu, X.-H.; Li, J.; Ma, S.-J.; Zheng, P.-H. Evidence for anaerobic ammonium oxidation process in freshwater sediments of aquaculture ponds. Environ. Sci. Pollut. Res. 2016, 23, 1344–1352. [Google Scholar] [CrossRef]
- Lu, S.; Liu, X.; Ma, Z.; Liu, Q.; Wu, Z.; Zeng, X.; Shi, X.; Gu, Z. Vertical segregation and phylogenetic characterization of ammonia-oxidizing bacteria and archaea in the sediment of a freshwater aquaculture pond. Front. Microbiol. 2016, 6, 1539. [Google Scholar] [CrossRef]
- Dai, L.; Liu, C.; Yu, L.; Song, C.; Peng, L.; Li, X.; Tao, L.; Li, G. Organic matter regulates ammonia-oxidizing bacterial and archaeal communities in the surface sediments of Ctenopharyngodon idellus aquaculture ponds. Front. Microbiol. 2018, 9, 2290. [Google Scholar] [CrossRef]
- Dong, X.; Greening, C.; Rattray, J.E.; Chakraborty, A.; Chuvochina, M.; Mayumi, D.; Dolfing, J.; Li, C.; Brooks, J.M.; Bernard, B.B. Metabolic potential of uncultured bacteria and archaea associated with petroleum seepage in deep-sea sediments. Nat. Commun. 2019, 10, 1816. [Google Scholar] [CrossRef]
- Bourquin, M.; Busi, S.B.; Fodelianakis, S.; Peter, H.; Washburne, A.; Kohler, T.J.; Ezzat, L.; Michoud, G.; Wilmes, P.; Battin, T.J. The microbiome of cryospheric ecosystems. Nat. Commun. 2022, 13, 3087. [Google Scholar] [CrossRef]
- Zhou, L.; Huang, S.; Gong, J.; Xu, P.; Huang, X. 500 metagenome-assembled microbial genomes from 30 subtropical estuaries in South China. Sci. Data 2022, 9, 310. [Google Scholar] [CrossRef]
- Rathour, R.; Gupta, J.; Mishra, A.; Rajeev, A.C.; Dupont, C.L.; Thakur, I.S. A comparative metagenomic study reveals microbial diversity and their role in the biogeochemical cycling of Pangong lake. Sci. Total Environ. 2020, 731, 139074. [Google Scholar] [CrossRef]
- Wu, L.; Zhang, Y.; Guo, X.; Ning, D.; Zhou, X.; Feng, J.; Yuan, M.M.; Liu, S.; Guo, J.; Gao, Z. Reduction of microbial diversity in grassland soil is driven by long-term climate warming. Nat. Microbiol. 2022, 7, 1054–1062. [Google Scholar] [CrossRef]
- Song, H.; Kerfahi, D.; Takahashi, K.; Nixon, S.L.; Tripathi, B.M.; Kim, H.; Tateno, R.; Adams, J. Metagenomic analysis reveals rapid development of soil biota on fresh volcanic ash. Sci. Rep. 2020, 10, 21419. [Google Scholar] [CrossRef]
- Kelly, C.N.; Schwaner, G.W.; Cumming, J.R.; Driscoll, T.P. Metagenomic reconstruction of nitrogen and carbon cycling pathways in forest soil: Influence of different hardwood tree species. Soil Biol. Biochem. 2021, 156, 108226. [Google Scholar] [CrossRef]
- Liu, S.; Wang, P.; Wang, X.; Chen, J. Ecological insights into the elevational biogeography of antibiotic resistance genes in a pristine river: Metagenomic analysis along the Yarlung Tsangpo River on the Tibetan Plateau. Environ. Pollut. 2021, 286, 117101. [Google Scholar] [CrossRef]
- Chen, J.; Yang, Y.; Ke, Y.; Chen, X.; Jiang, X.; Chen, C.; Xie, S. Sulfonamide-metabolizing microorganisms and mechanisms in antibiotic-contaminated wetland sediments revealed by stable isotope probing and metagenomics. Environ. Int. 2022, 165, 107332. [Google Scholar] [CrossRef]
- Martinez-Porchas, M.; Martinez-Cordova, L.R. World aquaculture: Environmental impacts and troubleshooting alternatives. Sci. World J. 2012, 2012, 389623. [Google Scholar] [CrossRef]
- Bao, S. Soil Agrochemical Analysis; China Agricultural Press: Beijing, China, 2000; Volume 67, p. 72. [Google Scholar]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Wood, D.E.; Salzberg, S.L. Kraken: Ultrafast metagenomic sequence classification using exact alignments. Genome Biol. 2014, 15, R46. [Google Scholar] [CrossRef]
- Wood, D.E.; Lu, J.; Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol. 2019, 20, 257. [Google Scholar] [CrossRef] [PubMed]
- Zhu, W.; Lomsadze, A.; Borodovsky, M. Ab initio gene identification in metagenomic sequences. Nucleic Acids Res. 2010, 38, e132. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, J.; Kim, M.S.; Koh, A.Y.; Xie, Y.; Zhan, X. FMAP: Functional mapping and analysis pipeline for metagenomics and metatranscriptomics studies. BMC Bioinform. 2016, 17, 257. [Google Scholar] [CrossRef] [PubMed]
- Franzosa, E.A.; McIver, L.J.; Rahnavard, G.; Thompson, L.R.; Schirmer, M.; Weingart, G.; Lipson, K.S.; Knight, R.; Caporaso, J.G.; Segata, N. Species-level functional profiling of metagenomes and metatranscriptomes. Nat. Methods 2018, 15, 962–968. [Google Scholar] [CrossRef] [PubMed]
- Vanacker, M.; Wezel, A.; Arthaud, F.; Guérin, M.; Robin, J. Determination of tipping points for aquatic plants and water quality parameters in fish pond systems: A multi-year approach. Ecol. Indic. 2016, 64, 39–48. [Google Scholar] [CrossRef]
- Nordin, A.; Högberg, P.; Näsholm, T. Soil nitrogen form and plant nitrogen uptake along a boreal forest productivity gradient. Oecologia 2001, 129, 125–132. [Google Scholar] [CrossRef]
- Si, G.; Peng, C.; Xu, X.; Xu, D.; Yuan, J.; Li, J. Effect of integrated rice-crayfish farming system on soil physico-chemical properties in waterlogged paddy soils. Zhongguo Shengtai Nongye Xuebao/Chin. J. Eco-Agric. 2017, 25, 61–68. [Google Scholar] [CrossRef]
- Hamilton, E.W., III; Frank, D.A. Can plants stimulate soil microbes and their own nutrient supply? Evidence from a grazing tolerant grass. Ecology 2001, 82, 2397–2402. [Google Scholar] [CrossRef]
- Xia, J.H.; Lin, G.; Fu, G.H.; Wan, Z.Y.; Lee, M.; Wang, L.; Liu, X.J.; Yue, G.H. The intestinal microbiome of fish under starvation. BMC Genom. 2014, 15, 266. [Google Scholar] [CrossRef]
- Kong, W.; Huang, S.; Shi, F.; Yang, Z.; Feng, Y.; Khatoon, Z.; Zhou, J.; Xiao, Y. Study on release kinetics of nitrogen and phosphorus from fish feed. Aquac. Res. 2020, 51, 3216–3229. [Google Scholar] [CrossRef]
- Huo, S.; He, Z.; Ma, C.; Zhang, H.; Xi, B.; Zhang, J.; Li, X.; Wu, F.; Liu, H. Spatio-temporal impacts of meteorological and geographic factors on the availability of nitrogen and phosphorus to algae in Chinese lakes. J. Hydrol. 2019, 572, 380–387. [Google Scholar] [CrossRef]
- Otten, T.; Xu, H.; Qin, B.; Zhu, G.; Paerl, H. Spatiotemporal patterns and ecophysiology of toxigenic Microcystis blooms in Lake Taihu, China: Implications for water quality management. Environ. Sci. Technol. 2012, 46, 3480–3488. [Google Scholar] [CrossRef]
- Kellermann, C.; Griebler, C. Thiobacillus thiophilus sp. nov., a chemolithoautotrophic, thiosulfate-oxidizing bacterium isolated from contaminated aquifer sediments. Int. J. Syst. Evol. Microbiol. 2009, 59, 583–588. [Google Scholar] [CrossRef]
- Hargreaves, J.A. Biofloc Production Systems for Aquaculture; Southern Regional Aquaculture Center: Stoneville, MS, USA, 2013; Volume 4503. [Google Scholar]
- Deng, M.; Hou, J.; Song, K.; Chen, J.; Gou, J.; Li, D.; He, X. Community metagenomic assembly reveals microbes that contribute to the vertical stratification of nitrogen cycling in an aquaculture pond. Aquaculture 2020, 520, 734911. [Google Scholar] [CrossRef]
- Zhu, H.; Yan, B.; Xu, Y.; Guan, J.; Liu, S. Removal of nitrogen and COD in horizontal subsurface flow constructed wetlands under different influent C/N ratios. Ecol. Eng. 2014, 63, 58–63. [Google Scholar] [CrossRef]
- Surey, R.; Lippold, E.; Heilek, S.; Sauheitl, L.; Henjes, S.; Horn, M.A.; Mueller, C.W.; Merbach, I.; Kaiser, K.; Böttcher, J. Differences in labile soil organic matter explain potential denitrification and denitrifying communities in a long-term fertilization experiment. Appl. Soil Ecol. 2020, 153, 103630. [Google Scholar] [CrossRef]
- Van Der Heijden, M.G.; Bardgett, R.D.; Van Straalen, N.M. The unseen majority: Soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol. Lett. 2008, 11, 296–310. [Google Scholar] [CrossRef]
- Bulgarelli, D.; Schlaeppi, K.; Spaepen, S.; Van Themaat, E.V.L.; Schulze-Lefert, P. Structure and functions of the bacterial microbiota of plants. Annu. Rev. Plant Biol. 2013, 64, 807–838. [Google Scholar] [CrossRef]
- Li, H.; Chi, Z.; Li, J.; Wu, H.; Yan, B. Bacterial community structure and function in soils from tidal freshwater wetlands in a Chinese delta: Potential impacts of salinity and nutrient. Sci. Total Environ. 2019, 696, 134029. [Google Scholar] [CrossRef]
- Zhang, T.; Cao, J.; Zhang, Y.; Fang, F.; Feng, Q.; Luo, J. Achieving efficient nitrite accumulation in glycerol-driven partial denitrification system: Insights of influencing factors, shift of microbial community and metabolic function. Bioresour. Technol. 2020, 315, 123844. [Google Scholar] [CrossRef]
- Li, Y.; Ling, J.; Chen, P.; Chen, J.; Dai, R.; Liao, J.; Yu, J.; Xu, Y. Pseudomonas mendocina LYX: A novel aerobic bacterium with advantage of removing nitrate high effectively by assimilation and dissimilation simultaneously. Front. Environ. Sci. Eng. 2021, 15, 57. [Google Scholar] [CrossRef]
- Flores, E.; Frías, J.E.; Rubio, L.M.; Herrero, A. Photosynthetic nitrate assimilation in cyanobacteria. Photosynth. Res. 2005, 83, 117–133. [Google Scholar] [CrossRef]
- Hao, W.; Zhang, J.; Duan, R.; Liang, P.; Li, M.; Qi, X.; Li, Q.; Liu, P.; Huang, X. Organic carbon coupling with sulfur reducer boosts sulfur based denitrification by Thiobacillus denitrificans. Sci. Total Environ. 2020, 748, 142445. [Google Scholar] [CrossRef] [PubMed]
- Beller, H.R.; Chain, P.S.; Letain, T.E.; Chakicherla, A.; Larimer, F.W.; Richardson, P.M.; Coleman, M.A.; Wood, A.P.; Kelly, D.P. The genome sequence of the obligately chemolithoautotrophic, facultatively anaerobic bacterium Thiobacillus denitrificans. J. Bacteriol. 2006, 188, 1473–1488. [Google Scholar] [CrossRef] [PubMed]
- Xiao, M.; Li, M.; Reynolds, C.S. Colony formation in the cyanobacterium Microcystis. Biol. Rev. 2018, 93, 1399–1420. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Qiu, R. Water eutrophication in China and the combating strategies. J. Chem. Technol. Biotechnol. Int. Res. Process Environ. Clean Technol. 2007, 82, 781–786. [Google Scholar] [CrossRef]





| Sample | NH4+ (mg kg−1) | NO3− (mg kg−1) | NO2− (mg kg−1) | TN (g kg−1) | DP (mg kg−1) | TP (g kg−1) | OM (g kg−1) | pH |
|---|---|---|---|---|---|---|---|---|
| FOA | 147.07 ± 13.35 b | 13.27 ± 2.76 a | 1.31 ± 0.19 a | 2.28 ± 0.06 a | 36.47 ± 2.61 a | 1.38 ± 0.07 a | 70.94 ± 0.94 a | 7.88 ± 0.03 a |
| FPA | 67.48 ± 7.07 c | 11.04 ± 0.70 a | 0.62 ± 0.09 b | 2.09 ± 0.08 b | 38.67 ± 6.70 a | 1.23 ± 0.07 b | 64.99 ± 1.39 b | 7.81 ± 0.06 a |
| FOB | 176.34 ± 7.89 a | 12.47 ± 2.98 a | 0.65 ± 0.07 b | 2.12 ± 0.05 b | 17.08 ± 1.97 b | 1.43 ± 0.03 a | 61.73 ± 0.64 c | 7.66 ± 0.06 b |
| FPB | 167.98 ± 3.76 a | 9.62 ± 2.83 a | 0.60 ± 0.07 b | 2.16 ± 0.05 b | 14.22 ± 1.66 b | 1.36 ± 0.06 a | 61.31 ± 1.66 c | 7.64 ± 0.05 b |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, Z.; Yao, Y.; Sun, M.; Li, G.; Zhu, J. Metagenomics Reveal Microbial Effects of Lotus Root–Fish Co-Culture on Nitrogen Cycling in Aquaculture Pond Sediments. Microorganisms 2022, 10, 1740. https://doi.org/10.3390/microorganisms10091740
Yang Z, Yao Y, Sun M, Li G, Zhu J. Metagenomics Reveal Microbial Effects of Lotus Root–Fish Co-Culture on Nitrogen Cycling in Aquaculture Pond Sediments. Microorganisms. 2022; 10(9):1740. https://doi.org/10.3390/microorganisms10091740
Chicago/Turabian StyleYang, Zhen, Yanhong Yao, Meng Sun, Gu Li, and Jianqiang Zhu. 2022. "Metagenomics Reveal Microbial Effects of Lotus Root–Fish Co-Culture on Nitrogen Cycling in Aquaculture Pond Sediments" Microorganisms 10, no. 9: 1740. https://doi.org/10.3390/microorganisms10091740






