1. Introduction
Urinary tract infections (UTIs) affect nearly 250 million people yearly and represent approximately 40% of infections worldwide. They account for 10–20% of nosocomial infections [
1,
2]. UTIs are associated with considerable morbidity and a large spectrum of clinical symptoms, ranging from asymptomatic bacteriuria to cystitis or septic shock, that can lead to life-threatening multiple-organ failure [
3]. Most UTIs have a bacterial origin, and the most frequent cause of the infection is
Enterobacteriaceae [
4]. The most commonly encountered
Enterobacteriaceae are
Escherichia coli,
Klebsiella pneumoniae and
Enterobacter cloacae [
5].
The ability of
Enterobacteriaceae to invade and persist in the uroepithelium depends on several virulence factors and their ability to form biofilms [
6]. Biofilm-forming bacteria are a common cause of recurrent and severe urinary tract infections and are generally multidrug-resistant bacteria [
7]. In addition to the formation of biofilms, resistance to empirical antimicrobial treatments has increased in recent years [
8], especially in Gram-negative bacteria [
9]. Several studies have shown an increase in antimicrobial resistance to the most commonly used antibiotics, such as ciprofloxacin and trimethoprim-sulfamethoxazole, in strains of
Enterobacteriaceae isolated from urinary tract infections [
10,
11]. Recent studies in Africa and Europe reported a substantial increase in Gram-negative bacteria from ESBL-producing urinary tract infections [
12,
13]. Indeed, the spread of ESBL-producing bacteria has made the empirical treatment of infections more difficult and has promoted resistance to beta-lactam antibiotics such as penicillin, cephalosporins and sometimes even carbapenems [
14].
The pathogenicity of
Enterobacteriaceae in urinary tract infections increases with the presence of virulence factors. Indeed,
Enterobacteriaceae strains harbor several virulence genes associated with serious or recurrent urinary tract infections [
15]. Among these genes, P fimbriae (
pap), S-fimbriae (
sfa), hemolysins (
hly), cytotoxic-necrotizing-factor (
cnf1) and Aerobactin (
iucD) are the most relevant [
16]. While
pap and
sfa genes are well known to promote docking, factors associated with the colonization of the host [
17], the
hly,
cnf1 and
fyuA genes, are mainly associated with intracellular survival, iron acquisition, immune system leakage, the inflammatory response and host tissue damage [
18,
19].
Effective management and treatment of urinary tract infections require an in-depth understanding of antimicrobial resistance, virulence genes and biofilm formation in strains of
Enterobacteriaceae isolated from urinary tract infections [
20,
21]. Understanding the link between biofilm formation, the presence of virulence genes and the distribution of antimicrobial resistance in strains of
Enterobacteriaceae implicated in urinary tract infections will also allow for more effective prevention and management strategies [
20]. Thus, the present study analyzed resistance profiles and virulence factors associated with
Enterobacteriaceae-related urinary tract infections in Benin.
4. Discussion
To better understand the pathogenicity of strains causing infections and develop new vaccines and therapeutic targets, it is necessary to identify the susceptibility to antibiotics, the factors associated with the formation of biofilms and the virulence factors of these strains [
7,
30]. These potential predictors help clinicians manage patients and anticipate the evolution of the infection in the host organization [
31]. In this study, we sought to determine the prevalence of
Enterobacteriaceae strains, their antibiotic resistance profiles, their ability to form biofilms and the presence of some urinary tract infection virulence genes in Benin.
During our study,
Enterobacteriaceae were isolated from patients mainly between 21 and 30 years old (22.23%). This age bias can be explained by the fact professional, cultural and sporting activities may contribute to the infection. The results of our study indicate the presence of
Enterobacteriaceae strains in a proportion of 26.41% in urine samples. Our results are lower than those obtained in Mali [
32], which found a prevalence of 76.7% of
Enterobacteriaceae in urine.
Enterobacteriaceae have also been shown to cause 84–87% of UTIs [
33]. This difference may be due to sample size, laboratory strain detection techniques, social demographics, climatic conditions, people’s levels of personal hygiene and healthcare-seeking habits.
Enterobacteriaceae are found in the urine because they can easily contaminate the urinary tract, especially in women, since they are normal flora of the large intestine [
7].
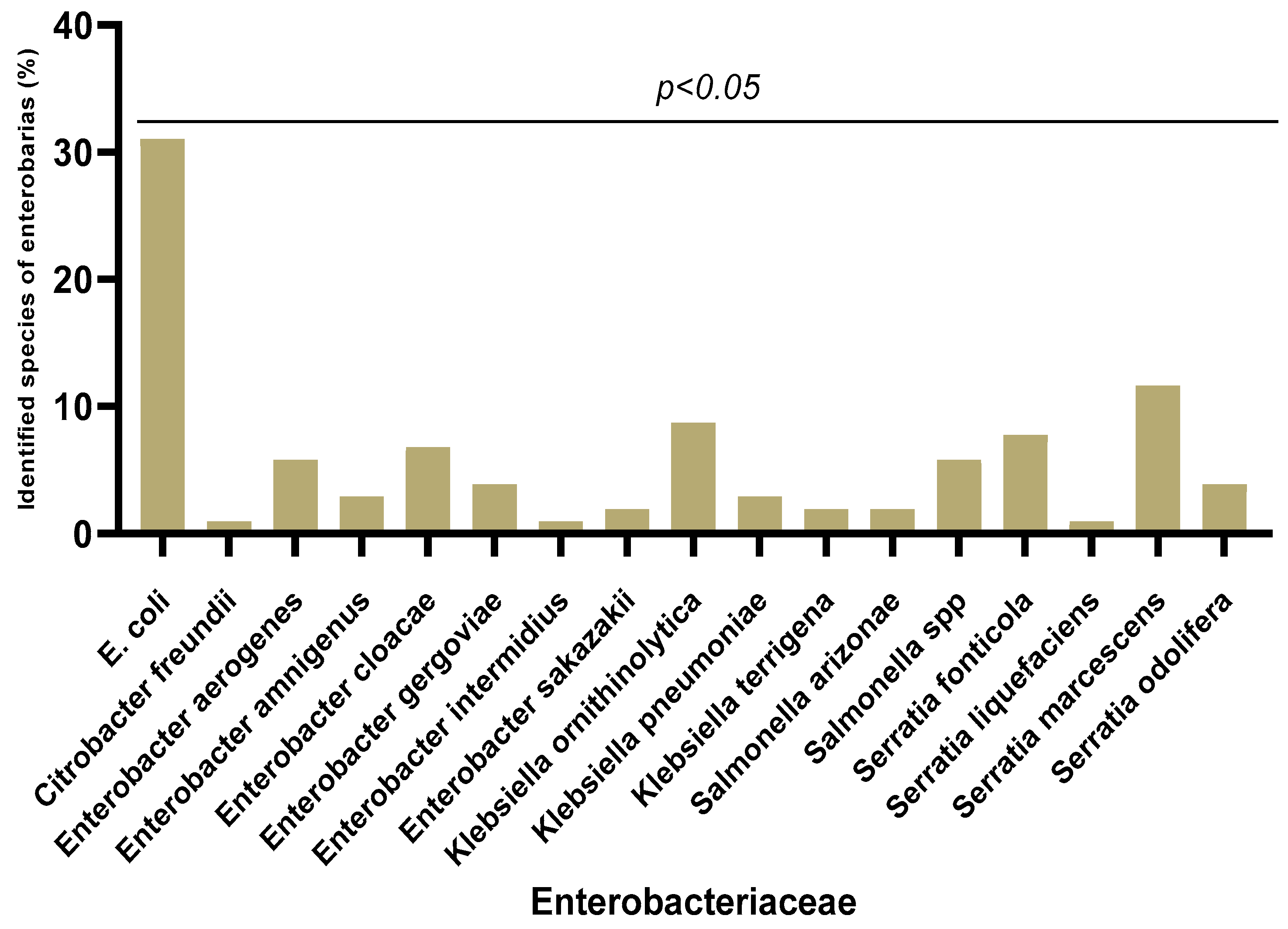
In our study, 18 different species of
Enterobacteriaceae were isolated, with a predominance of
Escherichia coli (30.97%), followed by
Serratia marcescens (11.65%) and
Klebsiella ornithinolytica (8.73%). On the other hand, the weakly represented species were
Serratia liquefaciens,
Citrobacter freundii and
Enterobacter intermidius (0.97%). As expected,
E. coli was the major
Enterobacterales species among the urinary tract samples, which is similar to previous works in other countries [
34,
35]. However, the rate of 30.97% in this study is lower than the 72% recorded in France [
36]. Traditionally,
E. coli has been the dominant uropathogen due to its expression of toxins, adhesins, pili, and fimbriae that allow it to adhere to the uroepithelium [
37]. These protect bacteria from urinary elimination and allow for bacterial multiplication and invasion of uroepithelial tissues. The presence of
S. marcescens as a second isolated species can be explained by the fact that it has a great affinity for the urinary tract [
38].
The resistance of
Enterobacteriaceae isolates varied according to the antibiotics. We observed variable resistance rates depending on the families of antibiotics and the species isolated. High resistance rates (≥80%) were obtained with molecules of the macrolide, cephalosporin and penicillin families. Resistance rates of more than 50% were obtained with molecules from the fluoroquinolone, aminoglycoside and tetracycline families. In a recent study conducted in Benin on surgical patients, a high resistance rate to many of the antibiotics tested was shown [
39]. However, our results are superior to those found in Senegal [
40] and close to those obtained in Algeria [
41]. Antibiotic resistance and the rapid spread of aminoglycosides and lactams such as cephalosporins and fluoroquinolones against uropathogenic bacteria compromise the clinical management of the infection and lead to a poor prognosis [
42]. The multiple-drug resistance (MDR) of pathogenic bacteria may be associated with severe morbidity in urinary tract infections, leading to a major global health problem [
43,
44,
45]. The non-regulation of the use of antibiotics in patients with access to over-the-counter prescriptions, the misuse of certain classes of antimicrobials, frequent self-medication with often random and inappropriate dosages, the premature discontinuation of treatment, the use of antibiotics as growth promoters in agriculture, the use of contraband molecules that are often less dosed or devoid of active ingredients, and unfavorable economic and social conditions are the main drivers that promote the emergence of bacteria that are multi-resistant to antibiotics [
46,
47]. In order to control the spread of this antibiotic resistance, since it represents a serious health problem in Benin, actions to raise awareness of the proper use of antibiotics coupled with monitoring of the acquisition of antibiotics must be implemented. Weak resistance to imipenem was detected. This trend was also found by Hashemi et al. [
48], with an
Enterobacteriaceae resistance rate to imipenem of around 19%. Carbapenems, therefore, remain, to this day, the most active molecules against uropathogenic
Enterobacteriaceae [
49].
The overall frequency of ESBL-producing
Enterobacteriaceae among uropathogenic strains in this study was 6%. A similar prevalence (6.05%) was estimated in Morocco in 2014 [
50]. Recently, the highest rate (56.2%) was reported in Benin among
Enterobacteriaceae samples collected at the Cotonou National Center Hubert Koutoukou Maga university Hospital, Benin [
51]. The differences observed from the study conducted in Benin may be due to the fact that their samples included various infections, whereas our sampling targeted only UTIs. In Europe, the resistance of
Enterobacteriaceae to third-generation cephalosporins ranged from 6.2 to 30.8% among bacterial isolates in 2019 [
52]. Muriuki et al. [
53] reports a similar finding for uropathogenic
E. coli in Kenya between 2015 and 2018. The production of ESBL is probably due to the often-empirical prescriptions of ß-lactams, particularly in ambulatory medicine, while awaiting the results of ECBU. The development of resistance to third-generation cephalosporins is a major cause of prolonged hospitalization of infected patients and limits treatment options agaimst the bacteria [
54]. A 25% prevalence of ESBL production, therefore, creates significant therapeutic problems and will limit or reduce treatment options [
55]. Our results on the production of ESBL by our strains of uropathogenic
Enterobacteriaceae are, therefore, reassuring, but should be monitored.
Of the 103
Enterobacteriaceae isolates tested for biofilm production, 40% formed a biofilm on the microplate. These results are similar to other previous studies [
56]. Biofilms provide a survival strategy for bacteria by positioning them to efficiently use available nutrients and prevent access to antimicrobial agents, antibodies and white blood cells [
57]. It has also been found that biofilms harbor a large number of enzymes that inactivate antibiotics, such as beta-lactamases, and thus, create an island of antimicrobial resistance [
58]. Biofilms, therefore, make our strains more virulent and multi-resistant. Recent studies have revealed that a reduction in oxygen tension in the bladder, combined with the presence of terminal electron receptors in the urine, facilitates the preferential expression of
E. coli [
59]. The expression of other factors, such as cytochrome bd quinol oxidase, promotes biofilm complexity and resistance to extracellular stressors by altering the abundance of extracellular matrix components [
59]. This biofilm formation may be due to curli (functional amyloid) fibers, which constitute the main protein component of many biofilms of Gram-negative bacteria [
60]. The presence of curli fibers in these biofilms provides a competitive advantage in mouse models of urinary tract infection by promoting adhesion to bladder epithelial cells [
61]. This adhesion is further increased by the presence of phospho-ethanolamine cellulose produced simultaneously by UPECs [
61].
Regarding genes encoding for virulence factors, in this study, the distribution of fimbriae was observed.
Enterobacteriaceae harbor the
sfa gene (25.15%). The
papG gene is present in 9.71% and the
focG gene in 8.74% of the
Enterobacteriaceae strains. These results are lower than those obtained in Romania [
62], in Mongolia [
63] and in Egypt [
64]. Fimbriae are required by the bacterium to promote the colonization of surfaces, which helps prevent urinary outflow and allows for infection by the bacterium, which may indicate their critical role in the production and progression of urinary infections. They also play an important role in the formation of biofilms [
65].
In our study, uropathogenic
Enterobacteriaceae secreted toxins such as α-hemolysin (
hly) and cytotoxic necrosis factor 1 (cnf1) with respective presence percentages of 2.91% and 15.53%. These results are similar to studies conducted in Romania (13%) [
61] and Mexico (15.4%) [
66]. These toxins promote the exfoliation of bladder cells and cell lysis, which makes available the iron and nutrients necessary for bacterial growth [
29]. Alpha-hemolysin has been associated with clinical severity in patients with UI and CNF1 with bladder inflammation [
67].
Genes encoding siderophores were detected in our study.
Enterobacteriaceae possessed genes coding for salmochelins
IronB (22.33%), followed by genes coding for aerobactin
iucD (21.36%) and, finally, those coding for yersiniabactin
fyuA (5.82%). Siderophores such as toxins allow the bacterium to mobilize iron. They are essential virulence factors in most pathogenic Gram-negative bacteria [
68].
In the present study, we also identified the presence of an autotransporter (
sat gene) in 6.79% of the isolated strains. This rate is much lower than the 31.1% found in Guadeloupe [
29]. Autotransporters can be self-secreted through the membrane of Gram-negative bacteria [
69] and modify the structure of the host cell. Thanks to these capacities, they make the strains more virulent.
The
sfa gene was the most detected among the virulence genes in the present study. This high prevalence could lead us to consider
sfa as a candidate for a potential vaccine. The difference in the prevalence of virulence genes between our studies and different studies abroad may be due to differences in sample size and methodology. Virulence factors are the product of different genes, which can be detected using the PCR method [
70,
71]. However, due to a possible mutation in the corresponding gene, PCR may not detect the presence of the gene [
31]. Therefore, although this phenomenon is rare, a negative PCR result does not necessarily equate to the absence of the corresponding gene [
31].
In our study, biofilm production is associated with the virulence gene
fyuA and gentamicin resistance. This observation has been made in various other studies. Thus, Stephenson and Brown [
72] reported that biofilm production was significantly associated with resistance to fluoroquinolones [
72]. In another study by Zamani et al. [
73], it was found that biofilm production in UPEC was significantly associated with the
Fim gene [
73]. Our study also found that the presence of
PapG,
iucD and
IronB virulence genes decreases biofilm production. On the other hand, in a survey carried out in Uganda [
17], biofilm production was not associated with any virulence gene or resistance to a particular antibiotic. Thus, further studies are required to better understand the relationship between virulence factors, antibiotic resistance and biofilm formation.