Exploring the Spatial Variation in the Microbiota and Bile Acid Metabolism of the Compound Stomach in Intensively Farmed Yaks
Abstract
1. Introduction
2. Materials and Methods
2.1. Experimental Animals and Sample Collection
2.2. UHPLC-MS/MS High-Throughput Target Quantitative Assay of Bile Acids
2.3. DNA Extraction and Sample Quality Control
2.4. Statistical Analysis
3. Results
3.1. Rumen, Reticulum, Omasum, and Abomasum 16S rRNA Revealed the Diversity and Structure of the Microbiome Associated with an Intensive Feeding System
3.2. Compositional Profiles of the Rumen, Reticulum, Omasum, and Abomasum Bacteria Altered by Intensive Feeding System
3.3. Comparative Analysis of Microbial Differences among the Four Stomach Compartments
3.4. Network Correlation Analysis of Bacteria in the Four Stomach Compartments
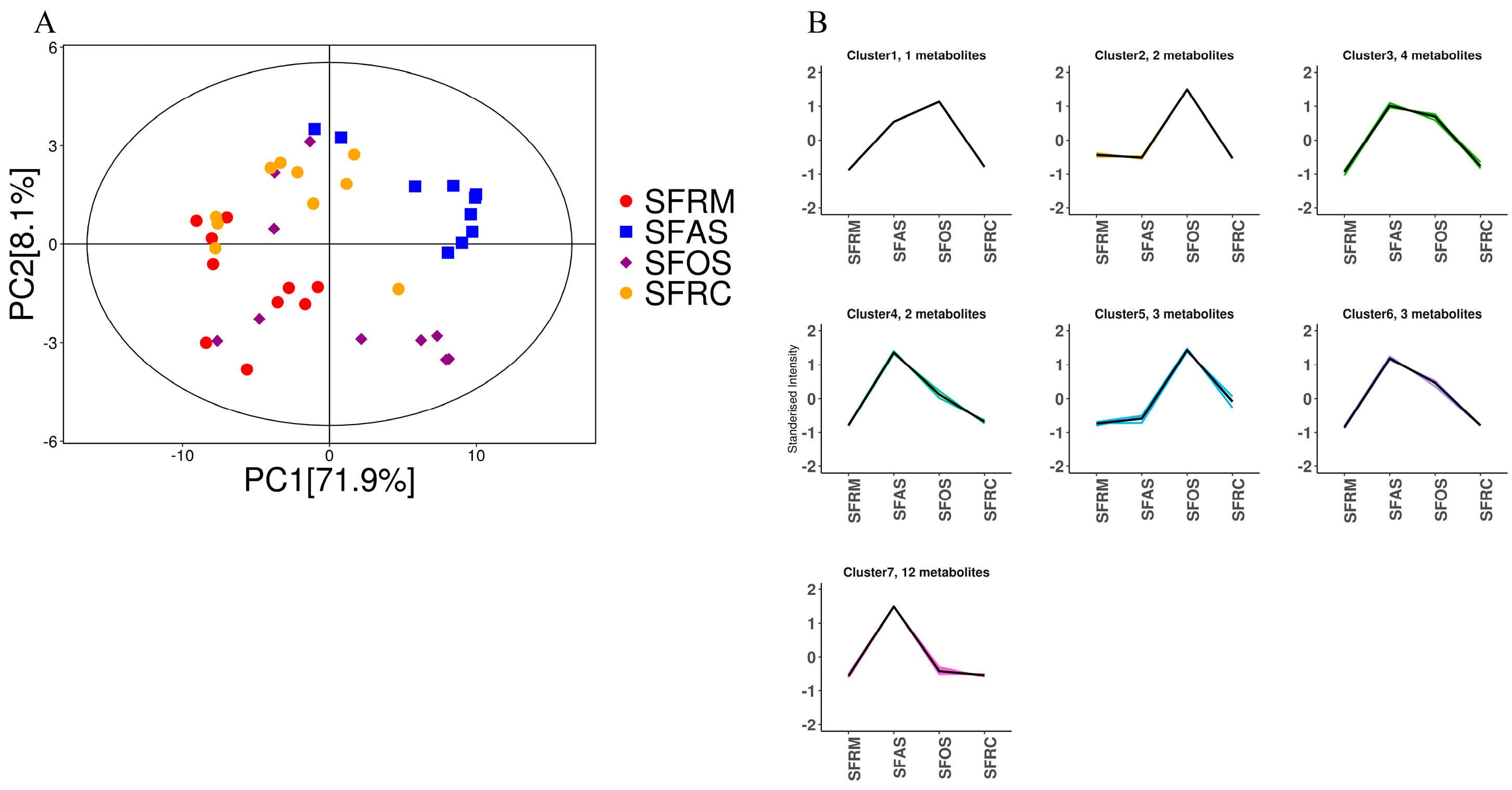
3.5. Bile Acid Analysis of Bacteria in the Four Stomach Compartments
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Dixit, S.; Kumar, S.; Sharma, R.; Banakar, P.S.; Singh, M.; Keshri, A.; Tyagi, A.K. Rumen multi-omics addressing diet-host-microbiome interplay in farm animals: A review. Anim. Biotechnol. 2023, 34, 3187–3205. [Google Scholar] [CrossRef] [PubMed]
- Falony, G.; Joossens, M.; Vieira-Silva, S.; Wang, J.; Darzi, Y.; Faust, K.; Kurilshikov, A.; Bonder, M.J.; Valles-Colomer, M.; Vandeputte, D.; et al. Population-level analysis of gut microbiome variation. Science 2016, 352, 560–564. [Google Scholar] [CrossRef]
- Kaur, H.; Kaur, G.; Gupta, T.; Mittal, D.; Ali, S.A. Integrating Omics Technologies for a Comprehensive Understanding of the Microbiome and Its Impact on Cattle Production. Biology 2023, 12, 1200. [Google Scholar] [CrossRef]
- Lei, Y.; Zhang, K.; Guo, M.; Li, G.; Li, C.; Li, B.; Yang, Y.; Chen, Y.; Wang, X. Exploring the Spatial-Temporal Microbiota of Compound Stomachs in a Pre-weaned Goat Model. Front. Microbiol. 2018, 9, 1846. [Google Scholar] [CrossRef]
- Mizrahi, I.; Wallace, R.J.; Morais, S. The rumen microbiome: Balancing food security and environmental impacts. Nat. Rev. Microbiol. 2021, 19, 553–566. [Google Scholar] [CrossRef]
- Zhou, X.; Gu, M.; Zhu, L.; Wu, D.; Yang, M.; Gao, Y.; Wang, X.; Bai, C.; Wei, Z.; Yang, L.; et al. Comparison of Microbial Community and Metabolites in Four Stomach Compartments of Myostatin-Gene-Edited and Non-edited Cattle. Front. Microbiol. 2022, 13, 844962. [Google Scholar] [CrossRef] [PubMed]
- Faust, K.; Raes, J. Microbial interactions: From networks to models. Nat. Rev. Microbiol. 2012, 10, 538–550. [Google Scholar] [CrossRef]
- McCann, J.C.; Wickersham, T.A.; Loor, J.J. High-throughput Methods Redefine the Rumen Microbiome and Its Relationship with Nutrition and Metabolism. Bioinform. Biol. Insights 2014, 8, 109–125. [Google Scholar] [CrossRef]
- Morgavi, D.P.; Forano, E.; Martin, C.; Newbold, C.J. Microbial ecosystem and methanogenesis in ruminants. Animal 2010, 4, 1024–1036. [Google Scholar] [CrossRef]
- Li, Q.S.; Wang, R.; Ma, Z.Y.; Zhang, X.M.; Jiao, J.Z.; Zhang, Z.G.; Ungerfeld, E.M.; Yi, K.L.; Zhang, B.Z.; Long, L.; et al. Dietary selection of metabolically distinct microorganisms drives hydrogen metabolism in ruminants. ISME J. 2022, 16, 2535–2546. [Google Scholar] [CrossRef]
- Shah, A.M.; Bano, I.; Qazi, I.H.; Matra, M.; Wanapat, M. “The Yak”-A remarkable animal living in a harsh environment: An overview of its feeding, growth, production performance, and contribution to food security. Front. Vet. Sci. 2023, 10, 1086985. [Google Scholar] [CrossRef]
- Xin, J.; Chai, Z.; Zhang, C.; Zhang, Q.; Zhu, Y.; Cao, H.; Zhong, J.; Ji, Q. Comparing the Microbial Community in Four Stomach of Dairy Cattle, Yellow Cattle and Three Yak Herds in Qinghai-Tibetan Plateau. Front. Microbiol. 2019, 10, 1547. [Google Scholar] [CrossRef]
- Xu, Q.; Ungerfeld, E.M.; Morgavi, D.P.; Waters, S.M.; Liu, J.; Du, W.; Zhao, S. Editorial: Rumen microbiome: Interacting with host genetics, dietary nutrients metabolism, animal production, and environment. Front. Microbiol. 2023, 14, 1267149. [Google Scholar] [CrossRef]
- Chai, J.; Liu, Z.; Wu, J.; Kang, Y.; Abdelsattar, M.M.; Zhao, W.; Wang, S.; Yang, S.; Deng, F.; Li, Y.; et al. Dietary beta-hydroxybutyric acid improves the growth performance of young ruminants based on rumen microbiota and volatile fatty acid biosynthesis. Front. Microbiol. 2023, 14, 1296116. [Google Scholar] [CrossRef]
- Zhang, Z.; Xu, D.; Wang, L.; Hao, J.; Wang, J.; Zhou, X.; Wang, W.; Qiu, Q.; Huang, X.; Zhou, J.; et al. Convergent Evolution of Rumen Microbiomes in High-Altitude Mammals. Curr. Biol. 2016, 26, 1873–1879. [Google Scholar] [CrossRef]
- Guo, N.; Wu, Q.; Shi, F.; Niu, J.; Zhang, T.; Degen, A.A.; Fang, Q.; Ding, L.; Shang, Z.; Zhang, Z.; et al. Seasonal dynamics of diet-gut microbiota interaction in adaptation of yaks to life at high altitude. NPJ Biofilms Microbiomes 2021, 7, 38. [Google Scholar] [CrossRef]
- Zhu, Y.; Sun, G.; Luosang-dunzhu; Li, X.; Luosang-zhaxi; Suolang-zhaxi; Suolang; Ciyang; Cidan-yangji; Basang-wangdui; et al. High energy level diet improves the growth performance and rumen fermentation of yaks in cold weather. Front. Vet. Sci. 2023, 10, 1212422. [Google Scholar] [CrossRef]
- Dao, R.; Wu, D.; Wang, H.; Jin, H.; Li, L.; Fu, X.; Sa, C.; Eerdunchaolu. Exploration of the Characteristics of Intestinal Microbiota and Metabolomics in Different Rat Models of Mongolian Medicine. Evid. Based Complement. Altern. Med. 2021, 2021, 5532069. [Google Scholar] [CrossRef]
- Keum, G.B.; Pandey, S.; Kim, E.S.; Doo, H.; Kwak, J.; Ryu, S.; Choi, Y.; Kang, J.; Kim, S.; Kim, H.B. Understanding the Diversity and Roles of the Ruminal Microbiome. J. Microbiol. 2024, 62, 217–230. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Ge, F.; Yao, X.; Guo, X.; Bao, P.; Ma, X.; Wu, X.; Chu, M.; Yan, P.; Liang, C. Microbiome and Metabolomics Reveal the Effects of Different Feeding Systems on the Growth and Ruminal Development of Yaks. Front. Microbiol. 2021, 12, 682989. [Google Scholar] [CrossRef]
- Li, Z.; Zhang, Z.; Xu, C.; Zhao, J.; Liu, H.; Fan, Z.; Yang, F.; Wright, A.D.; Li, G. Bacteria and methanogens differ along the gastrointestinal tract of Chinese roe deer (Capreolus pygargus). PLoS ONE 2014, 9, e114513. [Google Scholar] [CrossRef]
- Zeng, Y.; Zeng, D.; Ni, X.; Zhu, H.; Jian, P.; Zhou, Y.; Xu, S.; Lin, Y.; Li, Y.; Yin, Z.; et al. Microbial community compositions in the gastrointestinal tract of Chinese Mongolian sheep using Illumina MiSeq sequencing revealed high microbial diversity. AMB Express 2017, 7, 75. [Google Scholar] [CrossRef]
- Hu, R.; Zou, H.; Wang, Z.; Cao, B.; Peng, Q.; Jing, X.; Wang, Y.; Shao, Y.; Pei, Z.; Zhang, X.; et al. Nutritional Interventions Improved Rumen Functions and Promoted Compensatory Growth of Growth-Retarded Yaks as Revealed by Integrated Transcripts and Microbiome Analyses. Front. Microbiol. 2019, 10, 318. [Google Scholar] [CrossRef] [PubMed]
- Mao, S.; Zhang, M.; Liu, J.; Zhu, W. Characterising the bacterial microbiota across the gastrointestinal tracts of dairy cattle: Membership and potential function. Sci. Rep. 2015, 5, 16116. [Google Scholar] [CrossRef]
- Salvador, A.C.; Huda, M.N.; Arends, D.; Elsaadi, A.M.; Gacasan, C.A.; Brockmann, G.A.; Valdar, W.; Bennett, B.J.; Threadgill, D.W. Analysis of strain, sex, and diet-dependent modulation of gut microbiota reveals candidate keystone organisms driving microbial diversity in response to American and ketogenic diets. Microbiome 2023, 11, 220. [Google Scholar] [CrossRef]
- Li, Y.; Gao, J.; Xue, Y.; Sun, R.; Sun, X.; Sun, Z.; Liu, S.; Tan, Z.; Zhu, W.; Cheng, Y. Nutrient availability of roughages in isocaloric and isonitrogenous diets alters the bacterial networks in the whole gastrointestinal tract of Hu sheep. BMC Microbiol. 2023, 23, 70. [Google Scholar] [CrossRef]
- Guo, W.; Zhou, M.; Ma, T.; Bi, S.; Wang, W.; Zhang, Y.; Huang, X.; Guan, L.L.; Long, R. Survey of rumen microbiota of domestic grazing yak during different growth stages revealed novel maturation patterns of four key microbial groups and their dynamic interactions. Anim. Microbiome 2020, 2, 23. [Google Scholar] [CrossRef]
- Bradley, P.H.; Pollard, K.S. Proteobacteria explain significant functional variability in the human gut microbiome. Microbiome 2017, 5, 36. [Google Scholar] [CrossRef]
- Xie, F.; Jin, W.; Si, H.; Yuan, Y.; Tao, Y.; Liu, J.; Wang, X.; Yang, C.; Li, Q.; Yan, X.; et al. An integrated gene catalog and over 10,000 metagenome-assembled genomes from the gastrointestinal microbiome of ruminants. Microbiome 2021, 9, 137. [Google Scholar] [CrossRef]
- Just, S.; Mondot, S.; Ecker, J.; Wegner, K.; Rath, E.; Gau, L.; Streidl, T.; Hery-Arnaud, G.; Schmidt, S.; Lesker, T.R.; et al. The gut microbiota drives the impact of bile acids and fat source in diet on mouse metabolism. Microbiome 2018, 6, 134. [Google Scholar] [CrossRef]
- Jia, W.; Xie, G.; Jia, W. Bile acid-microbiota crosstalk in gastrointestinal inflammation and carcinogenesis. Nat. Rev. Gastroenterol. Hepatol. 2018, 15, 111–128. [Google Scholar] [CrossRef] [PubMed]
- Chavez-Talavera, O.; Tailleux, A.; Lefebvre, P.; Staels, B. Bile Acid Control of Metabolism and Inflammation in Obesity, Type 2 Diabetes, Dyslipidemia, and Nonalcoholic Fatty Liver Disease. Gastroenterology 2017, 152, 1679–1694. [Google Scholar] [CrossRef] [PubMed]
- Ghaffarzadegan, T.; Zhong, Y.; Fak Hallenius, F.; Nyman, M. Effects of barley variety, dietary fiber and beta-glucan content on bile acid composition in cecum of rats fed low- and high-fat diets. J. Nutr. Biochem. 2018, 53, 104–110. [Google Scholar] [CrossRef] [PubMed]
- Takei, H.; Narushima, S.; Suzuki, M.; Kakiyama, G.; Sasaki, T.; Murai, T.; Yamashiro, Y.; Nittono, H. Characterization of long-chain fatty acid-linked bile acids: A major conjugation form of 3beta-hydroxy bile acids in feces. J. Lipid Res. 2022, 63, 100275. [Google Scholar] [CrossRef]
- Du, Z.; Luo, Z.; Huang, Y.; Zhou, T.; Ma, L.; Wu, D.; Yao, X.; Shen, L.; Yu, S.; Yong, K.; et al. Screening for potential warning biomarkers in cows with ketosis based on host-microbiota co-metabolism analysis. Front. Microbiol. 2024, 15, 1373402. [Google Scholar] [CrossRef] [PubMed]





| Item | DM | CP | EE | Ash | NDF | ADF | ADL |
|---|---|---|---|---|---|---|---|
| Whole-plant ensiled corn | 67.95 | 9.27 | 1.17 | 12.98 | 61.32 | 37.52 | 2.72 |
| Concentrate | 89.64 | 12.29 | 2.44 | 7.90 | 41.41 | 10.84 | 5.24 |
| TMR | 55.05 | 10.48 | 1.68 | 10.97 | 53.21 | 26.81 | 2.98 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
He, S.; Yuan, Z.; Dai, S.; Wang, Z.; Zhao, S.; Zhang, B.; Mao, H.; Wu, D. Exploring the Spatial Variation in the Microbiota and Bile Acid Metabolism of the Compound Stomach in Intensively Farmed Yaks. Microorganisms 2024, 12, 1968. https://doi.org/10.3390/microorganisms12101968
He S, Yuan Z, Dai S, Wang Z, Zhao S, Zhang B, Mao H, Wu D. Exploring the Spatial Variation in the Microbiota and Bile Acid Metabolism of the Compound Stomach in Intensively Farmed Yaks. Microorganisms. 2024; 12(10):1968. https://doi.org/10.3390/microorganisms12101968
Chicago/Turabian StyleHe, Shichun, Zaimei Yuan, Sifan Dai, Zibei Wang, Shusheng Zhao, Bin Zhang, Huaming Mao, and Dongwang Wu. 2024. "Exploring the Spatial Variation in the Microbiota and Bile Acid Metabolism of the Compound Stomach in Intensively Farmed Yaks" Microorganisms 12, no. 10: 1968. https://doi.org/10.3390/microorganisms12101968
APA StyleHe, S., Yuan, Z., Dai, S., Wang, Z., Zhao, S., Zhang, B., Mao, H., & Wu, D. (2024). Exploring the Spatial Variation in the Microbiota and Bile Acid Metabolism of the Compound Stomach in Intensively Farmed Yaks. Microorganisms, 12(10), 1968. https://doi.org/10.3390/microorganisms12101968






